Abstract
Bone-eating Osedax worms can quickly colonize exposed bones and are important ecosystem engineers in whale fall communities, contributing to cause of bone degradation. This study shows that the deep SW Atlantic margin harbors many Osedax species. Using DNA barcoding, we found four putative new lineages as well as O. frankpressi Rouse, Goffredi, and Vrijenhoek, 2004 and O. braziliensis Fujiwara, Jimi, Sumida, Kawato, & Kitazato, 2019, with assemblages varying with depth. It is probable that the bathymetric distributions of these species are controlled by different water masses and their directions of flow. The haplotype network of Atlantic and Pacific O. frankpressi populations suggests segregation between populations, as is also seen in the high FST. However, the low p distance between both populations and the few substitution sites separating haplogroups from both regions (Atlantic and Pacific) could be evidence that populations of both basins are somehow close to each other. It is likely that whale fall habitats exist between both populations analyzed, connecting both basins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The bone-eating worm Osedax is a siboglinid annelid specialized in consuming organic compounds from vertebrate bones (collagen and lipids) through a symbiotic association with heterotrophic bacteria (Rouse et al. 2004; Goffredi et al. 2005, 2007; Jones et al. 2008; Vrijenhoek et al. 2008). Among its adaptations is the lack of a digestive system in adults and a remarkable sexual dimorphism, observed in all Osedax, except O. priapus Rouse, Wilson, Worsaae & Vrijenhoek, 2015 (Rouse et al. 2004). Females are large and attach to bones, while males are paedomorphic dwarves retaining larval characteristics, living anchored on the trunk and in the tubes of females (Rouse et al. 2004, 2009; Worsaae and Rouse 2010). The Osedax female body is divided in an ovisac and root systems that are inside the bone, with a protuding trunk, palps, and oviduct (Rouse et al. 2004, 2008, 2015, 2018; Glover et al. 2005, 2013; Amon et al. 2014). Different clades can be distinguished by the presence or absence of pinnules on the palps (Vrijenhoek et al. 2009; Glover et al. 2013; Amon et al. 2014). Other characteristics, such as the shape and color of patch, collar, palps, and/or the root system, can be used for species identification (Rouse et al. 2004, 2008, 2018; Glover et al. 2005, 2013; Fujikura et al. 2006; Amon et al. 2014).
Bone degradation by Osedax can shorten the duration of the whale fall communities and truncates’ successional stages, hindering the establishment of the sulfophilic stage in juvenile carcasses (Braby et al. 2007; Lundsten et al. 2010; Smith et al. 2015). The physical degradation of the bone provides conduits for sulfide from the inner parts, accelerating the rates of sulfide release and the consumption of organic compounds by microbial communities, thus abbreviating the sulfophilic stage (Braby et al. 2007; Treude et al. 2009; Higgs et al. 2011). The bioerosion activity of Osedax also increases the structural complexity of the bone matrix and alters the physical-chemical environment of its innermost part. This process enhances the abundance and biodiversity of endofauna, causing Osedax to be an ecosystem engineer of the whale fall community (Alfaro-Lucas et al. 2017).
Since the discovery of Osedax, 18 species within the genus have been described in NE Pacific (Rouse et al. 2004, 2008, 2015, 2018), five species in Southern Ocean (Glover et al. 2013; Amon et al. 2014), and only one species occurs in each of the following ocean basins: NW Pacific, N Atlantic, and SW Atlantic (Glover et al. 2005; Fujikura et al. 2006; Fujiwara et al. 2019), yielding a total of 26 species around the world. In addition, five putative new species remain unnamed (Salathé and Vrijenhoek 2012; Taboada et al. 2015; Rouse et al. 2018). The known bathymetric range of Osedax is 21- to 4204 m depth (Amon et al. 2014; Fujiwara et al. 2019) with temperatures usually low (between − 1 and 15 °C), even for the shallow-water species (Glover et al. 2005, 2013; Fujikura et al. 2006; Taboada et al. 2015). Phylogenetic studies show that Osedax species are grouped into six distinct clades (I–VI), which are related to the palp morphology (Vrijenhoek et al. 2009; Glover et al. 2013; Amon et al. 2014; Taboada et al. 2015; Rouse et al. 2015, 2018; Fujiwara et al. 2019). There is no evidence for geographic isolation of Osedax clades (Taboada et al. 2015; Rouse et al. 2018; Fujiwara et al. 2019), suggesting that dispersal events, rather than vicariance processes, are the most important mechanisms shaping the current distribution of Osedax (Glover et al. 2005).
While Osedax species descriptions have used morphological features on the prostomium bump/patch, collar, palps, and root system (Rouse et al. 2004, 2008, 2015, 2018; Glover et al. 2005, 2013; Fujikura et al. 2006) to differentiate species, these structure can exhibit large intraspecific variation, as is the case of O. braziliensis Fujiwara, Jimi, Sumida, Kawato and Kitazato, 2019, and can be affected by the preservation procedures of specimens, making it difficult to recognize species without the help of molecular taxonomy, using DNA barcoding (Amon et al. 2014; Fujiwara et al. 2019). Barcoding benefits from use of a reference library for comparison, which it is available for Osedax since molecular taxonomy had been used in all studies of the genus (Rouse et al. 2004, 2008, 2009, 2018; Glover et al. 2005, 2013; Fujikura et al. 2006; Vrijenhoek et al. 2009; Salathé and Vrijenhoek 2012; Amon et al. 2014; Taboada et al. 2015; Fujiwara et al. 2019).
In this study, we aimed to reveal the diversity of Osedax in whalebones implanted at multiple sites across three depths (550, 1500, and 3300 m) in the deep SW Atlantic Ocean using DNA barcoding, mitochondrial gene cytochrome-c oxidase subunit I, in order to (1) investigate bathymetric distinctions of Osedax assemblages and (2) place the SW Atlantic lineages in the global context of the Osedax diversity and distribution.
Material and methods
Study area and sample collection
The study area comprises Espírito Santo, Campos, and Santos Basins along the SW Atlantic Ocean between 21° and 28° S latitude at three depths (550, 1500, and 3330 m). The SW Atlantic margin is the habitat or a migratory route for at least 30 baleen or toothed whales, including the sub-Antarctic population of humpback whale (humpback whale, Megaptera novaeangliae (Borowski, 1781) which migrates from South Georgia Islands through Rio Grande Rise and northwards to the Abrolhos Bank (Zerbini et al. 1997, 2006; Santos et al. 2010; Wedekin et al. 2014). The increase of vessel traffic and coastal development, including gas and oil exploration, could drastically affect the whale populations along the Brazilian margin and indirectly the populations of Osedax in this ocean basin.
Three major water masses are found along the deep Brazilian margin: Antarctic Intermediate Water (AAIW), North Atlantic Deep Water (NADW), and Antarctic Bottom Water (AABW) (De Madron and Weatherly 1994; Campos et al. 1995; Silveira et al. 2000). The temperature of AAIW is higher than 3 °C, up to 6 °C, and the salinity range is between 34.2 and 34.6 (Silveira et al. 2000). The AAIW flows southward near to the east continental margin of Brazil between 500- and 1000-m depth, to the south of 25–28° S latitude (Müller et al. 1998; Silveira et al. 2000). The thermohaline limits of NADW are between 3 and 4 °C, with salinity > 34.6 and < 35 on the southeast Brazilian margin (De Madron and Weatherly 1994; Silveira et al. 2000). The NADW generally flows southward at depths of 1200–3000 m to ~ 32°S (McCartney 1993). The temperature of AABW, in the study region, is usually above 0 °C, up to 2 °C, with salinity below 35 (Stramma and England 1999). The AABW generally flows northward, via the Vema Channel, originating mainly as Lower Circumpolar Deep Water (De Madron and Weatherly 1994; Stramma and England 1999). This layer of AABW flows in the Brazilian Basin between 3100- and 4100-m depth (Hogg and Owens 1999).
Specimens of Osedax were sampled by deployment and recovery of whalebones using experimental autonomous structures (landers). We used thoracic vertebrae from a humpback whale (Megaptera novaeangliae). Vertebrae were collected on 23 October 2012 from a stranded carcass in the Pontal do Ipiranga beach (Linhares/ES—Brazil). Intervertebral discs and vertebra processes were removed using a handsaw. A transversal section was made in each vertebra and all pieces were kept frozen (− 20 °C) until deployment. Landers were aluminum three-sided, pyramid-shaped structures, with each face composed of three boxes lined by meshed bags (500 μm) and a PVC lid. Each lander was outfitted with an acoustic release (Teledyne Benthos 866A), where ballast was fixed, and glass sphere buoys (MacLane Labs Inc.) for flotation upon recovery. During the deployment, the lids were held open by a connection to the acoustic release, which closed the boxes when the acoustic release was triggered on the recovery cruise. For more details of the lander design and how bones were attached to the lander, see appendages 1 and 2 of Saeedi et al. 2019.
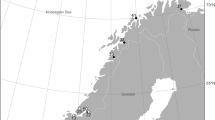
Landers were deployed at six sampling sites at two different depths (~ 1500- and ~ 3300-m depth, Fig. 1, Table 1) between 28 May and 06 June 2013 on board the R/V Alpha Crucis. The lander deployed at SP-1500 was recovered 16 months later on 09 October 2014 on board the R/V Alpha Delphini. Other landers were recovered between 18 and 28 May 2015 on board the Polar R/V Almirante Maximiano. Unfortunately, the lander RJ-1500 was lost during recovery. An extra lander (SP-550) was deployed in a pockmark field in the south section of Santos Basin at 550-m depth (Fig. 1, Table 1) on July 2016 from the R/V Alpha Crucis and recovered on May 2017 with the M/V Alucia.
Locations of lander deployment. The white circles indicate landers deployed and recovered; white circles with a cross indicate landers with bones without Osedax. The black circle indicates lander lost during the recovery; The yellow star indicates the locality of the natural whale fall studied by Sumida et al. (2016) with the first record of Osedax braziliensis in SW Atlantic Ocean
We found Osedax at four sampling sites (Table 1). All individuals were sorted on board and classified according to the presence or absence of pinnules on palps. Organisms were preserved in 96% molecular grade ethanol.
DNA extraction, amplification, and sequencing
DNA was extracted from a small piece of tissue from each specimen using the QIAGEN Blood & Tissue kit following the manufacturer’s protocol and eluting the DNA in 50–100 μl of ddH2O. The tissue for DNA extraction was preferentially from the thorax region or the whole animal for very small specimens.
We amplified ~ 550-bp fragment of the cytochrome c oxidase subunit-I gene (COI) using OsCO1f and OsCO1r primers (Glover et al. 2005). All PCRs contained 12.5 μL of GoTaq® Green Master Mix (Promega), 0.125 μL of each primer (20 μM), 2–4 μL DNA template (20–100 ng), and nuclease-free water (Promega) to reach up 25 μL of total volume. The thermal cycling profile for COI was 95 °C for 2 mins, 35 cycles of 94 °C for 60 s, 50 °C for 1 min, 72 °C, 1 min, and a final step of 72 °C for 7 mins. Amplicons of all genes were taken using bidirectional Sanger sequencing using a BigDye Terminator v3.1 cycle sequencing kit. PCR products were sequenced at HUG-CELL/USP (Human Genome and Stem Cell Research Center, a facility of the University of São Paulo).
Data analysis
The COI alignment (see sequences in Table 2) was performed using G-INS-I option in MAFFT v.7.309 (Katoh and Standley 2013). The alignment was inspected by translation using the invertebrate mitochondrial gene code and trimmed with 500 bp. We used neighbor-joining (NJ), maximum likelihood (ML), and Bayesian inference (BI) methods to infer the phylogenetic relationship of Osedax species. All trees were rooted with Oligobrachia haakonmosbiensis Smirnov, 2000 but we also included species of Vestimentifera and Sclerolinum (Table 2). According to a new phylogeny of Siboglinidae, the choice of a Frenulata species as outgroup is more appropriate, since Osedax is closely related to Vestimentifera and Sclerolinum (Li et al. 2015). The NJ tree topology was constructed under the TrN+G model with 1000 bootstrap replicates in MEGA v.7 (Kumar et al. 2016). The models used in ML and BI were chosen with PartitionFinder v.2 (Lanfear et al. 2016) using Bayesian information criterion. COI was partitioned by codon position. In ML, we used GTR+G, and in BI, 1st and 2nd positions were run with HYK+G and 3rd position with GTR+G. ML was implemented using RAxML v.8.2.7 (Stamatakis 2014). Statistical support of ML nodes was obtained using rapid bootstrap analysis and search for best-scoring ML in a single run (function—fa) of 5000 bootstraps. BI was implemented in MrBayes v.3.2 (Ronquist et al. 2012). The Markov chain Monte Carlo was simulated for 107 generations and sampled every 1000 generations; burn-in was set to 0.1%; two independent runs and four chains were implemented. We calculated average p distance within and between species using MEGA v.7. We also calculated p distance considering only the first two codon position of COI and only the third codon position. Usually, DNA barcoding studies use Kimura 2-parameter model to infer divergence in COI data. However, we chose to use uncorrected distances since K2P divergence commonly inflate the divergence between clades and p distance seems to be more appropriate to compare close related lineages (Srivathsan and Meier 2012). Saturation of COI was examined through saturation curves constructed in DAMBE v6.4.48 (Xia 2017).
We used a new COI alignment with 46 sequences of O. frankpressi Goffredi & Vrijenhoek, 2004 from the Atlantic and Pacific Oceans to investigate the population connectivity between ocean basins (Table 2). The haplotype network of O. frankpressi populations was done using the TCS method (Clement et al. 2000) in PopART (http://popart.otago.ac.nz.) (Leigh and Bryant 2015). In the TCS network, we split the O. frankpressi Atlantic population into two sites: SP-1500 and ES-1500. We calculated the total number of segregating and parsimony informative sites, number of haplotypes (h), haplotype diversity, and nucleotide diversity (π) in DNAsp 5.10.1 (Librado and Rozas 2009) for Atlantic and Pacific populations. We determined the Fu’s neutrality test (FS) (Fu 1997) and Tajima’s D neutrality test (Tajima 1989) as demographic analysis in each population. The populations were compared using the pairwise p distance to construct the FST values using 1000 permutations for significance to estimate the relative population size. A hierarchical analysis of molecular variance (AMOVA) between populations was conducted with the genetic distance. Neutrality tests and AMOVA were calculated using Arlequin 3.5.2.2 (Excoffier and Lischer 2010). All sequences analyzed in this study were deposited in GenBank and the accession numbers are in Table 2.
Results
We sampled 75 specimens of Osedax from four sampling stations and classified them, based on morphology, into two groups: individuals with pinnulated palps (23) or with nude palps (52). Based on DNA barcoding, we found six species-level lineages in the SW Atlantic, including four putative new species. Pinnulated specimens from the SW Atlantic were O. frankpressi Rouse, Goffredi & Vrijenhoek 2004, Osedax braziliensis Fujiwara, Jimi, Sumida, Kawato & Kitazato 2019 (Fig. 2b) or Osedax “BioSuOr-4” (Fig. 2c), and the nude palp group consisted of Osedax ”BioSuOr-1”, O. “BioSuOr-2”, and O. “BiosuOr-3” (Fig. 2a). The highest number of species occurred at SP-3300 with two nude-palp species: Osedax “BioSuOr-1” and Osedax “BioSuOr-2”, and one with pinnulated palps: Osedax braziliensis (Fig. 3). Only one species occurred at SP-1500 and SP-550, O. frankpressi and Osedax “BioSuOr-4”, respectively (Fig. 3). O. frankpressi also colonized bones implanted at ES-1500 occurring in the same bones with the nude-palp Osedax “BioSuOr-3” (Fig. 3).
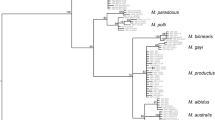
The final COI alignment consisted of 500 bp and 268 variable sites from which ~ 78.4% was parsimony informative. All terminal lineages were robustly supported by NJ and ML bootstrap replicates and posterior probability of BI (Fig. 4). Notwithstanding, only BI recovered robust supports in internal nodes (Fig. 4). Osedax “BioSuOr-1” and “BioSuOr-3” were joined with other nude-palp species in clade II, even though clade II was split in three sub-clades (Fig. 4). Osedax “BioSuOr-2” fell within clade III with low support (ML and NJ > 50%, BI 0.7). Osedax frankpressi and O. braziliensis were always robustly recovered as sister clades (NJ 100%, ML 90%, BI 1). The COI phylogeny clustered species of clades IV, V, and VI (Fig. 4).
The average interspecific p distances were 5.4–25.4% (Table 3). Interspecific divergences among the new SW Atlantic lineages were 16.6–20.4% (Table 3). Considering only the first two codon positions of COI, the distance was 2–7%, while the third codon position exhibited distances ~ 10 times higher than first two, between 14 and 62% (Suppl. Table 1). Intraspecific divergences of SW Atlantic lineages were 0.1–0.9% (Table 3). We observed that numbers of transitions and transversions were equivalent between them where COI divergences were low (Suppl. Fig. 1). However, with increasing COI divergence, transversions became higher than transitions (Suppl. Fig. 1).
The divergence between Atlantic and Pacific O. frankpressi populations was 3.1%, while intra-population divergence was 0.7% in the Atlantic and 0.3% in the Pacific (Table 3). The alignment of O. frankpressi populations consisted of 466 bp and 28 variable sites of which 20 were parsimony informative (Table 4). Between the two ocean basins, we found 20 COI haplotypes, 12 in the Pacific and eight in the Atlantic (Table 4, Fig. 5). No haplotype was shared between ocean basins, but in the Atlantic, one haplotype was shared between SP-1500 and ES-1500, ca. 720 km apart (Fig. 2). Haplotype diversity was slightly higher in Atlantic than in Pacific populations (Table 4). Both Tajima and Fu neutrality tests were negative for both Atlantic and Pacific populations, but only significant (p < 0.01) in Pacific populations. The largest molecular variance occurred between Pacific and Atlantic populations resulting in an FST of 0.88 (p < 0.001) (Table 5).
Discussion
DNA barcoding revealed four new lineages of Osedax inhabiting implanted whalebones in the deep SW Atlantic Ocean. Moreover, the distribution of O. frankpressi was expanded to Atlantic Ocean. O. braziliensis also found in bones implanted at SP3300, yielding a total of six Osedax species observed in the SW Atlantic. O. braziliensis was previously known from the SW Atlantic at a natural whale fall on the São Paulo Ridge at 4204 m depth (Sumida et al. 2016). The new occurrence reported here (at station SP3300), on the lower slope of South São Paulo Plateau at 3358-m depth, is about 200 km away from the original locality. We found only two small specimens of O. braziliensis, with trunk ca. 7 mm in length. Osedax braziliensis is one of the biggest Osedax species reaching 2.5 cm of trunk length (Fujiwara et al. 2019). Bones from the natural whale fall were probably resting on the bottom for 5–10 years and a set of the bones were densely colonized by O. braziliensis (> 40 specimens) being these bones highly degraded (~ 50% of the bone surface) (Sumida et al. 2016; Alfaro-Lucas et al. 2017). The low abundance, found in this study, is probably related to the short time span of the experiment (~ 1.8 year). Even though we did not measure the bone degradation in our experiment, our bones were almost intact, probably not reaching even 10% of the surface area (see Suppl. Fig. 2).
Of the four new Osedax lineages, and even O. frankpressi and O. braziliensis, recovered from the implanted whalebones in the SW Atlantic, no lineages were shared across the ~ 3300-m, ~ 1500-m, and ~ 550-m depth deployments. In our study, landers at ~ 1500-m depth were under the influence of the North Atlantic Deep Water (NADW), which flows southward over the upper and middle São Paulo Plateau, while SP-3300 was under the influence of the Antarctic Bottom Water (AABW) flowing northwards (De Madron and Weatherly 1994; Hogg and Owens 1999; Stramma and England 1999). The Antarctic Intermediate Water (AAIW) dominates the upper slope of São Paulo Plateau (Hogg and Owens 1999), flowing southward (Müller et al. 1998) in the region where SP-550 lander was deployed. The presence of these water masses was confirmed by temperature and salinity data (data not shown here). It is plausible that the absence of shared lineages among depths in our study is related to the influence of different water masses and the current direction. The original locality of O. braziliensis is also bathed by the AABW (Sumida et al. 2016; Fujiwara et al. 2019), reinforcing the idea that the water masses and current flows could control the distribution of Osedax species in SW Atlantic. Similarly, both landers at ~ 1500 m (SP-1500 and ES-1500), under the influence of NADW, share the occurrence of O. frankpressi.
Earlier studies on a deep region of Monterey Canyon (~ 400–2900-m depth) showed a broad bathymetric distribution of some Osedax species (Braby et al. 2007; Lundsten et al. 2010; Rouse et al. 2018). However, the wide bathymetric range may be associated with the deep-water circulation in Monterey Bay, since areas below 300-m depth are connected due to stratification of water column formed by a cyclonic circulation in the deep layer (> 300 m) and an anticyclonic at intermediate depths (< 300 m) (Breaker and Broenkow 1994). Antarctic Osedax species also have eurybathic distribution associated to a cold isothermal water column, such as O. crouchi and O. antarcticus, while O. deceptionensis shows wide distribution along the shallow Antarctic and Subantarctic habitats possibly the result of the Antarctic Circumpolar Current (Amon et al. 2014; Taboada et al. 2015). According to the literature and our data from SW Atlantic, the water masses and currents seem to be important in the distribution of Osedax species, but future studies need to be addressed to better understand this relationship.
This study revealed, for the first time, an inter-basin distribution of O. frankpressi, previously only recorded from the NE Pacific Ocean. There are five other species with trans-Pacific distribution (Rouse et al. 2018), O. rubiplumus also occurs in Antarctic waters (Smith et al. 2015) and O. deceptionensis seems to be widely distributed in shallow Antarctic and Subantarctic waters (Taboada et al. 2015). Some studies shows that small Osedax species, like O. mucofloris, O. packardorum, and O. japonicus, have small oocytes and reduced dispersal time and distance (Fujikura et al. 2006; Rouse et al. 2009; Miyamoto et al. 2013). On the other hand, species with large females and bigger oocytes, like O. frankpressi, may be long-distance dispersers (Rouse et al. 2009).
The Atlantic and Pacific populations of O. frankpressi had negative Tajima’s D and Fu’s FS, but only significant for Pacific populations (p ≤ 0.01). The results indicate a possible recent population expansion for the Pacific population, while for the Atlantic population is uncertain probably due to the low number of samples. Population expansion was reported for O. rubiplumus in Monterey Canyon (NE Pacific) and for O. rogersi and O. deceptionensis in Antarctica (Vrijenhoek et al. 2008; Amon et al. 2014; Taboada et al. 2015).
The low genetic diversity of Atlantic and Pacific populations could indicate recent founders, population bottlenecks, or selective sweeps. The bottleneck shape of the haplotype network and the significantly high FST (Figure 5 and Table 5) indicate segregation between populations of different ocean basins and that geographic distance may act as a barrier to gene flow. However, it is more plausible that the low genetic diversity in both regions and the high FST value are related to the low number of sites sampled for each population and/or the low number of individuals analyzed. Sampling gaps usually inflate FST values (Audzijonyte and Vrijenhoek 2010) explaining the high values found in this study. Smith and Baco (2003) estimated the mean nearest-neighbor distance between whale falls is 5 km for the Gray whale population in the California margin and is 12 km for the nine great whale species in the global ocean. The abundant supply of vertebrate bones facilitates a stepping-stone dispersion of whale fall species (Smith and Baco 2003; Glover et al. 2005) and explains the connection of Osedax species inhabiting the west and east sides of the Pacific Ocean (Rouse et al. 2018). In the same way, intermediate whale falls along the SE Pacific could connect the NE Pacific and SW Atlantic populations of O. frankpressi. Future studies in the SE Pacific could confirm if this region is also colonized by O. frankpressi supporting a connection between NE Pacific and SW Atlantic populations.
The Osedax phylogeny is not yet fully certain since some nodes are poorly supported, even with inclusion of several nuclear markers (Taboada et al. 2015; Rouse et al. 2015, 2018). For example, clade II joined almost all nude palp species, but the relationships within this clade are poorly supported (Rouse et al. 2018), and our phylogeny recovers clade II divided in three subclades (Fig. 2). According to our Bayesian phylogeny, we can confirm that two new nude palp lineages, Osedax “BioSuOr-1” and Osedax “BioSuOr-3” belong to clade II, closely related to O. docricketts/O. westernflyer/O. knutei and O. rogersi and O. lonnyi, respectively. The third Atlantic nude-palp lineage, Osedax “BioSuOr-2”, was related to clade III, a no-palp clade with only O. jabba. As Osedax “BioSuOr-2” bears two pairs of smooth palps, this position needs to be confirmed in future studies. In O. jabba, the root system is also different from other Osedax species; it is thin and filamentous, branching in all direction in the sediments and the fine ends is attached to surfaces of bone fragments (Rouse et al. 2018). We could not see the root system of Osedax “BioSuOr-2”, since they had lost the root system during the sorting process from the bones, but all specimens were well attached to the implanted vertebrae.
Clades IV, V, and VI were mixed in our phylogeny. Vrijenhoek et al. (2009) found that COI phylogeny did not resolve the position of some species in clades IV and V. However, their analyses did not include O. deceptionensis from clade VI, described only years later (Glover et al. 2013). The position of clade VI is not resolved, with some analyses showing this clade basal in Osedax phylogeny and in other studies clade VI is more related to clade I and/or II (Glover et al. 2013; Amon et al. 2014; Taboada et al. 2015; Rouse et al. 2015, 2018). The COI and H3 phylogeny shows that O. frankpressi (clade IV) falls close to O. roseus (clade V) (Vrijenhoek et al. 2009), which was also recovered by our analysis (Fig. 2). Osedax “BioSuOr-4”, a pinnulated palp lineage, is probably a member of clade V since the COI divergences of this lineage are slightly lower with species of clade V than with species of clade IV.
In conclusion, the results presented here revealed six Osedax species in the deep SW Atlantic whale falls, four of them being putative new species. Interestingly, other deep regions such as the NE and NW Pacific and the Southern Ocean also have a high numbers of Osedax species (Rouse et al. 2004, 2008, 2018; Vrijenhoek et al. 2009; Glover et al. 2013; Amon et al. 2014; Pradillon et al. unpublished), while shallow, cold-water regions are usually colonized by a few species (Glover et al. 2005, 2013; Fujikura et al. 2006; Taboada et al. 2015). This study also showed that bones implanted in different depths showed different species composition possibly related to the difference in water circulation in these depths. The presence of Osedax frankpressi across ocean basins (Pacific and Atlantic) infers that the provision of whalebones on the seafloor must be plentiful enough to facilitate the evolution and wide distribution for Osedax species.
References
Alfaro-Lucas JM, Shimabukuro M, Ferreira GD, Kitazato H, Fujiwara Y, Sumida PY (2017) Bone-eating Osedax worms (Annelida: Siboglinidae) regulate biodiversity of deep-sea whale-fall communities. Deep-Sea Res II 146:4–12. https://doi.org/10.1016/j.dsr2.2017.04.011
Amon DJ, Wiklund H, Dahlgren TG, Copley JT, Smith CR, Jamieson AJ, Glover AG (2014) Molecular taxonomy of Osedax (Annelida: Siboglinidae) in the Southern Ocean. Zool Scr 43(4):405–417. https://doi.org/10.1111/zsc.12057
Audzijonyte A, Vrijenhoek RC (2010) When gaps really are gaps: statistical phylogeography of hydrothermal vent invertebrates. Evolution 64(8):2369–2384. https://doi.org/10.1111/j.1558-5646.2010.00987.x
Braby CE, Rouse GW, Johnson SB, Jones WJ, Vrijenhoek RC (2007) Bathymetric and temporal variation among Osedax boneworms and associated megafauna on whale-falls in Monterey Bay, California. Deep Sea Res I 54:1773–1791. https://doi.org/10.1016/j.dsr.2007.05.014
Breaker LC, Broenkow WW (1994) The circulation of Monterey Bay and related processes. Oceanogr Mar Biol Annu Rev 32:1–64 ISSN: 0078-3218
Campos EJD, Miller JL, Müller TJ, Peterson RG (1995) Physical oceanography of the Southwest Atlantic Ocean. Oceanography 8(3):87–91. https://doi.org/10.5670/oceanog.1995.03
Clement MP, Posada D, Crandall K (2000) TCS: a computer program to estimate gene genealogies. https://doi.org/10.1046/j.1365-294x.2000.01020.x
De Madron XD, Weatherly G (1994) Circulation, transport and bottom boundary layers of the deep currents in Brazil Basin. J Mar Res 52:583–638. https://doi.org/10.1357/0022240943076975
Excoffier L, Lischer HE (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10(3):564–567. https://doi.org/10.1111/j.1755-0998.2010.02847.x
Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147:915–925
Fujikura K, Fujiwara Y, Kawato M (2006) A new species of Osedax (Annelida: Siboglinidae) associated with whale carcasses off Kyushu, Japan. Zool Sci 23(8):733–740. https://doi.org/10.2108/zsj.23.733
Fujiwara Y, Jimi N, Sumida PYG, Kawato M, Kitazato H (2019) New species of bone-eating worm Osedax from the abyssal South Atlantic Ocean (Annelida, Siboglinidae). ZooKeys 814:53–69. https://doi.org/10.3897/zookeys.814.28869
Glover AG, Källström B, Smith CR, Dahlgren TG (2005) World-wide whale worms? A new species of Osedax from the shallow north Atlantic. Proc R Soc 272:2587–2592. https://doi.org/10.1098/rspb.2005.3275
Glover AG, Wiklund H, Taboada S, Avila C, Cristobo J, Smith CR, Kemp KM, Jamieson AJ, Dahlgren TG (2013) Bone-eating worms from the Antarctic: the contrasting fate of whale and wood remains on the Southern Ocean seafloor. Proc R Soc B 280:20131390. https://doi.org/10.1098/rspb.2013.1390
Goffredi SK, Orphan VJ, Rouse GW, Jahnke L, Embaye T, Turk K, Lee R, Vrijenhoek RC (2005) Evolutionary innovation: a bone-eating marine symbiosis. Environ Microbiol 7:1369–1378. https://doi.org/10.1111/j.1462-2920.2005.00824.x
Goffredi SK, Johnson SB, Vrijenhoek RC (2007) Genetic diversity and potential function of microbial symbionts associated with newly discovered species of Osedax polychaete worms. Appl Environ Microbiol 73:2314–2323. https://doi.org/10.1128/AEM.01986-06
Higgs ND, Little CTS, Glover AG (2011) Bones as biofuel: a review of whale bone composition with implications for deep-sea biology and paleoanthropology. Proc R Soc B 278(1702):9–17. https://doi.org/10.1098/rspb.2010.1267
Hogg NG, Owens B (1999) Direct measurement of the deep circulation within the Brazil Basin. Deep-Sea Res II 46:335–353. https://doi.org/10.1016/S0967-0645(98)00097-6
Jones WJ, Johnson SB, Rouse GW, Vrijenhoek RC (2008) Marine worms (genus Osedax) colonize cow bones. Proc R Soc B 275:387–391. https://doi.org/10.1098/rspb.2007.1437
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30(4):772–780. https://doi.org/10.1093/molbev/mst010
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Lanfear R, Frandsen PB, Wright AM, Senfeld T, Calcott B (2016) PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol Biol Evol 34(3):772–773. https://doi.org/10.1093/molbev/msw260
Leigh JW, Bryant D (2015) Popart: full-feature software for haplotype network construction. Methods Ecol Evol 6(9):1110–1116. https://doi.org/10.1111/2041-210X.12410
Li Y, Kocot KM, Schander C, Santos SR, Thornhill DJ, Halanych KM (2015) Mitogenomics reveals phylogeny and repeated motifs in control regions of the deep-sea family Siboglinidae (Annelida). Mol Phylogenet Evol 85:221–229. https://doi.org/10.1016/j.ympev.2015.02.008
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452. https://doi.org/10.1093/bioinformatics/btp187
Lösekann T, Robador A, Niemann H, Knittel K, Boetius A, Dubilier N (2008) Endosymbioses between bacteria and deep-sea siboglinid tubeworms from an Arctic Cold Seep (Haakon Mosby Mud Volcano, Barents Sea). Environmental Microbiology 10 (12):3237–3254 https://doi.org/10.1111/j.1462-2920.2008.01712.x
Lundsten L, Schlining KL, Frasier K, Johnson SB, Kuhnz LA, Harvey JBJ, Clague G, Vrijenhoek RC (2010) Times-series analysis of six whale-fall communities in Monterey canyon, California, USA. Deep Sea Res I 57:1573–1584. https://doi.org/10.1016/j.dsr.2010.09.003
McCartney MS (1993) Crossing of the equator by the deep west boundary current in the western Atlantic Ocean. J Phys Oceanogr 23:1953–1974. https://doi.org/10.1175/1520-0485(1993)023<1953:COTEBT>2.0.CO;2
McMullin ER, Hourdez S, Schaeffer, SW and Fisher CR (2003) Phylogeny and biogeography of deep sea vestimentiferan tubeworms and their bacterial symbionts. Symbiosis 34, 1–41.
Miyamoto N, Yamamoto T, Yusa Y, Fujiwara Y (2013) Postembryonic development of the bone-eating worm Osedax japonicus. Naturwissenschaften 100:285–289. https://doi.org/10.1007/s00114-013-1024-7
Müller TJ, Ikeda Y, Zangenberg N, Nonato LV (1998) Direct measurements of western boundary currents off Brazil between 20°S and 28°S. J Geophys Res 103(C3):5429–5437. https://doi.org/10.1029/97JC03529
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes v. 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Rouse GW, Goffredi SK, Vrijenhoek RC (2004) Osedax: bone-eating marine worms with dwarf males. Science 30:668–671. https://doi.org/10.1126/science.1098650
Rouse GW, Worsaae K, Johnson SB, Jones WJ, Vrijenhoek RC (2008) Acquisition of dwarf male “harems” by recently settled females of Osedax roseus n. sp. (Siboglinida; Annelida). Biol Bull 214:67–82. https://doi.org/10.2307/25066661
Rouse GW, Wilson NG, Goffredi SK, Johnson SB, Smart T, Widmer C, Young CM, Vrijenhoek RC (2009) Spawning and development in Osedax boneworms (Siboglinidae, Annelida). Mar Biol 156:395–405. https://doi.org/10.1007/s00227-008-1091-z
Rouse GW, Wilson NG, Worsaae K, Vrijenhoek RC (2015) A dwarf male reversal in bone-eating worms. Curr Biol 25:1–6. https://doi.org/10.1016/j.cub.2014.11.032
Rouse GW, Goffredi SK, Johnson SB, Vrijenhoek RC (2018) An inordinate fondness for Osedax (Siboglinidae: Annelida): fourteen new species of bone worms from California. Zootaxa 4377(4):451–489. https://doi.org/10.11646/zootaxa.4377.4.1
Saeedi H, Bernardino AF, Shimabukuro M, Falchetto G, Sumida PYG (2019) Macrofaunal community structure and biodiversity patterns based on a wood-fall experiment in the deep South-west Atlantic. Deep Sea Res I 145:73–82. https://doi.org/10.1016/j.dsr.2019.01.008
Salathé RM, Vrijenhoek RC (2012) Temporal variation and lack of host specificity among bacterial endosymbionts of Osedax bone worms (Polychaeta: Siboglinidae). BMC Evol Biol 12:189. https://doi.org/10.1186/1471.2148.12.189
Santos MCO, Siciliano S, Vicente AFC, Alvarenga FS, Zampirolli E, Souza SP, Maranho A (2010) Cetacean records along São Paulo state coast southeastern Brazil. Braz J Oceanogr 58(2):123–142. https://doi.org/10.1590/S1679-87592010000200004
Silveira ICA, Schmidt ACK, Campos EJD, Godoi SS, Ikeda Y (2000) A corrente do Brazil ao largo da costa leste Brasileira. Rev Bras Oceanogr 48(2):171–183. https://doi.org/10.1590/S1413-77392000000200008
Smith CR, Baco AR (2003) Ecology of whale falls at the deep-sea floor. Oceanogr Mar Biol Annu Rev 41:311–354 ISSN: 0078-3218
Smith CR, Glover AG, Treude T, Higgs ND, Amon DJ (2015) Whale-fall ecosystems: recent insights into ecology, paleoecology, and evolution. Annu Rev Mar Sci 7:10.1–10.6. https://doi.org/10.1146/annurev-marine-010213-135144
Srivathsan A, Meier R (2012) On the inappropriate use of Kimura-2-parameter (K2P) divergences in the DNA-barcoding literature. Cladistics 28(2):190–194. https://doi.org/10.1111/j.1096-0031.2011.00370.x
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30(9):1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Stramma L, England M (1999) On the water masses and mean circulation of the South Atlantic Ocean. J Geophys Res 104(C9):20,863–20,883. https://doi.org/10.1029/1999JC900139
Sumida PYG, Alfaro-Lucas JM, Shimabukuro M, Kitazato H, Perez JAA, Soares-Gomes A, Toyofuku T, Lima AOS, Ara K, Fujiwara Y (2016) Deep-sea whale fall fauna from the Atlantic resembles that of the Pacific Ocean. Sci Rep 6:22139. https://doi.org/10.1038/srep22139
Taboada S, Riego A, Bas M, Arnedo MA, Cristobo J, Rouse GW, Avila C (2015) Bone-eating worms spread: insights into shallow-water Osedax (Annelida, Siboglinidae) from Antarctic, Subantarctic, and Mediterranean waters. PLoS One 10(11):e0140341. https://doi.org/10.1371/journal.pone.0140341
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123(3):585–595 ISSN: 0016-6731
Treude T, Smith CR, Wenzhöfer F, Carney E, Bernardino AF, Hannides AK, Krüger M, Boetius A (2009) Biogeochemistry of a deep-sea whale fall: sulfate reduction, sulfide efflux and methanogenesis. Mar Ecol Prog Ser 382:1–21. https://doi.org/10.3354/meps07972
Vrijenhoek RC, Johnson SB, Rouse GW (2008) Bone-eating Osedax females and their “harems” of dwarf males are recruited from a common larval pool. Mol Ecol 17:4535–4544. https://doi.org/10.1111/j.1365-294X.2008.03937.x
Vrijenhoek RC, Johnson SB, Rouse GW (2009) A remarkable diversity of bone-eating worms (Osedax; Siboglinidae; Annelida). BMC Biol 7(1):74. https://doi.org/10.1186/1741-7007-7-74
Wedekin LL, Rossi-Santos MR, Baracho C, Cypriano-Souza AL, Simões-Lopes PC (2014) Cetacean records along a coastal-offshore gradient in the Vitória-Trindade Chain, western South Atlantic Ocean. Braz JBiol 74(1):137–144. https://doi.org/10.1590/1519-6984.21812
Worsaae K, Rouse GW (2010) The simplicity of males: dwarf males of four species of Osedax (Siboglinidae: Annelida) investigated by confocal laser scanning microscopy. J Morphol 271:127–142. https://doi.org/10.1002/jmor.10786
Xia X (2017) DAMBE6: new tools for microbial genomics, phylogenetics, and molecular evolution. J Hered 108(4):431–437. https://doi.org/10.1093/jhered/esx033
Zerbini AN, Secchi ER, Siciliano S, Simões-Lopes PC (1997) A review of the occurrence and distribution of whales of the genus Balaenoptera along the Brazilian coast. Report International Whaling Commission 47:407–417 Available at: https://archive.iwc.int/pages/search.php?search=!collection33&bc_from=themes. Accessed 14 Aug 2017
Zerbini AN, Andriolo A, Heide-Jørgensen MP, Pizzorno JL, Maia YG, VanBlaricom GR, DeMaster DP, Simões-Lopes P, Moreira S, Bethlem C (2006) Satellite-monitored movements of humpback whales Megaptera novaeangliae in the Southwest Atlantic Ocean. Mar Ecol Prog Ser 313:295–304. https://doi.org/10.3354/meps313295
Acknowledgments
We would like to thank the Masters and crews of R/V Alpha Crucis and R/V Alpha Delphini from IOUSP during the deployment and the recovering of SP-1500 lander. We are also indebted to the Brazilian Navy and the Master and crew of PRV Almirante Maximiano for the help during the recovering of the other landers. We also thank the editor, Dr. Craig R. Smith, and the two anonymous reviewers for the comments on the manuscript. MS was supported by CAPES/Proex PhD scholarship.
Funding
This study was funded by São Paulo Research Foundation (FAPESP) into the Research Program on Biodiversity Characterization, Conservation, Restoration and Sustainable Use (BIOTA Program)—grant number 2011/50185-1.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed by the authors.
Sampling and field studies
All necessary permits for sampling have been obtained by the authors from the competent authorities.
Data availability statement
All data generated or analysed during this study are included in this published article and its supplementary information files. All new COI sequences of Osedax were deposited in GenBank and the accession numbers are detailed in Table 2.
Additional information
Communicated by C. Smith
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Figure S1
(PNG 582 kb)
Supplementary Figure S2
(PNG 13551 kb)
Supplementary Table S1
(XLSX 53 kb)
Rights and permissions
About this article
Cite this article
Shimabukuro, M., Sumida, P.Y.G. Diversity of bone-eating Osedax worms on the deep Atlantic whale falls—bathymetric variation and inter-basin distributions. Mar. Biodivers. 49, 2587–2599 (2019). https://doi.org/10.1007/s12526-019-00988-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12526-019-00988-2