Abstract
To explore the relationship between fruit developmental periods and the protein datasets of Alpinia oxyphylla, a label-free quantitative proteomic profiling analysis of the A. oxyphylla fruits sampled at the Earlystage, Metaphase and Advanced stage was carried out using the liquid chromatography combined with tandem mass spectrometry. A total of 19,219 peptide fragments and 4946 quantitative proteins were obtained. Annotation and enrichment analysis of these peptides and proteins were performed using the Gene Ontology (GO) and the Kyoto Encyclopedia of Genes and Genomes (KEGG) databases with bioinformatics software. The results showed that the differentially expressed proteins were mainly involved in metabolic and cellular processes, cellular component organization or biogenesis and response to the stimulus. Three significantly enriched metabolic pathways were non-alcoholic fatty liver disease, vibrio cholera infection and valine, leucine and isoleucine degradation. The significantly enriched proteins were NADH dehydrogenase, putative vacuolar proton translocation ATPase and putative acyltransferase component. The proteomic profiles of the fruit samples from the Advanced and the Metaphase stages differed little, while the difference in proteomic components between the Earlystage and the Advanced groups was significant. This study should lay the theoretical foundation for the effective utilization of A. oxyphylla germplasm resource in the treatment of human diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alpinia oxyphylla, one of the famous traditional Chinese medicines and healthy food is mainly distributed in South China, especially in Hainan province, its resources are very rich. The fruit of A. oxyphylla called Yizhi, is widely used to treat abdominal pain, dyspepsia, poor memory, diarrhea and kidney asthenia (Pharmacopoeia Committee 2005; Tewari et al. 1999, Kubo et al. 1995). Many studies have confirmed that Yizhi possesses significant pharmacological roles, the protective effect of ethanol extract from Yizhi on glutamate-induced neuronal apoptosis was examined in primary cultured mouse cortical neurons (Yu et al. 2003), and of these extractions, the n-hexane and ethyl acetate fractions showed anti-angiogenic potentials in both in vivo and in vitro models of zebrafish (He et al. 2010). Some researchers reported that sesquiterpenes from A. oxyphylla essential oils could effectively promote drug absorption across and/or into the skin (Fang et al. 2003). Recently, a simple and rapid ultra-performance liquid chromatography-tandem mass spectrometry (UPLC-MS/MS) coupled with a one-step liquid–liquid extraction method was developed and validated for the simultaneous determination of two flavonoids (chrysin and tectochrysin) from Yizhi extract in rat plasma (Zhao et al. 2018), and the underlying mechanisms have attracted the attention of scientists (Yan et al. 2016; Shi et al. 2015; Liu et al. 2014). It should be noted that the ethnopharmacological uses, phytochemistry, biological activities, and therapeutic applications of A. oxyphylla have just been fully reviewed (Zhang et al. 2018). Overall, the traditional usage of A. oxyphylla fruits as folk medicine has been confirmed to be both effective (scientifically validated) and efficient (feasible and useful in clinical practice) by modern pharmacology researches.
Although impressive progress have been achieved in the isolation and purification of bioactive components from a native plant or fruit of A. oxyphylla (Xu et al. 2012a, b; An et al. 2006), and the trial and validation of the Yizhi’s potential applications in a variety of experimental animal models (Zhang et al. 2012, 2013), some novel pharmacological actions are worth excavating and the molecular mechanisms of clinical efficacies, such as anti-cancer and neuroprotective effects remain unclear (Lin et al. 2013; Wang et al. 2013). More in vivo, in vitro and clinical studies to confirm the mode of action are strongly needed. The development of whole fructus A. oxyphylla or the combination of several effective components from Yizhi to exert neuroprotective effects has become a hot research topic. Nowadays, the multiple-omics analysis approaches began to be used in the pharmacological study of A. oxyphylla fruit and showed a great application prospect.
Comparative proteomics analysis of Yizhi under different conditions may provide a novel therapeutic strategy for human diseases. The complete chloroplast genome of A. oxyphylla as an important edible and traditional herbal medicine has just been sequenced, analyzed, and compared to five species in the Zingiberaceae family (Gao et al. 2019). Several years ago, a novel neuroprotective natural product, oxyphylla A [(R)-4-(2-hydroxy-5-methylphenyl)-5-methylhexanoic acid] from the fruit of Alpinia oxyphylla for use in Parkinson’s disease was discovered and made functional characterization through LC/MS-based Multivariate Data Analysis-Guided Fractionation (Li et al. 2016a, b). Researchers integrated metabolomics, chemometrics, and pharmacological strategy led to the efficient discovery of novel bioactive ingredients from A. oxyphylla while avoiding the non-targeting, labor-intensive steps usually required for the identification of bioactive compounds. Moreover, the successful development of a synthetic route toward oxyphylla A should lead to its availability on a large scale for further functional development and pathological studies. To explore the underlying pharmacological mechanisms of A. oxyphylla fruits i.e., Yizhi at the molecular level, in this study, mass spectrometry-based label-free quantitative proteomics of Yizhi samples at three developmental stages was performed.
Materials and Methods
Materials and Sampling Time
Alpinia oxyphylla fruits were obtained from the central part of Baisha City, Hainan province. The entire fruit development stages were defined as pre-mature (1st to 20th of April), early mature (late April to early May), and late mature (middle-to-late May). The healthy fruits at these development stages were randomly harvested from the same plant and frozen in liquid nitrogen for subsequent protein work. In this study, a total of nine fruit samples of Alpinia oxyphylla with three duplications in three developmental periods were subjected to the label-free quantitative proteomics analysis. These fruit samples were designated as the Earlystage-1 to Earlystage-3, the Metaphase-1 to Metaphase 3, and the Advanced-1 to Advanced-3 in the experiment.
Extraction of Proteins and Enzymatic Hydrolysis of Peptide Fractions
All experimental steps were conducted at 4 °C. The proteins of Alpinia oxyphylla were extracted from the frozen tissue powders milled in liquid nitrogen by using the SDT lysis buffer (4% SDS (w/v), 100 mM Tris-HCl, pH 7.6, 0.1 M DTT), and the protein concentration was quantified by the bicinchoninic acid (BCA) method. Appropriate amounts of proteins in each sample were subjected to the trypsin enzymatic hydrolysis by using the filter-aided sample preparation (FASP) method (Wiśniewski et al. 2009). All the peptide samples were desalted onto a Sep-Pak C18 cartridge (Waters Associates Inc., Milford, MA, 01757 USA), freeze-dried, and re-dissolved with 40 μl of 0.1% formic acid solution. The peptide concentration was determined using spectrophotometry (OD, 280 nm) (Unicam SP500 series 2).
LC–MS/MS Analysis
All peptide samples were separated on a nanoflow HPLC instrument (Easy-nLC 1000, Thermo Fisher Scientific). HPLC parameters include the mobile phase A (0.1% (v/v) of formic acid in water), and the mobile phase B (84% (v/v) of acetonitrile and 0.1% (v/v) of formic acid in water). The chromatographic column was firstly equilibrated with 95% of the HPLC buffer A, i.e. 0.1% (v/v) of formic acid in the water, and subsequently, technical replicates (3 × ∼2.5 μg) of each sample were loaded onto an Acclaim PepMap100 nanoViper C18 trap column (100 μm inner diameter, 2 cm length; Thermo Scientific) by an automated flow injection system, and separated with a constant flow of 300 nL/min onto an analytical EASY column (75 μm × 10 cm, C18-A2, 3 μm, Thermo Scientific).
A Q-Exactive mass spectrometer was used in positive ion mode to detect the peptide fractions after the chromatographic separation. The mass spectrometric analysis was performed as follows: Parent ion scanning with a range of 300–1800 m/z, an Orbitrap resolution of 70,000 (at m/z 200), a target automatic gain control (AGC) value of 1e6, maximum IT of 50 ms and dynamic exclusion of 60.0 s.
The mass-to-charge ratio of peptide and polypeptide fragments was collected as follows: 20 fragment maps [MS2 scan, higher-energy collisional dissociation (HCD)] were obtained after each full scan. The precursor ion of each targeted peptide was isolated using a 2 m/z unit window. Fragmentation was performed with a normalized collision energy of 30 eV, and the secondary mass spectrometry resolution was 17,500 (at m/z 200).
Identification and Quantitation of Proteins
The original RAW file obtained by LC–MS/MS was imported into MaxQuant software (version 1.5.3.17) for database searching. The main library parameters were set as in Table 1. Protein quantification was carried out via MaxQuant’s label-free quantification (LFQ) algorithm which combines and adjusts peptide intensities into a protein intensity value (Cox et al. 2014). Results were filtered to a 1% FDR at the levels of protein and peptide with MaxQuant (Cox and Mann 2008).
Bioinformatic Analysis
Bioinformatic analysis of proteomics data from the different developmental stages of the Alpinia oxyphylla fruits was conducted. The functional annotation of proteins was performed using the Blast2GO database program (https://geneontology.org) (Götz et al. 2008), while the Kyoto Encyclopedia of Genes and Genomes (KEGG) database (https://www.genome.jp/kegg) combined with the software KASS (KEGG Annotation Server) were used to the identification and classification of the detected proteins. The Fisher’s exact test was used to analyze the statistical significance of each pathway and certain GO term protein enrichments, and the target proteins were eventually subjected to the analyses of GO term category and KEGG pathway (Kanehisa et al. 2012; Ashburner et al. 2000).
Clustering Analysis of the Target Protein Datasets
A normalization of quantitative information of the target protein datasets was firstly processed, and the values were limited within the interval of − 1 to 1. Subsequently, the Complexheatmap R package (R version 3.4) was used for the clustering of differentially expressed proteins in Alpinia oxyphylla fruits, and the hierarchically clustered heat maps were generated based on the Euclidean distance and Average Linkage Clustering (ALC) algorithm.
Results and Discussion
Accuracy and Precision of the Identification of Peptide Fragments
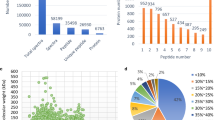
To ensure the good mass deviation and yield the high-quality MS1 and MS2 data in LC-MS/MS assay, a high-performance benchtop quadrupole Orbitrap mass spectrometer was used for the mass spectrometry-based proteomics analysis in this study. The result showed that the mass error distribution of all the identified peptide fragments was under 10 ppm (Supplementry Fig. 1a), and the Andromeda score distribution along with the number of identified peptides was fully revealed by the peptide search engine Andromeda that integrated into the MaxQuant environment (Cox et al. 2011). Overall, a total of 19,219 peptide fragments were identified in samples by MS analysis, and the median Andromeda identification score was 86.94, with about 84.09% over a score of 60, suggesting a high quality of the experimental data (Supplementry Fig. 1b).
Molecular weights of the most identified proteins ranged from 10 to 50 kDa, and low molecular weight proteins (< 10 kDa) accounted for about 10%. Overall, proteins with molecular weights less than 50 kDa accounted for nearly 80% of the total proteins, indicating a narrow molecular weight distribution of proteins (Supplementry Fig. 1c). Peptides of 7–19 amino acids in length were dominant in the identified peptides (Supplementry Fig. 1d), and the count of proteins gradually decreased along with the increment of the percentage of peptide coverage (Supplementry Fig. 1e). Finally, the peptide count distribution was shown in supplementry Fig. 1f. All the results indicated that the identification of peptide fragments was accurate and credible.
Identification and Comparison of Protein Datasets
A total of 4946 proteins were successfully identified in the fruit samples of Alpinia oxyphylla by mass spectrometry, and 3917 core proteins were found in all samples. There were 134, 179 and 22 specific proteins detected in the different development stages of Alpinia oxyphylla fruit, respectively (Fig. 1). The proteins whose expression changes more than 2 times or less than 0.5 times (P < 0.05) are recorded as differentially expressed proteins in this study. The comparisons of protein profiles were conducted between fruit samples at different developmental stages (Table 2). The results revealed that with the ripening of the fruit, the types and abundance of protein in the fruit greatly varied compared to that of the early stage. The general trend was that the expression of more and more proteins was significantly reduced.
In group comparisons, the quantitative results of proteins were shown in volcano plots by analyzing the fold changes of differential protein expression combined with the corresponding p values. Comparisons between groups of Metaphase and Earlystage, Advanced and Earlystage, Advanced and Metaphase were performed (Fig. 2a–c). For each comparison, all the significantly differential expression proteins were subjected to clustering analysis and the results were eventually exhibited in heat maps (Fig. 3a–c).
GO and KEGG Analyses of Differentially Expressed Proteins
The Gene Ontology (GO) functional analysis of proteins was quantified for fruit samples of Alpinia oxyphylla considering three aspects: biological processes, molecular functions and cell components (Ashburner et al. 2000). Fisher’s exact test which is based on protein counts was used for enrichment analysis, and the top 20 significant GO terms identified between sample groups and the corresponding P values were shown in Fig. 4. The results revealed that a significant enrichment of cell components mainly concerns intracellular membrane-bounded organelle and catalytic complex. The affected molecular functions are mainly composed of SNAP receptor activity and nucleocytoplasmic transporter activity. Finally, significant enrichment of the biological processes mainly affects the positive regulation of macromolecule and cellular biosynthetic/metabolic process, nitrogen compound metabolic process and the ribonucleoprotein complex localization.
Gene ontology (GO) analysis results for significant protein enrichment between all A. oxyphylla fruit groups. The picture displays the Top 20 most significantly enriched entries in the three categories, biological processes (BP), molecular function (MF) and cell component (CC) (p < 0.05, one-way ANOVA)
A comparison of the differential proteins between groups of the Advanced and the Earlystage by GO term enrichment analysis revealed that the significant differences mainly include the organophosphate and carbohydrate derivative biosynthetic processes (Biological process category), the nutrient reservoir activity (Molecular function category) and the small ribosomal subunit (Cell component category) (Fig. 5a). Similarly, the differential protein analyses based on the GO enrichment results between groups of the Advanced vs the Metaphase, and the Metaphase vs the Earlystage were performed too (Fig. 5b, c).
Gene ontology (GO) analysis results for significant protein enrichment between two A. oxyphylla fruit groups. The picture shows the most significantly enriched entries in the three categories, biological processes (BP), molecular function (MF) and cell component (CC) (p < 0.05, one-way ANOVA). a The Advanced group vs the Earlystage group; b the Advanced group vs the Metaphase group; c the Metaphase group vs the Earlystage group
The liquid chromatography/mass spectrometry has been successfully applied to identify and quantify the principal compounds from A. oxyphylla fruit in recent years (Li et al. 2016a, b; Sun et al. 2016). New natural products with promising pharmacological functions are constantly isolated and characterized (Li et al. 2016a, b; Hou et al. 2015; Lv et al. 2011). Some researchers investigated the accumulation profiles of multiple components between pericarp and seed of A. oxyphylla capsular fruits gathered from different production regions and found that nootkatone is predominantly distributed in the seeds and the flavonoids and diarylheptanoids are mainly deposited in the capsules. Moreover, the content levels of the tested secondary metabolites occurring in the capsules varied greatly among different production regions (Chen et al. 2014). Other scientists reported that there was a large variation in the contents of 10 common nucleobases and nucleosides among the fruits of A. oxyphylla collected from different cultivation regions, and especially they found some samples from the same growing region exhibited quite different content levels of nucleobases and nucleosides, which implies that besides the factor of geography, harvest time may also affect the contents of these components (Song et al. 2014). In general, the effect of harvest time on the contents of bioactive compounds and the chemical fingerprint profile in fruits of A. oxyphylla has been paid more and more attention in recent years (Miao et al. 2015; Li et al. 2013a, b). However, the effect of different harvest times on protein expression profiles in fruits of A. oxyphylla has not been reported. In this study, the differences in proteomic profiles of the A. oxyphylla fruits harvested at different developmental stages were detected by a label-free quantitative proteomic profiling analysis. Considering the fact that harvesting time has a serious impact on the development and maturity of fruit and seed, it is not difficult for us to understand the obvious changes in protein compositions and contents of the A. oxyphylla fruits among sample groups. In this study, the quantitative determination of proteomic profiles of the A. oxyphylla fruits harvested at three developmental stages was conducted. The significantly differential proteins between all sample groups are enriched into the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways. The KEGG classification results indicated that the differential expressed proteins are mainly concentrated in the processes of oxidative phosphorylation, synaptic vesicle cycle and lysosome (Fig. 6). Some studies have shown that the extracts of A. oxyphylla fruit (AOF) can stimulate or participate in the process of oxidative phosphorylation of cells. For example, the ethanol crude extracts of A. oxyphylla fruit, especially its ethyl acetate fraction (EF) and petroleum ether-soluble fraction (PE) have been confirmed to possess potent antioxidant activities (Bian et al. 2013; Wang et al. 2013). Previous investigation showed that fractions of the petroleum ether (PE) extract layer of AOF led to a decrease in phosphorylation of anti-apoptotic kinases ERK and an increase in phosphorylation of pro-apoptotic kinase JNK and p38. HPLC analysis pointed out five main components (9-hydroxynootkatone, oxyphyllacinol, yakuchinone A, yakuchinone B and tectochrysin) which may contribute to the anti-proliferative activity of PE (Zhang et al. 2015). It was found that the AOF extract significantly triggers the phosphorylated insulin-like growth factor-1 receptor-phosphatidylinositol 3-kinase/serine-threonine kinase pathway in RSC96 Schwann cells and up-regulates the cell cycle regulatory proteins cyclin A, cyclin D1 and cyclin E (Chang et al. 2017). Recent studies have suggested that the dysfunction of synaptic vesicle recycling at presynaptic terminals may contribute to the onset of Parkinson’s disease (PD) (Chapman 2014; Esposito et al. 2011). The anti-Parkinsonian activities of the AOF extracts such as protocatechuic acid and oxyphylla A were tested on PD models in vitro and in vivo (Li et al. 2016a, b; Li et al. 2013a, b). Similarly, pharmacological mechanisms underlying the neuroprotective effects of terpene, a major compound of A. oxyphylla extract on Alzheimer’s disease have also been explored (Xu et al. 2020). Many findings have indicated that clearance of α-synuclein by the autophagy-lysosome pathway (ALP) plays a key role in many neurodegenerative conditions (Batelli et al. 2011; Ebrahimi-Fakhari et al. 2011; Lee et al. 2004). A. oxyphylla contains diverse bioactive constituents and has various pharmacological activities including neuroprotective, anticancer, anti-oxidant properties and so on (Zhang et al. 2018; Li et al. 2013a, b). According to the data we have, the active substances that have been found are all chemical compounds, but there is no report that the potential pharmacology function has been explored from the protein level, comprehensively analyzing the changes of protein abundance and variety in fruits of A. oxyphylla with harvest time. The reason for this situation may be that the active substances in the fruit of A. oxyphylla are generally extracted with organic solvents, and the solubility of proteins in those organic phases is relatively low, which is easy to precipitate or denature, so the proteins cannot be detected by the conventional extraction test. Of these enriched KEGG pathways, the phenylpropanoid biosynthesis is related to plant defense reactions via preformed or inducible physical and chemical barriers against infection to signal molecules that involved in local and systemic signaling for defense gene induction (Taheri and Tarighi 2010; Vogt 2010; Dixon et al. 2002). The phenylpropanoid pathway is required for the biosynthesis of lignin and other important compounds such as flavonoids, coumarins and lignans in plants. The polyphenols including lignin and flavonoids can effectively prevent the invasion of pathogens and furthermore participate in the plant-pathogen interaction (Zabala et al. 2006).
Protein–Protein Interaction (PPI) Analysis
The most significantly enriched proteins with high connectivity and the significant enrichment of the metabolic pathways between all sample groups were obtained by PPI analysis. In the protein–protein interaction (PPI) network map (Fig. 7), the three metabolic pathways that were significantly enriched were Non-Alcoholic Fatty Liver Disease (NAFLD), Vibrio cholera infection and valine, leucine, and isoleucine degradation. A. oxyphylla extract (AOE) prevents diabetes in mice by modulating gut microbiota and changing the expression of specific miRNAs involved in biologically significant signaling pathways (Xie et al. 2018; Sang et al. 2017). Consequently, the associated NAFLD is also prevented due to the decrease in free fatty acids delivered to the liver. The A. oxyphylla fruit (AOF) has been used for treating diarrhea and related gastrointestinal disorders for thousands of years in China, and a number of traditional Chinese medicine formulae provide AOF as an alternative herbal treatment for diarrhea (Wang et al. 2015). The results of PPI analysis show that AOF has the potential to treat Vibrio cholera infection, and novel active substances with anti-diarrhea and anti-Vibrio cholera infection properties, including proteins, are expected to be isolated and identified in the forthcoming future. The three branched-chain amino acids (BCAA), Valine, leucine and isoleucine are all essential for human growth (Kindt and Halvorsen 1980). These amino acids affect muscle recovery and the immune system and are closely related to metabolic and physiological functions (Zhang et al., 2017; Calder 2006). Thus, in the present study, the PPI analysis suggests that the degradation of identified amino acids, i.e. valine, leucine and isoleucine may play an important role in the pharmacological effects of A. oxyphylla. The most significantly enriched proteins in these metabolic pathways were the NADH dehydrogenase (unigene 108543_i1, ko04932), putative vacuolar proton translocation ATPase (unigene 101787_i1, ko05110) and putative acyltransferase component (unigene 055317_i1, ko00280). NADH dehydrogenase is an enzyme that catalyzes the transfer of electrons from NADH to coenzyme Q in the inner membrane of mitochondria. It is the entry enzyme of oxidative phosphorylation in mitochondria. The function of a specific electron-transferring-chain containing NADH dehydrogenase, cytochrome reductase and cytochrome oxidase is to link electron transfer with proton translocation out of the mitochondrion. In doing so, they generate a transmembraneous proton motive force which subsequently drives ATP synthesis by the H+-ATPase (Weiss et al. 1991). At present, our laboratory has completed the sequencing of the chloroplast genome of A. oxyphylla, and the results reveal that the phototrophic component of NADH dehydrogenase (ndhB and ndhC), photosystem II (psbZ) and ATP synthase (atpE and atpF) exhibit adaptive evolution under different environments, and the strength of light is an important trigger for the adaptation at the chloroplast level (Gao et al. 2019). Therefore, we speculate that the differences in light strength due to different harvest times may result in the significant variations of NADH dehydrogenase abundance among sampling groups in this study. Analysis of the genetic diversity of natural populations of A. oxyphylla using Inter-Simple Sequence Repeat (ISSR) markers revealed an obvious genetic diversity in natural populations, with much higher genetic variation within populations than between populations. The geographic distribution of samples coincided closely with the distribution of genetic diversity within A. oxyphylla, indicating a weak gene flow (Wang et al. 2012). The results of proteomic profiling analysis in this study with a combination of other researchers’ reports on genetic diversity detection of A. oxyphylla populations may provide valuable guidance regarding the conservation and genetic improvement of this herb.
A total of 25 proteins were found to be significantly differentially expressed in the Metaphase group when compared to that of the Early-stage group. Of these proteins, 8 proteins were up-regulated and 17 proteins were down-regulated. The up-regulated protein designated as unigene066055 (aldehyde dehydrogenase (NAD) activity) was supposed to be involved in the pentose phosphate pathway (PPP) by enrichment analysis. Of the down-regulated proteins, the protein named unigene103767 was considered to be related to spliceosome and systemic lupus erythematosus (SLE), and it should be noted in particular that the expression of a specific protein (gene name: unigene117427, oxidoreductase activity) that involved in the phenylpropanoid biosynthesis was significantly down-regulated in the Metaphase stage. The pentose phosphate pathway (PPP) is required for the synthesis of ribonucleotides and is a major source of NADPH. In addition, PPP plays a pivotal role in helping glycolytic cancer cells to meet their anabolic demands and combat oxidative stress (Patra and Hay 2014). SLE is a prototypic autoimmune disease characterized by the production of antibodies to components of the cell nucleus in association with a diverse array of clinical manifestations (Mok and Lau 2003). To our knowledge, there is no report on the treatment of SLE with A. oxyphylla fruit extract. The identified protein (encoded by the unigene103767) may play a potential role in the treatment of SLE. Phenylpropanoids contribute to all aspects of plant responses towards biotic and abiotic stimuli (Vogt 2010; Dixon et al. 1996). The decrease of its synthesis may indicate that the resistance of A. oxyphylla plants to environmental stress in Metaphase period is lower than that in Earlystage period. In a word, the detailed information of these differential proteins was listed in the supplementary Table S1.
The comparison result of the protein expression levels between the Advanced and the Earlystage groups revealed that there were 75 significantly differential proteins, i.e. 27 up-regulated and 48 down-regulated proteins in this study. Of the down-regulated proteins, the protein (unigene103767, ko05322) was thought to be related to systemic lupus erythematosus (SLE); the protein (unigene056408, ko04210, cysteine-type peptidase activity) was considered to be involved in apoptosis, nucleotide-binding oligomerization domain-like (NOD-like) receptor signaling pathway and renin secretion; the protein (unigene117427, ko00940) was supposed to participate in phenylpropanoid biosynthesis, and the protein (unigene108543, ko05016) was related to Huntington’s disease and non-alcoholic fatty liver disease (NAFLD). Overall, there is no doubt that the protein (unigene103767) in the fruit of A. oxyphylla had the highest expression level in the Earlystage group. Therefore, we speculate that the application of Earlystage-fruit extracts of A. oxyphylla in the treatment of SLE may achieve better therapeutic efficacy. Apoptosis is a controlled process of cell death occurring when cells face irreversible stress (Xu et al. 2012a, b). In the extrinsic apoptosis pathway, components of the death-inducing signaling complex (DISC) including apoptosis-related cysteine peptidases (CASP), CASP8 and CASP10 are activated upon stimuli. Both intrinsic and extrinsic apoptotic pathways converge on the level of CASP3 activation, which in turn cleaves various intra-cellular substrates and cause the morphological changes observed in apoptotic cells (Youle and Strasser 2008). Previous quantitative analysis of global proteome indicated that dispelling the inhibitor of cysteine-type endopeptidase and activated cell apoptosis may be another apoptosis approach in acute myeloid leukemia (AML), which may be used as therapy target of AML (Zhu et al. 2016). In the past decades, many findings have suggested that A. oxyphylla possesses potential chemopreventive and antitumorigenic activities. For instance, suppression of mouse skin tumor promotion and induction of apoptosis in HL-60 cells by A. oxyphylla was reported in 1998 (Lee et al. 1998). NOD-like receptor is a kind of cytoplasmic pattern recognition receptor, which plays an important role in an innate immune response. After being activated, it can induce the release of various inflammatory factors through a series of signaling pathways. The quantitative proteome in suberoylanilide hydroxamic acid (SAHA) and valproic acid (VPA) treated AML HL60 cells was extensively studied, and the results showed that upon SAHA treatment, NOD-like receptor signaling pathway was enriched (Zhu et al. 2016). We think that both cysteine-type peptidase (unigene056408, ko04210) and the protein (unigene 108543, ko05016) can be used as potential biomarkers in the pharmacological study of A. oxyphylla. In the up-regulated proteins, two proteins (unigene103022, ko05418; unigene041900, ko04910) should be noted that they were closely linked to atherosclerosis and insulin signaling pathways, respectively. The ethanol extract of A. oxyphylla fruit, especially the ethyl acetate fraction (EF) was found to possess potent antioxidant and anticancer activities (Wang et al. 2013). Angiotensin II (Ang-II) plays an important role in cardiovascular diseases such as atherosclerosis. It has been demonstrated that AOF significantly inhibits Ang- II induced cardiac pathological remodeling-related pathways in H9c2 cardiomyoblast cells (Tsai et al. 2016; Chang et al. 2013). A. oxyphylla extract prevents diabetes in mice by modulating gut microbiota and changes microRNA expression profiles in db-/db-mouse livers (Xie et al. 2018; Sang et al. 2017). Taken together, we speculate that the two proteins (unigene103022 and unigene041900) identified in this study may play an important role in the regulation of the physiological function of insulin. The supplementary Table S2 shows the detailed information of these differential proteins.
A total of 19 significantly differentially expressed proteins were identified when compared the Advanced group to the Metaphase group, which includes 8 up-regulated and 11 down-regulated proteins. Two specific proteins (i.e. unigene030025, ko05012; unigene114790, ko05168, serine/threonine kinase) were considered to be related to the neurodegenerative diseases such as Parkinson’s disease, Alzheimer’s disease, and Huntington’s disease, and herpes simplex infection, respectively. The serine/threonine kinase mainly phosphorylates the serine or threonine in the downstream signal protein, transmits the extracellular signal into the cell, and then achieves a variety of biological functions by affecting gene transcription. It was reported that treatment with extract of AOF can trigger the phosphorylated insulin-like growth factor-1 receptor-phosphatidylinositol 3-kinase/serine-threonine kinase pathway, and up-regulate the proliferating cell nuclear antigen in a dose-dependent manner. AOF extract activates IGFR-PI3K/Akt signaling to induce Schwann cell proliferation and sciatic nerve regeneration (Chang et al. 2017). The exploration and characterization of novel bioactive substances with neuroprotective properties in AOF will undoubtedly benefit the development of natural therapy for the prevention and treatment of neurodegenerative diseases. The detailed information of these differential proteins was listed in the supplementary Table S3. In general, the proteomic profiles of the fruits from the Advanced and the Metaphase stages differed little, while the difference in proteomic components between the Earlystage and the Advanced groups was relatively significant.
A. oxyphylla has been used as both a food and medicinal substance in China for centuries (Zhao et al. 2016), and it contained diverse active constituents and had various pharmacological activities. Numerous studies have confirmed the efficacy of A. oxyphylla extracts in the treatment of neurodegenerative diseases (Wang et al. 2018; Jiang et al. 2013; Koo et al. 2004), systemic lupus erythematosus (SLE) (Khodaei and Alizadeh 2017), antioxidant (Bian et al. 2013), and anti-cancer (Song et al. 2014). In the present work, we analyzed the proteomic profiles of A. oxyphylla fruits at three developmental stages by LC–MS/MS analysis and successfully identified abundant differentially expressed proteins. Some target proteins can be used for the further validation of pharmacological efficacy and protective effects by animal disease models. The results of this study should definitely facilitate the selection of A. oxyphylla fruit collection period for different human diseases, the isolation and identification of new active constituents in A. oxyphylla fruits, and benefit for a deep understanding of its pharmacological effects.
Change history
06 August 2020
The original article can be found online.
References
An LJ, Guan S, Shi GF, Bao YM, Duan YL, Jiang B (2006) Protocatechuic acid from Alpinia oxyphylla against MPP+-induced neurotoxicity in PC12 cells. Food Chem Toxicol 44(3):436–443
Ashburner M, Ball CA, Blake JA et al (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25(5):25–29
Batelli S, Peverelli E, Rodilossi S, Forloni G, Albani D (2011) Macroautophagy and the proteasome are differently involved in the degradation of alpha-synuclein wild type and mutated A30P in an in vitro inducible model (PC12/TetOn). Neuroscience 195:128–137
Bian QY, Wang SY, Xu LJ, Chan CO, Mok DKW, Chen SB (2013) Two new antioxidant diarylheptanoids from the fruits of Alpinia oxyphylla. J Asian Nat Prod Res 15(10):1094–1099
Calder PC (2006) Branched-chain amino acids and immunity. J Nutr 136(1):288S–293S
Chang YM, Velmurugan BK, Kuo WW, Chen YS, Ho TJ, Tsai CT, Ye CX, Tsai CH, Tsai FJ, Huang CY (2013) Inhibitory effect of alpinate Oxyphyllae fructus extracts on Ang II-induced cardiac pathological remodeling-related pathways in H9c2 cardiomyoblast cells. Biomed 3:148-152
Chang YM, Chang HH, Tsai CC, Lin HJ, Ho TJ, Ye CX, Chiu PL, Chen YS, Chen RJ, Huang CY, Lin CC (2017) Alpinia oxyphylla Miq. Fruit extract activates IGFR-PI3K/Akt signaling to induce Schwann cell proliferation and sciatic nerve regeneration. BMC Complem Altern M 17(1):84
Chapman MA (2014) Interactions between cell adhesion and the synaptic vesicle cycle in Parkinson’s disease. Med Hypotheses 83(2):203–207
Chen F, Li HL, Tan YF, Guan WW, Zhang JQ, Li YH, Zhao YS, Qin ZM (2014) Different accumulation profiles of multiple components between pericarp and seed of Alpinia oxyphylla capsular fruit as determined by UFLC-MS/MS. Molecules 19:4510–4523
Cox J, Mann M (2008) MaxQuant enables high peptide identification rates, individualized p.p.b. range mass accuracies and proteome-wide protein quantification. Nat Biotechnol 26(12):1367–1372
Cox J, Neuhauser N, Michalski A, Scheltema RA, Olsen JV, Mann M (2011) Andromeda: a peptide search engine integrated into the MaxQuant environment. J Proteome Res 10(4):1794–1805
Cox J, Hein MY, Luber CA, Paron I, Nagaraj N, Mann M (2014) Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol Cell Proteomics 13(9):2513–2526
Dixon RA, Lamb CJ, Masoud S, Sewalt VJH, Paiva NL (1996) Metabolic engineering: prospects for crop improvement through the genetic manipulation of phenylpropanoid biosynthesis and defense responses-a review. Gene 179(1):61–71
Dixon RA, Achnine L, Kota P, Liu CJ, Reddy MSS, Wang LJ (2002) The phenylpropanoid pathway and plant defense-a genomics perspective. Mol Plant Pathol 3(5):371–390
Ebrahimi-Fakhari D, Cantuti-Castelvetri I, Fan Z, Rockenstein E, Masliah E, Hyman BT, McLean P, Unni VK (2011) Distinct roles in vivo for the ubiquitin-proteasome system and the autophagy-lysosomal pathway in the degradation of alpha-synuclein. J Neurosci 31:14508–14520
Esposito G, Clara FA, Verstreken P (2011) Synaptic vesicle trafficking and Parkinson’s disease. Dev Neurobiol 72(1):134–144
Fang JY, Leu YL, Hwang TL, Cheng HC, Hung CF (2003) Development of sesquiterpenes from Alpinia oxyphylla as novel skin permeation enhancers. Eur J Pharm Sci 19:253–262
Gao B, Yuan L, Tang T, Hou J, Pan K, Wei N (2019) The complete chloroplast genome sequence of Alpinia oxyphylla Miq. and comparison analysis within the Zingiberaceae family. PLoS ONE 14(6):e0218817
Götz S, García-Gómez JM, Terol J, Williams TD, Nagaraj SH, Nueda MJ, Robles M, Talón M, Dopazo J, Conesa A (2008) High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res 36(10):3420–3435
He ZH, Ge W, Yue GGL, Lau CBS, He MF, But PPH (2010) Anti-angiogenic effects of the fruit of Alpinia oxyphylla. J Ethnopharmaco 132:443–449
Hou L, Ding G, Guo BL, Huang WH, Zhang XJ, Sun ZY, Shi XF (2015) New sesquiterpenoids and a diterpenoid from Alpinia oxyphylla. Molecules 20:1551–1559
Jiang B, Wang WJ, Li MP, Huang XJ, Huang F, Gao H, Sun PH, He MF, Jiang ZJ, Zhang XQ, Ye WC (2013) New eudesmane sesquiterpenes from Alpinia oxyphylla and determination of their inhibitory effects on microglia. Bioorg Med Chem Lett 23(13):3879–3883
Kanehisa M, Goto S, Sato Y, Furumichi M, Tanabe M (2012) KEGG for integration and interpretation of largescale molecular data sets. Nucleic Acids Res 40(D1):D109–D114
Khodaei H, Alizadeh M (2017) Inhibition of IL-4 but not IFN-γ production by splenocytes of mice immunized with ovalbumin after oral administration of 5-hydroxymethylfurfural. Food Agr Immuno 28(1):27–34
Kindt E, Halvorsen S (1980) The need of essential amino acids in children: an evaluation based on the intake of phenylalanine, tyrosine, leucine, isoleucine, and valine in children with phenylketonuria, tyrosine amino transferase defect, and maple syrup urine disease. Am J Clin Nutr 33(2):279–286
Koo BS, Lee WC, Chang YC, Kim CH (2004) Protective effects of Alpinae oxyphyllae fructus (Alpinia oxyphylla MIQ) water-extracts on neurons from ischemic damage and neuronal cell toxicity. Phytotherapy Res 18(2):142–148
Kubo M, Matsuda H, Suo T, Yamanaka J, Sakanaka M, Yoshimura M (1995) Study on Alpiniae Fructus. I. Pharmacological evidence of efficacy of Alpiniae Fructus on ancient herbal literature. Yakugaku Zasshi 115:852–862
Lee E, Park KK, Lee JM, Chun KS, Kang JY, Lee SS, Surh YJ (1998) Suppression of mouse skin tumor promotion and induction of apoptosis in HL-60 cells by Alpinia oxyphylla Miquel (Zingiberaceae). Carcinogenesis 19(8):1377–1381
Lee HJ, Khoshaghideh F, Patel S, Lee SJ (2004) Clearance of alpha-synuclein oligomeric intermediates via the lysosomal degradation pathway. J Neurosci 24:1888–1896
Li XZ, Zhang SN, Liu SM, Lu F (2013a) Recent advances in herbal medicines treating Parkinson’s disease. Fitoterapia 84:273–285
Li YH, Chen F, Wang JF, Wang Y, Zhang JQ, Guo T (2013b) Analysis of nine compounds from Alpinia oxyphylla fruit at different harvest time using UFLC-MS/MS and an extraction method optimized by orthogonal design. Chem Cent J 7:134
Li GH, Zhang ZJ, Quan Q, Jiang RW, Szeto SSW, Yuan S, Wong WT, Lam HHC, Lee SMY, Chu IK (2016a) Discovery, synthesis, and functional characterization of a novel neuroprotective natural product from the fruit of Alpinia oxyphylla for use in Parkinson’s disease through LC/MS-based multivariate data analysis-guided fractionation. J Proteome Res 15(8):2595–2606
Li YH, Tan YF, Wei N, Zhang JQ (2016b) Diuretic and anti-diuretic bioactivity of the seed and shell extracts of Alpinia oxyphylla fruit. Afr J Tradit Complement Altern Med 13(5):25–32
Lin RJ, Yen CM, Chou TH, Chiang FY, Wang GH, Tseng YP, Wang L, Huang TW, Wang HC, Chan LP, Ding HY, Liang CH (2013) Antioxidant, anti-adipocyte differentiation, antitumor activity and anthelmintic activities against Anisakis simplex and Hymenolepis nana of yakuchinone A from Alpinia oxyphylla. BMC Complem Altern M 13:237
Liu AJ, Zhao X, Li H, Liu Z, Liu B, Mao X, Guo L, Bi K, Jia Y (2014) 5-hydroxymethylfurfural, an antioxidant agent from Alpinia oxyphylla Miq. Improves cognitive impairment in Aβ1-42 mouse model of Alzheimer′s disease. Int Immunopharmacol 23:719–725
Lv XQ, Luo JG, Wang XB, Wang JS, Luo J, Kong LY (2011) Four new sesquiterpenoids from the fruits of Alpinia oxyphylla. Chem Pharm Bull 59(3):402–406
Miao Q, Kong WJ, Zhao XS, Yang SH, Yang MH (2015) GC-FID coupled with chemometrics for quantitative and chemical fingerprinting analysis of Alpinia oxyphylla oil. J Pharmaceut Biomed 102:436–442
Mok CC, Lau CS (2003) Pathogenesis of systemic lupus erythematosus. J Clin Pathol 56:481–490
Patra KC, Hay N (2014) The pentose phosphate pathway and cancer. Trends Biochem Sci 39(8):347–354
Pharmacopoeia Committee (2005) Pharmacopoeia of the People’s Republic of China, vol I. Chemical Industry Press, Beijing, pp 104, 165, 202, 204
Sang SG, Xiao M, Ni YL, Feng GZ, Xiong XH, Luo ZY, Du GK, Xie YQ (2017) Alpinia oxyphylla Miq. extract changes microRNA expression profiles in db-/db-mouse lives. Int J Clin Exp Med 10(10):14447–14457
Shi SH, Zhao X, Liu AJ, Liu B, Li H, Wu B, Bi KS, Jia Y (2015) Protective effect of n-butanol extract from Alpinia oxyphylla on learning and memory impairments. Physiol Behav 139:13–20
Song WJ, Li YH, Wang JG, Li ZY, Zhang JQ (2014) Characterization of nucleobases and nucleosides in the fruit of Alpinia oxyphylla collected from different cultivation regions. Drug Test Anal 6(3):239–245
Sun Z, Kong XZ, Zuo LH, Kang J, Hou L, Zhang XJ (2016) Rapid extraction and determination of 25 bioactive constituents in Alpinia oxyphylla using microwave extraction with ultra-high performance liquid chromatography with tandem mass spectrometry. J Sep Sci 39(3):603–610
Taheri P, Tarighi S (2010) Riboflavin induces resistance in rice against Rhizoctonia solani via jasmonate-mediated priming of phenylpropanoid pathway. J Physiol 167(3):201–208
Tewari A, Plant AK, Mengi N, Patra NK (1999) A review on Alpinia species: chemical, biocidal and pharmacological aspects. J Med Aroma Plant Sci 21:1155–1168
Tsai CT, Chang YM, Lin SL, Chen YS, Yeh YL, Padma VV, Tsai CC, Chen RJ, Ho JJ, Huang CY (2016) Alpinate Oxyphyllae fructus inhibits IGFII-related signaling pathway to attenuate Ang II-induced pathological hypertrophy in H9c2 cardiomyoblasts. J Med Food 19(3):300–309
Vogt T (2010) Phenylpropanoid biosynthesis. Mol Plant 3:2–20
Wang HY, Liu XJ, Wen MF, Pan K, Zou ML, Lu C, Liu SS, Wang WQ (2012) Analysis of the genetic diversity of natural populations of Alpinia oxyphylla Miquel using inter-simple sequence repeat markers. Crop Sci 52:1767–1775
Wang CZ, Yuan HH, Bao XL, Lan MB (2013) In vitro antioxidant and cytotoxic properties of ethanol extract of Alpinia oxyphylla fruits. Pharm Biol 51(11):1419–1425
Wang S, Zhao Y, Zhang JQ, Huang XX, Wang YF, Xu XT, Zheng B, Zhou X, Tian HJ, Liu L, Mei QB (2015) Antidiarrheal effect of Alpinia oxyphylla Miq. (Zingiberaceae) in experimental mice and its possible mechanism of action. J Ethnopharmacol 168:182–190
Wang MS, Bi WC, Fan KY, Li TD, Yan TX, Xiao F, He BS, Bi KS, Jia Y (2018) Ameliorating effect of Alpinia oxyphylla-Schisandra chinensis herb pair on cognitive impairment in a mouse model of Alzheimer’s disease. Biomed Pharmacother 97(1):128–135
Weiss H, Friedrich T, Hofhaus G, Preis D (1991) The respiratory-chain NADH dehydrogenase (complex I) of mitochondria. In: Christen P, Hofmann E (eds) EJB reviews 1991. Springer, Berlin, Heidelberg
Wiśniewski JR, Zougman A, Nagaraj N, Mann M (2009) Universal sample preparation method for proteome analysis. Nat Methods 6(5):359–362
Xie Y, Xiao M, Ni Y, Jiang S, Feng G, Sang S, Du G (2018) Alpinia oxyphylla Miq. extract prevents diabetes in mice by modulating gut microbiota. J Diabetes Res 2018:1–10
Xu JJ, Ji CJ, Zhang YM, Su J, Li Y, Tan NH (2012a) Inhibitory activity of eudesmane sesquiterpenes from Alpinia oxyphylla on production of nitric oxide. Bioorg Med Chem Lett 22(4):1660–1663
Xu JZ, Wang YF, Tan XR, Jing HJ (2012b) MicroRNAs in autophagy and their emerging roles in crosstalk with apoptosis. Autophagy 8(6):873–882
Xu J, Wang F, Guo JJ, Xu CS, Cao YZ, Fang ZL, Wang QW (2020) Pharmacological mechanisms underlying the neuroprotective effects of Alpinia oxyphylla Miq. on Alzheimer’s disease. Int J Mol Sci 21:2071
Yan TX, Wu B, Liao ZZ, Liu B, Zhan X, Bi KS, Jia Y (2016) Brain-derived neurotrophic factor signaling mediates the antidepressant-like effect of the total flavonoids of Alpiniae oxyphyllae Fructus in chronic unpredictable mild stress mice. Phytother Res 30:1493–1502
Youle RJ, Strasser A (2008) The BCL-2 protein family: opposing activities that mediate cell death. Nat Rev Mol Cell Biol 9:47–59
Yu XY, An LJ, Wang YQ, Zhao H, Gao CZ (2003) Neuroprotective effect of Alpinia oxyphylla Miq. fruits against glutamate-induced apoptosis in cortical neurons. Toxicol Lett 14:205–212
Zabala G, Zou J, Tuteja J, Gonzalez DO, Clough SJ, Vodkin LO (2006) Transcriptome changes in the phenylpropanoid pathway of Glycine max in response to Pseudomonas syringae infection. BMC Plant Biol 6(1):26
Zhang Z, Cheang LCV, Wang M, Li GH, Chu IK, Lin ZX, Lee SMY (2012) Ethanolic extract of fructus Alpinia oxyphylla protects against 6-hydroxydopamine-induced damage of PC12 cells in vitro and dopaminergic neurons in zebrafish. Cell Mol Neurobiol 32:27–40
Zhang JQ, Wang S, Li YH, Xu P, Chen F, Tan YF, Duan JN (2013) Anti-diarrheal constituents of Alpinia oxyphylla. Fitoterapia 89:149–156
Zhang Q, Cui C, Chen CQ, Hu XL, Liu YH, Fan YH, Meng WH, Zhao QC (2015) Anti-proliferative and pro-apoptotic activities of Alpinia oxyphylla on HepG2 cells through ROS-mediated signaling pathway. J Ethnopharmacol 169:99–108
Zhang SH, Zeng XF, Ren M, Mao XB, Qiao SY (2017) Novel metabolic and physiological functions of branched chain amino acids: a review. J Animal Sci Biotechnol 8:10
Zhang Q, Zheng YL, Hu XJ, Hu XL, Lv WW, Lv D, Chen JJ, Wu ML, Song QC, Shentu JZ (2018) Ethnopharmacological uses, phytochemistry, biological activities, and therapeutic applications of Alpinia oxyphylla Miquel: a review. J Ethnopharmaco 224:149–168
Zhao XS, Wei JH, Shu XY, Kong WJ, Yang MH (2016) Multi-elements determination in medical and edible Alpinia oxyphylla and Morinda officinalis and their decoctions by ICP-MS. Chemosphere 164:430–435
Zhao X, Su X, Liu C, Jia Y (2018) Simultaneous determination of chrysin and tectochrysin from Alpinia oxyphylla fruits by UPLC-MS/MS and its application to a comparative pharmacokinetic study in normal and dementia rats. Molecules 23:1702
Zhu X, Liu X, Cheng Z, Zhu J, Xu L, Wang F, Qi W, Yan J, Liu N, Sun Z, Liu H, Peng X, Hao Y, Zheng N, Wu Q (2016) Quantitative analysis of global proteome and lysine acetylome reveal the differential impacts of VPA and SAHA on HL60 cells. Sci Rep 6:19926
Acknowledgements
This work was funded by the National Natural Science Foundation of China (No. 81560611).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dai, Y., Yuan, L., Fu, J. et al. Label-Free Quantitative Proteomic Profiling Identifies Potential Active Components to Exert Pharmacological Effects in the Fruit of Alpinia oxyphylla by Mass Spectrometry. J. Plant Biol. 63, 297–310 (2020). https://doi.org/10.1007/s12374-020-09251-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12374-020-09251-9