Abstract
Crustaceans are one of the most widespread and speciose groups of marine organisms, fulfilling multiple ecological roles in numerous ecosystems. On coral reefs, many crustacean species form associations with scleractinian corals. Although the Red Sea is considered a biodiversity hotspot, few studies examined the diversity of coral-associated crustacean communities to date. In this study, 460 decapod crustaceans were recovered from 67 coral colonies of the three branching genera Acropora, Pocillopora and Stylophora in the central Saudi Arabian Red Sea. Crabs and shrimps were morphologically identified to the lowest taxonomic level possible, and portions of the mitochondrial COI and 16S rRNA genes were amplified with the objective of assessing their diversity and phylogenetic relationships. Finally, patterns of co-occurrence were evaluated to investigate the presence of species-specific symbiotic epifauna on different host corals. Overall, we recovered four families, five genera, and nine species of Red Sea crabs, nested into 11 molecular clades, and two families, eight genera and 11 species of shrimps, grouped within 12 lineages. Crabs of the species Trapezia tigrina were found to be exclusively associated with Pocillopora corals, while Tetralia crabs and the shrimps Jocaste japonica and Harpilius lutescens only occurred on Acropora colonies, providing evidence that potential loss of host corals due to local and global impacts could lead to consequent shifts in the symbiotic communities on reefs and to the loss of certain associated taxa. This study represents an advancement towards the understanding and molecular characterization of coral-associated benthic communities in the Red Sea and lays the ground for further research assessing the patterns of biodiversity, evolution, and ecological preferences of these organisms in the area.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tropical coral reefs are known to be one of the most diverse and productive ecosystems on Earth (Eddy et al. 2018), harbouring a wide variety of organisms and providing habitat, shelter, and food to several species (Wagner et al. 2020). These ecosystems host nearly one quarter of the total marine biodiversity, with estimates of up to millions of species, yet only a part of them is formally described (Knowlton et al. 2010; Stella et al. 2011a; Fisher et al. 2015; Hoeksema 2017). While academic research notoriously focused on estimating hard coral and fish biodiversity patterns, other groups of marine organisms have been largely overlooked (Plaisance et al. 2009; Stella et al. 2011a). Most coral reef biodiversity is attributed to the cryptofauna, composed of diverse but understudied invertebrate taxa, for which estimates indicate that around 900 species commonly associate with scleractinian corals (Stella et al. 2011a). For instance, it has been shown that symbionts residing on several host corals highly contribute to the overall reef biodiversity (Molodtsova et al. 2016; van der Schoot and Hoeksema 2024). Coral-associated invertebrates can establish a variety of obligate (i.e. one or both organisms entirely depend on the other for their survival) or facultative (i.e. the organisms involved can generally also survive independently or on other substrates) interactions with their hosts (Gittenberger and Gittenberger 2011; Rouzé et al. 2014; Ivanenko et al. 2018; Maggioni et al. 2022). However, the drivers for the establishment and maintenance of such assemblages are yet to be clarified alongside their vulnerability under various local and global threats including climate change (Gates and Ainsworth 2011; Gibson et al. 2011; Stella et al. 2022). In particular, the composite architecture of branching corals provides a variety of habitat, food, and refugia for invertebrate epifauna, mostly belonging to the phyla Arthropoda and Mollusca, some of which, in turn, offer their hosts protection from predators and cleaning from sediment, developing mutualistic relationships (e.g. coral guard crabs) (Sheppard et al. 2009; Stella et al. 2011a, 2011b; Enochs and Manzello 2012).
Among decapod crustaceans, diverse faunal assemblages with a wide array of specialisations have been observed to depend on the host coral for feeding and refuge (Abele 1976; Vytopil and Willis 2001), while contributing to maintain coral health (Stewart et al. 2006). For example, species of guard crabs belonging to the genus Trapezia Latreille, 1828, are known to protect their host from predators, such as the crown of thorns starfish Acanthaster planci (Linnaeus, 1758) (Glynn 1980; Pratchett 2001; McKeon and Moore 2014), while the shrimp Alpheus lottini Guerin, 1829, has been observed to defend corals from corallivorous molluscs, such as those ascribed to Drupella Thiele, 1925 (McKeon et al. 2012). Moreover, coral-associated decapod crustaceans, such as tetraliid crabs, are known to act as cleaners for their hosts (Stier et al. 2010; Limviriyakul et al. 2016).
In recent years, some initiatives such as the Census of Marine Life (http://www.coml.org), the Moorea Biocode Project (http://bscit.berkeley.edu/biocode), and the Santo expedition in Vanuatu (http://www.santo2006.org) have prioritised the characterization of species using integrated taxonomic approaches. However, research on the epibenthic fauna inhabiting coral reefs in the Saudi Arabian Red Sea is still lagging behind (Edwards and Emberton 1980; Spiridonov and Neumann 2008; Plaisance et al. 2011; Berumen et al. 2013; Britayev et al. 2017). Recently, some studies have focused on the morphology and taxonomy of crabs associated with scleractinians from various regions of the Red Sea (Spiridonov and Neumann 2007; Werding and Hiller 2007; Brösing et al. 2014; Britayev et al. 2017), while other research applied genetic tools to explore invertebrate communities on Automated Reef Monitoring Systems (ARMS) (Al-Rshaidat et al. 2016; Pearman et al. 2018; Carvalho et al. 2019; Villalobos et al. 2022). Nevertheless, studies on coral-associated decapods in the area are still few considering that the region is a recognised marine biodiversity hotspot for multiple groups of metazoans, hosting one of the highest rates of endemism in the world (Briggs 1974; DiBattista et al. 2015; Berumen et al. 2019). Although drivers of evolution of marine organisms in the basin are still debated, the composite geological history of the area, past sea level fluctuations affecting its isolation, and its unique environmental conditions, including extreme temperatures and high levels of salinity, may have influenced the patterns of biodiversity in the Red Sea (DiBattista et al. 2013; Berumen et al. 2019). In this context, the characterization of coral-associated communities is especially important considering that the increasing habitat loss could lead to changes in the structure of the symbiotic community and, eventually, affect highly specialised organisms (Hoegh-Guldberg et al. 2017).
The aim of this study is to characterize the communities of decapod crustaceans living in between the branching corals Acropora Oken, 1815, Pocillopora verrucosa (Ellis & Solander, 1786), and Stylophora pistillata (Esper, 1792), in the area of the Farasan Banks, Saudi Arabia, in the central Red Sea. We applied an integrated morphological and molecular approach to define the identity and evolutionary relationships of the ectosymbionts (Baeza 2015) that we found living on the tissues of the branching corals. Moreover, we investigated the association patterns of the retrieved crustaceans with the three coral genera to verify if patterns of host specificity and rates of co-occurrence could be detected and to understand whether the potential loss of the hosts could ultimately drive the loss of specific associated taxa, when threatened under a climate change scenario.
Materials and methods
Sampling and morphological identification
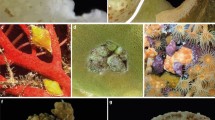
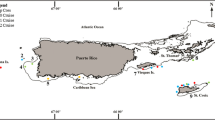
Sampling for the present study took place in May 2017 at 13 reef sites in the area of the Farasan Banks, Saudi Arabia, central Red Sea (Fig. 1a, b). A total of 67 branching coral colonies, about 20 cm in diameter, belonging to Acropora spp. (n = 21), P. verrucosa (n = 18), and S. pistillata (n = 28), were collected by SCUBA diving between 1 and 30 m depth. Before sampling, each coral colony was covered with a plastic zip-lock bag to minimise the loss of associated ectosymbionts. Coral colonies were photographed underwater using a Canon Powershot G15 digital camera in an Ikelite underwater housing and collected using hammer and chisel. The coral colonies were then sorted aboard the research vessel MV Dream Master (Saudi Arabia), identified based on the physical sample and the in situ pictures, and inspected to retrieve ectosymbiotic taxa. Decapod crustaceans were then extracted from the colonies, placed in falcon tubes, labelled, and preserved in 97% ethanol for further analyses.
Crabs and shrimps were morphologically identified at King Abdullah University of Science and Technology (KAUST, Thuwal, Saudi Arabia), where each specimen was separated from the others, photographed under a stereomicroscope, and labelled prior to storage in 95% ethanol. Decapod crustaceans were identified to the lowest possible taxonomic level based on diagnostic morphological traits (e.g. Galil 1987; Castro 1997; Castro et al. 2004; McKeon and Moore 2014; Castro 2015; Rouzé et al. 2017) and by consulting expert taxonomists. Specimens are stored at KAUST.
DNA extraction, amplification, and sequencing
Total genomic DNA was extracted from all collected symbionts. The last pereiopod of each specimen was sub-sampled from each individual crab and shrimp for DNA extraction using a Dneasy® Blood and Tissue kit (Qiagen Inc., Hilden, Germany) according to the manufacturer’s protocol. Extracted DNA quantity and quality were assessed using a NanoDrop® 2000c spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA).
Two regions were amplified using polymerase chain reactions (PCRs). A portion of the cythochrome c oxydase subunit I gene (COI) was amplified using the primers LCO1490 (5′ – GGT CAA CAA ATC ATA AAG ATA TTG G – 3′) and HCO2198 (5′ – TGA TTT TTT GGT CAC CCT GAA GTT TA – 3′) (Folmer et al. 1994) and a portion of the 16S rRNA gene (16S) using the primers 16H10 (5′ – AAT CCT TTC GTA CTA AA – 3′) (Schubart 2009) and 16L2 (5′ – TGC CTG TTT ATC AAA AAC AT – 3′) (Schubart et al. 2000). Reactions were performed in a final volume of 15 µL obtained with 1.2 µL of DNA, 1.5 µL of each primer (10 µM), 3.3 µL of H2O, and 7.5 µL of 2 × Multiplex PCR Master Mix (Qiagen, Hilden, Germany). The following temperature conditions were used for the amplification of COI: 95 °C for 15 min, followed by 30 cycles of 94 °C for 30 s, 46 °C for 1 min, and 72 °C for 1 min, followed by a final extension at 72 °C for 7 min. The temperature profile used for the amplification of 16S was as following: 95 °C for 15 min, followed by 39 cycles of 95 °C for 5 s, 47 °C for 1 min, and 72 °C for 1 min, followed by a final extension at 72 °C for 10 min.
All PCR products were purified adding 2 µL lllustra™ ExoProStar™ (Global Life Sciences Solutions Operations UK Ltd, Buckinghamshire, UK) to 5 µL of amplified DNA in a final volume of 7 µL followed by incubation for 15 min at 37 °C and for 15 min at 80 °C. COI and 16S purified products were sequenced in both forward and reverse directions using an ABI 3730xl DNA analyser (Applied Biosystems, Waltham, MA, USA) at KAUST BioSciences Core Laboratories (Thuwal, Saudi Arabia).
Molecular data analyses
Forward and reverse sequences were assembled and edited using Geneious® v.2021.2.2 (Biomatters Ltd., Auckland, New Zealand). Previously deposited sequences available on public databases (GenBank and BOLD) (Ratnasingham and Hebert 2007) were included to the newly produced dataset. Multiple sequence alignments were performed using MAFFT v.7.490 (Katoh and Standley 2013) with the E-INS-I option. Alignments were manually inspected and edited using AliView v.1.28 (Larsson 2014). Newly obtained sequences were deposited in GenBank database (Accession numbers: Online Resource 1).
Phylogenetic trees were inferred using maximum likelihood (ML) with RAxML v.2 (Stamatakis 2014) and Bayesian inference (BI) with MrBayes v.3.2.6 (Ronquist et al. 2012) on the CIPRES server (Miller et al. 2010). Prior to running phylogenetic analyses, appropriate evolutionary models were selected using jModelTest2 (Darriba et al. 2012) on the CIPRES server (Miller et al. 2010), resulting in the model GTR + I + G for COI and GTR + G for 16S. Maximum likelihood analyses were performed using default parameters and 1000 bootstrap replicates. For Bayesian analyses, two independent runs for four Markov chains were conducted for 10 million generations, with trees sampled every 1000th generation, and burn-in was set to 25%. Inter- and intraspecific genetic distances were calculated using MEGA v.11 with 1000 bootstrap replicates (Tamura et al. 2021. See Online Resource 2).
Statistical analyses
The occurrence of decapod crustaceans on their host corals and the patterns of association between different species of crabs and shrimps were assessed through correlation analyses performed using the R package corrplot (Wei and Simko 2021). Graphs were designed using the R packages ggplot2 (Wickham 2016), corrplot (Wei and Simko 2021) and ggpubr (Kassambara 2023). All statistical analyses were performed using Rstudio v.4.2.2 (R Core Team 2022).
Results
Morphological diversity and identification of coral-associated decapods
A total of 66 coral colonies out of the 67 collected were inhabited by decapod crustacean symbionts. A total number of 460 decapod crustacean individuals, including 301 crabs and 159 shrimps, was retrieved, representing six families, 14 genera, and 23 morphospecies (Online Resource 1). The total number of decapod crustaceans (crabs and shrimps) residing within each coral colony ranged from 2 to 11 in Acropora spp. colonies (mean = 4.6 ± 2.4 standard error SE), from 0 to 26 in P. verrucosa colonies (mean = 8.6 ± 7.5 SE), and from 1 to 25 in S. pistillata colonies (mean = 7.4 ± 6.7 SE) (Fig. 2).
Most of the specimens collected were morphologically identified to species level, while poorly preserved samples were only identified to genus level. Morphological analyses revealed that the collected symbiotic crabs could be ascribed to the three families: Trapeziidae Miers, 1886, Tetraliidae Castro, Ng and Ahyong, 2004, and Xanthidae MacLeay, 1838. Crabs belonging to the family Trapeziidae included the four morphospecies: Trapezia bidentata (Forskål, 1775), Trapezia cymodoce (Herbst, 1801), Trapezia guttata Rüppell, 1830, and Trapezia tigrina Eydoux and Souleyet, 1842. Representatives of the family Tetraliidae belonged to the three different morphospecies Tetralia cavimana Heller, 1860, Tetralia glaberrima (Herbst, 1790), and Tetralia nigrolineata Serène & Pham, 1957. Non-strictly coral-symbiotic crabs (i.e. taxa that can also be free-living independently of the host coral) included representatives of the family Xanthidae, namely Luniella spinipes (Heller, 1860) and Actaea spp., and of the family Pilumnidae Samouelle, 1819, namely the genus Pilumnus Leach, 1816.
The symbiotic shrimps collected belonged to two families: Alpheidae Rafinesque, 1815, and Palaemonidae Rafinesque, 1815. The family Alpheidae was mostly represented by the species Alpheus lottini Guérin, 1829, and the genus Synalpheus Spence Bate, 1888. Shrimps belonging to the family Palaemonidae included the species Harpiliopsis depressa (Simpson, 1860) and Periclimenes madreporae (Bruce, 1969) and the genera Jocaste Holthuis, 1952, Exoclimenella Bruce, 1995, and Harpilius Dana, 1852.
Alignments and sequence data
For the crabs, we successfully amplified 150 COI sequences and 148 16S sequences. Newly obtained sequences were combined with 107 COI and 64 16S sequences previously deposited in GenBank and BOLD (Online Resource 1). The COI alignment comprised 528 bp, including 248 conserved and 280 variable sites, while the 16S a total of 503 bp, with 211 conserved and 283 variable sites. For the shrimps, we obtained 104 COI and 78 16S sequences. Newly generated sequences were combined with 43 and 26 sequences from GenBank for COI and 16S, respectively (Online Resource 1). The COI alignment comprised 640 bp, with 312 conserved and 328 variable sites, while the 16S alignment was composed of 505 bp, including 186 conserved and 307 variable sites. For the remaining 131 specimens, either they did not successfully amplify with any of the two markers or the obtained sequences were not readable.
Phylogenetic analyses
Both ML and BI phylogenetic reconstructions resolved the same major clades for both crabs and shrimps. However, the BI trees provided a better resolution of the phylogenetic hypotheses presented. Hence, we reported the BI topologies (Figs. 3 and 4), including BI posterior probabilities and ML bootstrap values at nodes.
Bayesian inference phylogenetic reconstruction of the symbiotic crabs in association with branching corals of the genera Acropora, Pocillopora, and Stylophora sampled in the Central Red Sea based on two molecular markers: a COI (Alpheus lottini was selected as outgroup) and b 16S (Alpheidae sp. was chosen as outgroup). Node values correspond to Bayesian posterior probability (≥ 0.7) and maximum likelihood bootstrap values (≥ 70%). Taxa for which new sequences were obtained in this study are in bold
Bayesian inference phylogenetic reconstruction of the symbiotic shrimps in association with branching corals of the genera Acropora, Pocillopora, and Stylophora sampled in the Central Red Sea based on two molecular markers: a COI (Trapezia cymodoce was selected as outgroup) b 16S (Trapezia cymodoce was chosen as outgroup). Node values correspond to Bayesian posterior probability (≥ 0.7) and maximum likelihood bootstrap values (≥ 70%). Taxa for which new sequences were obtained in this study are in bold
The crab sequences analysed fell within a total of 33 clades in the COI phylogenetic hypotheses and within 21 molecular clades in the 16S reconstruction. In particular, for the COI, crabs associated with coral colonies from the Red Sea were included in seven molecular clades (clade III, clade V, clade VII, clade XI, clade XVII, clade XXVI, clade XXVIII) and for the 16S within 11 different molecular clades (clade III, clade V, clade VII, clade XI, clade XVII, clade XXVI, clade XXVIII, clade XXXIII, clade XXXVI, clade XXXVII, clade XXXVIII). In the COI reconstruction, the genus Trapezia nested into four clades, clade III (T. guttata), clade V (T. bidentata), clade VII (T. tigrine), and clade XI (T. cymodoce) (Fig. 3a). These four clades included sequences from different locations (Red Sea and Reunion Island for T. guttata (clade III), Red Sea and New Caledonia for T. cymodoce (clade XI), Red Sea, French Polynesia, and Mexico for T. tigrina (clade VII)). In the 16S reconstruction instead, representatives of the genus Trapezia clustered into six molecular clades, namely clade III (T. guttata), clade V (T. bidentata L1), clade XXXVII (T. bidentata L2), clade VII (T. tigrina), clade XI (T. cymodoce), and clade XXXVIII (Trapezia sp.) (Fig. 3b). Five of these six clades also included sequences from different locations (Red Sea, New Caledonia, and Reunion Island for T. guttata (clade III), Red Sea and Reunion Island for T. bidentata L1 (clade V), New Caledonia and Palmyra Atoll for T. bidentata L2 (clade XXXVII), and Red Sea and Philippines for T. tigrina (clade VII)). One clade (clade XXXVIII (Trapezia sp.)) only included sequences from the Red Sea. Thus, the 16S reconstruction allowed to distinguish two lineages of T. bidentata (L1 (clade V) and L2 (clade XXXVII)), which instead clustered into a single clade in the COI reconstruction (clade V), and to further report an additional lineage of Trapezia sp. (clade XXXVIII), which grouped two specimens from our Red Sea dataset which sequences were not readable when amplified with the COI marker. With regard to the genus Tetralia, morphological analyses revealed the presence of three morphospecies among our samples, namely T. cavimana, T. glaberrima, and T. nigrolineata. Yet, all the Red Sea material fell within the same molecular clade (clade XVII) in both the COI and the 16S trees independently of the morphology. The COI reconstruction revealed two additional clades for this genus, grouping previously deposited sequences from New Caledonia and Papua New Guinea, which were not available for the 16S marker, namely clade XVIII (T. ocucaerulea) and clade XIX (Trapezia sp.), respectively (Fig. 3b). Finally, non-symbiotic crabs from the Red Sea fell within two clades in the COI phylogenetic tree (Fig. 3a), clade XXVI (Luniella spinipes) and clade XXVIII (Actaea sp.), and within four clades in the 16S reconstruction (Fig. 3b), namely, clade XXVI (Luniella spinipes), clade XXVIII (Actaea sp.), clade XXXIII (Pilumnus sp.), and clade XXXVI (Pilumnus sp.).
When considering shrimps sequences, the COI analyses revealed the presence of 32 molecular clades, while the 16S reconstruction identified 20 molecular clades. The material from the Red Sea examined for the present study fell within 12 molecular clades (clade I, clade IV, clade VIII, clade XI, clade XII, clade XIV, clade XVI, clade XVII, clade XIX, clade XXI, clade XXIII, clade XXIV) when amplified with the COI marker and within eight molecular clades (clade I, clade VIII, clade XI, clade XII, clade XVII, clade XXIII, clade XXIV, clade XXXIV) when considering the 16S phylogeny reconstructions. Species of the genus Alpheus fell within three clades based on the COI locus, namely clade I (A. lottini) (clade A sensu Williams et al. (2002) and Van Wormhoudt et al. (2019)), clade IV (Alpheus bucephalus Coutière, 1905), and clade VIII (Alpheus bucephaloides Nobili, 1905) (Fig. 4a). However, the genus Alpheus was only represented by two clades from the Red Sea in the tree inferred from 16S: clade I (A. lottini) and clade VIII (A. bucephaloides) (Fig. 4b). In both phylogeny reconstructions, two clades included representatives of the genus Synalpheus, namely clade XXIII (Synalpheus triunguiculatus (Paulson, 1875)) and clade XXIV (Synalpheus charon (Heller, 1861)) (Fig. 4a, b). Although the genera and species of shrimps retrieved in this study belonged to two families, namely Alpheidae and Palaemonidae, the phylogenetic relationships between the two families could not be further clarified by the molecular analyses performed for this study (Fig. 4; Online Resource 4). Moreover, both COI and 16S phylogenetic analyses revealed three additional molecular clades of symbiotic shrimps from the Red Sea material, namely clade XI (Harpiliopsis depressa), clade XII (Jocaste japonica) (Ortmann, 1890), and clade XVII (Palaemonella pottsi) (Borradaile, 1915) (Fig. 4a, b). Finally, while the COI trees identified one molecular clade for the species Harpilius lutescens (clade XIX), one clade for the genus Exoclimenella (clade XIV), and two clades of the genus Periclimenes (clade XVI and XXI) (Fig. 4a), the 16S phylogeny reconstructions only revealed one clade for the species P. madreporae (clade XXXIV) (Fig. 4b).
Composition and species co-occurrence of decapod ectosymbiont communities
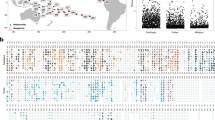
Crabs belonging to the species T. guttata mainly occurred within S. pistillata colonies (98%; n = 136), while T. tigrina was observed to be exclusively associated with P. verrucosa (100%; n = 21) (Fig. 5a). Trapezia bidentata and T. cymodoce were observed to be associated with both P. verrucosa (80% and 17%, respectively) and S. pistillata (20% and 83%, respectively) corals (Fig. 5a). Crabs belonging to the genus Tetralia were solely associated with Acropora colonies (100%) (Fig. 5a). Alpheus shrimps and Harpiliopsis depressa were commonly associated with both P. verrucosa (53% and 72%, respectively) and S. pistillata (47% and 28%, respectively) (Fig. 5a). The shrimps J. japonica and Harpilius lutescens were exclusively found on Acropora colonies (100%) (Fig. 5a). Lastly, representatives of the genera Cuapetes Clark, 1919, and Periclimenes Costa, 1844, which are not commonly known as Acropora symbionts (Stella et al. 2011a; Horká et al. 2016; Frolová et al. 2022), were also retrieved from the sampled Acropora colonies.
a Presence-absence matrix showing the occurrence of the 460 collected decapod crustaceans ectosymbionts with the three different branching host coral taxa. b Correlation plot showing the co-occurrence of decapod crustaceans ectosymbionts. In the bottom bar, “1” indicates a positive linear correlation between the two species considered, thus the presence of the two species of decapod crustaceans at the same time within the same host coral; “0” and “-1” indicate no linear correlation and negative linear correlation between two species, respectively, hence suggesting that the two associated crustaceans never occur with the same host coral. Colour depth and size of the circles indicate the strength of the correlation. Decapods that were only identified to genus level or not recognised as symbionts of the branching scleractinian corals considered were excluded from the correlation analysis
When investigating the co-occurrence of different species of decapod crustaceans within the same coral colonies, a correlation analyses showed a strong and significant correlation (p ≤ 0.001) in the presence of T. bidentata with T. tigrina (r = 0.67) and Harpiliopsis depressa (r = 0.62), T. tigrina with A. lottini (r = 0.4) and H. depressa (r = 0.55), A. bucephalus with A. bucephaloides (r = 0.89), H. depressa with P. pottsi (r = 0.41), and S. triunguiculatus with S. charon (r = 0.48) (Fig. 5b; Online Resource 3). A significant positive correlation (p ≤ 0.01) was also found when considering the presence of A. lottini with T. bidentata (r = 0.36), T. cymodoce (r = 0.31), and H. depressa (r = 0.36), T. cymodoce with S. charon (r = 0.33), and P. pottsi with S. triunguiculatus (r = 0.32) (Fig. 5b; Online Resource 3). Other species pairs, although sporadically observed together, did not show any significant correlation in their co-occurrence within the hosts (Fig. 5b; Online Resource 3).
Discussion
Decapod crustaceans represent most of the coral-associated fauna reported in the literature (Alonso-Domínguez et al. 2022). Although limited research is available on crustacean presence and abundance in the Red Sea and to our knowledge the association of Decapoda with branching corals in this region was not previously investigated through a molecular approach, the species occurrence observed in the present study is consistent with findings reported from different areas of the world (Rouzé et al. 2017; Pisapia et al. 2020). When analysing the biodiversity of decapod crustaceans associated with P. verrucosa, our findings confirmed those of Britayev et al. (2017), who observed the presence of four species of the crab genus Trapezia (T. bidentata, T. tigrina, T. guttata, T. cymodoce) and of three shrimps (A. lottini, H. depressa, and P. madreporae) along the northern Saudi Arabian coast of the Red Sea. Moreover, along the Sudanese coastline of the Red Sea, Edwards and Emberton (1980) reported the occurrence of the crab T. guttata and the shrimps A. bucephaloides, S. charon, and H. depressa as symbionts of S. pistillata colonies, which is in line with the patterns of ectosymbiont occurrence observed in the present study. Our results were also in agreement with those of Pisapia et al. (2020), who reported Trapezia species to be the most abundant taxa with the coral family Pocilloporidae Gray, 1840 in Moorea, French Polynesia. Trapeziidae crabs observed in our study were similar to those reported by Rouzé et al. (2017) from New Caledonia, highlighting the presence of the species T. guttata (clade I), T. bidentata (clade V), T. tigrina (clade VII), and T. cymodoce (clade XI). Our specimens of the shrimp species A. lottini were all included within the lineage of A. lottini L1, described by Rouzé et al. (2017) from New Caledonia and Reunion Island. Although a second lineage of A. lottini was previously observed in New Caledonia and the Pacific Ocean (A. lottini L2, sensu Rouzé et al. (2017); clade B sensu Knowlton and Weigt (1997) and Van Wormhoudt et al. (2019)), none of our specimens appeared to be related to such lineage. This suggests that the A. lottini L1 lineage may be a widespread taxon distributed from the Red Sea to the Pacific Ocean, while the A. lottini L2 lineage may have a narrower geographic distribution. However, due to a lack of enough comparative material encompassing all biogeographical regions, the divergence of Red Sea lineages could not be ascertained. In future studies, next-generation sequencing approaches (e.g. the target enrichment of Ultra Conserved Elements and Exons) (Wolfe et al. 2019) could elucidate species boundaries and the actual geographical distribution of the different lineages in the Indo-Pacific region.
When looking at different host coral species, we observed variation in the composition of the decapod crustacean communities. While, on average, P. verrucosa coral colonies had a comparable number of associated crabs and shrimps individuals, the latter represented a smaller portion of symbiotic individuals in comparison to Acropora corals and less than half of the observations considering S. pistillata hosts (Fig. 2). Both alpheid shrimps and trapeziid crabs are recognised to be highly dependent on their hosts, showing species-specific patterns of association with the corals (Vytopil and Willis 2001; Stella et al. 2010), as well as a high territoriality and species-specific ecological traits (Lassig 1977; Rouzé et al. 2017). Accordingly, our study showed associations between Alpheus and Trapezia species and pocilloporid corals, in particular considering the snapping shrimp A. lottini and the crabs T. tigrina, T. cymodoce, and T. bidentata. The specific association between the lineage of A. lottini (L1) and T. cymodoce was also reported by Rouzé et al. (2017) from New Caledonia, suggesting that this shrimp species may provide a beneficial contribution to T. cymodoce by cleaning their chelipeds (Lassig 1977). The presence of both trapeziid crabs and alpheid shrimps on Pocillopora colonies was also observed by Huber (1987) and Castro (1996), highlighting their success in excluding other crustacean taxa with similar demands from their ecological niches, which could explain the species-specificity of their occurrence (Chomitz et al. 2023). Moreover, the co-occurrence of alpheid shrimps with other Trapezia species was reported by Stier et al. (2012), confirming the synergistic effects of multiple ectosymbionts occurring on branching corals (Billick and Case 1994; McKeon et al. 2012). Interestingly, Hoeksema and Fransen (2011) found that various shrimps species belonging to Palaemonidae and Thoridae Kingsley, 1878, co-occurred in the scleractinian Heliofungia actiniformis (Quoy and Gaimard, 1833) by living in different parts of the coral host, an aspect that unfortunately we did not investigate during our underwater sampling. As such, species of crabs and shrimps may co-occur on coral colonies to combine their defensive strategies and enhance the chances of survival of their hosts against predators (e.g. corallivorous starfish and gastropods) (McKeon et al. 2012). While their co-occurrence did not appear to be significantly correlated, Tetralia crabs were observed to share their habitat within Acropora colonies with the palaemonid shrimps H. lutescens and J. japonica, possibly due to their common substrate preference (Limviriyakul et al. 2016). Rouzé et al. (2017) also highlighted that the strength of the interactions between crabs and shrimps species may vary depending on their geographical locality. Hence, such behaviours and patterns of symbiont associations should be further investigated in the Red Sea and the wider Indo-Pacific region, to better elucidate their interactions.
The patterns of association of decapod crustaceans highlighted in our study are particularly significant not only when looking at the interactions between crustacean taxa but also when analysing the occurrence of symbionts with the hosts. While the data here reported refer to a single sampling effort, coral-associated fauna may be subject to temporal variation (Alvarado and Vargas-Castillo 2012), and further sampling could be needed to test whether the decapod community composition associated with branching corals changes through time and under particular conditions (e.g. seasonality). Although the coral colonies collected for this study were not bleached, nor presented evidence of partial mortality, sampling was performed after the 2015–2016 bleaching event, which affected coral reefs globally and in the Red Sea (Monroe et al. 2018), and could have impacted the associated benthic communities as well (Britayev et al. 2023).
The association of certain crustacean species (e.g. the genus Tetralia and the species Trapezia tigrina, Jocaste japonica, and Harpilius lutescens) with specific corals takes on relevance considering the current status of coral reef ecosystems. The challenges coral reefs are facing in a scenario of climate change (Hoegh-Guldberg et al. 2017; Hughes et al. 2018a), ocean acidification (Pandolfi et al. 2011; Andersson and Gledhill 2012), and anthropogenic stressors (Burke et al. 2011; Hughes et al. 2018b) could in fact have implications not only on the host corals, but also on the associated benthic communities, threatening their biodiversity (Hoeksema 2017). Accordingly, the loss of significant coral taxa could lead to habitat depletion for obligate symbionts and shifts in the structure of reef ecosystems and communities, eventually leading to the extinction of highly specialised symbionts occurring within a limited range of hosts. For instance, Stella et al. (2011b) demonstrated that bleaching of Pocillopora damicornis (Linnaeus, 1758) colonies in Lizard Island, Australia, negatively affected the occurrence of the obligate symbionts T. cymodoce crabs, significantly lowering their density and fecundity within a few weeks.
As the frequency of marine heatwaves and bleaching events has increased in the Red Sea (Genevier et al. 2019), leading to mortality of branching corals, according to our results, this could lead to the loss of habitat availability for associated decapod crustaceans and consequent decrease in the occurrence and abundance of host-specific genera (Britayev et al. 2023). While Furby et al. (2013) reported the families Acroporidae Verrill, 1901, and Pocilloporidae to be the most abundant coral genera in shallow-water reefs of the central Saudi Arabian Red Sea, they found 33% of the Acropora colonies to be affected by the 2010 bleaching event, compared to 19% of the Pocillopora coverage. For instance, the higher susceptibility documented for acroporids during major bleaching events in the central Red Sea (Furby et al. 2013; Monroe et al. 2018) could lead to a loss of Tetralia crabs and of shrimps of the species J. japonica and H. lutescens, which in our study were solely observed in association with Acropora colonies (Fig. 5). Considering that Robitzch et al. (2015) found a single population of P. verrucosa in our study area, mortality of these corals could negatively affect trapeziid crabs and alpheid shrimps living in between their branches, as shown by our results (Fig. 5). Such loss of associated organisms would ultimately affect their functional roles (e.g. cleaning and protection of the hosts), thus leading to cascade effects negatively influencing entire reef ecosystems. Nevertheless, patterns of ectosymbiont assemblages and occurrence in different study areas could align or shift depending on the evolutionary lineage of the host corals, even considering single species (Rouzé et al. 2017), thus leading to different scenarios in areas beyond the Red Sea.
Conclusions
The present study represents an example towards the understanding of the diversity of decapod crustaceans associated with branching corals in the central Red Sea and serves as a baseline for further research on the molecular diversity of decapod crustaceans, as well as providing a reference barcoding dataset for future studies applying methodologies such as ARMS and environmental DNA (eDNA). Since investigating the occurrence and abundance of marine organisms is fundamental to assess their patterns of biodiversity and evolution (Bowen et al. 2013), as well as their patterns of co-occurrence, this study provided insights into the potential loss of decapod crustaceans associated with branching corals in the context of bleaching and host mortality, highlighting how the loss of certain coral taxa in the Red Sea could lead to the extinction of highly specialized symbionts.
References
Abele LG (1976) Comparative species composition and relative abundance of decapod crustaceans in marine habitats of Panama. Mar Biol 38:263–278. https://doi.org/10.1007/BF00388939
Alonso-Domínguez A, Ayón-Parente M, Hendrickx M, Ríos-Jara E, Vargas-Ponce O, Esqueda-González M, Rodríguez-Zaragoza F (2022) Taxonomic diversity of decapod and stomatopod crustaceans associated with pocilloporid corals in the central Mexican pacific. Diversity 14:72. https://doi.org/10.3390/d14020072
Al-Rshaidat MMD, Snider A, Rosebraugh S, Devine AM, Devine TD, Plaisance L, Knowlton N, Leray M (2016) Deep COI sequencing of standardized benthic samples unveils overlooked diversity of Jordanian coral reefs in the northern Red Sea. Genome 59:724–737. https://doi.org/10.1139/gen-2015-0208
Alvarado JJ, Vargas-Castillo R (2012) Invertebrados asociados al coral constructor de arrecifes Pocillopora damicornis en Playa Blanca, Bahía Culebra, Costa Rica. Rev Biol Trop 60:77–92. https://doi.org/10.15517/rbt.v60i2.19965
Andersson AJ, Gledhill D (2012) Ocean acidification and coral reefs: effects on breakdown, dissolution, and net ecosystem calcification. Annu Rev Mar Sci 5:321–348. https://doi.org/10.1146/annurev-marine-121211-172241
Baeza JA (2015) Crustaceans as symbionts: an overview of their diversity, host use, and lifestyles. In: Thiel M, Watling L (eds) Lifestyles and feeding biology. The Natural History of the Crustacea, vol 2. Oxford University Press, Oxford
Barua A, Afrin T, Akhand AA, Ahmed MS (2021) Molecular characterization and phylogenetic analysis of crabs (crustacea: Decapoda: Brachyura) based on mitochondrial COI and 16S rRNA genes. Conserv Genet Resour 13:291–301. https://doi.org/10.1007/s12686-021-01212-9
Berumen M, Hoey A, Bass W, Bouwmeester J, Catania D, Cochran J, Khalil M, Miyake S, Mughal M, Spaet J, Saenz-Agudel P (2013) The status of coral reef ecology research in the Red Sea. Coral Reefs 32:737–748. https://doi.org/10.1007/s00338-013-1055-8
Berumen ML, Arrigoni R, Bouwmeester J, Terraneo TI, Benzoni F (2019) Corals of the Red Sea. In: Voolstra CR, Berumen ML (eds) Coral reefs of the Red Sea. Springer, Cham, pp 123–155
Billick I, Case TJ (1994) Higher-order interactions in ecological communities – what are they and how can they be detected. Ecology 75:1529–1543. https://doi.org/10.2307/1939614
Bowen BW, Rocha LA, Toonen RJ, Karl SA (2013) The origins of tropical marine biodiversity. Trends Ecol Evol 28:359–366. https://doi.org/10.1016/j.tree.2013.01.018
Briggs JC (1974) Marine zoogeography. McGraw-Hill Companies, New York, USA
Britayev T, Spiridonov V, Deart Y, El-Sherbiny M (2017) Biodiversity of the community associated with Pocillopora verrucosa (Scleractinia: Pocilloporidae) in the Red Sea. Mar Biodivers 47:1093–1109. https://doi.org/10.1007/s12526-017-0759-3
Britayev T, Petrochenko RA, Burmistrova YA, Nguyen TH, Lishchenko FV (2023) Density and bleaching of corals and their relationship to the coral symbiotic community. Diversity 15:465–468. https://doi.org/10.3390/d15030456
Brösing A, Al-Aidaroos AM, Tuerkay M (2014) The Red Sea species of Cymo de Haan, 1833 Decapoda, Brachyura, Xanthidae, associates of scleractinian corals. Zootaxa 3779:195–214. https://doi.org/10.11646/zootaxa.3779.2.5
Burke L, Reytar K, Spalding M, Perry A (2011) Reefs at risk revisited. World Resources Institute, Washington, DC
Carvalho S, Aylagas E, Villalobos R, Kattan Y, Berumen M, Pearman J (2019) Beyond the visual: using metabarcoding to characterize the hidden reef cryptobiome. Proc Royal Soc B-Biol Sci 286:20182697. https://doi.org/10.1098/Frspb.2018.2697
Castro P (1978) Movements between coral colonies in Trapezia ferruginea (Crustacea: Brachyura), an obligate symbiont of scleractinian corals. Mar Biol 46:237–245. https://doi.org/10.1007/BF00390685
Castro P (1988) Animal symbioses in coral reef communities: a review. Symbiosis 5:161–184
Castro P, Ng P, Ahyong S (2004) Phylogeny and systematics of the Trapeziidae Miers, 1886 (Crustacea: Brachyura), with the description of a new family. Zootaxa 643: 1. https://doi.org/10.11646/zootaxa.643.1.1
Castro P (1997) Trapeziid crabs (Brachyura:Xanthoidea:Trapeziidae) of New Caledonia, eastern Australia, and the Coral Sea. In: Richer De Forges B. (ed.), Les fonds meubles des lagons de Nouvelle-Calédonie (sédimentologie, benthos). Études & Thèses, volume 3. ORSTOM, Paris, pp. 59–107
Castro P (2015) Symbiotic Brachyura. In Castro P, Davie P, Guinot D, Schram FR, von Vaupel Klein JC (eds.), Decapoda: Brachyura, treatise on zoology – anatomy, taxonomy, biology, volume 9C-I. Brill, Leiden and Boston
Chomitz BR, Kleypas JA, Cortés J, Alvarado JJ (2023) Change in the composition of fauna associated with Pocillopora spp. (Scleractinia Pocilloporidae) following transplantation. Rev Biol Trop 71:e54882. https://doi.org/10.15517/rev.biol.trop..v71iS1.54882
Costa FO, deWaard JR, Boutillier J, Ratnasingham S, Dooh RT, Hajibabaei M, Hebert PDN (2007) Biological identifications through DNA barcodes: the case of the crustacea. Can J Fish Aquat 64:272–295. https://doi.org/10.1139/f07-008
Darriba D, Taboada G, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772–772. https://doi.org/10.1038/nmeth.2109
DiBattista JD, Berumen ML, Gaither MR, Rocha LA, Eble JA, Choat JH, Craig MT, Skillings DJ, Bowen BW (2013) After continents divide: comparative phylogeography of reef fishes from the Red Sea and Indian Ocean. J Biogeogr 40:1170–1181. https://doi.org/10.1111/jbi.12068
DiBattista JD, Roberts MB, Bouwmeester J, Bowen BW, Coker DJ, Lozano-Cortés DF, Choat JH, Gaither MR, Hobbs JPA, Khalil MT, Kochzius M, Myers RF, Paulay G, Robitzch VSN, Saenz-Agudelo P, Salas E, Sinclair-Taylor TH, Toonen RJ, Westneat MW, Williams ST, Berumen ML (2015) A review of contemporary patterns of endemism for shallow water reef fauna in the Red Sea. J Biogeogr 43:423–439. https://doi.org/10.1111/jbi.12649
Eddy TD, Cheung WWL, Bruno JF (2018) Historical baselines of coral cover on tropical reefs as estimated by expert opinion. PeerJ 6:e4308. https://doi.org/10.7717/peerj.4308
Edwards A, Emberton H (1980) Crustacea associated with the scleractinian coral, Stylophora pistillata (Esper), in the Sudanese Red Sea. J Exp Mar Biol Ecol 42:225–240. https://doi.org/10.1016/0022-0981(80)90178-1
Enochs I, Manzello D (2012) Responses of cryptofaunal species richness and trophic potential to coral reef habitat degradation. Diversity 4:94–104. https://doi.org/10.3390/d4010094
Fisher R, O’Leary RA, Low-Choy S, Mengersen K, Knowlton N, Brainard RE, Caley MJ (2015) Species richness on coral reefs and the pursuit of convergent global estimates. Curr Biol 25:500–505. https://doi.org/10.1016/j.cub.2014.12.022
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R (1994) DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotech 3:294–299
Frolová P, Horká I, Ďuriš Z (2022) Molecular phylogeny and historical biogeography of marine palaemonid shrimps (Palaemonidae: Palaemonella-Cuapetes group). Sci Rep 12:15237. https://doi.org/10.1038/s41598-022-19372-5
Furby KA, Bouwmeester J, Berumen ML (2013) Susceptibility of central Red Sea corals during a major bleaching event. Coral Reefs 32:505–513. https://doi.org/10.1007/s00338-012-0998-5
Galil B (1976) The adaptive functional structure of mucus-gathering setae intrapezid crabs symbiotic with corals. Symbiosis 4:75–86
Galil B (1987) Trapeziidae (Decapoda, Brachyura, Xanthoidea) of the Red Sea. Isr J Zool 34:159–182
Gates RD, Ainsworth TD (2011) The nature and taxonomic composition of coral symbiomes as drivers of performance limits in scleractinian corals. J Exp Mar Biol Ecol 408:94–101. https://doi.org/10.1016/j.jembe.2011.07.029
Genevier LGC, Jamil T, Raitsos DE, Krokos G, Hoteit I (2019) Marine heatwaves reveal coral reef zones susceptible to bleaching in the Red Sea. Glob Chang Biol 25:2338–2351. https://doi.org/10.1111/gcb.14652
Gibson R, Atkinson R, Gordon J, Smith I, Hughes D (2011) Coral-associated invertebrates: diversity, ecological importance and vulnerability to disturbance. Oceanogr Mar Biol 49:43–104
Gittenberger A, Gittenberger E (2011) Cryptic, adaptive radiation of endoparasitic snails: sibling species of Leptoconchus (Gastropoda: Coralliophilidae) in corals. Org Divers Evol 11:21–41. https://doi.org/10.1007/s13127-011-0039-1
Glynn PW (1980) Defense by symbiotic crustacea of host corals elicited by chemical cues from predator. Oecologia 47:287–290. https://doi.org/10.1007/bf00398518
Glynn PW (2013) Fine-scale interspecific interactions of coral reefs: functional roles of small and cryptic metazoans. In: Lang MA, Marinelli RL, Roberts SJ, Taylor PR (eds) Research and discoveries: the revolution of science through scuba. Smithsonian Institution Scholarly Press, Washington D.C., pp 229–248
Head C, Bonsall M, Jenkins T, Koldewey H, Pratchett M, Taylor M, Rogers A (2018) Exceptional biodiversity of the cryptofaunal decapods in the Chagos Archipelago, central Indian Ocean. Mar Poll Bull 135:636–647. https://doi.org/10.1016/j.marpolbul.2018.07.063
Hoegh-Guldberg O, Poloczanska ES, Skirving W, Dove S (2017) Coral reef ecosystems under climate change and ocean acidification. Front Mar Sci 4:158. https://doi.org/10.3389/fmars.2017.00158
Hoeksema BW, Fransen CHJM (2011) Space partitioning by symbiotic shrimp species cohabitating in the mushroom coral Heliofungia actiniformis at Semporna, eastern Sabah. Coral Reefs 30:519. https://doi.org/10.1007/s00338-011-0736-4
Hoeksema BW (2017) The hidden biodiversity of tropical coral reefs. Biodiversity 18:8–12. https://doi.org/10.1080/14888386.2017.1307787
Horká I, De Grave S, Fransen CHJM, Petrusek A, Ďuriš Z (2016) Multiple host switching events shape the evolution of symbiotic palaemonid shrimps (Crustacea: Decapoda). Sci Rep 6:26486. https://doi.org/10.1038/srep26486
Huber ME (1987) Aggressive behavior of Trapezia intermedia Miers and T. digitalis Latreille (Brachyura: Xanthidae). J Crustacean Biol 7:238–248. https://doi.org/10.2307/1548604
Hughes TP, Kerry JT, Baird AH, Connolly SR, Dietzel A, Eakin CM, Heron SF, Hoey AS, Hoogenboom MO, Liu G, McWilliam MJ, Pears RJ, Pratchett MS, Skirving WJ, Stella JS, Torda G (2018a) Global warming transforms coral reef assemblages. Nature 556:492–496. https://doi.org/10.1038/s41586-018-0041-2
Hughes TP, Barnes ML, Bellwood DR, Cinner JE, Cumming GS, Jackson JB, Kleypas J, van de Leemput IA, Lough JM, Morrison TH, Palumbi SR, van Nes EH, Scheffer M (2018b) Coral reefs in the Anthropocene. Nature 546:82–90. https://doi.org/10.1038/nature22901
Ivanenko VN, Hoeksema BW, Mudrova SV, Nikitin MA, Martínez A, Rimskaya-Korsakova NN, Berumen ML, Fontaneto D (2018) Lack of host specificity of copepod crustaceans associated with mushroom corals in the Red Sea. Mol Phylogenet Evol 127:770–780. https://doi.org/10.1016/j.ympev.2018.06.024
Kassambara A (2023) ggpubr: “ggplot2” based publication ready plots. https://rpkgs.datanovia.com/ggpubr/
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Knowlton N, Weigt LA (1997) Species of marine invertebrates: a comparison of the biological and phylogenetic species concepts. In: Claridge MF, Dawah AH, Wilson MR (eds) Species, the units of biodiversity. Springer, Cham, pp 199–220
Knowlton N, Brainard RE, Fisher R, Moews M, Plaisance L, Caley MJ (2010) Coral reef biodiversity. In: McIntyre AD (ed) Life in the World’s oceans: diversity, distribution, and abundance. Wiley-Blackwell, Hoboken, NJ USA, pp 65–77
Larsson A (2014) AliView: a fast and lightweight alignment viewer and editor for large data sets. Bioinformatics 30:3276–3278. https://doi.org/10.1093/bioinformatics/btu531
Lassig BR (1977) Communication and coexistence in a coral community. Mar Biol 42:85–92. https://doi.org/10.1007/BF00392016
Limviriyakul P, Tseng L-C, Shih T-W, Hwang J-S (2016) Host selection and preferences of coral symbiotic crab Tetralia rubridactyla. J Exp Mar Biol Ecol 485:24–34. https://doi.org/10.1016/j.jembe.2016.08.001
Maggioni D, Arrigoni R, Seveso D, Galli P, Berumen ML, Denis V, Hoeksema BW, Huang D, Manca F, Pica D, Puce S, Reimer JD, Montano S (2022) Evolution and biogeography of the Zanclea-Scleractinia symbiosis. Coral Reefs 41:779–795. https://doi.org/10.1007/s00338-020-02010-9
McKeon C, Moore J (2014) Species and size diversity in protective services offered by coral guard-crabs. PeerJ 2:e574. https://doi.org/10.7717/peerj.574
McKeon C, Stier A, McIlroy S, Bolker B (2012) Multiple defender effects: synergistic coral defense by mutualist crustaceans. Oecologia 169:1095–1103. https://doi.org/10.1007/s00442-012-2275-2
Miller MA, Pfeiffer W, Schwarts T (2010) Creating the CIPRES Science Gateway for inference of large phylogenetic trees. 2010 Gateway Computing Environments Workshop (GCE), New Orleans, LA, USAhttps://doi.org/10.1109/GCE.2010.5676129
Molodtsova TN, Britayev TA, Martin D (2016) Cnidarians and their polychaete symbionts. In: Goffredo S, Dubinsky Z (eds) The Cnidaria, past, present and future. Springer, Cham. https://doi.org/10.1007/978-3-319-31305-4_25
Monroe AA, Ziegler M, Roik A, Röthig T, Hardenstine RS, Emms MA, Jensen T, Voolstra CR, Berumen ML (2018) In situ observations of coral bleaching in the central Saudi Arabian Red Sea during the 2015/2016 global coral bleaching event. PLoS ONE 13:0195814. https://doi.org/10.1371/journal.pone.0195814
Pandolfi JM, Connolly SR, Marshall DJ, Cohen AL (2011) Projecting coral reef futures under global warming and ocean acidification. Science 333:418–422. https://doi.org/10.1126/science.1204794
Pearman JK, Leray M, Villalobos R, Machida RJ, Berumen ML, Knowlton N, Carvalho S (2018) Cross-shelf investigation of coral reef cryptic benthic organisms reveals diversity patterns of the hidden majority. Sci Rep 8:8090. https://doi.org/10.1038/s41598-018-26332-5
Pisapia C, Stella J, Silbiger N, Carpenter R (2020) Epifaunal invertebrate assemblages associated with branching Pocilloporids in Moorea. French Polynesia Peerj 8:9364. https://doi.org/10.7717/peerj.9364
Plaisance L, Knowlton N, Paulay G, Meyer C (2009) Reef-associated crustacean fauna: biodiversity estimates using semi-quantitative sampling and DNA barcoding. Coral Reefs 28:977–986. https://doi.org/10.1007/s00338-009-0543-3
Plaisance L, Caley M, Brainard R, Knowlton N (2011) The diversity of coral reefs: what are we missing? PLoS ONE 6:25026. https://doi.org/10.1371/journal.pone.0025026
Pratchett MS (2001) Influence of coral symbionts on feeding preferences of crown-of-thorns starfish Acanthaster planci in the western Pacific. Mar Ecol Prog Ser 214:111–119. https://doi.org/10.3354/meps214111
R Core Team (2022) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Ratnasingham S, Hebert PDN (2007) BOLD: the barcode of life data system (www.barcodinglife.org). Mol Ecol Notes 7:355–364. https://doi.org/10.1111/j.1471-8286.2007.01678.x
Robitzch V, Banguera-Hinestroza E, Sawall Y, Al-Sofyani A, Voolstra CR (2015) Absence of genetic differentiation in the coral Pocillopora verrucosa along environmental gradients of the Saudi Arabian Red Sea. Front Mar Sci 2:5. https://doi.org/10.3389/fmars.2015.00005
Ronquist F, Teslenko M, van der Mark P, Ayres D, Darling A, Höhna S, Larget B, Liu L, Suchard M, Huelsenbeck J (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Rouzé H, Lecellier G, Mills SC, Planes S, Berteaux-Lecellier V, Stewart H (2014) Juvenile Trapezia spp. crabs can increase juvenile host coral survival by protection from predation. Mar Ecol Prog Ser 515:151–159. https://doi.org/10.3354/meps10970
Rouzé H, Leray M, Magalon H, Penin L, Gélin P, Knowlton N, Fauvelot C (2017) Molecular characterization reveals the complexity of previously overlooked coral-exosymbiont interactions and the implications for coral-guild ecology. Sci Rep 7:44923. https://doi.org/10.1038/srep44923
Schubart CD (2009) Mitochondrial DNA and decapod phylogenies: the importance of pseudogenes and primer optimization. In: Martin JW, Crandall KA, Felder DL (eds) Crustacean issues 18, decapod crustacean phylogenetics. CRC Press, Taylor & Francis Group, pp 47–65
Schubart CD, Neigel JE, Felder DL (2000) Use of mitochondrial 16S rRNA gene for phylogenetic and population studies of Crustacea. In: von Vaupel Klein JC, Schram F (eds) Crustacean issues 12, The biodiversity crisis and Crustacea. Balkema, Rotterdam, pp 817–830
Sheppard CRC, Davy SK, Pilling GM (2009) The biology of corals reefs (the biology of habitats series). Oxford University Press, Oxford
Spiridonov V, Neumann V (2008) Coral-inhabiting swimming crabs (Crustacea, Decapoda, Portunidae) of the Sudanese Red Sea. Org Divers Evol 8:170.e1-170.e19. https://doi.org/10.1016/j.ode.2007.06.005
Stamatakis A (2014) RAxML Version 8: a tool for phylogenetic analysis and post-analysis of large phylogenesis. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Stella JS, Jones G, Pratchett M (2010) Variation in the structure of epifaunal invertebrate assemblages among coral hosts. Coral Reefs 29:957–973. https://doi.org/10.1007/s00338-010-0648-8
Stella JS, Pratchett MS, Hutchings PA, Jones GP (2011a) Coral-associated invertebrates: diversity, ecological importance and vulnerability to disturbance. Oceanogr Mar Biol 49:43–104. https://doi.org/10.1201/B11009-4
Stella JS, Munday PL, Jones GP (2011b) Effects of coral bleaching on the obligate coral-dwelling crab Trapezia cymodoce. Coral Reefs 30:719–727. https://doi.org/10.1007/s00338-011-0748-0
Stella JS, Wolfe K, Roff G, Rogers A, Priest M, Golbuu Y, Mumby PJ (2022) Functional and phylogenetic responses of motile cryptofauna to habitat degradation. J Anim Ecol 91:2203–2219. https://doi.org/10.1111/1365-2656.13809
Stewart H, Holbrook S, Schmitt R, Brooks A (2006) Symbiotic crabs maintain coral health by clearing sediments. Coral Reefs 25:609–615. https://doi.org/10.1007/s00338-006-0132-7
Stier AC, McKeon CS, Osenberg CW, Shima JS (2010) Guard crabs alleviate deleterious effects of vermetid snails on a branching coral. Coral Reefs 29:1019–1022
Stier AC, Gil MA, McKeon CS, Lemer S, Leray M, Mills SC, Osenberg CW (2012) Housekeeping mutualisms: do more symbionts facilitate host performance? PLoS ONE 7:e32079. https://doi.org/10.1371/journal.pone.0032079
Tamura K, Stecher G, Kumar S (2021) MEGA 11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
van der Schoot RJ, Hoeksema BW (2024) Host specificity of coral-associated fauna and its relevance for coral reef biodiversity. Int J Parasitol 54:65–88. https://doi.org/10.1016/j.ijpara.2023.09.002
Van Wormhoudt A, Adjeroud M, Rouzé H, Leray M (2019) Recent and old duplications in crustaceans “Internal Transcribed Spacer 1”: structural and phylogenetic implications. Mol Biol Rep 46:5185–5195. https://doi.org/10.1007/s11033-019-04976-4
Villalobos R, Aylagas E, Pearman JK, Curdia J, Lozano-Cortés D, Coker DJ, Jones B, Berumen ML, Carvalho S (2022) Inter-annual variability patterns of reef cryptobiota in the central Red Sea across a shelf gradient. Sci Rep 12:16944. https://doi.org/10.1038/s41598-022-21304-2
Vytopil E, Willis B (2001) Epifaunal community structure in Acropora spp. (Scleractinia) on the Great Barrier Reef: implications of coral morphology and habitat complexity. Corals Reefs 20:281–288. https://doi.org/10.1007/s003380100172
Wagner D, Friedlander A, Pyle R, Brooks C, Gjerde K, Wilhelm T (2020) Coral reefs of the high seas: hidden biodiversity hotspots in need of protection. Front Mar Sci 7:567428. https://doi.org/10.3389/fmars.2020.567428
Wei T, Simko V (2021) R package “corrplot”: visualization of a correlation matrix (version 0.92). https://github.com/taiyun/corrplot
Werding B, Hiller A (2007) The Porcellanidae (Crustacea: Decapoda: Anomura) of the Red Sea with description of a new species of Petrolisthes. Zootaxa 1460:1–24. https://doi.org/10.11646/zootaxa.1460.1.1
Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer-Verlag, New York. https://ggplot2.tidyverse.org
Williams ST, Jara J, Gomez E, Knowlton N (2002) The marine Indo-West Pacific break: contrasting the resolving power of mitochondrial and nuclear genes. Integr Comp Biol 42:941–952
Wolfe JM, Breinholt JW, Crandall KA, Lemmon AR, Lemmon EM, Timm LE, Siddall ME, Bracken-Grissom HD (2019) A phylogenomic framework, evolutionary timeline and genomic resources for comparative studies of decapod crustaceans. Proc R Soc B 286:20190079. https://doi.org/10.1098/rspb.2019.0079
Acknowledgements
This research was completed in accordance with the policies and procedures at KAUST (King Abdullah University of Science and Technology). Permissions relevant for KAUST to undertake the research have been obtained from the relevant governmental agencies in the Kingdom of Saudi Arabia. Authors wish to thank the captain and crew of the MV Dream Master, the KAUST Coastal and Marine Resources Core Laboratory, and AK Gusti (KAUST) for fieldwork logistics in the Red Sea. Additionally, we wish to thank three anonymous reviewers for their valuable comments as well as the journal editor for handling the submission and providing useful comments that helped improving the quality of the manuscript.
Funding
This study was supported by KAUST (baseline research funds to ML Berumen and F Benzoni). LM was supported by additional funding from the KAUST Visiting Student Research Program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethical approval
No animal testing was performed during this study. This study was conducted in compliance with all relevant policies and procedures of King Abdullah University of Science and Technology (KAUST).
Sampling and field studies
All necessary permits for sampling and observational field studies have been obtained by the authors from the competent authorities and are mentioned in the acknowledgements. The study is compliant with CBD and Nagoya protocols.
Data availability
Data generated and/or analysed during the current study are included in this published article and its supplementary information files. Sequences obtained in this study were deposited in GenBank database (OR857734-OR857835; OR857584-OR857733; OR852973-OR853050; OR852824-OR852972); accession numbers are available as Online Resource of the present manuscript.
Author contribution
LM: Formal analysis; investigation; data curation; writing, original draft; writing, review and editing; visualization. TIT: Conceptualization; methodology; validation; investigation; data curation; writing, original draft; writing, review and editing; supervision; project administration. RA: Conceptualization; investigation; sampling; data collection; writing, review and editing; supervision. DM: Conceptualization, investigation, sampling. MT: Sampling. AA: Validation, investigation. RL: Validation, investigation. MP: Sampling. MLB: Resources, funding acquisition. FB: Methodology; resources; writing, review and editing; supervision; project administration; funding acquisition. All authors have read and agreed to the published version of the manuscript.
Additional information
Communicated by B. W. Hoeksema
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Macrina, L., Terraneo, T.I., Arrigoni, R. et al. Molecular diversity and patterns of co-occurrence of decapod crustaceans associated with branching corals in the central Red Sea. Mar. Biodivers. 54, 65 (2024). https://doi.org/10.1007/s12526-024-01457-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12526-024-01457-1