Abstract
The population genetic structure of a species is frequently shaped by biogeographic barriers that prevent the movement of individuals from one genetic population to another. Although Octopus mimus Gould, 1852 was reported to inhabit the shallow marine waters of the Peruvian province, recent studies suggest its synonymy with O. hubbsorum Berry, 1953 from the Panamanian and the Cortez provinces where it is the main octopus species caught in the artisanal fishery. The aim of the present study was to assess the genetic variability of O. mimus and to determine the presence of population structure across three sampled regions of the eastern Pacific Ocean using the mitochondrial marker cytochrome c oxidase subunit I. The hypothesis that the biogeographic boundary between the Panamanian and Peruvian provinces acts as a genetic break was further tested and recognized by the conformation of two phylogenetic clades and two haplogroups. Results indicate that these lineages are demographically independent populations of O. mimus and do not represent different species as evidenced by the Generalized Mixed Yule Coalescent method. The oceanographic processes have apparently allowed enough genetic flow along the Pacific coast yielding low but significant levels of genetic structure (FST = 0.16, p < 0.05) as seen in some fish populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
There are six biogeographic provinces in the eastern Pacific: California, Cortez, Panamanian, Galapagos, Peru-Chilean, and Juan Fernández (Briggs and Bowen 2012). All these provinces are defined by high endemism rates in a wide range of marine biotas and by genetic barriers in species boundaries. In previous genetic studies, the assessment of population structure in marine species has been addressed across the eastern Pacific, thus highlighting the importance of biogeographic barriers (Brante et al. 2012; Cárdenas et al. 2009; Ewers-Suacedo et al. 2016; Haye et al. 2014; Saarman et al. 2010). A study of Hippocampus ingens Girard, 1858 (Saarman et al. 2010) encompassed Cortez, Panamanian, and Peruvian provinces and evidenced low genetic differentiation among populations.
The Cortez province spans the Gulf of California. The Panamanian province extends from the mouth of the Gulf of California, Mexico, south to the Gulf of Guayaquil, Ecuador, where the Peruvian province begins down to the Taitao Peninsula (Briggs and Bowen 2012). The mixing of tropical and temperate waters and the convergence of the Humboldt and Panamanian currents in the Gulf of Guayaquil may interrupt gene flow between northern and southern organisms, while dynamic processes such as the seasonal shift of the Intertropical Convergence Zone and “El Niño” events could facilitate the dispersion of larvae (Saarman et al. 2010). Tropical invertebrates including cephalopods from the Panamanian Province have been recorded in Peru, an incursion thought to be associated to the El Niño event (Cardoso and Hochberg 2013; Paredes et al. 1998, 2004).
Octopus mimus Gould, 1852 inhabits shallow waters of the eastern Pacific including the three provinces mentioned above (Pliego-Cárdenas et al. 2016; Söller et al. 2000; Warnke et al. 2002). Söller et al. (2000) and later Warnke et al. (2002) using one mitochondrial gene (COIII) supported the distinction of O. mimus from O. vulgaris Cuvier, 1798 and indicated that the former was present off the coast of Central America. Later on, using sequences of three mitochondrial genes (COI, COIII and r16S), Pliego-Cárdenas et al. (2014) reported that Octopus hubbsorum Berry, 1953 and O. mimus belong to the same lineage. This finding suggests that they are synonymous and represent the continuum of a single species of Octopus present in both Central and South America western coasts. Subsequently, Pliego-Cárdenas et al. (2016) using the same three mtDNA markers showed that the populations of O. mimus from the eastern tropical Pacific and from the southeastern Pacific belong to distinct clades. They argued that different environmental conditions (warm North Pacific currents vs cold South Pacific currents) of the marine biogeographic provinces could be associated with the distinction of the two O. mimus groups, i.e., Panamanian and Peruvian, as a consequence of intraspecific genetic structure rather than of a speciation process. Thus, we refer to O. hubbsorum as O. mimus hereafter.
Octopus mimus is the main octopus fished along the eastern Pacific (Markaida and Gilly 2016). The fishery is artisanal, and octopuses are caught using “hooka” method (Sauer et al. 2019). Regional fishery management for this species includes minimum catch size and closed seasons in Mexico, Peru and Chile (Sauer et al. 2019). In Ecuador, the fishery has no official status (Markaida et al. 2018), and in Mexico, a national management plan for O. mimus is being developed (Diario Oficial de la Federación 2017).
The genetic population structure of Octopus mimus has been investigated separately in the northeastern (Domínguez-Contreras et al. 2018) and southeastern Pacific (Pardo-Gandarillas et al. 2018), using seven non-specific microsatellite loci and sequences of COIII, respectively. Both studies reported that O. mimus shows low levels of genetic diversity associated with the following: (a) a planktonic paralarval stage which can last for several days before the octopus settle on seabed, from 30 to 50 days in some species of Octopus (Villanueva and Norman 2008); (b) high fecundity, up to 200,000 oocytes per female (Warnke 1999; Pliego-Cárdenas et al. 2011); and (c) changes in oceanographic conditions (temperature and currents). However, each of the two study areas was restricted to a single biogeographic province (the Cortez province for the northeastern Pacific and the Peruvian province for the southeastern Pacific). Therefore, the aim of this study was to examine the genetic variability and structure of O. mimus across the three biogeographic provinces of the eastern Pacific: Cortez, Panamanian, and Peruvian. We hypothesize that oceanographic processes have allowed enough gene flow along the eastern Pacific coast yielding low levels of genetic structure as seen in some reef fishes (Muss et al. 2001; Saarman et al. 2010). Identifying population differentiation through genetic divergence will help to establish regional conservation management units and will give meaningful information to elaborate sustainability plans for O. mimus.
Materials and methods
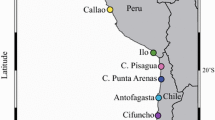
Arm tissue samples of Octopus mimus were obtained from 28 localities covering four coastal countries of the eastern tropical Pacific: Mexico, Colombia, Ecuador, and Peru (see Fig. 1). Each locality was assigned to one of the biogeographic provinces (Cortez, Panamanian, or Peruvian). In total, 171 individuals were collected using fishery-dependent sampling between June 2010 and May–December 2014 (see Supplementary Material). All samples were stored in 96% ethyl alcohol and kept at − 4 °C for subsequent molecular analysis at the Laboratorio de Genética y Biología Molecular of the Universidad Autónoma Metropolitana, Unidad Iztapalapa, México.
Map showing the 28 localities where arm tissue samples of O. mimus were obtained. The sampling covered four coastal countries of the eastern tropical Pacific, Mexico, Colombia, Ecuador, and Peru, and three biogeographic provinces, Cortez (green stars), Panamanian (red squares), and Peruvian (blue circles). Black dashed lines indicate the provinces boundaries according to Briggs and Bowen (2012)
Genomic DNA was extracted from arm muscle tissue and was subsequently purified using the PROMEGA Wizard SV Genomic DNA Purification System Kit following the established manufacturer procedure. DNA amplifications were carried out through polymerase chain reaction (PCR) using the mitochondrial cytochrome c oxidase subunit I (COI) primers, H2198: 5′-TAA ACT TCA GGG TGA CCA AAA AAT CA-3′ and L1490: 5′-GGT CAA CAA ATC ATA AAG ATA TTG G-3′ (Folmer et al. 1994), under a 25-μl total volume reaction, comprising 5× PCR buffer (Promega), 25 mM MgCl2 (Thermo Scientific), 2 mM dNTPs (Thermo Scientific), 50 U Taq DNA polymerase (Bio-Rad), and 0.4 μM of each primer. PCR conditions consisted of 30 thermal cycles including the initial denaturation at 94 °C for 45 s, annealing at 49 °C for 45 s and final extension at 72 °C for 90 s. PCR products were run electrophoretically on a 1.5% agarose gel on TAE 0.5× buffer and were sequenced in both directions at Macrogen, Inc. (Seoul, Korea). Sequences of several species of Octopus as well as O. mimus from Chile (see Table 1) were retrieved from GenBank and aligned to the sampled sequences through the Clustal W method (Thompson et al. 1994) in MEGA v7.2 (Kumar et al. 2015). The alignment was verified with the respective amino acid translation.
Genetic diversity was determined per province. The estimated parameters were the number of segregating sites (S), haplotypes (K), nucleotide diversity (π), and haplotype diversity (Hd) (DnaSP v5. Librado and Rozas 2009). The spatial distribution of genetic variation among the three provinces was examined using a hierarchical analysis of molecular variance (AMOVA) with significance levels set at α = 0.05 and 10,000 random permutations as implemented in Arlequin (Excoffier and Lischer 2010). This method estimates the intraspecific genetic structure using haplotype frequencies (Excoffier et al. 1992). Genetic differentiation between pairs of regions was evaluated a posteriori by calculating pairwise FST (Excoffier and Lischer 2010). The genealogical relationship network among haplotypes was constructed using the median-joining method (Bandelt et al. 1999) in Network v5.0. A phylogenetic analysis was conducted in MrBayes v3.2. (Ronquist et al. 2012) using the GTR+G model, chosen by jModelTest (Darriba et al. 2012), and Octopus insularis Leite & Haimovici in Leite et al. (2008) as outgroup (Pliego-Cárdenas et al. 2016). The Bayesian analysis was run for 10 million generations and sampling every 1000 generations. TRACER v1.6 was used to determine acceptable burn-in and to ensure the analysis had reached the stationarity (Rambaut et al. 2018).
We investigated whether the Octopus mimus clades recovered in the phylogenetic analysis are independent lineages or conspecific. To test this hypothesis, we used (a) the Generalized Mixed Yule Coalescent method (Fujisawa and Barraclough 2013), hereafter referred to as GMYC, which is a coalescent-based species delimitation method, and (b) a data set including 11 Octopus species (O. bimaculatus Verrill, 1883; O. bimaculoides Pickford & McConnaughey, 1949; O. hummelincki Adam, 1936; O. insularis; O. maya Voss & Solís-Ramírez, 1966, O. mimus, O. oculifer (Hoyle, 1904) O. sinensis D´Orbigny, 1834 In Férussac and D´Orbigny, 1834–1848O. tetricus Gould, 1852; O. vulgaris and O. vulgaris type II D´Orbigny, 1840) and 40 sequences (see Table 1). The GMYC method is frequently used to delimit species based on a single locus. It is a likelihood method used to distinguish between inter- and intraspecific processes in a phylogeny by finding the maximum likelihood of a genealogical model (Fujisawa and Barraclough 2013). As good GMYC performance depends on several assumptions, we followed the recommendations of Talavera et al. (2013). Although our data set has singletons (a single haplotype for a species), this does not affect the good identification of species (Talavera et al. 2013). To include the most divergent haplotypes, the sampling covers most of the geographical range of the species. The ultrametric tree was created in BEAST v1.10 (Suchard et al. 2018) using a strict clock, a coalescent process, and a general time reversible (GTR) model, which best fit to the data as determined with the Akaike Information Criterion scores implemented in JModeltest. The chain ran for 15 million generations, sampled every 1000th generation. Tracer v1.6 was used to determine the convergence of chains. The analysis was performed two times, and log and tree files were combined with LogCombiner. Trees were then summarized using TreeAnotator discarding 10% of trees as burn-in. The GMYC model was tested using the splits package (Ezard et al. 2009) and a single threshold implemented in R.
Results
The 171 resolved sequences (605 bp) yielded 19 haplotypes, and a total of 20 polymorphic sites were identified. The populations from the three biogeographic provinces resolved from intermediate to relatively high haplotype diversity (Table 2; Hd = 0.495–0.707) and low nucleotide diversities (π = 0.0009–0.0042), with the Peruvian province showing the highest levels (Hd = 0.707; π = 0.0042) and the Panamanian, the lowest (Hd = 0.495; π = 0.0013). Overall AMOVA genetic differentiation using the FST index indicates low but significant differentiation (FST = 0.16, p < 0.05), with most of the variation occurring within populations (83%) and 17% among groups (Table 3). Furthermore, the pairwise FST values support the hypothesis of population differentiation between the Panamanian and Peruvian provinces (FST = 0.16, p > 0.05), and between the Cortez and the Peruvian provinces (FST = 0.13, p > 0.05), but a differentiated population between the Cortez and the Panamanian provinces was not supported (Table 4).
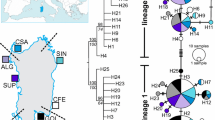
The haplotype network showed the distribution of haplotypes in two well-defined groups (Fig. 2). Haplogroup 1 contains haplotypes from the three regions but with higher number of samples from the Panamanian region; therefore, hereafter, this haplogroup is referred as the Panamanian group. Haplogroup 2 contains predominantly specimens from the Peruvian province and only two specimens from Ecuador, thus, from hereafter is referred as the Peruvian group. These haplogroups are separated from each other by mutations at three sites. The most frequent haplotype, contained in the Panamanian group (H1, 55%), comprised 91 specimens from 22 sampled localities from the three provinces. It is followed by haplotypes 2, 3, and 4 (H2, H3, and H4, respectively) with frequencies > 5%. The haplotypes from Chile (GenBank accession numbers: GU355923-GU355926) were identical to H2 and H3 from the Peruvian group.
All haplotypes clustered together in one well-supported single monophyletic clade (Fig. 3) and were organized in two main Octopus mimus subclades, one formed by specimens from the three regions and the other clustered specimens from the Panamanian (two individuals) and the Peruvian provinces. This finding is in agreement with the haplotype network.
The GMYC model recovered seven ML clusters (confidence interval 6–8) and 11 ML entities (confidence interval 10–13, p < 0.05) that correspond to the 11 Octopus species and support the conspecificity of the two lineages of Octopus mimus (Panamanian and Peruvian) (Fig. 4).
Discussion
Genetic diversity levels found in Octopus mimus along the three provinces are comparable to those observed for other Octopus species. Mean nucleotide diversity for O. mimus in this study (0.0036) is the same to that reported for this species in the Peruvian region (0.003) using COIII (Pardo-Gandarillas et al. 2018), and slightly lower than that for O. vulgaris using COI (π = 0.005–0.021, Sales et al. 2013). According to Pardo-Gandarillas et al. (2018), low levels of genetic diversity for O. mimus can be associated to demographic events caused by historic oceanographic changes.
It is well known that biogeographic boundaries act as genetic barriers precluding the gene flow between populations (Keigwin 1982). Two biogeographic barriers have been described along the tropical eastern Pacific Ocean, which subdivide populations of fishes and mollusks (Muss et al. 2001). These correspond to the transition zones among the biogeographic Cortez-Panamanian and Panamanian-Peruvian provinces (García-De León et al. 2018; Saarman et al. 2010). The mouth of the Gulf of California is characterized by strong salinity and seasonal temperature changes (Castro et al. 2000) that act like a barrier for some species but not for others (Bernardi et al. 2003; García-De Léon et al. 2018; Hurtado et al. 2007). The absence of genetic differentiation between the Cortez and the Panamanian populations could be caused by several factors, as a small sample size representing the Cortez province as well as by oceanographic processes. Fishes and mollusks show a similar pattern. Bernardi et al. (2003) found no differentiation between the populations of some species of fishes from the Gulf of California and the Pacific Ocean due to their dispersal potential. Similarly, Hurtado et al. (2007) attributed the panmixia of the snail Nerita funiculata Menke, 1850 along the eastern tropical Pacific to the longevity of its larvae which can be dispersed from Panama to the Gulf of California (about 5000 km). However, the use of more variable molecular markers in O. mimus, such as microsatellites and a larger sample size in the region, might help to resolve this issue in further studies.
Contrary to the lack of distinction between lineages from the Cortez and the Panamanian provinces, pairwise FST values showed a low but significant genetic structure of the populations of Octopus mimus separated by the Panamanian-Peruvian biogeographic border, which is concordant to the hypothesis of oceanographic barriers to dispersal. It is possible that the association of the Pacific Ocean currents and the sea surface temperature (SST) of the region are enough to act like genetic barrier and to delimit two O. mimus populations. The Panama tropical current moving southward contrasts with the cold Humboldt current moving northward; thus, the SST in the Panamanian province typically exceed 20 °C while in the Peruvian province the temperature is, usually, lower than 20° C (Hickman Jr. 2009; Montecinos et al. 2003).
The relatively low level of population differentiation detected between the populations of Octopus mimus is likely due to oceanic processes in the Pacific (Muss et al. 2001; Saarman et al. 2010) such as the El Niño Southern Oscillation (Cardoso and Hochberg 2013), which could favor the dispersal of paralarvae, which are tolerant to the warm temperatures (up to 24 °C) of El Niño (Warnke 1999). Cardoso and Hochberg (2013) recorded squids from the Panamanian province in Peruvian waters, and this incursion was attributed to tropical waters invasion into the Peruvian province during the El Niño event (Paredes et al. 1998, 2004). This oceanic process might explain the presence of the same octopus haplotypes in the Panamanian and the Peruvian provinces (Marin et al. 2018; this study).
Another example of an Octopus inhabiting more than one province is O. insularis, which has a wide distribution along the Western Atlantic, from Brazil to the Gulf of Mexico, spanning the Caribbean and the Brazilian biogeographic provinces (Briggs and Bowen 2012; Lima et al. 2017). The extensive distribution range of O. insularis is linked to the long-distance dispersal of paralarvae (González-Gómez et al. 2018), which allows gene flow among distant populations. Flores-Valle et al. (2018) support this hypothesis by finding octopuses from Mexico and Brazil sharing the same haplotype. In benthic octopuses with planktonic paralarvae, dispersion is ensured by ocean current transport far away from hatching localities. The paralarval stage of benthic octopuses, such as O. mimus, typically lasts from 3 weeks to 6 months (Villanueva and Norman 2008).
The use of the GMYC method in the present study allowed the identification of two genetic groups that coalesce in the same species group, Octopus mimus, and not as independent lineages, supporting the earlier findings by Pliego-Cárdenas et al. (2016). Nevertheless, it has to be considered that this study used a single marker, and, although the GMYC method is suitable when using a single locus (Fujisawa and Barraclough 2013) in species delimitation studies (Monaghan et al. 2009; Talavera et al. 2013; Vidigal et al. 2018), a multilocus analysis together with a morphological study could provide additional support to the COI sequences, since the phylogenetic tree cannot represent the evolutionary history of the species but, instead, represents the history of a single gene (Fujita et al. 2012).
Fishery management
As previously stated, there are no management plans for Octopus mimus fishery although in some countries seasonal closures and a minimum catch size have been implemented (Sauer et al. 2019). In Mexico, the closed season and minimum catch size have been implemented in a single Biosphere Reserve from the Gulf of California but not for the Mexican Pacific (Diario Oficial de la Federación 2017). The results from this study allow us to suggest that Octopus mimus should encompass two main fishery management units along eastern tropical Pacific (ETP), at least until data from microsatellite analysis become available. The occurrence of other known biogeographic barriers along ETP should be tested such as the border between the Gulf of California and the Mexican Pacific where a genetic break has been reported for the North Pacific hake and a threatened seahorse species (García-De Léon et al. 2018; Saarman et al. 2010). Efforts to study other aspects of population genetics and to investigate regional genetic structure will lead to (a) improved management plans and (b) a better understanding of how fisheries impact this species.
References
Acosta-Jofré M, Sahade R, Laudien J, Chiappero MB (2012) A contribution to the understanding of phylogenetic relationships among species of the genus Octopus (Octopodidae: Cephalopoda). Sci Mar 76(2):311–318
Adam W (1936) Notes sur les cephalopodes. VI. Une nouvelle espece d'Octopus (Octopus hummelincki) des Indes Occidentales Neerlandaises. Bull Mus r hist nat Belg 12(40):1–3
Amor MD, Norman MD, Cameron HE, Strugnell JM (2014) Allopatric speciation within a cryptic species complex of Australasian octopuses. PLoS ONE 9(6):e98982
Bandelt HJ, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16(1):37–48
Bernardi G, Findley L, Rocha-Olivares A (2003) Vicariance and dispersal across Baja California in disjunct marine fish populations. Evol 57(7):1599–1609
Berry SS (1953) Preliminary diagnoses of six west American species of octopus. Leafl in Malacol 1(10):51–58
Brante A, Fernández M, Viard F (2012) Phylogeography and biogeography concordance in the marine gastropod Crepipatella dilatata (Calyptraeidae) along the Southeastern Pacific Coast. J Hered 103:630–637
Briggs JC, Bowen BW (2012) A realignment of marine biogeographic with particular reference to fish distributions. J Biogeogr 39:12–30
Cárdenas L, Castilla JC, Viard F (2009) A phylogeographical analysis across three biogeographical provinces of the south-eastern Pacific: the case of the marine gastropod Concholepas concholepas. J Biogeogr 36:969–981
Cardoso F, Hochberg FG (2013) Revision of the genus Lolliguncula Steenstrup, 1881 (Cephalopoda: Loliginidae) off the Pacific Coast of South America. Rev Peru Biol 20(2):129–136
Castro R, Mascarenhas AS, Durazo R, Collins CA (2000) Seasonal variation of the temperature and salinity at the entrance to the Gulf of California, Mexico. Cienc Mar 26(4):561–583
Cuvier GL (1798) Tableau élementaire de l'Histoire Naturelle des Animaux. Paris, Baudouin. 710 pp
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9(8):772
Diario Oficial de la Federación (2017). Carta Nacional Pesquera. 11/06/2018
Domínguez-Contreras JF, Munguia-Vega A, Ceballos-Vazquez BP, Arellano-Martínez M, García-Rodriguez FJ, Culver M, Reyes-Bonilla H (2018) Life histories predict genetic diversity and population structure within three species of octopus targeted by small-scale fisheries in Northwest Mexico. Peer J 6:e4295
D´Orbigny A (1834) Histoire naturelle générale et particulière des céphalopodes acétabulifères. Tome 1. In: Férussac AE, D´Orbigny A (eds) Histoire naturelle générale et particulière des céphalopodes acétabulifères vivants et fossiles. Baillière, Paris, pp 68-69
D´Orbigny A (1840) Histoire naturelle générale et particulière des céphalopodes acétabulifères. Tome 1. In: Férussac AE, D´Orbigny A (eds) Histoire naturelle générale et particulière des céphalopodes acétabulifères vivants et fossiles. Baillière, Paris, pp 29-30
Ewers-Suacedo C, Pringle JM, Sepúlveda HH, Byers JE, Navarrete SA, Wares JP (2016) The oceanic concordance of phylogeography and biogeography: a case study in Notochthamalus. Ecol Evol 6(13):4403–4420
Excoffier L, Lischer HE (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10(3):564–567
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genet 131(2):479–491
Ezard T, Fujisawa T, Barraclough T (2009) Splits: species’ limits by threshold statistics. R package version 1.0. Available: http://R-Forge.R-project.org/projects/splits/. Accessed on April 4, 2019
Férussac AE, d´Orbigny A (1834-1848) Histoire naturelle générale et particulière des céphalopodes acétabulifères vivants et fossiles. Baillière, Paris
Flores-Valle A, Pliego-Cárdenas R, Jiménez-Badillo ML, Arredondo-Figueroa JL, Barriga-Sosa IA (2018) First record of Octopus insularis Leite and Haimovici, 2008 in the octopus fishery of a marine protected area in the Gulf of Mexico. J Shellfish Res 37(1):221–227
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R (1994) DNA primers for amplification of mitochondrial cytochrome c oxidase subunit from diverse metazoan invertebrates. Mol Mar Biol Biotechnol 3(5):294–299
Fujisawa T, Barraclough TG (2013) Delimiting species using single-locus data and the Generalized Mixed Yule Coalescent approach: a revised method and evaluation on simulated data sets. Syst Biol 62(5):707–724
Fujita MK, Leaché AD, Burbrink FT, McGuire JA, Moritz C (2012) Coalescent-based species delimitation in an integrative taxonomy. Trends Ecol Evol 27(9):480–488
García-De León FJ, Galván-Tirado C, Velasco LS, Silva-Segundo C, Guzmán RH, Barriga-Sosa IDLA, Jaimes PD, Canino M, Cruz-Hernández P (2018) Role of oceanography in shaping the genetic structure in the North Pacific hake Merluccius productus. Plos ONE 13(3):e0194646
González-Gómez R, Barriga-Sosa IA, Pliego-Cárdenas R, Jiménez-Badillo L, Markaida U, Meiners-Mandujano C et al (2018) An integrative taxonomic approach reveals Octopus insularis as the dominant species in the Veracruz Reef System (southwestern Gulf of Mexico). Peer J 6:e6015
Gould AA (1852) Reports of the United States Exploring Expedition. 1838-1842. Vol. I. Mollusca and Shells. Philadelphia. 12:1–510
Haye PA, Segovia NI, Muñoz-Herrera NC, Gálvez FE, Martínez A, Meynard A et al (2014) Phylogeographic structure in benthic marine invertebrates of the Southeast Pacific Coast of Chile with differing dispersal potential. PLoS ONE 9(2):e88613
Hickman CP Jr (2009) Evolutionary responses of marine invertebrates to insular isolation in Galapagos. Galapagos Res 66:32–42
Hoyle WE (1904) Reports on the Cephalopoda (expedition to the west coast of central America to the Galapagos, to the west coast of Mexico, and in the Gulf of California, in charge of Alexander Agassiz, carried on by the U. S. Fish Commission Steamer •Albatross• during 1891). Bull Mus Comp Zool 43(1):1−31
Hurtado LA, Frey M, Gaube P, Pfeiler E, Markow TA (2007) Geographical subdivision, demographic history and gene flow in two sympatric species of intertidal snails, Nerita scabricosta and Nerita funiculata, from the tropical eastern Pacific. Mar Biol 151:1863–1873
Juárez OE, Rosas C, Arena-Ortiz ML (2012) Phylogenetic relationships of Octopus maya revealed by mtDNA sequences. Cienc Mar 38(3):563–575
Kaneko N, Kubodera T, Iguchis K (2011) Taxonomic study of shallow-water octopuses (Cephalopoda: Octopodidae) in Japan and adjacent waters using mitochondrial genes with perspectives on octopus DNA barcoding. Malacologia 54(2):97–108
Keigwin L (1982) Isotopic paleoceanography of the Caribbean and East Pacific: role of Panama uplift in late neogene time. Science 217(4557):350–353
Kumar S, Stecher G, Tamura K (2015) MEGA7: molecular evolutionary genetics analysis version 7.0. for bigger data sets. Mol Biol Evol 33(7):1870–1874
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinform 25:1451–1452
Lima FD, Bervel-Filho WM, Leite TS, Rosas C, Lima SM (2017) Occurrence of Octopus insularis Leite and Haimovici, 2008 in the Tropical Northwestern Atlantic and implications of species misidentification to octopus fisheries management. Mar Biodivers 47(3):7123–7734
Marin A, Serna J, Robles C, Ramírez B, Reyes-Flores LE, Zelada-Mázmela E et al (2018) A glimpse into the genetic diversity of the Peruvian seafood sector: unveiling species substitution, mislabeling and trade of threatened species. PLoS ONE 13(11):e0206596
Markaida U, Gilly WF (2016) Cephalopods of Pacific Latin America. Fish Res 173(2):113–121
Markaida U, Flores L, Arias E, Mora E (2018) Reproduction and population structure of Octopus mimus fished in a marine protected area of Ecuador. J Mar Biol Assoc UK 98(6):1383–1389
Monaghan MT, Wild R, Elliot M, Fujisawa T, Balke M, Inward DJ et al (2009) Accelerated species inventory on Madagascar using coalescent-based models of species delineation. Syst Biol 58(3):298–311
Montecinos A, Purca S, Pizarro O (2003) Interannual to interdecadal sea surface temperature variability along the western coast of South America. Geophys Res Lett 30(11):1150
Muss A, Robertson DR, Stepien CA, Wirtz P, Bowen BW (2001) Phylogeography of Ophioblennius: the role of ocean currents and geography in reef evolution. Evolution 55(3):561–572
Pardo-Gandarillas MC, Ibañez CM, Yamashiro C, Méndez MA, Poulin E (2018) Demographic inference and genetic diversity of Octopus mimus (Cephalopoda: Octopodidae) throughout the Humboldt Current System. Hydrobiologia 808(1):125–135
Paredes C, Tarazona J, Canahuire E, Romero L, Cornejo O, Cardoso F (1998) Presencia de moluscos tropicales de la Provincia Panameña en la costa central del Peru y su relacion con los eventos “el Niño”. Rev Peru Biol 5(2):123–128
Paredes C, Cardoso F, Tarazona J (2004) Distribución temporal de moluscos y crustáceos tropicales en la Provincia Peruana y su relación con los eventos El Niño. Rev Peru Biol 11(2):213–218
Pickford GE, McConnaughey BH (1949) The Octopus bimaculatus problem: a study in sibling species. Bull Bingham Oceanogr Coll 12(4):1–66
Pliego-Cárdenas R, García-Domínguez FA, Ceballos-Vázquez BP, Villalejo-Fuerte M, Arellano-Martínez M (2011) Reproductive aspects of Octopus hubbsorum (Cephalopoda: Octopodidae) from Espíritu Santo Island, southern Gulf of California, Mexico. Cienc Mar 37(1):23–32
Pliego-Cárdenas R, Hochberg F, García De León F, Barriga-Sosa I (2014) Close genetic relationships between two American octopuses: Octopus hubbsorum Berry, 1953, and Octopus mimus Gould, 1852. J Shellfish Res 33(1):293–303
Pliego-Cárdenas R, Flores L, Markaida U, Barriga-Sosa IA, Mora E, Arias E (2016) Genetic evidence of the presence of Octopus mimus in the artisanal fisheries of octopus in Santa Elena Peninsula, Ecuador. Am Malacol Bull 34(1):51–55
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarisation in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67(5):901–904
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S et al (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61(3):539–542
Saarman NP, Louie KD, Hamilton H (2010) Genetic differentiation across eastern Pacific oceanographic barriers in the threatened seahorse Hippocampus ingens. Conserv Genet 11:1989–2000
Sales J, Rego P, Hilsdorf A, Moreira A, Haimovici M, Tomás AR et al (2013) Phylogeographical features of Octopus vulgaris and Octopus insularis in the Southeastern Atlantic based on the analysis of mitochondrial markers. J Shellfish Res 32(2):325–339
Sauer WHH, Gleadall IG, Downey-Breedt N, Doubleday Z, Gillespie G, Haimovici M, Ibáñez CM, Katugin ON et al (2019) World octopus fisheries. Rev Fish Sci Aquac. https://doi.org/10.1080/23308249.2019.1680603
Söller R, Warnke K, Saint-Paul U, Blohm D (2000) Sequence divergence of mitochondrial DNA indicates cryptic biodiversity in Octopus vulgaris and supports the taxonomic distinctiveness of Octopus mimus (Cephalopoda: Octopodidae). Mar Biol 139:29–35
Strugnell JM, Norman MD, Vecchione M, Guzik M, Allcock AL (2014) The ink sac clouds octopod evolutionary history. Hydrobiologia 725:215–235
Suchard MA, Lemey P, Baele G, Ayres DL, Drummond AJ, Rambaut A (2018) Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evol 4(1):vey016
Talavera G, Dinca V, Vila R (2013) Factors affecting species delimitations with the GMYC model: insights from a butterfly survey. Methods Ecol Evol 4:1101–1110
Thompson JD, Higgins GD, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22(22):4673–4680
Verrill AE (1883) Descriptions of two species of octopus from California. Bull Mus Comp Zool 11(6):117–123
Vidigal TH, Coscarelli D, Paixao H, Bernardes S, Montresor LC, Pepato AR (2018) Integrative taxonomy of the neotropical genus Omalonix (Elasmognatha: Succineidae). Zool Scr 47:174–186
Villanueva R, Norman MD (2008) Biology of the planktonic stages of benthic octopuses. Oceanogr Mar Biol 46:105–202
Voss GL, Solís-Ramírez M (1966) Octopus maya, a new species from the Bay of Campeche, Mexico. Bull Mar Sci 16(3):615–625
Warnke K (1999) Observations on the embryonic development of Octopus mimus (Mollusca: Cephalopoda) from northern Chile. The Veliger 42(3):211–217
Warnke K, Söller R, Blohm D, Saint-Paul U (2002) Assessment of the phylogenetic relationship between Octopus vulgaris Cuvier, 1797 and O. mimus Gould 1852, using mitochondrial 16S rDNA in combination with morphological characters. Abh Geol B-A 57:401–405
Acknowledgments
RPC and SMP thank Consejo Nacional de Ciencia y Tecnología (CONACyT) for the scholarships received. UM thanks SENESCYT for supporting his research at the Instituto Nacional de Pesca of Ecuador through the PROMETEO Program. We thank Frederick Hochberg for donating samples from Colombia and Jose A. García for his assistance in editing the English version of the manuscript. We also thank to two anonymous reviewers and the editor for constructive comments.
Funding
This study was supported by Universidad Autónoma Metropolitana with grant nos. 147.09.01, 147.09.04, and 147.09.07 to IDLABS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed by the authors.
Sampling and field studies
All samples were obtained by fishermen who have the necessary permits for sampling.
Data availability
The data sets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Author contribution
RPC, UM, and IBS conceived and designed research. RPC, SMP, and MAA conducted experiments. RPC and IBS contributed new reagents or analytical tools. RPC and SMP analyzed data. RPC, IBS, and SMP wrote the manuscript. All authors read and approved the manuscript.
Additional information
Communicated by M. Vecchione
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(XLSX 12 kb)
Rights and permissions
About this article
Cite this article
Pliego-Cardenas, R., Prieto, S.M., Apon, M.A. et al. Genetic structure of Octopus mimus Gould, 1852 along three biogeographic marine provinces. Mar. Biodivers. 50, 46 (2020). https://doi.org/10.1007/s12526-020-01078-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12526-020-01078-4