Abstract
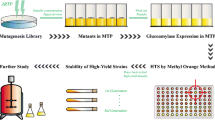
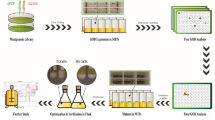
α-Glucosidase is a crucial enzyme for the production of isomaltooligosaccharide. In this study, a novel method comprising eosin Y (EY) and α-d-methylglucoside (AMG) in glass plates was tested for the primary screening of α-glucosidaseproducing strains. First, α-glucosidase-producing Aspergillus niger strains were selected on plates containing EY and AMG based on transparent zone formation resulting from the solubilization of EY by the hydrolyzed product. Conventional methods that use trypan blue (TB) and p-nitrophenyl-α-d-glucopyranoside (pPNP) as indicators were then compared with the new strategy. The results showed that EY-containing plates provide the advantages of low price and higher specificity for the screening of α-glucosidase-producing strains. We then evaluated the correlation between the hydrolytic activity of α-glucosidase and diffusion distance, and found that good linearity could be established within a 6–75 U/ml enzyme concentration range. Finally, the hydrolytic and transglycosylation activities of α-glucosidase obtained from the target isolates were determined by EY plate assay and 3,5-dinitrosalicylic acid-Saccharomyces cerevisiae assay, respectively. The results showed that the diameter of the transparent zone varied among isolates was positively correlated with α-glucosidase hydrolytic activity, while good linearity could also be established between α-glucosidase transglycosylation activity and non-fermentable reducing sugars content. With this strategy, 7 Aspergillus niger mutants with high yield of α-glucosidase from 200 obvious single colonies on the primary screen plate were obtained.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Basu, A., Mutturi, S., and Prapulla, S.G. 2016. Production of isomaltooligosaccharides (IMO) using simultaneous saccharification and transglucosylation from starch and sustainable sources. Process Biochem.51, 1464–1471.
Bissaro, B., Monsan, P., Faure, R., and O’Donohue, M.J. 2015. Glycosynthesis in a waterworld: new insight into the molecular basis of transglycosylation in retaining glycoside hydrolases. Biochem. J.467, 17–35.
Charron, M.J., Dubin, R.A., and Michels, C.A. 1986. Structural and functional analysis of the MAL1 locus of Saccharomyces cerevisiae. Mol. Cell Biol.6, 3891–3899.
Chen, G.G., Li, W., Zhang, Y.K., Qin, Y.L., Wu, K.Y., and Liang, Z.Q. 2011. A high-throughput method for screening of Aspergillus niger mutants with high transglycosylation activity by detecting nonfermentable reducing sugar. World J. Microbiol. Biotechnol.27, 1519–1523.
Coleri, A., Cokmus, C., Ozcan, B., Akkoc, N., and Akcelik, M. 2009. Isolations of α-glucosidase-producing thermophilic bacilli from hot springs of Turkey. Mikrobiologiia78, 68–78.
Dej-Adisai, S. and Pitakbut, T. 2015. Determination of α-glucosidase inhibitory activity from selected Fabaceae plants. Pak. J. Pharm. Sci.28, 1679–1683.
Fernández-Arrojo, L., Marín, D., Gómez De Segura, A., Linde, D., Alcalde, M., Gutiérrez-Alonso, P., Ghazi, I., Plou, F.J., Fernández-Lobato, M., and Ballesteros, A.O. 2007. Transformation of maltose into prebiotic isomaltooligosaccharides by a novel α-glucosidase from Xantophyllomyces dendrorhous. Process Biochem.42, 1530–1536.
Ganzlin, M. and Rinas, U. 2008. In-depth analysis of the Aspergillus niger glucoamylase (glaA) promoter performance using high-throughput screening and controlled bioreactor cultivation techniques. J. Biotechnol.135, 266–271.
Gutiérrez-Alonso, P., Gimeno-Pérez, M., Ramírez-Escudero, M., Plou, F.J., Sanz-Aparicio, J., and Fernández-Lobato, M. 2016. Molecular characterization and heterologous expression of a Xanthophyllomyces dendrorhous α-glucosidase with potential for prebiotics production. J. Appl. Microbiol.100, 3125–3135.
Hleap, J.S. and Blouin, C. 2018. The response to selection in Glycoside Hydrolase Family 13 structures: A comparative quantitative genetics approach. PLoS One13, e0196135.
Hostinová, E., Solovicová, A., and Gašperík, J. 2005. Cloning and expression of a gene for an α-glucosidase from Saccharomycopsis fibuligera homologous to family GH31 of yeast glucoamylases. Appl. Microbiol. Biotechnol.69, 51–56.
Hu, X., Xiao, Y., Wu, J., and Ma, L. 2011. Evaluation of polyhydroxybenzophenones as α-glucosidase inhibitors. Arch. Pharm.344, 71–77.
Jauregui, H.O., Hayner, N.T., Driscoll, J.L., Williams-Holland, R., Lipsky, M.H., and Galletti, P.M. 1981. Trypan blue dye uptake and lactate dehydrogenase in adult rat hepatocytes — Freshly isolated cells, cell suspensions, and primary monolayer cultures. In Vitro17, 1100–1110.
Jin, Y., Peng, L.Q., and Zhao, A.L. 2018. Hyperoxia induces the apoptosis of alveolar epithelial cells and changes of pulmonary surfactant proteins. Eur. Rev. Med. Pharmacol. Sci.22, 492–497.
Jones, G.R., Cundall, R.B., Murray, D., and Duddell, D.A. 1984. Eosin Y — Macromolecule Complexes. Part 2. — Interactions between eosin Y and polycations, a cationic surfactant and proteins. J. Chem. Soc. Faraday Trans. 280, 1201–1213.
Krohn, B.M. and Lindsay, J.A. 1991. Purification and characterization of a thermostable α-glucosidase from a Bacillus subtilis high-temperature growth transformant. Curr. Microbiol.22, 273–278.
Krolicka, M., Hinz, S.W.A., Koetsier, M.J., Eggink, G., van den Broek, L.A.M., and Boeriu, C.G. 2018. β-N-Acetylglucosaminidase MthNAG from Myceliophthora thermophila C1, a thermostable enzyme for production of N-acetylglucosamine from chitin. Appl. Microbiol. Biotechnol.102, 7441–7454.
Larsbrink, J., Izumi, A., Hemsworth, G.R., Davies, G.J., and Brumer, H. 2012. Structural enzymology of Cellvibrio japonicus Agd31B protein reveals α-transglucosylase activity in glycoside hydrolase family 31. J. Biol. Chem.287, 43288–43299.
Lee, S.S., He, S., and Withers, S.G. 2001. Identification of the catalytic nucleophile of the Family 31 α-glucosidase from Aspergillus niger via trapping of a 5-fluoroglycosyl-enzyme intermediate. Biochem. J.359, 381–386.
Li, Y., Bu, M., Chen, P., Li, X., Chen, C., Gao, G., Feng, Y., Han, W., and Zhang, Z. 2018. Characterization of a thermophilic monosaccharide stimulated β-glucosidase from Acidothermus cellulolyticus. Chem. Res. Chin. Univ.34, 212–220.
Li, D., Liang, L., Tang, Y., Fu, L., Xiao, S., and Yuan, Q. 2019. Direct and single-step sensing of primary ovarian cancers related glycosidases. Chin. Chem. Lett.30, 1013–1016.
Lin, F., Fan, W., and Wise, G.E. 1991. Eosin Y staining of proteins in polyacrylamide gels. Anal. Biochem.196, 279–283.
Ma, X.D., Ke, T., Xiong, L., Yan, H., and Ma, L.X. 2007. A new plate method for screening of polysaccharide-degrading enzymes and their producing microorganisms. Acta Microbiol. Sin.47, 1102–1104.
Ma, X.D., Ma, L.X., Xue, Z.F., and Zhou, J.C. 2000. A new method to determine α-amylase activity and its producing bacteria. J. Huazhong Agri. Univ.19, 456–460.
Margesin, R., Gander, S., Zacke, G., Gounot, A.M., and Schinner, F. 2003. Hydrocarbon degradation and enzyme activities of cold-adapted bacteria and yeasts. Extremophiles7, 451–458.
Marín, D., Linde, D., and Fernández-Lobato, M. 2006. Purification and biochemical characterization of an α-glucosidase from Xanthophyllomyces dendrorhous. Yeast23, 117–125.
Nakao, M., Nakayama, T., Harada, M., Kakudo, A., Ikemoto, H., Kobayashi, S., and Shibano, Y. 1994. Purification and characterization of a Bacillus sp. SAM1606 thermostable α-glucosidase with transglucosylation activity. Appl. Microbiol. Biotechnol.41, 337–343.
Nishikawa, M. and Ogawa, K. 2002. Distribution of microbes producing antimicrobial epsilon-poly-L-lysine polymers in soil microflora determined by a novel method. Appl. Environ. Microbiol.68, 3575–3581.
Nwokoro, O. and Anthonia, O. 2015. Studies on the production of alkaline α-amylase from Bacillus subtilis CB-18. Acta Sci. Pol. Technol. Aliment.14, 71–75.
Pan, Y.C. and Lee, W.C. 2005. Production of high-purity isomaltooligosaccharides syrup by the enzymatic conversion of transglucosidase and fermentation of yeast cells. Biotechnol. Bioeng.89, 797–804.
Saha, B.C. and Zeikus, J.G. 1991. Characterization of thermostable α-glucosidase from Clostridium thermohydrosulfuricum 39E. Appl. Microbiol. Biotechnol.35, 568–571.
Shimba, N., Shinagawa, M., Hoshino, W., Yamaguchi, H., Yamada, N., and Suzuki, E. 2009. Monitoring the hydrolysis and transglycosylation activity of α-glucosidase from Aspergillus niger by nuclear magnetic resonance spectroscopy and mass spectrometry. Anal. Biochem.393, 23–28.
Song, K.M., Okuyama, M., Nishimura, M., Tagami, T., Mori, H., and Kimura, A. 2013. Aromatic residue on β→α loop 1 in the catalytic domain is important to the transglycosylation specificity of glycoside hydrolase family 31 α-glucosidase. Biosci. Biotechnol. Biochem.77, 1759–1765.
Strober, W. 2001. Trypan blue exclusion test of cell viability. Curr. Protoc. Immunol.21, A.3B.1–A.3B.2.
Suzuki, Y., Kishigami, T., and Abe, S. 1976. Production of extracellular α-glucosidase by a thermophilic Bacillus species. Appl. Environ. Microbiol.31, 807–812.
Tagami, T., Yamashita, K., Okuyama, M., Mori, H., Yao, M., and Kimura, A. 2015. Structural advantage of sugar beet α-glucosidase to stabilize the Michaelis complex with long-chain substrate. J. Biol. Chem.290, 1796–1803.
Tanaka, Y., Aki, T., Hidaka, Y., Furuya, Y., Kawamoto, S., Shigeta, S., Ono, K., and Suzuki, O. 2002. Purification and characterization of a novel fungal α-glucosidase from Mortierella alliacea with high starch-hydrolytic activity. Biosci. Biotechnol. Biochem.66, 2415–2423.
Tang, C., Qian, Z., Qian, Y., Huang, Y., Zhao, M., Ao, H., and Feng, H. 2017. A fluorometric and real-time assay for α-glucosidase activity through supramolecular self-assembly and its application for inhibitor screening. Sens. Actuators B Chem.245, 282–289.
Tennant, J.R. 1964. Evaluation of the trypan blue technique for determination of cell viability. Transplantation2, 685–694.
Zeng, W., Lin, Y., Qi, Z., He, Y., Wang, D., Chen, G., and Liang, Z. 2013. An integrated high-throughput strategy for rapid screening of poly (γ-glutamic acid)-producing bacteria. Appl. Microbiol. Biotechnol.97, 2163–2172.
Zhang, Y.K., Li, W., Wu, K.Y., Chen, G.G., and Liang, Z.Q. 2011. Purification and characterization of an intracellular α-glucosidase with high transglycosylation activity from A. niger M-1. Prep. Biochem. Biotechnol.41, 201–217.
Zhang, L., Su, Y., Zheng, Y., Jiang, Z., Shi, J., Zhu, Y., and Jiang, Y. 2010. Sandwich-structured enzyme membrane reactor for efficient conversion of maltose into isomaltooligosaccharides. Bioresour. Technol.101, 9144–9149.
Zhou, C., Xue, Y., Zhang, Y., Zeng, Y., and Ma, Y. 2009. Recombinant expression and characterization of Thermoanaerobacter tengcongensis thermostable α-glucosidase with regioselectivity for high yield isomaltooligosaccharides synthesis. J. Microbiol. Biotechnol.19, 1547–1556.
Acknowledgments
This work was supported by the National Natural Science Foundation of China (31560448, 31760452); the Natural Science Foundation of Guangxi (2016GXNSFAA380130,).
Author information
Authors and Affiliations
Corresponding author
Additional information
Conflict of Interest
The authors declare that have not any conflict of interest in this work.
Rights and permissions
About this article
Cite this article
Zhou, B., Huang, N., Zeng, W. et al. Development of a strategy for the screening of α-glucosidase-producing microorganisms. J Microbiol. 58, 163–172 (2020). https://doi.org/10.1007/s12275-020-9267-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-020-9267-4