Abstract
The treatment response of acute lymphoblastic leukemia (ALL) depends on the percentage of lymphoblasts, cytogenetic aberrations, and altered gene expression. The analysis of the gene expression is applicable for determination of risk stratification and prognosis of cancers. c-MYC, P14ARF, MDM2, and P53 play a vital role in cell survival through a functional network. This study aimed to investigate the expression of these genes, also their correlation with immunophenotypic subtypes of ALL and the percentage of blasts. Real-time PCR was performed for the expression analysis of P53, MDM2, c-MYC, and P14ARF in the bone marrow or peripheral blood samples of 52 ALL patients and 13 normal samples as controls. The morphological analysis and flow cytometry were carried out to examine the phenotypes and percentage of lymphoblasts. The decreased expression levels of P53 and MDM2 were seen in ALL patients compared with control group. In T cell subgroup of ALL the expression of P14ARF gene was more decreased among other subgroups. The expression of MDM2 was decreased in ALL patients who were under the age of 16. Based on our study, the interaction between P53 and MDM2 might be more complex and different from reports published in previous studies. Our findings showed that MDM2 is not negatively correlated with P53, at least in our samples. It can be very effective on the current and future studies to use different techniques for analysis of genome, transcriptome, and proteome in definitive risk stratification and prognosis determination.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Acute lymphoblastic leukemia (ALL) is the most frequent leukemia in children [1] and accounts for a quarter of pediatric and adult malignancies. Its survival rate has been dramatically increased to 90% thanks to the research progress recently made in the last few years [2]. The clinical manifestations, age of patients, gene expression profile, the count of white blood cells and initial response to chemotherapy are used for the risk stratification of patients diagnosed with ALL [3]. However, these items cannot be solely regarded as prognostic factors, as the recurrence rate of the disease is about 20% in children [4]. Understanding the mechanisms underlying the disease pathogenesis would shed light on therapeutic strategies adapted for patients with ALL [5]. Determination of transcriptional alterations and characterizations in individuals afflicted with ALL could help physicians to gain an insight into the risk stratification and selection of the best drug with proper doses [4]. For instance, analysis of the P53 transcription profile has been broadly employed for the risk stratification of acute myeloid leukemia [6]. Studies indicated that the P53/MDM2 (or HDM2, Mouse double minute 2 homolog) axis is involved in the cell cycle, proliferation, apoptosis and genomic stability of the cells [7] and the presence of polymorphisms in the P53/MDM2 axis predisposes adults to develop ALL [8]. Activation of the P53 protein is an early apoptotic sign in response to chemotherapeutic agents. So, the rate of P53 activation could be considered a useful index for disease prognosis [9].Unlike most human cancers, P53 is found in wild-type form in ALL, but patients with relapsed and refractory ALL show cytogenetic mutations. However, P53 abnormality is considered a risk factor in both T- and B-ALL and results in very poor survival in adult ALL [9, 10]. The function of P53 is delicately regulated by P14ARF (Alternate Reading Frame) and MDM2. Proteasomal degradation of P53 is primed by MDM2 protein through P53-MDM2 binding while the MDM2 protein is inhibited by P14ARF [7]. The overexpression of MDM2, as well as the ablation of ARF, has been frequently reported in ALL. Studies have indicated that t(12;21)(p13;q22)[ETV6-RUNX1] is the most frequent chromosomal translocation in pediatric ALL in which the P53 signaling pathway is disrupted as a result of MDM2overexpression [7]. This phenomenon shows that the overexpression of MDM2 causes a poor early response to treatment [11]. c-MYC is an oncoprotein, exerting a positive impact on cell growth and proliferation and has an inhibitory effect on the functionality of P53. The overexpression of c-MYC could lead to the differentiation arrest independent of the stimulation of proliferation [12, 13]. P14ARF is a tumor suppressor that is induced by mitogenic stimulation mediated by MYC and Ras. Increased c-MYC activity results in the activation of P14ARF which in turn inhibits the function of MDM2. Also, there is a P53/MDM2 feedback loop in which P53 is capable of activating the MDM2 protein, but MDM2 inactivates the P53 [12]. According to the effects of c-MYC on P14ARF and considering the reciprocal functions of P53 and MDM2 on each other, this study aimed to evaluate the expression of P53, MDM2, P14ARF and c-MYC genes in patients affected by ALL in comparison with control group. The correlation of the gene expression, immunophenotypic subtypes and percent of blast cells in patients was also investigated.
Materials and Methods
Sample Collection
Fifty-two samples including 38 bone marrow samples (BM) and also 14 peripheral blood samples (PB) were obtained from newly diagnosed patients with ALL. The samples were collected from three distinct hospitals, namely Taleghani, Imam Khomeini, and Mofid which all are located in Tehran city. Patients have not undergone chemotherapy, and the diagnosis of malignancy was made by expert oncologists. On the other hand, 13 BM (n:8) and PB (n:5) samples were selected as control group. The control BM samples were collected from patients with lymphoma in whom bone marrow and peripheral blood were not involved by malignant cells. The study was approved by the Ethics Committee of the Shahid Beheshti University of Medical Sciences, and informed consent was obtained from all patients and control group. Considering the age of onset, patients with ALL were divided into two subgroups as follows: under the age of 16 and above the age of 16. The BM and PB samples were collected in ethylene diamine tetra acetic acid (EDTA) -containing tubes. The mononuclear cells were purified by Ficoll–Paque gradient centrifugation.
Translocation Analysis
The real-time polymerase chain reaction (real-time PCR) method was used for the study of four frequent translocations including t(12;21) (TEL/AML1), t(9;22) (BCR/ABL1), t(1;19) (TCF3/PBX1) and t(4;11) (KT2A/AFF1) in samples of all participants performed by the ABI (Thermo-Fisher, USA) instrument.
Immunophenotypic Analysis
Based on flow cytometry analysis of CD markers (listed in supplemental data), the lymphocytes of patients with ALL were divided into T-ALL, Pro-B, Early Pre-B, Pre-B and Mature-B ALL, according to the immunophenotypic subtypes. The PB and BM samples were stained with anti-CD markers’ antibodies (Dako, Denmark) and incubated in the dark at room temperature for 15 min. Then, the red blood cells were lysed by lysis buffer, and the resultant supernatant was discarded. The stained cells were washed and re-suspended by phosphate-buffered saline (PBS). Finally, cells were analyzed by a flow cytometry instrument (Attune, USA).
Gene Expression Analysis
Total RNA (Ribo Nucleic Acid) was extracted from PB and BM samples using the RNeasy kit (Qiagen, USA). NanoDrop (Thermo scientific, USA) was used to assess the quality of extracted RNA and then the extracted RNA was employed for the complementary deoxyribonucleic acid (cDNA) synthesis by the T primed first strand kit (thermo-scientific, USA). PCR and electrophoresis were done for primers to achieve annealing temperature. Electrophoresis of gradient PCR products for P53 and MDM2 and melt curve for MDM2 and P14 are shown in supplementary data. The analysis of gene expression was carried out by the ABI Step-One Plus thermocycler (Applied Biosystems) in duplicate, and the obtained data were analyzed by the 2−ΔΔCT method. The primers are listed in supplementary data.
Statistical Analysis
The differences between the values of ALL and control group were analyzed by the T test. Mann–Whitney U test and one-way ANOVA were used by the SPSS software version 24 for comparison among subgroups. The level of statistical significance was accepted when the p-value was less than 0.05.
Results
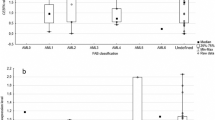
Sixty-five samples were analyzed (52 and 13 samples from ALL patients and control group respectively) and their demographic characteristics were depicted in Table 1. The results showed that there was no significant difference between the age of ALL patients and control group. Among patients with ALL, 61.5% of individuals were above 16 years of age and 38.5% under the age of 16. Chromosomal translocations namely t(9;22), t(1;19), t(12;21), t(4;11) occurred in six, three, six, and one (totally 16) patients, respectively. The frequencies of ALL immunophenotypic subtypes are shown in Table 2. Differences of gene expression among various immunophenotypic subtypes of ALL were also done and results are shown in supplementary information. The morphological analysis revealed that in most of the patients with ALL the blast cells constitute about 80–100% of cell population.
The gene expression analysis showed that the expression levels of P53 and MDM2 were decreased in ALL patients compared with control group (Fig. 1). According to the percentage of lymphoblast, the patients were divided into four groups; however, the rate of gene expression for each of the genes mentioned above was not significantly different among the groups as shown in Supplementary data. The correlation of the gene expression and the translocations was assessed, but the number of samples that underwent translocations was not sufficient to draw any conclusion from the obtained values. However, our findings showed that the rate of MDM21 expression in t(12;21)-positive cells was considerably lower than cells positive for t(9;22). Accordingly, the expression rate of P14ARF gene was significantly decreased in t(12;21)-positive cells compared with cells positive for t(1;19) (Fig. 2). Since there was only one sample with t(4;11), this was excluded from statistical analysis. The analysis of gene expression in all subgroups of ALL patients showed a significant reduction in the expression level of P14ARF gene in T-ALL samples (Supplementary data). Of note, a significant decline in the expression of MDM2 in patients who were above 16 years old was observed when compared with those under the age of 16.
Discussion
The expression of P53 gene is associated with the cell cycle of myeloblasts and the cellular internal and external messages in acute myeloid leukemia (AML) [14]. As well as cytogenetic instability and mutation in proto-oncogenes due to chemical toxic materials [15, 16] which can affect P53 expression [17]. It has been thought that the percentage of lymphoblasts is linked with the status of the cell cycle in ALL patients. However,we showed that the expression rate of P53 gene in cell population with more than 80% blast cells is insignificantly higher than those with less than 80% blasts which might be due to the compensatory mechanism, as the expression of tumor suppressors has been associated with the cell differentiation in AML [18]. Clonal aberrations have been frequently reported in 57–82% and 60–79% of children and adult cases diagnosed with ALL, respectively [19]. The prevalence of translocations in ALL is shown in Suplementary data. An increase in the expression of MDM2 in t(12;21)-positive cells could result in a reduction in the activity of P53 protein. In opposed to our results, it has been reported that MDM2 expression increases in ALL, particularly in blast cells positive for t(12;21) [20]. It has been postulated that MDM2 can decrease the functionality of P53 protein in four ways:
-
1.
MDM2 is capable of binding directly to P53 gene and inhibiting its transcription [11].
-
2.
MDM2 facilitates the degradation of P53 protein through the ubiquitination process [11].
-
3.
MDM2 binds to P53 and facilitates the egress of P53 protein from the nucleus [11].
-
4.
MDM2 distrupts the translation of P53 protein [21].
It is evident that one of the biological functions of MDM2 is to decrease P53 expression. However, the overexpression of P53 is not mandatory when the expression of MDM2 is low and vice versa; because the downregulation of P53 (as a result of genetic mutations or methylation) might result in the downregulation of MDM2 (P53 is a positive regulator of MDM2). In other words, P53 serves as a transcription factor for MDM2 and creates an auto-regulatory feedback loop [22]. After 12 h of glucocorticoid (GC) therapy, an increase in the expression of both P53 and MDM2 would be evident in ALL cells. It is likely that the elevation of MDM2 expression is secondary to the increased level of P53 and thought to be caused by a poor response to initial GC therapy [11]. Therefore, it would be plausible that a decrease in the expression of MDM2 could be secondary to the reduction of P53 in our study.
The overexpression of MDM2 has been reported as average in 7% of human cancers (in 20% and 16% of soft tissue carcinoma and osteosarcoma cases, respectively and a little percent of leukemia, lymphoma, and pancreatic carcinoma). Also, there is a negative correlation between P53 mutation and MDM2 expression [21]. Increased MDM2 activity in t(9;22) positive ALL can be due to the higher expression of MDM2 in this translocation [23], as our results showed in comparison with its expression in t(12;21), though the number of translocation positive samples was not enough for definitive statistical analysis. Although the rate of P53 mutation is low in ALL [24], the downregulation of MDM2 might stem from P53 mutation in our samples, requiring further analysis to be validated. Analysis of the MDM2 expression, chromosomal translocations, immunophenotypic subtypes, and blast cell count may be useful for determination of prognosis, risk stratification, prevention of relapse and side effects of therapy in ALL but the current research revealed contradictory results about the activity of MDM2 in ALL. It seems that the evaluation of gene expression would not be enough for making a decision on the function of MDM2 and fate of cells in ALL. On the other hand, there are some targeted therapies in ALL against MDM2 pathway and MDM2 effects on P53, such as Nilotinib (tyrosine kinase and MDM2 inhibitor) and Nutlin (by antagonizing Mdm2-p53 binding role) [7], and a randomized preclinical on NVP-CGM097 as an inhibitor of MDM2 in B-ALL [25]. Thus cognition of the expression rate of MDM2 and P53 may be helpful in the decision to use of the drugs.
c-MYC is considered as an amplifier agent for most of the active genes through multiple pathways. The overexpression of c-MYC enhances cell cycle progression, nucleotide biosynthesis, energy production and protein synthesis, and could also lead to the inhibition of the differentiation process [26]. In addition to Burkitt lymphoma, the translocation of c-MYC is also reported in 6% of cases with T-ALL. Also, rearrangement of c-MYC has been observed in 2-5% of patients diagnosed with ALL [26]. In our study, c-MYC expression was increased in ALL patients in comparison with the control group, however such an increase was not statistically significant. c-MYC has been recognized as a first proto-oncogene, but it is the first oncogene that activates the P14ARF in a tumor-suppressive manner [27]. The activation of P14ARF mediated by c-MYC results in the inactivation of the inhibitory effect of MDM2 protein on P53. On the other hand, P53 is capable of inhibiting c-MYC at the transcription level. Thus, P53 and c-MYC are able to control each other, as well as the proliferation and survival of the cells [28].
P14ARF is a checkpoint protein by P53-independent function and is effective on the expression of c-MYC [29]. Increased expression of P14ARFversus reduced expression of c-MYC in our study suggests that P14ARF has inhibitory effects on the expression of c-MYC. In mice knocked out for the P19ARF gene (P19ARF is the murine homolog of human P14ARF), ALL does not develop; however, the t(12;21) along with the ablation of P14ARF could lead to the development of ALL in mice [30]. The present study revealed significant downregulation of P14ARF in t(12;21) positive samples that can indicate the deletion of P14ARF gene. Also significant downregulation of P14ARF in T-ALL was seen in comparison with other immunophenotypic subtypes of ALL in our study. P14ARFgene deletion leads to a reduction in the activity of P53 and occurrs in 20% and 70% of cases diagnosed with B-ALL and T-ALL, respectively [30, 31]. Loss of p16INK4A and p14ARF in T cells and activated NOTCH signaling are the main mechanisms of T-ALL pathogenesis [31].
As a conclusion, it appears that the nature of the correlation between P53 and MDM2 proteins, as well as their effects on each other, might be different from reports published in previous studies. Our findings showed that MDM2 is not negatively correlated with P53, at least in our samples. Therefore, further studies with different techniques at the levels of genome, transcriptome, and proteome are warranted to shed light on the precise mechanisms underlying the relationship between these two genes.
References
Allahbakhshian MF, Kamel M, Mehrpouri M, Heris R, Hamidpour M, Salari S et al (2018) The expression of interferon gamma (IFN-γ) and interleukin 6 (IL6) in patients with acute lymphoblastic leukemia (ALL). Pathol Oncol Res POR. https://doi.org/10.1007/s12253-018-0536-z
Tomizawa D, Kiyokawa N (2017) Acute lymphoblastic leukemia, hematological disorders in children. Springer, Berlin, pp 33–60
Terwilliger T, Abdul-Hay M (2017) Acute lymphoblastic leukemia: a comprehensive review and 2017 update. Blood Cancer J 7(6):e577
Bhojwani D, Kang H, Menezes RX, Yang W, Sather H, Moskowitz NP et al (2008) Gene expression signatures predictive of early response and outcome in high-risk childhood acute lymphoblastic leukemia: a Children’s Oncology Group Study. J Clin Oncol 26(27):4376–4384
Khosravi MR, Mohammadi MH, Khadem P, Lashkari S, Aghaeenezhad H, Gharehbaghian A et al (2018) Evaluation of P21Cip1 and P27Kip1 expression in de novo acute lymphoblastic leukemia patients. Biomed Res Ther 5(7):2518–2527
Ahmadzadeh A, Mohammadi MH, Mezginezhad F, Nezhad HA, Parkhideh S, Khosravi M et al (2018) The expression of the TP53 gene in various classes of acute myeloid leukemia. World Cancer Res J 5(4):e1178
Trino S, De Luca L, Laurenzana I, Caivano A, Del Vecchio L, Martinelli G et al (2016) P53-MDM2 pathway: evidences for a new targeted therapeutic approach in B-acute lymphoblastic leukemia. Front Pharmacol 7:491–497
Chen J, Zhu B, Chen J, Li Y (2013) Genetic variations in MDM2 and P53 genes confer risk for adult acute lymphoblastic leukemia in a Chinese population. DNA Cell Biol 32(7):414–419
Bainer RO, Trendowski MR, Cheng C, Pei D, Yang W, Paugh SW et al (2017) A p53-regulated apoptotic gene signature predicts treatment response and outcome in pediatric acute lymphoblastic leukemia. Cancer Manag Res 9:397–410
Salmoiraghi S, Montalvo MLG, Ubiali G, Tosi M, Peruta B, Zanghi P et al (2016) Mutations of TP53 gene in adult acute lymphoblastic leukemia at diagnosis do not affect the achievement of hematologic response but correlate with early relapse and very poor survival. Haematologica. 101(6):e245
Ociepa T, Maloney E, Kamieńska E, Wysocki M, Kurylak A, Matysiak M et al (2010) Simultaneous assessment of p53 and MDM2 expression in leukemic cells in response to initial prednisone therapy in children with acute lymphoblastic leukemia. Pol J Pathol 61(4):199–205
Allen A, Gill K, Hoehn D, Sulis M, Bhagat G, Alobeid B (2014) C-myc protein expression in B-cell acute lymphoblastic leukemia, prognostic significance? Leuk Res 38(9):1061–1066
Rafiee M, Keramati MR, Ayatollahi H, Sadeghian MH, Barzegar M, Asgharzadeh A et al (2016) Down-regulation of ribosomal S6 kinase RPS6KA6 in acute myeloid leukemia patients. Cell J (Yakhteh) 18(2):159–164
Müller-Tidow C, Metzelder S, Buerger H, Packeisen J, Ganser A, Heil G et al (2004) Expression of the p14 ARF tumor suppressor predicts survival in acute myeloid leukemia. Leukemia 18(4):720–726
Ayatollahi H, Rafiee M, Keramati M-R, Balali-Mood M, Asgharzadeh A, Sadeghian MH et al (2015) Lack of FLT3-TKD835 gene mutation in toxicity of sulfur mustard in Iranian veterans. Iran J Basic Med Sci 18(9):862–866
Mahvi A, Mardani G, Ghasemi-Dehkordi P, Saffari-Chaleshtori J, Hashemzadeh-Chaleshtori M, Allahbakhshian-Farsani M et al (2015) Effects of phenanthrene and pyrene on cytogenetic stability of human dermal fibroblasts using alkaline comet assay technique. Proc Natl Acad Sci India Sect B Biol Sci 85(4):1055–1063
Salvadori DMF, da Silva GN (2013) Genetic instability in normal-appearing and tumor urothelium cells and the role of the TP53 gene in the toxicogenomic effects of antineoplastic drugs. Advances in the scientific evaluation of bladder cancer and molecular basis for diagnosis and treatment. IntechOpen, Rijeka
Salarpour F, Goudarzipour K, Mohammadi MH, Ahmadzadeh A, Faraahi S, Farsani MA (2017) Evaluation of CCAAT/enhancer binding protein (C/EBP) alpha (CEBPA) and runt-related transcription factor 1 (RUNX1) expression in patients with de novo acute myeloid leukemia. Ann Hum Genet 81(6):276–283
Mrozek K, Harper DP, Aplan PD (2009) Cytogenetics and molecular genetics of acute lymphoblastic leukemia. Hematol/Oncol Clin 23(5):991–1010
Kaindl U, Morak M, Portsmouth C, Mecklenbräuker A, Kauer M, Zeginigg M et al (2014) Blocking ETV6/RUNX1-induced MDM2 overexpression by Nutlin-3 reactivates p53 signaling in childhood leukemia. Leukemia 28(3):600–608
Bohlman S, Manfredi JJ (2014) p53-independent effects of Mdm2. Mutant p53 and MDM2 in cancer. Springer, Berlin, pp 235–246
Agrawal A, Yang J, Murphy RF, Agrawal DK (2006) Regulation of the p14ARF-Mdm2-p53 pathway: an overview in breast cancer. Exp Mol Pathol 81(2):115–122
Bernt KM, Hunger SP (2014) Current concepts in pediatric Philadelphia chromosome-positive acute lymphoblastic leukemia. Front Oncol 4:54
Iacobucci I, Erriquez D, Ferrari A, Papayannidis C, Venturi C, Trino S et al (2012) Down-regulation of BMI-1 Is a new marker of sensitivity to Mdm2 inhibition in B-acute lymphoblastic leukemia. Blood 120(21):2522
Townsend EC, DeSouza T, Murakami MA, Montero J, Stevenson K, Christie AL et al (2015) The MDM2 inhibitor NVP-CGM097 is highly active in a randomized preclinical trial of B-cell acute lymphoblastic leukemia patient derived xenografts. Blood 126(23):797
Cortiguera Ruiz MG, Batlle López A, Albajar Molera M, Delgado Villar MD, León Serrano J (2015) MYC as therapeutic target in leukemia and lymphoma. Blood Lymph Cancer Targets Ther 5:75–91
Amente S, Gargano B, Varrone F, Ruggiero L, Dominguez-Sola D, Lania L et al (2006) p14ARF directly interacts with Myc through the Myc BoxII domain. Cancer Biol Ther 5(3):287–291
Madapura HS, Salamon D, Wiman KG, Lain S, Klein E, Nagy N (2016) cMyc-p53 feedback mechanism regulates the dynamics of T lymphocytes in the immune response. Cell Cycle 15(9):1267–1275
Sarkar D, Fisher PB (2006) Regulation of Myc function by ARF: checkpoint for Myc-induced oncogenesis. Cancer Biol Ther 5(6):693–695
Bernardin F, Yang Y, Cleaves R, Zahurak M, Cheng L, Civin CI et al (2002) TEL-AML1, expressed from t (12; 21) in human acute lymphocytic leukemia, induces acute leukemia in mice. Can Res 62(14):3904–3908
Van Vlierberghe P, Ferrando A (2012) The molecular basis of T cell acute lymphoblastic leukemia. J Clin Investig 122(10):3398–3406
Acknowledgements
The authors would like to thank all patients for their participation in this study. This study was supported by a research grant provided by the Shahid Beheshti University of Medical Sciences.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Allahbakhshian Farsani, M., Rafiee, M., Aghaee Nezhad, H. et al. The Expression of P53, MDM2, c-myc, and P14ARF Genes in Newly Diagnosed Acute Lymphoblastic Leukemia Patients. Indian J Hematol Blood Transfus 36, 277–283 (2020). https://doi.org/10.1007/s12288-019-01214-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12288-019-01214-6