Abstract
The oxidative stress response was studied in five different clones of Saccharum spontaneum (IMP-564, IS76-216, Pamba, SES-90 and Taiwan) by analyzing the physiological and biochemical parameters and gene expression pattern. The sixty days-old plants were sprayed with different concentrations of 30% H2O2 (300 ppm, 500 ppm and 1000 ppm) for consecutive three days in glass house. Adaptive response was recorded in terms of chlorophyll fluorescence, chlorophyll stability index (CSI), activities of peroxidase (POX) and super oxide (SOD), lipid peroxidation (LPO), protein and proline content at 48 h and 72 h of H2O2 treatment. The IMP -564 clone recorded high SOD and lower lipid peroxidation levels. Based on chlorophyll fluorescence, CSI, peroxidase, SOD and lipid peroxidation values, and the clone SES-90 recorded positive trend for stress tolerance. CSI showed significant difference between genotypes and different concentrations of H2O2. As the genotypes IMP-564 and SES-90 showed positive response to oxidative stress tolerance, the S. spontaneum genotype SES-90 was selected for comparative gene expression studies with Erianthus sp clones IJ76-389 and IK76-91. Different concentrations of H2O2 and different time intervals (control, 500 ppm and 1000 ppm for 48 h and 72 h) were used for gene expression studies. The stress responsive genes viz. Ascorbate peroxidase gene (Apx) Nam, Ataf1 and Cuc2 gene (NAC), Ethylene response factor gene (Erf), Glutathione S-transferases gene (Gst), Myeloblastosis antisense gene (Mybas) and Catalase gene (Cat) showed differential expression among the genotypes of S. spontaneum and Erianthus sp. Among the genotypes, SES 90 and IK76-91 showed better tolerance against oxidative stress and the expression levels of stress responsive genes in the genotype IK76-91 was higher than the genotype SES 90.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sugarcane is a commercial crop of India and accounts for more than 70 percent of world sugar production. India is the second largest producer of sugar in the world market, closely following Brazil. Dramatically changing environment poses serious direct and indirect threat to the sustainable agricultural production leading to global food insecurity. The fate of sugarcane productivity is of no difference from other crops. Environmental stresses such as drought, salinity, chilling, metal toxicity, and UV-B (short Ultra Violet) radiation as well as pathogens attack lead to enhanced generation of reactive oxygen species (ROS) in plants due to disruption of cellular homeostasis (Sharma and Dubey 2007; Hu et al. 2008; Tanou et al. 2009; Maheswari and Dubey 2009; Han et al. 2009; Srivastava and Dubey 2011; Mishra et al. 2011; Hamim et al. 2017; Ferreira et al. 2017). Environmental stresses, such as salinity, drought and extreme temperature increase the ROS levels in plant cells. Over production of ROS in the system will ultimately result in oxidative stress. This can lead to the peroxidation of lipids, damage to nucleic acid, oxidation of proteins, enzyme inhibition, DNA mutation and finally to cell death (Sharma and Dubey ; Gill and Tuteja 2010; Mitler et al. 2011; Kumari et al. 2007,2015). ROS like H2O2 is more stable and can diffuse directly from chloroplasts to the nuclei (Exposito-Rodriguez et al. 2017), thus influencing gene expression in the nuclei both directly and indirectly.
To adapt to the environmental stresses, plants undergo various physiological and metabolic processes. The major molecular response is the activation of stress-responsive genes and synthesis of diverse functional proteins through a complex signal transduction network (Shinozaki et al. 2003; Hirayama and Shinozaki 2010). Some of the most common responses for abiotic stress tolerance in plants are overproduction of osmo-protectants or osmolytes (such as sucrose, betaines, and proline), cellular metabolic changes (defense-related secondary metabolite production, proteolytic activity, adenylate energy charge, ionic homeostasis, redox state regulation, and antioxidant enzymes activation), anatomical and morphological changes in plant tissues.
Various enzymes like superoxide dismutase (SOD), peroxidase (POX), ascorbate peroxidase (APX), catalase (CAT) and non-enzymatic antioxidants such as tocopherols, ascorbic acid (AsA), glutathione (GSH) and genes like Erf and NAC are involved in regulation of signal transduction and gene expression during stress response and play different roles in detoxifying ROS (Manimekalai et al. 2018a; b). Another group which regulate stress response are a number of transcription factors (Shinozaki et al. 2003; Yamaguchi-Shinozaki and Shinozaki, 2006; Cattivelli et al. 2008), which regulate various stress-inducible genes cooperatively or separately, and may constitute gene networks. Stress induced transcription factors belong to DREB (Dehydration Response Elements Binding) protein family, zinc finger protein family, MYB (Myeloblastosis) family, bZlP (basic leucine zipper domain) family, WRKY family (named after the conserved WRKYGQK amino acid sequence), and NAC family. These transcription factors regulate stress-inducible genes thereby initiating stress responses and establish plant’s response to tolerate the stress. Over expression of these transcription factors genes enhanced the drought, salt, and cold tolerance in many crops (Ito et al. 2006; Nakashima and Shinozaki 2006; Zeng et al. 2008; Ambawat et al. 2013; Hong et al. 2016; Roy 2015).
Saccharum spontaneum, a species involved in hybridization program with sugarcane show tolerance to biotic/abiotic stresses. Since past decades, the wild species has contributed the tolerant genes to cultivated sugarcane through introgressive breeding. Several reports explained the use of hydrogen peroxide and methyl viologen in simulating oxidative stress in plants (Upadhyaya et al. 2007; Vijayalakshmi et al. 2014). Vijayalakshmi et al. (2014) used different concentrations of H2O2 (0, 0.05, 0.1, 0.15, 0.2 mM) to induce oxidative stress in rice. Deng et al. (2012) induced the oxidative stress in sweet potato by exogenous application of H2O2 (up to 2.5 mM) into the adventitious root culture. In the present study, Saccharum spontaneum genotypes were screened for the adaptive physiological, biochemical traits and expression of stress responsive genes. The study helped in identifying the stress responsive genotypes of S. spontaneum under oxidative stress conditions and uses them for higher level of gene to gene interaction studies.
Materials and Methods
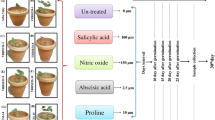
A pot culture experiment was designed to measure the physiological parameters in S.spontaneum genotypes. The potting mixture was prepared with sand, loam and farmyard manure in the ratio 2:1:1. The clones were selected from the already available germplasm present in Sugarcane Breeding Institute, Coimbatore, based on the earlier screening reports, viability of buds and tillering capacity. Sixty days old plants of five genotypes (IMP-564, IS76-216, Pamba, SES-90 and Taiwan) were used for the experiment. Three different concentrations (300 ppm, 500 ppm and 1000 ppm) of 30% hydrogen peroxide (commercially available in the market) were used as treatments. Three replications were maintained for different treatment. One set from each genotype was retained as control by spraying with normal water. The spraying was done consecutively for three days (24 h, 48 h and 72 h) at 8 am. Physiological parameters like photochemical efficiency (CF), chlorophyll stability index (CSI), lipid peroxidation, protein and proline content and enzyme assays for peroxidase and super oxide dismutase (SOD) for 48 h and 72 h of H2O2 treatment were followed as per the protocol described by Manimekalai et al. 2018a). The data were analyzed using the software JMP 9.0.
For gene expression studies, data mining through literature survey was performed and seven multiple stress responsive genes Hsp (Heat Shock Proteins gene), Apx (Ascorbate peroxidase gene), NAC (defined by Nam, Ataf1 and Cuc2 genes), Erf (Ethylene response factor gene), Gst (Glutathione S-transferases gene), Mybas (Myeloblastosis antisense gene) and Cat (Catalase) were selected. The experiment conducted in a randomized complete block design with four treatments and two replications, consisting of 0 ppm (control), 300, 500 and 1000 ppm of 30% hydrogen peroxide. The plants were maintained in a glass house in pots. Plant samples were collected on 2nd (48 h) and 3rd (72 h) days of treatment and were further processed for expression studies.
RNA isolation was done using TRIzol reagent (Invitrogen), cDNA conversion with cDNA synthesis kit (Thermofisher scientific), and gene expression analysis with SYBR green assay (Applied Biosystems) as per the protocol (Manimekalai et al. 2018b). The fold change of expression was calculated in terms of relative quantification (Rq) using Livak’s method and the values were analyzed using statistical methods (Livak and Schmittgen 2001).
Results and Discussion
Abiotic stress factors like drought, salinity, water-logging, and heavy metal toxicity lead to oxidative stress in plants (You and Chang 2015). Generally the wild species respond to such factors in different ways to make it adaptable. At the molecular level, differential expression of certain stress responsive contributes to the tolerance (Li et al. 2019). Exogenous application of hydrogen peroxide was proven to induce oxidative stress in rice and sweet potato (Vijayalakshmi et al. 2014; Deng et al. 2012), and we used it in the present study to induce the oxidative stress in S. spontaneum and Erianthus sp.
Morphologically, though not much remarkable variation was seen at lower concentration of hydrogen peroxide (300 ppm), but slight yellowing of leaves (Fig. 1b and e) and leaf tip drying and rolling at higher concentrations (Fig. 2c) were observed. The physiological and biochemical assays were performed to screen the clones of Saccharum spontaneum. Gene expression studies offered the opportunity to compare the physiological and biochemical responses with molecular response.
Physiological and Biochemical Responses to the Oxidative Stress
The three-way ANOVA on physiological and biochemical parameters (chlorophyll fluorescence, peroxidase, superoxide dismutase, lipid peroxidation, protein and proline) showed highly significant differences among the genotypes (p < 0.001) for SOD, POX, LPO, while changes in proline was significant at p < 0.05. The change in the soluble protein content was non-significant among the genotypes. The effect of different concentrations of hydrogen peroxide on levels of SOD was significant (p < 0.001) and POX (p < 0.05), while other traits were non-significant. The duration of stress treatment (T) showed significant effect on protein and LPO (p < 0.05). The interaction effect of VxS and VxT had significant effect on LPO and SOD, POX, proline, LPO, respectively. The SxT and the higher order of interaction VxSxT had no significant effect on the studied parameters. Thus, genotypes, concentration of hydrogen peroxide and duration of treatment and their interactions on the various biochemical traits were showed different levels of response (Table 1). The main effect of genotypes, concentrations of hydrogen peroxide and the interaction effect (VxS) were significant (p < 0.001) for CSI (Table 2).
Enzyme Activities—SOD and POX
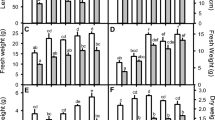
Super Oxide Dismutase (SOD) is a key anti-oxidative enzyme neutralizes the activated species of oxygen (Van Toai and Bolles 1991). It is the first defense against reactive oxygen species which converts super oxides to H2O2. Increased SOD activity of certain genotypes can be taken as an indirect indication of enhanced stress tolerance (Arisi et al. 1998). Lots of other studies corroborate the role of SOD during stress. Specific activity of SOD increased, under prolonged oxidative stress induced by methyl viologen (Chagas et al. 2008; Bowler et al. 1994) and in the present study too, SOD showed high activity in clones IMP-564 and SES-90 (Fig. 3a). IMP 564 showed increased SOD activity with increase in concentration at both 48 and 72 h. But the activity was found to decrease on prolonged treatment (1000 ppm) 72 h of H2O2 treatment with 500 ppm in IMP 564 showed 24% increase with respect to control. At 48 h, 1000 ppm of H2O2 it showed 18% increase over its control. IS76-216 showed an increase in SOD activity at 300 ppm but failed to accumulate at higher concentration of H2O2. ANOVA showed that the genotypes (V) and concentration of H2O2 (S) were highly significant on superoxide dismutase (SOD), but exposure time was non-significant. However, the interaction between VxT was significant with SOD. The other interaction viz, V × S, S × T, V × S × T were non-significant.
Peroxidase (POX) activity increases as a protective response in plants during environmental stresses (Pujari and Chanda 2002). All clones in this study showed an increased POX activity at 500 ppm H2O2 for 48 h (Fig. 3 b). Except for the clone SES-90, there was a decrease in POX activity with higher concentration of H2O2 (1000 ppm). SES-90 showed an increase of 23% activity at 72 h treatment. IMP 564 showed a drastic decrease in POX activity on high concentration of H2O2 (1000 ppm) with respect to its 500 ppm (25% reduction). Pamba and Taiwan did not show any significant SOD activity, rather there was decreased enzyme activity. Statistical analysis revealed that the main effect V, S and the interaction V × T showed significance on POX, while the other level of interactions were not significant.
Lipid Peroxidation
Lipid peroxidation is directly proportional to the damage to the plants. A low LPO value signifies tolerance response of the plants. Higher levels of H2O2 accompanied by an increase in lipid peroxidation were observed in young sugarcane plants during the initial growth phase under severe water stress (Boaretto et al. 2014). Within the clone SES-90, the plants that were treated with 500 ppm of H2O2 for 48 h and 1000 ppm of H2O2 for 48 and 72 h showed lower LPO values than with 300 ppm (Fig. 4 a). This denotes the induction of stress tolerance upon prolonged exposure to H2O2. During 72 h of H2O2 exposure, the clones IS76-216, Pamba, and Taiwan were found to be the most affected, demonstrating their susceptibility. Pamba showed a higher lipid peroxidation (50% increase over control) on prolonged exposure (72 h with 1000 ppm). Similar enhancement on LPO was observed in maize (Xu et al. 2011), Leymus chinensis and Stipa grandis (Xu and Zhou. 2006; Xu et al. 2009 and Song et al. 2015).
Soluble Protein Content
Higher protein content is correlated with higher RuBisCo activity and high photosynthetic efficiency under stress conditions. The abiotic stress results in enzyme inhibition and disruption of pathways in protein synthesis (Paramjeet et al. 2014). In the present study, all genotypes showed a marked decrease in the protein content with prolonged exposure to H2O2. All genotypes showed increased soluble protein content at 1000 ppm except the clone Taiwan (Fig. 4b). However, the variation is non-significant among the genotypes and interaction effect except time (T).
Proline Content
Proline content increases substantially under abiotic stress conditions. Organisms acquire oxidative stress tolerance by accumulating proline. Proline accumulates in the cytosol and vacuole during stress and was shown to protect plant cells against damages caused by 1O2 or HO· (Hong et al. 2016; Matysik et al. 2002). It acts as an osmolyte, a chemical chaperon and as a direct scavenger of ROS (Liang et al. 2013). In our study, SES-90 showed higher accumulation of proline 500 ppm H2O2, and had 40% increase in proline content at 1000 ppm (Fig. 4c). The main effect V and the interaction VxT are significant for proline content, while the other level of interactions were non-significant.
Chlorophyll Fluorescence
Chlorophyll fluorescence is determined using the Fv/Fm (Variable fluorescence/Maximum fluorescence) ratio. A lower Fv/Fm value indicates the plants are under stress (Maxwell and Johnson 2000). There was no much difference with respect to chlorophyll fluorescence. However, the genotype Pamba showed increase (8.0% in chlorophyll fluorescence at 500 ppm over its control. IMP 564 too showed a moderate increase in fluorescence (Fig. 5).
Chlorophyll Stability Index
CSI is an indicator of maintenance of photosynthetic pigments. Reactive oxygen species causes damage to chloroplast thereby reduces the chlorophyll stability. Higher CSI value indicates higher stress tolerance. A study in rice by Mohan et al. (2000) indicated that chlorophyll content was more affected in susceptible varieties than in tolerant varieties under salt stress. In the present study, IS76-216 and Pamba showed a relatively low index when compared to the control (Fig. 6). SES-90 showed higher chlorophyll stability index at 500 ppm of H2O2 than other clones (31.8% increase over control). Similarly, higher CSI was observed in tolerant varieties of sugarcane under heat stress (Kohila and Gomathi 2018) and drought stress (Sanghera and Kumar 2018). ANOVA for CSI showed significant difference between genotypes and concentrations of H2O2 and hence, this parameter is particularly of use in screening larger number genotypes. Overall analysis showed that the clones IMP-564 and SES-90 exhibited desirable levels of physiological and biochemical parameters which may contribute the tolerance.
Relative Gene Expression under Oxidative Stress
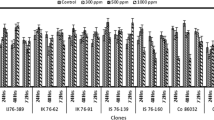
Expression pattern of seven multiple stress responsive genes was analyzed under different hydrogen peroxide concentrations and at various time intervals. The expression of genes under stress was analyzed with respective to its control. The expression of key genes implicated in stress was reported here.
NAC Gene
NAC (NAM (No apical meristem), ATAF 1/2 (Arabidopsis transcription activator factor 1/2) and CUC2 (cup shaped cotyledon)) family consists of several genes (> 100), and they are reported to be involved in diverse functions in plants especially biotic and abiotic stress tolerance. Fang et al. (2015) reported that the NAC transcription factor gene, SNAC3 functions as a positive regulator under high temperature and drought stress in rice. The gene enhanced the abiotic stress tolerance by modulating H2O2 homeostasis state through controlling the expression of ROS- associated genes. Phylogenetic analysis of NAC transcription factor genes by Manimekalai et al. (2016) revealed the presence of multiple stress tolerant NAC genes in sugarcane orthologous to sorghum, wheat and maize. Current study showed an increased expression at 500 ppm H2O2 in Erianthus sp. genotypes (IJ76- 389 and IK76-91) and S. spontaneum under both 48 h and 72 h exposure (Fig. 7); however, the level of expression is comparatively lesser in S.spontaneum.
Myb Transcription Factor Gene
Most MYB proteins function as transcription factors with varying numbers of MYB domain repeats conferring their ability to bind DNA. They are widely distributed in plants and implicated in the ABA response (Ambavat et al. 2013). In the present study, the expression levels were higher at 500 ppm (Fig. 8). This was in congruence with the studies by Roy (2015) where Myb genes showed higher expression under the epigenetic regulation of stress in Arabidopsis. The genotype IJ76-389 had higher expression levels upon prolonged exposure (72 h) and at higher concentration of H2O2 (1000 ppm). Our previous study (Manimekalai et al. 2018b) also indicated an increase in gene expression for Mybas under higher concentration of H2O2 stress. Relatively, the expression levels of Mybas genes were higher in Erianthus sp. than S. spontaneum.
Erf (Ethylene Response Factors) Gene
The expression of Erf genes is responsive to several abiotic stress factors. The up-regulation of Erf gene was reported against environmental stress conditions (Mizoi et al. 2012; Cui et al. 2021). Quantitative PCR study by Wang et al. (2015) revealed over expression of JcERF2 gene by multiple stresses like abscisic acid, drought and salt stress in tobacco. Gene expression varies with different genotypes (Fig. 9). SES-90 showed a dramatic increase in gene expression at 500 ppm at 48 h, further the expression ceases after 72 h which could be implicated its involvement in signalling pathway leading to tolerance.
Apx (Ascorbate Peroxidase) Gene
Apx gene, known for its scavenging nature, seems to express on prolonged exposure to stress. Our study showed the over expression of Apx gene during 72 h of H2O2 treatment (Fig. 10). At 1000 ppm, both the species showed an increased expression under stress conditions. Likewise, studies in barley leaves (Harb et al. 2016) revealed higher expression of Apx gene upon prolonged stress conditions.
Cat (Catalase) Gene
CAT is an enzymatic antioxidant that catalyses the conversion of H2O2 into H2O and O2, thereby detoxifying the ROS present in plant cells (Mhamdi et al. 2010). Harb et al. (2016) reported that the catalase gene was up-regulated after 9 days of drought treatment. The scavenging nature of catalase gene is well established through different studies. In the current study, both E. arundinaceus and S. spontaneum showed an increase in expression at 500 ppm treatment at 48 h exposure but during 72 h the expression was reduced (Fig. 11). The lower Cat gene levels lead to reduced CAT enzyme activities (Cavalcanti et al. 2004). Chakraborty and Bhattacharjee (2015) have also observed a decreased CAT activity in rice under cold stress and CAT activity declined in cowpea under salinity stress (Cavalcanti et al. 2004).
Hsp (Heat Shock Proteins) Genes
HSPs are molecular chaperones involved in membrane and protein stability during stress conditions. In Sugarcane, the Hsp gene from E. arundinaceus conferred enhanced water and salinity stress tolerance (Augustine et al. 2015). In the present study, 48 h of 500 ppm H2O2 showed an increase in expression of Hsp gene in the genotypes IK 76–91 and SES-90 than at 72 h (Fig. 12).
Gst (Glutathione S-transferases) Gene
Glutathione S-transferases are ubiquitous enzymes that respond to xenobiotic and oxidative compounds. Studies suggest that expression of GSTs is induced by various abiotic stresses such as drought (Xu et al. 2015), oxidative stress (Levine et al. 1994; Chen et al. 2012), salt stress (Jia et al. 2016) and multiple abiotic stresses (Ding et al. 2017). In our study, all the genotypes showed higher Gst expression when exposed to 500 ppm after 48 h (Fig. 13). But on prolonged exposure (72 h), the expression levels receded.
Conclusion
Physiological parameters (chlorophyll stability index and lipid peroxidation) and biochemical parameters (superoxide dismutase, peroxidase and proline content) are significantly different among the S.spontaneum genotypes. Based on these parameters, the genotypes IMP-564 and SES-90 are considered as tolerant to oxidative stress. The genes such as NAC and Apx are actively transcribed during oxidative stress which contributes to tolerance under stress conditions either by acting as signal transduction or by efficiently scavenging reactive oxygen species. The comparison of expression pattern of multiple stress responsive genes showed higher levels of expression in Erianthus sp. clone IK76-91 than the S. spontaneum clone SES 90 under oxidative stress.
Change history
07 November 2022
A Correction to this paper has been published: https://doi.org/10.1007/s12355-022-01218-z
Abbreviations
- ROS:
-
Reactive oxygen species
- SOD:
-
Superoxide dismutase
- POX:
-
Peroxidase
- LPO:
-
Lipid peroxidation
- CF:
-
Chlorophyll fluorescence
- Fv/Fm:
-
Variable fluorescence/Maximum fluorescence
- CSI:
-
Chlorophyll stability index
- TFs:
-
Transcription factors
- qRT-PCR:
-
Quantitative Reverse Transcriptase—Polymerase Chain Reaction
- H2O2 :
-
Hydrogen peroxide
References
Ambawat, S., P. Sharma, N.R. Yadav, and C.Y. Ram. 2013. MYB transcription factor genes as regulators for plant responses: An overview. Physiology and Molecular Biology of Plants 19 (3): 307–321. https://doi.org/10.1007/s12298-013-0179-1.
Arisi, A.M., G. Cornic, L. Jouanin, and C.H. Foyer. 1998. Overexpression of iron superoxide dismutase in transformed poplar modifies the regulation of photosynthesis at low CO2 partial pressures or following exposure to the prooxidantherbicide methyl viologen. Plant Physiology 117: 565–574.
Augustine, S.M., J.A. Narayan, D.P. Syamaladevi, C. Appunu, M. Chakravarthi, and V. Ravichandran. 2015. Erianthus arundinaceus HSP70 (EaHSP70) over expression increases drought and salinity tolerance in sugarcane (Saccharum spp. hybrid). Plant Science 232: 23–34. https://doi.org/10.1016/j.plantsci.2014.12.012.
Boaretto, L.F., G. Carvalho, L. Borgo, S. Creste, M.G. Landell, and P. Mazzafera. 2014. Water stress reveals differential antioxidant responses of tolerant and non-tolerant sugarcane genotypes. Plant Physiology and Biochemistry 74: 165–175. https://doi.org/10.1016/j.plaphy.2013.11.016.
Bowler, C., W. Van Camp, M. Van Montagu, and D. Inzé. 1994. Superoxide dismutases in plants. Critical Reviews in Plant Science 13: 199–218.
Cattivelli, L., F. Rizza, F.W. Badeck, E. Mazzucotelli, A.M. Mastrangelo, E. Francia, C. Mare, A. Tondelli, and A.M. Stanca. 2008. Drought tolerance improvement in crop plants: an integrated view from breeding to genomics. Field Crops Research 105: 1–14.
Cavalcanti, F.R., J.T.A. Oliveira, A.S. Martins-Miranda, R.A. Viégas, and J.A.G. Silveira. 2004. Superoxide dismutase, catalase and peroxidase activities do not confer protection against oxidative damage in salt-stressed cowpea leaves. The New Phytologist 163 (3): 563–571.
Chagas, R.M., J.A.G. Silveira, R.V. Ribeiro, V.A. Vitorello, and H. Carrer. 2008. Photochemical damage and comparative performance of superoxide dismutase and ascorbate peroxidase in sugarcane leaves exposed to paraquat-induced oxidative stress. Pesticide Biochemistry and Physiology 90: 181–188.
Chakraborty, A., and S. Bhattacharjee. 2015. Differential competence of redox-regulatory mechanism under extremes of temperature determines growth performances and cross tolerance in two indica rice cultivars. Journal of plant physiology 176: 65–77. https://doi.org/10.1016/j.jplph.2014.10.016.
Chen, J.H., H.W. Jiang, E.J. Hsieh, H.Y. Chen, C.T. Chien, H.L. Hsieh, et al. 2012. Drought and salt stress tolerance of an Arabidopsis glutathione S-transferase U17 knockout mutant are attributed to the combined effect of glutathione and abscisic acid. Plant Physiology 158: 340–351. https://doi.org/10.1104/pp.111.181875.
Cui, M., M. Haider, P. Chai, J. Guo, P. Du, H. Li, W. Dong, B. Huang, Z. Zheng, L. Shi, X. Zhang, and S. Han. 2021. Genome-wide identification and expression analysis of AP2/ERF transcription factor related to drought stress in cultivated peanut (Arachis hypogaea L.). Frontiers in Genetics 12: 750–761. https://doi.org/10.3389/fgene.2021.750761.
Deng, X.P., Y.J. Cheng, X.B. Wu, S.S. Kwak, W. Chen, and A.E. Eneji. 2012. Exogenous hydrogen peroxide positively influences root growth and metabolism in leaves of sweet potato seedlings. Australian Journal of Crop Science. 6: 1572–1578.
Ding, N., A. Wang, X. Zhang, Y. Wu, R. Wang, H. Cui, et al. 2017. Identification and analysis of glutathione S-transferase gene family in sweet potato reveal divergent GST-mediated networks in aboveground and underground tissues in response to abiotic stresses. BMC Plant Biology 17: e225. https://doi.org/10.1186/s12870-017-1179-z.
Exposito-Rodriguez, M., P.P. Laissue, G. Yvon-Durocher, N. Smirnoff, and P.M. Mullineaux. 2017. Photosynthesis-dependent H2O2 transfer from chloroplasts to nuclei provides a high-light signalling mechanism. Nature Communications 8 (1): 49. https://doi.org/10.1038/s41467-017-00074-w.
Fang, Y., K. Liao, H. Du, Y. Xu, H. Song, and X. Li. 2015. A stress- responsive NAC transcription factor SNAC3 confers heat and drought tolerance through modulation of reactive oxygen species in rice. Journal of Experimental Botany 66: 6803. https://doi.org/10.1093/jxb/erv386.
Ferreira, T., M.S. Tsunada, D. Bassi, P. Araújo, L. Mattiello, and G.V. Guidelli. 2017. Sugarcane Water Stress Tolerance Mechanisms and Its Implications on Developing Biotechnology Solutions. Frontiers in Plant Science 8: 1077. https://doi.org/10.3389/fpls.2017.01077.
Gill, S.S., and N. Tuteja. 2010. Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiology and Biochemistry 48: 909–930. https://doi.org/10.1016/j.plaphy.2010.08.016.
Hamim, Violita, and Triadiati and Miftahudin. 2017. Oxidative stress and photosynthesis reduction of cultivated (Glycine max L.) and wild soybean (G.tomentella L.) exposed to drought and paraquat. Asian Journal of Plant Science 16: 65–77.
Han, C., Q. Liu, and Y. Yang. 2009. Short-term effects of experimental warming and enhanced ultraviolet-B radiation on photosynthesis and antioxidant defense of Picea asperata seedlings. Plant Growth Regulation 58: 153–162. https://doi.org/10.1007/s10725-009-9363-2.
Harb, A., D. Awad, and N. Samarah. 2016. Gene expression and activity of antioxidant enzymes in barley (Hordeum vulgare L.) under controlled severe drought. Journal of Plant Interactions. 10: 109–116. https://doi.org/10.1080/17429145.2015.1033023.
Hirayama, T., and K. Shinozaki. 2010. Research on plant abiotic stress responses in the post-genome era: Past, present and future. Plant Journal 61: 1041–1052. https://doi.org/10.1111/j.1365-313X.2010.04124.x.
Hong, Y., H. Zhang, L. Huang, L. Dayong, and F. Song. 2016. Over expression of a stress-Responsive NAC transcription Factor Gene ONAC022 improves Drought and salt Tolerance in Rice. Frontiers in Plant Science 10: 3389.
Hu, W.H., X.S. Song, and K. Shi. 2008. Changes in electron transport, superoxide dismutase and ascorbate peroxidase isoenzymes in chloroplasts and mitochondria of cucumber leaves as influenced by chilling. Photosynthetica 46: 581. https://doi.org/10.1007/s11099-008-0098-5.
Ito, Y., K. Koji, M. Kyonoshin, T. Teruaki, K. Masatomo, S. Motoaki, S. Kazuo, and Y.S. Kazuko. 2006. Functional Analysis of Rice DREB1/CBF-type Transcription Factors Involved in Cold-responsive Gene Expression in Transgenic Rice. Plant and Cell Physiology 47: 141–153.
Jia, B., M. Sun, X. Sun, R. Li, Z. Wang, J. Wu, et al. 2016. Overexpression of GsGSTU13 and SCMRP in Medicago sativa confers increased salt–alkaline tolerance and methionine content. Physiologia Plantarum 156: 176–189. https://doi.org/10.1111/ppl.12350.
Kate, M., and N.J. Giles. 2000. Chlorophyll fluorescence—a practical guide. Journal of Experimental Botany 51: 659–668. https://doi.org/10.1093/jexbot/51.345.659.
Kohila, S., and R. Gomathi. 2018. Adaptive physiological and biochemical response of sugarcane genotypes to high-temperature stress. Indian Journal of Plant Physiology 23 (2): 245–260.
Kumari, A., P. Das, A.K. Parida, and P.K. Agarwal. 2015. Proteomics, metabolomics, and ionomics perspectives of salinity tolerance in halophytes. Frontiers in Plant Science 6: 537. https://doi.org/10.3389/fpls.2015.00537.
Levine, A., R. Tenhaken, R. Dixon, and C. Lamb. 1994. H2O2 from the oxidative burst orchestrates the plant hypersensitive disease resistance response. Cell 79: 583–593. https://doi.org/10.1016/0092-8674(94)90544-4.
Li, J., C. Chen, J. Wei, Y. Pan, C. Su, and X. Zhang. 2019. SpPKE1, a multiple stress-responsive gene confers salt tolerance in tomato and tobacco. International Journal of Molecular Sciences 20 (10): 2478. https://doi.org/10.3390/ijms20102478.
Liang, X., L.U. Zhang, S.K. Natarajan, and D.F. Becker. 2013. Proline mechanism of stress survival. Antioxidants and Redox Signaling 19: 998–1011. https://doi.org/10.1089/ars.2012.5074.
Livak, K.J., and T.D. Schmittgen. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2DDC(T) method. Methods 25: 402–408.
Maheshwari, R., and R.S. Dubey. 2009. Nickel-induced oxidative stress and the role of antioxidant defence in rice seedlings. Plant Growth Regulation 59: 37–49. https://doi.org/10.1007/s10725-009-9386-8.
Manimekalai, R., N. Jini, M. Gokul, A. Selvi, R. Gomathi, and R. Arun kumar. 2018a. Biochemical and physiological response to oxidative stress in cultivated sugarcane and wild genera. Indian Journal of Plant Physiology 23: 1–10. https://doi.org/10.1007/s40502-018-0368-6.
Manimekalai, R., N. Jini, M. Gokul, A. Selvi, K. Pradheep, and R. Gomathi. 2018b. Hydrogen peroxide induced oxidative stress in sugarcane and response expression pattern of stress responsive genes through quantitative RT-PCR. Sugar Tech 20: 681–691. https://doi.org/10.1007/s12355-018-0604-4.
Manimekalai, R., N. Jini, M. Gokul, A. Selvi, A. Meena, R. Gomathi, and B. Ram. 2016. Genome wide analysis of NAC gene family ‘sequences’ in sugarcane and its comparative phylogenetic relationship with rice, sorghum, maize and Arabidopsis for prediction of stress associated NAC genes. Agri Gene 3: 1–11. https://doi.org/10.1016/j.aggene.2016.10.003.
Matysik, J., B. Bhalu. Alia, and P. Mohanty. 2002. Molecular mechanisms of quenching of reactive oxygen species by proline under stress in plants. Current Science 82: 525–532.
Mhamdi, A., G. Queval, S. Chaouch, S. Vanderauwera, B.F. Van, and G. Noctor. 2010. Catalase function in plants: A focus on Arabidopsis mutants as stress-mimic models. Journal of Experimental Botany 61 (15): 4197–4220. https://doi.org/10.1093/jxb/erq282.
Mishra, S., A.B. Jha, and R.S. Dubey. 2011. Arsenite treatment induces oxidative stress, upregulates antioxidant system, and causes phytochelatin synthesis in rice seedlings. Protoplasma 248 (3): 565–577. https://doi.org/10.1007/s00709-010-0210-0.
Mittler, R., S. Vanderauwera, N. Suzuki, G. Miller, V.B. Tognetti, and K. Vandepoele. 2011. ROS signaling: The new wave? Trends in Plant Science 16: 300–309. https://doi.org/10.1016/j.tplants.2011.03.007.
Mizoi, J., K. Shinozaki, and K. Yamaguchi-Shinozaki. 2012. AP2/ERF family transcription factors in plant abiotic stress responses. Biochimica et biophysica acta 1819 (2): 86–96. https://doi.org/10.1016/j.bbagrm.2011.08.004.
Mohan, M.M., S.L. Narayana, and S.M. Ibrahim. 2000. Chlorophyll Stability Index (CSI): Its impact on salt tolerance in rice. International Rice Research Notes 25: 38–39.
Nakashima, K., and K. Yamaguchi-Shinozaki. 2006. Regulons in osmotic stress responsive and cold stress-responsive gene expression in plants. Physiologia Plantarum 126: 62–71.
Noctor, G., and C. Foyer. 1998. Ascorbate and glutathione: Keeping active oxygen under control. Annu Reviwe of Plant Physiology 49: 249–279. https://doi.org/10.1146/annurev.arplant.49.1.249.
Paramjeet, K., N. Sharma, M. Kumar and S. Kaur. 2014. Changes in growth and antioxidative enzyme activities in Vicia faba L. seedlings under chromium stress. Indian Journal of Plant Physiology 19(2), 101–106
Pujari, D.S., and S.V. Chanda. 2002. Effect of salinity stress on growth, peroxidase and IAA oxidase activities in vigna seedlings. Acta Physiologia Plantarum 24 (4): 435–439.
Roy, S. 2015. Function of MYB domain transcription factors in abiotic stress and epigenetic control of stress response in plant genome. Plant Signalling and Behaviour 10: 1559–2324. https://doi.org/10.1080/15592324.2015.1117723.
Sanghera, G.S., and A. Kumar. 2018. Recent perspectives towards enhancing drought tolerance in sugarcane. Journal of Plant Science Researh 34 (1): 23–34.
Sharma, P., and R.S. Dubey. 2007. Involvement of oxidative stress and role of antioxidative defense system in growing rice seedlings exposed to toxic concentrations of aluminum. Plant Cell Reports 26 (11): 2027–2038. https://doi.org/10.1007/s00299-007-0416-6.
Shinozaki, K., K. Yamaguchi-Shinozaki, and M. Seki. 2003. Regulatory network of gene expression in the drought and cold stress responses. Current Opinion in Plant Biology 6: 410–417. https://doi.org/10.1016/s1369-5266(03)00092-x
Song, X., Y. Wang, and X. Lv. 2015. Responses of plant biomass, photosynthesis and lipidperoxidation to warming and precipitation change in two dominant species (Stipa grandis and Leymus chinensis) from North China Grasslands. Ecology and Evolution 6 (6): 1871–1882. https://doi.org/10.1002/ece3.1982.
Srivastava, S., and R.S. Dubey. 2011. Manganese-excess induces oxidative stress, lowers the pool of antioxidants and elevates activities of key antioxidative enzymes in rice seedlings. Plant Growth Regulation 64: 1–16. https://doi.org/10.1007/s10725-010-9526-1.
Tanou, G., A. Molassiotis, and G. Diamantidis. 2009. Induction of reactive oxygen species and necrotic death-like destruction in strawberry leaves by salinity. Environmental and Experimental Botany 65 (2–3): 270–281. https://doi.org/10.1016/j.envexpbot.2008.09.005.
Upadhyaya, H., M.H. Khan, S.K. Panda, and S.K. Dhyani. 2007. Hydrogen peroxide induces oxidative stress in detached leaves of Oryza sativa L. Plant Physiology 33 (1–2): 83–95.
Van Toai, T.T., and C.S. Bolles. 1991. Post-anoxic injury in soybean (Glycine max) seedlings. Plant Physiology 9: 588–592.
Vijayalakshmi, D., S. Srividhya, S. Muthulakshmi, and R. Satishraj. 2014. Induction of Oxidative Stress by Hydrogen Peroxide Treatment in Rice Genotypes to Study the Osmolyte Accumulation Pattern and Antioxidant Capacity. Journal of Stress Physiology and Biochemistry 10 (3): 37–46.
Wang, X., H. Han, J. Yan, F. Chen, and W. Wei. 2015. A new AP2/ERF transcription factor from the oil plant Jatropha curcas confers salt and drought tolerance to transgenic tobacco. Applied Biochemistry and Biotechnology 176 (2): 582–597. https://doi.org/10.1007/s12010-015-1597-z.
Xu, W., K. Cui, A. Xu, L. Nie, J. Huang, and S. Peng. 2015. Drought stress condition increases root to shoot ratio via alteration of carbohydrate partitioning and enzymatic activity in rice seedlings. Acta Physiologiae Plantarum 37: 9. https://doi.org/10.1007/s11738-014-1760-0.
Xu, Z., and G.S. Zhou. 2006. Combined effects of waterstress and high temperature on photosynthesis, nitrogenmetabolism and lipid peroxidation of a perennial grass Leymus chinensis. Planta 224: 1080–1090.
Xu, Z., G.S. Zhou, and H. Shimizu. 2009. Effects of soildrought with nocturnal warming on leaf stomatal traits andmesophyll cell ultrastructure of a perennial grass. Crop Science 49: 1843–1851.
Xu, Z., G.S. Zhou, G. Han, and Y. Li. 2011. Photosynthetic potential and its association with lipid peroxidation inresponse to high temperature at different leaf ages in maize. Journal of Plant Growth Regulation 30: 41–50.
Yamaguchi-Shinozaki, K., and K. Shinozaki. 2006. Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annual review of plant biology 57: 781–803. https://doi.org/10.1146/annurev.arplant.57.032905.105444.
You, J., and Z. Chang. 2015. ROS Regulation during abiotic stress response in crop plants. Frontiers in Plant Science. 6: 1092. https://doi.org/10.3389/fpls.2015.01092.
Zheng, X., B. Chen, G. Lu, and B. Han. 2008. Over expression of a NAC transcription factor enhances rice drought and salt tolerance. Biochemical and Biophysical Research Communication 379: 985–989.
Zonglie, H., K. Lakkineni, Z. Zhang, and D.S. Verma. 2000. Removal of feedback inhibition of Δ1-pyrroline-5-carboxylate synthetase results in increased proline accumulation and protection of plants from osmotic stress. Plant Physiology 122: 1129–1136. https://doi.org/10.1104/pp.122.4.1129.
Author information
Authors and Affiliations
Contributions
RM planned and executed the work; JN conducted the experiments and prepared the MS; AS edited the MS, and AK analyzed the data.
Corresponding author
Ethics declarations
Conflict of interest
Authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Narayanan, J., Manimekalai, R., Selvi, A. et al. Physiological, Biochemical and Molecular Responses to Oxidative Stress in Saccharum spontaneum. Sugar Tech 25, 282–293 (2023). https://doi.org/10.1007/s12355-022-01189-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12355-022-01189-1