Abstract
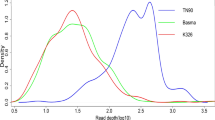
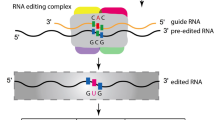
In plants, RNA editing is a post-transcriptional modification of transcripts and commonly occurs in plastids and mitochondria. In the case of flowering plants, not only PPR but also non-PPR proteins, such as MORF/RIP, ORRM and OZ partake in the diverse RNA editing complex. In Arabidopsis thaliana, 12 types of RNA editing patterns have been predicted in nuclear, chloroplast, and mitochondrial transcripts. In this study, tissue-specific RNA editing events were detected in gene families involved in RNA editing. Different Arabidopsis tissues at variable developmental stages, including 4-, 8-, and 12-d-old seedlings and 16-, 21-, 27-, and 32-d-old leaves, stems, stipes and roots, were collected and used for this study. Nine types of RNA editing events, including C-to-U, U-to-C, A-to-I, A-to-C, A-to-U, G-to-A, G-to-C, U-to-A and U-to-G, were identified in target genes. Most of the editing events occurred in seedlings and leaves and a few in stem tissues. Extensive U-to-C editing (60%) and A-to-G editing (54%) was detected in 12-d-old seedlings and 21-d-old leaves, respectively. This is the first experimental report of tissue-specific nuclear RNA editing events found in plants. This study will provide important information for revealing the mechanism of RNA editing in plants.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Barkan A, Small I 2014 Pentatricopeptide repeat proteins in plants. Annu Rev Plant Biol 65:415–442
Chen TC, Liu YC, Wang X, Wu CH, Huang CH, Chang CC 2017 Whole plastid transcriptomes reveal abundant RNA editing sites and differential editing status in Phalaenopsis aphrodite subsp. formosana. Bot Stud 58:1–14
Cheng S, Gutmann B, Zhong X, Ye Y, Fisher MF, Bai F, Castleden I, Song Y, Song B, Huang J, Liu X, Xu X, Lim BL, Bond CS, Yiu SM, Small I 2016 Redefining the structural motifs that determine RNA binding and RNA editing by pentatricopeptide repeat proteins in land plants. Plant J 85:532–547
Delannoy E, Stanley WA, Bond CS, Small ID (2007) Pentatricopeptide repeat (PPR) proteins as sequence-specificity factors in post-transcriptional processes in organelles. Biochem Soc Trans 35:1643–1647
Drescher A, Hupfer H, Nickel C, Albertazzi F, Hohmann U, Herrmann R, Maier R (2002) C-to-U conversion in the intercistronic ndhI/ndhG RNA of plastids from monocot plants: Conventional editing in an unconventional small reading frame? Mol Genet Genomics 267:262–269
Eggington JM, Greene T, Bass BL 2011 Predicting sites of ADAR editing in double-stranded RNA. Nat Commun 2:1–9
Goodman RA, MacBeth MR, Beal PA 2012 ADAR Proteins: structure and catalytic mechanism. Curr Top Microbiol Immunol 353:1–33
Guillaumot D, Lopez-Obando M, Baudry K, Avon A, Rigaill G, Falcon de Longevialle A, Broche B, Takenaka M, Berthomé R De Jaeger G, Delannoy E, Lurin C 2017 Two interacting PPR proteins are major Arabidopsis editing factors in plastid and mitochondria. Proc Natl Acad Sci USA 114:8877–8882
Hammani K, Giegé P 2014 RNA metabolism in plant mitochondria. Trends Plant Sci 19:380–389
He P, Huang S, Xiao G, Zhang Y, Yu J 2016 Abundant RNA editing sites of chloroplast protein-coding genes in Ginkgo biloba and an evolutionary pattern analysis. BMC Plant Biol 16:1–12
Ichinose M, Sugita M 2016 RNA editing and its molecular mechanism in plant organelles. Genes 8:1–15
Jean-Claude Farré, Cindy Aknin, Alejandro Araya BC 2012 RNA editing in mitochondrial for splicing Trans -Introns is required for splicing. PLoS One 7:1–10
Kent WJ 2002 BLAT -The BLAST -like alignment tool. Genome Res 12:656–664
Kuttan A, Bass BL 2012 Mechanistic insights into editing-site specificity of ADARs. Proc Natl Acad Sci USA 109:E3295–E3304
Luo GH, Li XH, Han ZJ, Zhang ZC, Yang Q, Guo HF, Fang JC 2016 Transition and transversion mutations are biased towards GC in transposons of Chilo suppressalis. Genes 7:1–12
Lurin C, Andrés C, Aubourg S, Bellaoui M, Bitton F, Bruyère C, Caboche M, Debast C, Gualberto J, Hoffmann B, Lecharny A, Le Ret M, Martin-Magniette ML, Mireau H, Peeters N, Renou JP, Szurek B, Taconnat L, Small I 2004 Genome-wide analysis of Arabidopsis pentatricopeptide repeat proteins reveals their essential role in organelle biogenesis. Plant Cell 16:2089–2103
Meng Y, Chen D, Jin Y, Mao C, Wu P, Chen M 2010 RNA editing of nuclear transcripts in Arabidopsis thaliana. BMC Genomics 11:1–7
Rodrigues NF, Christoff AP, da Fonseca GC, Kulcheski FR, Margis R 2017 Unveiling chloroplast RNA editing events using next generation small RNA sequencing data. Front Plant Sci 8:1–13
Shikanai T 2015 RNA editing in plants: Machinery and flexibility of site recognition. Biochim Biophys Acta 1847:779–785
Takenaka M, Zehrmann A, Verbitskiy D, Härtel B, Brennicke A 2013 RNA Editing in Plants and Its Evolution. Annu Rev Genet 47:335–352
Tseng CC, Lee CJ, Chung YT, Sung TY, Hsieh MH 2013 Differential regulation of Arabidopsis plastid gene expression and RNA editing in non-photosynthetic tissues. Plant J 82:375–392
Wagoner JA, Sun T, Lin L, Hanson MR 2015 Cytidine deaminase motifs within the DYW domain of Two pentatricopeptide repeat-containing proteins are required for site-specific chloroplast RNA editing. J Biol Chem 290:2957–2968
Wu B, Chen H, Shao J, Zhang H, Wu K, Liu C 2017 Identification of symmetrical RNA editing events in the mitochondria of Salvia miltiorrhiza by strand-specific RNA sequencing. Sci Rep 7:1–11
Yagi Y, Nakamura T, Small I 2014 The potential for manipulating RNA with pentatricopeptide repeat proteins. Plant J 78:772–782
Zeng WH, Liao SC, Chang CC 2007 Identification of RNA editing sites in chloroplast transcripts of Phalaenopsis aphrodite and comparative analysis with those of other seed plants. Plant Cell Physiol 48:362–368
Acknowledgements
Part of this work was supported by Grants-in-Aid for Scientific Research from the Japan Society for the Promotion of Science (26670167, 17H02204 and 18K19288).
Author information
Authors and Affiliations
Contributions
TT, UQ and MTAA designed research; UQ and MTAA performed research, analyzed the data and wrote the paper; TT, UQ and MTAA revised the manuscript. All authors have agreed on the contents of the manuscript and declare no conflict of interests.
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Qulsum, U., Azad, M.T.A. & Tsukahara, T. Analysis of Tissue-specific RNA Editing Events of Genes Involved in RNA Editing in Arabidopsis thaliana. J. Plant Biol. 62, 351–358 (2019). https://doi.org/10.1007/s12374-018-0452-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12374-018-0452-5