Abstract
Leaf growth and development are primarily driven by cell proliferation and expansion. Among a number of genes involved in the regulation of cell proliferation, the GROWTH-REGULATING FACTOR (GRF) transcription factors have been established to act as positive regulators of cell proliferation and leaf growth in angiosperms. While the Arabidopsis thaliana GRF family comprises nine members, not all members of the family have been experimentally confirmed for the positive role, not only due to no or only slight changes in leaf size of corresponding single mutants, but also due to unavailability of multiple mutants to overcome the obstacle. Furthermore, some discrepancies and confusion in their roles have been disclosed in the literature. Here, we systemically prepared a series of such multiple mutants and confirmed that all GRF members, except for GRF8, acted as positive regulators of cell proliferation and leaf growth. We also systematically examined the spatio-temporal distribution patterns of all nine GRF proteins in the leaf organ, and found that their distribution patterns were highly reminiscent of the behavior of the cell cycle arrest front. We therefore propose that GRFs play an important role in shaping the arrest front and growth patterns of leaves.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant leaves are the primary organs that perform photosynthesis and thus convert sun light into chemical energy as a form of carbohydrates. The carbohydrates serve as nutrient sources not only for plants to grow and develop, but also for almost all living organisms on earth, including humans, highlighting the importance of understanding leaf growth and development. The leaf primordium develops from a group of cells at the flank of the shoot apical meristem and takes a finger-tip-like shape initially. Afterward, it first undergoes vigorous cell division and develops into a flattened lamina structure (Donnelly et al. 1999; Tsukaya 2013; Sarvepalli et al. 2019). Over time, the lamina enlarges to a mature leaf via a coordinated involvement of cell division and expansion. The final size of leaf organs is primarily under the control of genetic programs that determines the rate and duration of cell proliferation and expansion. In Arabidopsis thaliana (hereafter, Arabidopsis), the leaf primordial cells are highly prolific or meristematic with limited cell expansion (Donelly et al. 1999; Autran et al., 2002; Lee et al., 2009). Yet, the situation is reversed as the primordium grows further: the cells in the distal region of the leaf lamina start to lose their meristematic potential, initiating the differentiation program manifested by vigorous cell expansion; the non-meristematic region of the lamina expands to the base gradually; and, eventually, the whole lamina loses cell proliferation activities, except in some tissues, reaching its final size with fully expanded cells. The transition boundary between the meristematic cells and differentiating cells seemingly moves in a tip-to-base direction, or basipetally, displaying a moving front, so-called the cell cycle arrest front.
The cell division occurring in the leaf primordium is tightly regulated by the core cell cycle regulator, the complex comprising cyclins (CYCs) and CYC-dependent kinases (CDKs). The CYC–CDK complexes are highly cell cycle phase-specific (for reviews, see Vercruysse et al. 2020). In brief, A-type CYCs (CYCAs) and D-type CYCs (CYCDs) are mainly involved in regulation of the G1 progression and G1/S transition, while B-type CYCs (CYCBs) predominantly drive the G2/M transition and progression through mitosis. Similarly, A-type CDKs (CDKAs) exert their function during G1/S and G2/M transitions, and B-type CDKs (CDKB) mainly during the G2/M transition and progression through mitosis.
A number of genes involved in regulation of cell proliferation have been identified (Vercruysse et al. 2020). Among them, the GROWTH-REGULATING FACTOR (GRF) and GRF-INTERACTING FACTOR (GIF) transcriptional complex have been known to be essential for plant growth and development (for reviews, Kim and Tsukaya 2015; Omidbakhshfard et al. 2015; Liebsch and Palatnik 2020). The GRF–GIF duo promotes the cell cycle activities of lateral organs, such as leaves and floral organs, at their primordial and actively growing stages, and thus contribute to determination of their final size and shape. GRFs are transcription factors that are conserved in streptophytes, including land plants (Kim 2019; Fonini et al. 2020). The Arabidopsis thaliana GRF family is consisted of nine members. Meanwhile, GIFs are transcription co-activators that are highly conserved in the major lineages of eukaryotic organisms, including land plants, and the Arabidopsis thaliana GIF family comprises three members. It has been shown that the members of the two families in Arabidopsis and other model plants form a transcriptional complex in almost all possible combinations (Kim and Kende 2004; Horiguchi et al. 2005; Liang et al. 2014; Liu et al. 2014; Duan et al. 2016; Zhang et al. 2018).
The role of the GRF–GIF duo as a positive regulator of cell proliferation has been best studied in Arabidopsis and further shown, in large, to be conserved in other model plants, such as rice and maize, although, in certain organs and conditions, the duo can regulate both cell number and size (Kim and Tsukaya 2015; Omidbakhshfard et al. 2015; Kim 2019; Liebsch and Palatnik 2020). Most of single, loss-of-function mutants of GRFs displayed no or only minute defects in leaf growth, whereas multiple combinations of those single mutations caused an obvious reduction in leaf growth, suggesting that GRFs seem to be functionally redundant. In more detail, single mutants of grf1—grf4 displayed the size and shape of the wild-type leaves, whereas their double, triple, and quadruple mutants showed gradual reductions in cell proliferation activities, producing smaller and narrower leaves than those of the wild type (Kim et al. 2003; Kim and Kende 2004; Kim and Lee 2006). The grf5 single mutant produced slightly small leaves due to a reduction in cell number (Horiguchi et al. 2005). The grf7 single mutant also had smaller leaves than the wild type, although we failed to recognize the difference in the same single mutant allele (see below), and the grf8 single mutant did not show any difference in leaf size compared with the wild type (Kim et al. 2012), although GRF8 has been, by implication, presumed to be a positive regulator of cell proliferation (Tsukaya 2021). In contrast, the grf9 single mutant displayed a slightly bigger leaves with more cells than the wild type, indicating that GRF9 acts as negative regulators of cell proliferation and leaf growth (Omidbakhshfard et al. 2018). It should be noticed, however, that other reports showed no significant changes in leaf size of the same grf9 mutant allele (Horiguchi et al. 2005; Liang et al. 2014).
The cellular function and biological roles of GRFs in Arabidopsis have been established through analyses of grf multiple mutants and micro-RNA396 (miR396) overexpressors (Liu et al. 2009; Rodriguez et al. 2010; Lee et al. 2018). Arabidopsis miR396 targets almost all GRF mRNAs with the exceptions of GRF5 and GRF6 mRNAs, inducing their cleavage and degradation. As such, overexpression of the MIR396 genes (MIR396a and MIR396b) by the viral 35S promoter downregulated GRF expression as a whole and thus produced distinctively small and narrow leaves (Liu et al. 2009; Rodriguez et al. 2010). 35S::MIR396b plants showed reductions in both the cell number and expression level of CYCB1;1::GUS, a molecular marker for the G2/M transition and mitosis (Rodriguez et al. 2010).
Among the three members of the GIF family, GIF1 (aka ANGUSTIFOLIA3, AN3) plays a prominent role in leaf growth, since gif1 and an3 single mutants produce distinctively small and narrow leaves due to a reduction in cell proliferation (Kim and Kende 2004; Horiguchi et al. 2005). Although gif2 and gif3 single mutants show no visible defects, the mutations synergistically enhance the defective leaf phenotype of gif1, revealing functional redundancy between them: the gif1 gif2 gif3 triple mutant produced tiny leaves with a modicum of mesophyll cells due to down-regulation of cell cycle-related genes, such as CYCs and a CDK (Lee et al. 2009). In addition, when combined together, gif1, an3, grf, and 35S::MIR396b severely compromise leaf growth, giving rise to the notion of the GRF–GIF–miR396 regulatory module (Kim and Kende 2004; Rodriguez et al. 2010).
Although the cellular mechanism by which the GRF–GIF duo regulates leaf growth has been largely understood, the role or impact of all individual GRFs has not been experimentally demonstrated. Besides, the inconsistency pertinent to the role of GRF7 and GRF9 in leaf growth has been disclosed, as mentioned above. In the present study, we perform a systematic assessment of the role and impact of each member of the GRF family by constructing multiple grf mutants and demonstrate that most of the GRF members act as positive regulators of cell proliferation contributing to regulation of the final size and shape of the leaf organ. We also discuss the correlation between GRF action and the behavior of the cell cycle arrest front.
Results
The gif1 gif2 35S::MIR396b Triple Mutant Almost Completely Compromises Leaf Growth
It was previously shown that the gif1 grf1 grf2 triple mutations (simply, gif1 grf1/2) resulted in a severe reduction in leaf size and cell number (Kim and Kende 2004), and that an3 and 35S::MIR396b also caused synergistic deterioration in leaf growth (Rodriguez et al. 2010). Here, we extended the genetic interaction between 35S::MIR396b and all gif mutations (Fig. 1). 35S::MIR396b, gif1, gif1/2, and gif1/3 mutants produced small and narrow leaves with small numbers of mesophyll cells, which is obviously indicative of the positive role of both the GRF and all GIF members in cell proliferation and leaf growth, as reported previously (Horiguchi et al. 2005; Lee et al. 2009; Rodriguez et al. 2010). Furthermore, introduction of gif1 single and gif1/3 double mutations into 35S::MIR396b markedly reduced leaf size and cell number. Moreover, gif1/2 35S::MIR396b almost completely compromised leaf growth and cell proliferation. In addition to the leaf size, higher the order of mutational combinations was, narrower the leaf shape was (leaf narrowness was indicated by the leaf index, i.e., a ratio of leaf length divided by leaf width). The results clearly showed the synergistic interaction between all gif mutations and 35S::MIR396b.
Leaf phenotypes of gif and 35S::MIR396b mutants. A Leaf phenotypes of 25-day-old plants: upper panel, whole plants and first-to-fourth rosette leaves; lower, palisade cells in the maximum-width region. Scale bars indicate 1 cm and 25 μm, respectively. B Area, length, and width of the first pair of leaves (columns) as well as leaf index (closed circles). C, Calculated numbers of palisade cells, which were derived from leaf area divided by cell area as well as counted numbers of cells aligned along a transverse axis in the maximum-width region. The number 396 indicates 35S::MIR396b; 1 × 396, gif1 35S::MIR396b; 12 × 396, gif1 gif2 35S::MIR396b; 13 × 396, gif1 gif3 35S::MIR396b. n = 10 for leaf area, n = 100 for palisade cell parameters. Error bars indicate S.E. values
Apart from the changes in cell number of the mutant leaves, their cell areas increased with inverse correlation to cell numbers (Fig. 1C), which was referred to a ‘compensatory’ syndrome occurring in response to a substantial reduction in cell number (Tsukaya 2002). Of note, however, the compensatory increases in cell area were not to fully make up for reductions in leaf size resulting from defective cell proliferation.
Most of the GRF Members Act as Positive Regulators of Cell Proliferation and Leaf Growth
It is conceivable that 35S::MIR396b should exert its influence through miR396-targeted GRFs. In fact, the target GRFs are downregulated by 35S::MIR396b, and even non-target members, such as GRF5 and GRF6, are likewise downregulated by 35S::MIR396b, probably because the autoactivation loop of the GRF–GIF transcriptional complex is impaired by miR396 overexpression (Liu et al. 2009; Rodriguez et al. 2010; Debernardi et al. 2014). Nevertheless, it has not been fully understood which members of the GRF family contribute to leaf growth and to what extent (Vercruysse et al. 2020). To clarify the involvement and impact of individual GRFs in leaf growth, we constructed a series of multiple grf mutants in the Columbia accession through a long process of cross-pollination. Single mutants used are followings: grf1-3, grf2-1, grf3-1, grf4-1, grf5-2, grf7-1, grf8-1, and grf9-1 (the grf6 mutant is not available at present). They are all derived from T-DNA insertion (for detailed information about their mutational nature, see Lee et al. 2018; hereafter, the numbers for allele designation were left out for simplicity, unless necessary). It should be noted that, since the original grf2 mutation (designated as grf2-0) was derived from the Wassilewskija (Ws) accession, the grf1/2/3 triple mutant in this study was obtained after five back-crosses with the Columbia (Col) accession. The grf2 single mutant analyzed in this study was segregated out from one more cross between the cleaned triple mutant and wild-type Col and designated as grf2-1. All single mutants did not show any visible changes in leaf size and shape, although careful quantification reveals slight reductions in some single mutants, such as grf1, grf3, and grf5 (Figs. 2 and 3). On the other hand, grf1/3, grf1/4, and grf1/5 double mutants developed noticeably smaller and narrower leaves with fewer cells than those of their parental single mutants. The grf1/2/3 triple mutant also showed smaller and narrower leaves than its constituent single mutants, but not smaller than the grf1/3 double mutant, probably because of the mixed genetic background even after five back-crosses with Col. We previously demonstrated gradual suppression of leaf growth owing to additions of grf1-1, grf2-0, and grf3-0, which are all in Ws (Kim et al. 2003; Kim and Kende 2004). The results all together indicate that GRF1–GRF5 promote leaf growth and cell proliferation. The leaf size of grf1/7 and grf1/3/7 was not different from those of grf1 and grf1/3, respectively, indicating that the grf7 mutation, albeit being a null allele (Kim et al. 2012), did not affect leaf growth at the particular combinations. At the other combinations (grf1/5/7 and grf1/3/5/7), however, the grf7 mutation revealed its additive influence, resulting in a little more reductions in both leaf size and cell number compared with grf1/5 and grf1/3/5, respectively, and moreover, its influence was significantly manifested in the grf1/3/4/5/7 quintuple mutant compared with grf1/3/4/5. The change in leaf narrowness was also prominent in the grf1/3/4/5/7 quintuple mutant (Figs. 2 and 3). It is clear therefore that GRF7 contributes to cell proliferation and thus leaf growth, though with less impact compared with GRF1–GRF5.
Quantitative parameters of grf mutants. A, B Size of the first pair of leaves (columns) and leaf index (closed circles). C Calculated numbers of palisade cells, which were derived from leaf area divided by cell area. Error bars indicate S.E. values. Values of each group with the same letters were not significantly different (P < 0.05; one-way analysis of variance with the Tukey test)
The positive role of GRF9 in leaf growth was also disclosed: although the grf9 single mutant showed almost no reductions in leaf growth, grf1/3/4/5/9 had substantially smaller and narrower leaves with fewer cells than grf1/3/4/5 (Figs. 2 and 3). On the other hand, any contribution to leaf growth by GRF8 was not detected at the levels of grf8 single and grf1/5/7/8 quadruple mutants when compared with grf1/5/7 (Fig. 3A). Even the one-order higher multiple mutant, grf1/4/5/7/8, showed no difference in leaf size compared with that of grf1/4/5/7 (Fig. 3B).
Additionally, significant reductions in cell number by cumulative addition of mutations were accompanied by increases in cell area in an inversely proportional manner, which is a manifestation of the compensatory syndrome (Fig. 3C).
Distribution Patterns of GRF::GUS Fusion Proteins During Leaf Growth
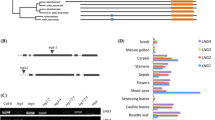
Expression patterns of some GRF genes in leaves have been examined using the β-glucuronidase (GUS) marker gene fused to their promoters or whole genomic sequences (Kim et al. 2003; Horiguchi et al. 2005; Rodriguez et al. 2010; Liang et al. 2014; Beltramino et al. 2021). In the present study, we systematically prepared transgenic plants expressing the GUS gene fused to the whole genomic sequences of all nine GRFs, including the ~ 2 kb-long promoter as well as exons and introns, and performed the GUS assay. Among all, the leaves of PGRF1::GRF1::GUS (simply, GRF1::GUS) and GRF2::GUS displayed prominent signals (Fig. 4). The signals were strongly detected all over the whole primordial leaves at 3 and 5 days after germination (DAG). Afterward, the signals began to fade away from the leaf tip, being restricted to the basal region over time, and finally disappeared almost completely at 11 DAG. The staining patterns of GRF3::GUS–GRF9::GUS were, in large, similar to those of GRF1::GUS and GRF2::GUS, though weaker than those. Unlike the others, however, GRF4::GUS and GRF6::GUS started to display the signal at 5 DAG, and GRF4::GUS appeared at the distal region of leaves at 5 DAG. On the other hand, the staining signal of GRF8::GUS was barely detected, indicating that no detectable effect of the grf8 mutation on leaf growth may be due to the extremely weak expression of GRF8 in the leaf. In summary, the results suggest that expression of most of GRF genes is active at the earliest primordial stages, after which the expression is rapidly repressed in a tip-to-base direction.
Expression Patterns of the Core Cell Cycle Genes
It has been well established that the CYCB1;1::GUS fusion gene specifically marks the cells that are in the G2/M transition and mitosis (Donnelly et al. 1999; Vercruysse et al. 2020). We employed the marker gene to investigate changes in the mitotic activities during leaf growth. In the wild type, CYCB1;1::GUS expression was detected in a high level all over the first pair of the wild-type primordial leaves at 5 DAG, and then started to fade away from the tip, being restricted to the base (Fig. 5). Finally, the expression completely disappeared at 11 DAG, although it persisted in the veins. In contrast, the levels of CYCB1;1::GUS expression were very low in grf1/3/5 and grf1/3/4/5 mutants. Moreover, its expression front moved downwards earlier than that of the wild type, resulting in almost no signals at 9 DAG. CYCB1;1::GUS expression was more impaired in grf1/3/4/5 than in grf1/3/5, revealing an additive role of GRF4 in regulation of CYCB1;1::GUS expression. The expression patterns of CYCB1;1::GUS in gif1 and 35S::MIR396b were similar to those in grf multiple mutants, resulting in no signals at 9 DAG, except for the veins. CYCB1;1::GUS expression was further impaired by gif1 35S::MIR396b double mutations.
It is noteworthy that the development of the vascular structures of all the mutants were severely impaired. Most of the mutants did not develop the tertiary veins, and the gif1 35S:MIR396 double mutant developed only the primary vein.
Discussion
Most of GRF Members Act as Positive Regulators of Cell Proliferation During Leaf Growth
Since the discovery of GRF transcription factors in deepwater rice a couple of decades ago, many researchers have elucidated the roles and significance of GRFs in various aspects of plant growth and development (Van der Knaap et al. 2000; Kim and Tsukaya 2015; Omidbakhshfard et al. 2015). Among those, the role of GRF in leaf growth has been most extensively studied in Arabidopsis, producing evidence for the notion that GRFs are positive regulators of cell proliferation and thus determine the final size and shape of leaves. As mentioned above, however, not all members of the GRF family have been empirically tested not only due to the difficulty in phenotypic analysis of corresponding single mutants, but also due to unavailability of multiple mutants. Here, we prepared a series of such multiple mutants and demonstrated that all GRFs with the exception of GRF8 acted as positive regulators of cell proliferation and leaf growth (Figs. 2 and 3).
We previously demonstrated that GRF1–GRF3 acted as positive regulators of cell proliferation in leaf growth by analyzing the grf1-1/2-0/3-0 triple mutant, which was in the Ws accession (Kim and Kende 2004; Kim and Lee 2006), although their cellular role was wrongly attributed to cell expansion in the initial analysis (Kim et al. 2003). Here, we once more confirmed the previous results by analyzing the grf1-3/2-1/3-1 triple mutant, which was nearly isogenic to the Col accession. The result would help to dispose of the confusion caused by the initial analysis. We also previously concluded that GRF4 acted as a positive regulator of cell proliferation by analyzing the grf1-1/2-0/3-0/4-1 quadruple mutant (Kim and Lee 2006). The quadruple mutant was, however, derived from the cross between grf4-1 in Col and the grf1-1/2-0/3-0 triple mutant in Ws, and thus, the conclusion was obscured due to the drawback of the mixed genetic background. Here, we clearly demonstrated the role of GRF4 in cell proliferation and leaf growth by comparing multiple mutants, which are all in Col (Figs. 2, 3 and 5). Analysis of multiple mutants containing the grf5 mutation revealed the robust contribution of GRF5 to cell proliferation and leaf growth (Figs. 2 and 3), as previously shown by Horiguchi et al. (2005). Recently, Beltramino et al. (2021) showed the additive effect of grf3 and grf5 on cell proliferation and leaf growth.
We noticed that some of our data pertinent to GRF7 and GRF9 were in discrepancy with previous ones. Kim et al (2012) reported that the grf7-1 single mutant and GRF7-specific RNA-interference lines produced smaller leaves than the wild type, although the authors did not analyze cellular parameters. To the contrary, we failed to recognize any significant changes in leaf size of the same single mutant allele and were able to detect an obvious reduction in leaf size and cell number only when the grf1/3/4/5 and grf1/3/4/5/7 multiple mutants were compared. The discrepancy might stem from different conditions of plant culture: the previous and this studies analyzed plants grown on agar plates and soil, respectively, suggesting a possible reconciliation.
Unfortunately, however, the discrepancy between the results from Omidbakhshfard et al. (2018) and this study seems hardly to be resolved. The previous study showed that grf9-1 and grf9-2 single mutants had slightly bigger leaves with more cells than those of the wild type, suggesting that the wild-type GRF9 acts as a negative regulator of cell proliferation. Unlike in the case of grf7 mutants, the growth conditions, including seed stratification and photoperiod regime, were similar in both studies. The authors also meticulously demonstrated that the negative effect of GRF9 on cell proliferation was, at least in part, mediated by the transcriptional activation of a target gene, which encodes the OBP3-RESPONSIVE GENE 3 transcription factor acting as a negative regulator of cell proliferation. To the contrary, this study revealed that the same grf9-1 single mutant as used in the previous study was not bigger than the wild type and that GRF9 acted as a positive regulator of cell proliferation (Figs. 2 and 3). Horiguchi et al. (2005) observed a slight increase in leaf size of the grf9-1 mutant, though not supported by statistics, and Liang et al. (2014) found no contribution of the grf9-1 mutation to leaf growth at the level of the grf7/8/9 triple mutations.
The GRF–GIF–miR396 Module May Play a Pivotal Role in Shaping the Behavioral Features of the Arrest Front
Leaf growth results from coordinated processes of cell proliferation and expansion, which is spatiotemporally orchestrated through the activities of growth-promoting and growth-repressing regulators (Sarvepalli et al. 2019). Some of those regulators are expressed in gradients as a result of their own transcriptional activities and/or post-transcriptional control by their upstream regulators, such as miRs. Therefore, those regulatory players underpin the spatio-temporal modulation of cellular properties and behaviors, resulting in a growth gradient. The basipetal gradient, in which the arrest front progresses in a tip-to-base direction, is commonly found in most monocot and eudicot plants, although other types of growth gradient have been defined for other eudicot plants (Das Gupta et al. 2015; Nelissen et al. 2016). Arabidopsis leaves display the basipetal growth gradient (Sarvepalli et al. 2019). Initially, cell proliferation vigorously occurs throughout the emerging leaf primordia in an exponential manner, which is revealed by the CYCB1;1::GUS marker and kinematic analyses of leaf growth (Donnelly et al. 1999; Autran et al. 2002; Ferjani et al. 2007; Lee et al. 2009). As the primordia grow further, the meristematic activities started to disappear from their tips, forming the arrest front (Donnelly et al. 1999; Kazama et al. 2010). Cells residing in the distal region of the arrest front stop to divide and start to expand vigorously, whereas cells in the proximal region continue to divide actively. When the arrest front touches the base, almost all cells in the leaf blade, except for vascular and meristemoid cells, shut down its meristematic program, giving way to the differentiation program and leading to the exponential expansion of cell volume and leaf size. Similarly, the expressions of most GRFs were strong throughout the emerging primordia, then started to disappear in a tip-to-base direction, and were almost completely shut down at 9 DAG (Fig. 4), indicating that GRF expression displays a basipetal gradient. On the other hand, miR396 levels displayed an acropetal gradient, i.e., high in the distal region and low in the proximal region (Rodriguez et al. 2010; Beltramino et al. 2018). Furthermore, both grf multiple mutations and 35S::MIR396b not only reduced the number of CYCB1;1::GUS-positive cells in the primordia, but also caused a precocious movement of the CYCB1;1::GUS boundary (Fig. 5; Rodriguez et al. 2010). The results indicate that the GRF–miR396 module regulates both the rate and duration of cell cycling and also suggest that the module may be involved in regulation of the formation and/or maintenance of the arrest front. The notion is supported by the fact that overexpression of GRF5 significantly delayed the downward movement of the arrest front (Vercruyssen et al. 2014). A previous study proposed that the repression of GRF expression by miR396 may contribute to the behavior of the arrest front (Rodriguez et al. 2010). The present study provides further evidence, supporting the proposal. In addition, the expression gradients of the GRF-miR396 module during leaf growth seem to be conserved in other species with the basipetal growth pattern, highlighting the importance of the module with respect to the behavior of the arrest front (Das Gupta et al. 2015).
The notion may hold true for the GIF family, since the gif1 mutation reduced the number of CYCB1;1::GUS-positive cells in the primordia and caused a precocious movement of the CYCB1;1::GUS boundary (Fig. 5). Expression patterns of GIFs closely overlapped the behavior of the arrest front (Horiguchi et al. 2005; Lee and Kim, 2014; Kawade et al. 2017). Kinematic analyses on leaf growth of gif 1/2/3 and an3 mutants revealed that GIFs regulate both the rate and duration of cell cycling (Ferjani et al. 2007; Lee et al. 2009). Activation of GIF1/AN3 delayed the downward movement of the arrest front (Vercruyssen et al. 2014). Taken together, we propose that the GRF–GIF–miR396 module may play a pivotal role in shaping the behavioral features of the arrest front. Kazama et al (2010) hypothesized a KLUH-derived mobile growth factor to explain the behavior of the arrest front. It is therefore tempting to speculate that the GRF–GIF–miR396 module may mediate the action of the putative mobile growth factor. Recently, it was shown that GIF1/AN3 proteins diffused through the plasmodesmata and established a gradient from a base-to-tip direction, suggesting that it might act as a mobile growth factor (Kawade et al. 2017).
Of particular note is the finding that grf, gif1, and 35S::MIR396b mutants did not develop the tertiary and quaternary veins, and gif1 35S::MIR396b did not develop even the secondary veins (Fig. 5). It was reported that the cotyledons of the grf1/2/3/4 quadruple mutant also had malformed veins (Kim and Lee 2006). The results are in good agreement with the observation that most of GRFs are expressed in leaf veins (Fig. 4). The leaves of the gif1/2/3 triple mutant developed only the primary veins, indistinguishable from that of gif1 35S::MIR396 (data not shown). The results indicate that the GRF–GIF–miR396 module plays an essential role in the vasculature development, inviting more systematic and detailed analyses to understand how the module regulates cellular and molecular processes involved in the vasculature development.
Materials and Methods
Plant Material and Growth Conditions
The Arabidopsis thaliana seeds were sown on autoclaved wet soil (Mix5, Sunshine, USA), stratified at 4℃ for 3 days, and transferred to a growth room at 23℃ under a photoperiod of 16-h light/8-h darkness, which was marked as 0 DAG. Soil was periodically supplemented with nutrients (Bio Garden, E&G, Korea). 35S::MIR396b and GRF2pro:: GRF2::GUS seeds were kind gifts from Dr. Palatnik, and CYCB1;1::GUS seeds from Dr. Celenza.
Construction of Multiple Mutants
The construction process of multiple mutants was previously described (Lee et al. 2018), and some additional multiple mutants were prepared in this study by crosses between preexisting multiple mutants.
Construction of P GRF:: GRF::GUS
Most of PGRF::GRF::GUS constructs were previously described (Lee et al. 2018). PGRF8::GRF8::GUS was constructed in this study using the In-Fusion Advantage PCR cloning kit according to the manufacturer’s instruction (Takara Bio, USA). The primers used for amplification of the GRF8 genomic fragment, including the ~ 2 kb-long promoter and 5’-untranslated region (UTR) as well as exons and introns, but except the stop codon and 3’ UTR, were JHK660 and JHK663 (5ʹ-GCAGGTCGACTCTATAATGCGAGGCTGAAGGTGT-3ʹ and 5ʹ-GACCACCCGGGGATCC TGTGTAGCTTGAGCTTCT-3ʹ, respectively). In-Fusion enzymes joined the PCR products and pBI101.1 linearized with HindIII and BamHI to be in frame with GUS. These recombinant plasmids were confirmed by sequencing and introduced into Arabidopsis plants by the Agrobacterium tumefaciens-mediated transformation method (Clough and Bent 1998). Dozens of independent transgenic lines were selected on MS agar plates (0.5 × Murashige-Skoog salts, 1% sucrose, 0.8% phytoagar, 50 μg/ml kanamycin), and T2 lines were subjected to the GUS assay.
GUS Assay
The GUS staining procedure was performed according to Rodrigues-Pousada et al. (1993) with a slight modification, and photographs were obtained using a light microscope (Eclipse NI-U, Nikon, Japan).
Measurement of Dimensional Parameters of Leaves
Digital images of detached leaves were acquired using a scanner. Area, length, and width of the first pair of leaves were determined with the image-analyzing program SCIONIMAGE (Scion Corp.).
Number and Size of Palisade Cells
The first pair of leaves were fixed with ethanol/acetic acid (6:1) for 4 h, and were washed with 100% ethanol three times and 70% ethanol once, after which they were cleared in the Visikol solution (USA) and mounted on slide glass. The microscopic images were obtained by using DIC microscope (Eclipse NI-U, Nikon, Japan). The cell numbers were determined by counting the palisade cells aligned along a transverse axis in the maximum-width region of leaves (Fig. 1) or derived from leaf area divided by cell area (Fig. 3). To determine cell area, ten cells grouped in the halfway from the midvein to the leaf margin at the widest point were analyzed with the SCIONIMAGE software.
References
Autran D, Jonak C, Belcram K, Beemster GTS, Kronenberger J, Grandjean O et al (2002) Cell numbers and leaf development in Arabidopsis: a functional analysis of the STRUWWELPETER gene. EMBO J 21:6036–6049
Beltramino M, Ercoli MF, Debernardi JM, Goldy C, Rojas AML, Nota F et al (2018) Robust increase of leaf size by Arabidopsis thaliana GRF3-like transcription factors under different growth conditions. Sci Rep 8:13447
Beltramino M, Debernardi JM, Ferela A, Palatnik JF (2021) ARF2 represses expression of plant GRF transcription factors in a complementary mechanism to microRNA miR396. Plant Physiol 185:1798–1812
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Das Gupta M, Nath U (2015) Divergence in patterns of leaf growth polarity is associated with the expression divergence of miR396. Plant Cell 27:2785–2799
Debernardi JM, Mecchia MA, Vercruyssen L, Smaczniak C, Kaufmann K, Inzé D et al (2014) Post-transcriptional control of GRF transcription factors by microRNA miR396 and GIF co-activator affects leaf size and longevity. Plant J 79:413–426
Donnelly PM, Bonetta D, Tsukaya H, Dengler RE, Dengler NG (1999) Cell cycling and cell enlargement in developing leaves of Arabidopsis. Dev Biol 215:407–419
Duan P, Ni S, Wang J, Zhang B, Xu R, Wang Y et al (2016) Regulation of OsGRF4 by OsmiR396 controls grain size and yield in rice. Nature Plants 2:1
Ferjani A, Yano S, Horiguchi G, Tsukaya H (2007) Analysis of leaf development in fugu mutants of Arabidopsis reveals three compensation modes that modulate cell expansion in determinate organs. Plant Physiol 144:988–999
Fonini LS, Lazzarotto F, Barros PM, Cabreira-Cagliari C, Martins MAB, Saibo NJM et al (2020) Molecular evolution and diversification of the GRF transcription factor family. Genet Mol Biol 43:e20200080
Horiguchi G, Kim G, Tsukaya H (2005) The transcription factor AtGRF5 and the transcription coactivator AN3 regulate cell proliferation in leaf primordia of Arabidopsis thaliana. Plant J 43:68–78
Kawade K, Tanimoto H, Horiguchi G, Tsukaya H (2017) Spatially different tissue-scale diffusivity shapes ANGUSTIFOLIA3 gradient in growing leaves. Biophysical J 113:1109–1120
Kazama T, Ichihashi Y, Murata S, Tsukaya H (2010) The mechanism of cell cycle arrest front progression explained by a KLUH/CYP78A5-dependent mobile growth factor in developing leaves of Arabidopsis thaliana. Plant Cell Physiol 51:1046–1054
Kim JH (2019) Biological roles and an evolutionary sketch of the GRF–GIF transcriptional complex in plants. BMB Rep 52:227–238
Kim JH, Kende H (2004) A transcriptional coactivator, AtGIF1, is involved in regulating leaf growth and morphology in Arabidopsis. Proc Natl Acad Sci USA 101:13374–13379
Kim JH, Lee BH (2006) GROWTH-REGULATING FACTOR4 of Arabidopsis thaliana is required for development of leaves, cotyledons, and shoot apical meristem. J Plant Biol 49:463–468
Kim JH, Tsukaya H (2015) Regulation of plant growth and development by the growth-regulating factor and GRF-interacting factor duo. J Exp Bot 66:6093–6107
Kim JH, Choi D, Kende H (2003) The AtGRF family of putative transcription factors is involved in leaf and cotyledon growth in Arabidopsis. Plant J 36:94–104
Kim JS, Mizoi J, Kidokoro S, Maruyama K, Nakajima J, Nakashima K et al (2012) Arabidopsis GROWTH-REGULATING FACTOR7 functions as a transcriptional repressor of abscisic acid- and osmotic stress-responsive genes, including DREB2A. Plant Cell 24:3393–3405
Lee BH, Kim JH (2014) Spatio-temporal distribution patterns of GRF-INTERACTING FACTOR expression and leaf size control. Plant Signal Behav 9:e29697
Lee BH, Ko JH, Lee S, Lee Y, Pak JH, Kim JH (2009) The Arabidopsis GRF-INTERACTING FACTOR gene family performs an overlapping function in determining organ size as well as multiple developmental properties. Plant Physiol 151:655–668
Lee SJ, Lee BH, Jung JH, Park SK, Song JT, Kim JH (2018) GROWTH-REGULATING FACTOR and GRF-INTERACTING FACTOR specify meristematic cells of gynoecia and anthers. Plant Physiol 176:717–729
Liang G, He H, Li Y, Wang F, Yu D (2014) Molecular mechanism of microRNA396 mediating pistil development in Arabidopsis. Plant Physiol 164:249–258
Liebsch D, Palatnik J (2020) MicroRNA miR396, GRF transcription factors and GIF co-regulators: a conserved plant growth regulatory module with potential for breeding and biotechnology. Curr Opin Plant Biol 53:31–42
Liu D, Song Y, Chen Z, Yu D (2009) Ectopic expression of miR396 suppresses GRF target gene expression and alters leaf growth in Arabidopsis. Physiol Plant 136:223–236
Liu H, Guo S, Xu Y, Li C, Zhang Z, Zhang D et al (2014) OsmiR396d-regulated OsGRFs function in floral organogenesis in rice through binding to their targets OsJMJ706 and OsCR4. Plant Physiol 165:160–174
Nelissen H, Gonzalez N, Inze´ D, (2016) Leaf growth in dicots and monocots: So different yet so alike. Curr Opin Plant Biol 33:72–76
Omidbakhshfard MA, Fujikura U, Olas JJ, Xue GP, Balazadeh S, Mueller-Roeber B (2018) GROWTH-REGULATING FACTOR 9 negatively regulates Arabidopsis leaf growth by controlling ORG3 and restricting cell proliferation in leaf primordia. PLoS Genet 14:e1007484
Omidbakhshfard MA, Proost S, Fujikura U, Mueller-Roeber B (2015) Growth-Regulating Factors (GRFs): A small transcription factor family with important functions in plant biology. Mol Plant 8:998–1010
Rodrigues-Pousada RA, De Rycke R, Dedonder A, Van Caeneghem W, Engler G, Van Montagu M et al (1993) The Arabidopsis 1-aminocyclopropane-1-carboxylate synthase gene 1 is expressed during early development. Plant Cell 5:897–911
Rodriguez RE, Mecchia MA, Debernardi JM, Schommer C, Weigel D, Palatnik JF (2010) Control of cell proliferation in Arabidopsis thaliana by microRNA miR396. Development 137:103–112
Sarvepalli K, Gupta MD, Challa KR, Nath U (2019) Molecular cartography of leaf development—role of transcription factors. Curr Opin Plant Biol 47:22–31
Tsukaya H (2002) Interpretation of mutants in leaf morphology: genetic evidence for a compensatory system in leaf morphogenesis that provides a new link between cell and organismal theory. Int Rev Cytol 217:1–39
Tsukaya H (2013) Leaf Development the Arabidopsis Book 11:e0163
Tsukaya H (2021) The leaf meristem enigma: the relationship between the plate meristem and the marginal meristem. Plant Cell 33:3194–3206
Van der Knaap E, Kim JH, Kende H (2000) A novel gibberellin-induced gene from rice and its potential regulatory role in stem growth. Plant Physiol 122:695–704
Vercruysse J, Baekelandt A, Gozalez N, Inzé D (2020) Molecular networks regulating cell division during Arabidopsis leaf growth. J Exp Bot 71:2365–2378
Vercruyssen L, Verkest A, Gonzalez N, Heyndrickx KS, Eeckhout D, Han SK et al (2014) ANGUSTIFOLIA3 binds to SWI/SNF chromatin remodeling complexes to regulate transcription during Arabidopsis leaf development. Plant Cell 26:210–229
Zhang D, Sun W, Singh R, Zheng Y, Cao Z, Li M et al (2018) GRF-interacting factor1 regulates shoot architecture and meristem determinacy in maize. Plant Cell 30:360–374
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF-2018R1D1A1B07050016). We thank Dr. Palatnik for 35S::MIR396b and GRF2pro:: GRF2::GUS seeds; Dr. Celenza for CYCB1;1::GUS seeds; and the Arabidopsis Biological Resource Center for grf mutant seeds.
Author information
Authors and Affiliations
Contributions
BHL and JHK conceived and designed the research plan. BHL, J-HJ, S-JL, and T-TM contributed to the construction of grf multiple mutants and GRF::GUS transgenic lines. G-HL and BHL prepared multiple mutants containing CYCB1;1::GUS lines and performed most of the experiments. BHL and JHK wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no conflict of interest.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lee, GH., Lee, B.H., Jung, JH. et al. Systematic Assessment of the Positive Role of Arabidopsis thaliana GROWTH-REGULATING FACTORs in Regulation of Cell Proliferation During Leaf Growth. J. Plant Biol. 65, 413–422 (2022). https://doi.org/10.1007/s12374-022-09366-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12374-022-09366-1