Abstract
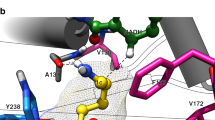
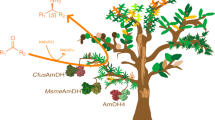
Amine dehydrogenases (AmDHs) are one of the key emerging enzymes used in the synthesis of various amines with the expense of only one ammonium ion as an amino donor, thereby, generates only water molecule as a by-product. Currently, most available AmDHs have been created through protein engineering using the existing natural L-amino acid dehydrogenase, and native AmDHs are rarely reported. In this study, a novel native AmDH from Laribacter hongkongensis (LhAmDH) was identified based on the GenBank database using a sequence-driven approach. LhAmDH showed a good activity towards various carbonyl compounds such as cyclohexanone (170 mU/mg) and isovaleraldehyde (214 mU/mg). The reductive amination of model substrate, cyclohexanone (up to 100 mM) into cyclohexylamine was successfully performed in LhAmDH and FDH system with > 99% conversion using Escherichia coli whole-cell as well as purified enzymes. Furthermore, three enzymes cascade (ω-transaminase, LhAmDH, and FDH) was designed to produce chiral amine from the corresponding ketone using inexpensive ammonium formate as sole sacrificial agent. The active site of LhAmDH residues were predicted using the protein structure homology model building program SWISS-MODEL server. In the docking analysis, cyclohexanone is well-orientated with −5.4 kcal/mol of binding energy and 3.16 Å distance from side chain of E104, which is a key residue for interacting ammonia and substrate. This LhAmDH model can be used as a promising template to produce chiral amines through semi-rational design.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Patil, M. D., G. Grogan, A. Bommarius, and H. Yun (2018) Recent advances in ω-transaminase-mediated biocatalysis for the enantioselective synthesis of chiral amines. Catalysts. 8: 254.
Nugent, T. C. and M. El-Shazly (2010) Chiral amine synthesis — Recent developments and trends for enamide reduction, reductive amination, and imine reduction. Adv. Synth. Catal. 352: 753–819.
Jeon, H., S. Yoon, M. Ahsan, S. Sung, G. H. Kim, U. Sundaramoorthy, S. K. Rhee, and H. Yun (2017) The kinetic resolution of racemic amines using a whole-cell biocatalyst co-expressing amine dehydrogenase and NADH oxidase. Catalysts. 7: 251.
Patil, M. D., G. Grogan, A. Bommarius, and H. Yun (2018) Oxidoreductase-catalyzed synthesis of chiral amines. ACS Catal. 8: 10985–11015.
Knaus, T., W. Böhmer, and F. G. Mutti (2017) Amine dehydrogenases: efficient biocatalysts for the reductive amination of carbonyl compounds. Green Chem. 19: 453–463.
Tseliou, V., T. Knaus, M. F. Masman, M. L. Corrado, and F. G. Mutti (2019) Generation of amine dehydrogenases with increased catalytic performance and substrate scope from e-deaminating L-Lysine dehydrogenase. Nat. Commun. 10: 3717.
Bommarius, A. (2019) Amine dehydrogenases occur in nature. Nat. Catal. 2: 288–289.
Itoh, N., C. Yachi, and T. Kudome (2000) Determining a novel NAD+-dependent amine dehydrogenase with a broad substrate range from Streptomyces virginiae IFO 12827: purification and characterization. J. Mol. Catal. B Enzym. 10: 281–290.
Mayol, O., K. Bastard, L. Beloti, A. Frese, J. P. Turkenburg, J. L. Petit, A. Mariage, A. Debard, V. Pellouin, A. Perret, V. de Berardinis, A. Zaparucha, G. Grogan, and C. Vergne-Vaxelaire (2019) A family of native amine dehydrogenases for the asymmetric reductive amination of ketones. Nat. Catal. 2: 324–333.
Mayol, O., S. David, E. Darii, A. Debard, A. Mariage, V. Pellouin, J. L. Petit, M. Salanoubat, V. de Berardinis, A. Zaparucha, and C. Vergne-Vaxelaire (2016) Asymmetric reductive amination by a wild-type amine dehydrogenase from the thermophilic bacteria Petrotoga mobilis. Catal. Sci. Technol. 6: 7421–7428.
Ducrot, L., M. Bennett, G. Grogan, and C. Vergne-Vaxelaire (2021) NAD(P)H-dependent enzymes for reductive amination: Active site description and carbonyl-containing compound spectrum. Adv. Synth. Catal. 363: 328–351.
Na, A. R., D. Choi, and H. Cho (2020) Synthesis, structural analysis, and biological evaluation of novel ((2,4-dioxothiazolidin-5-ylidene)methyl)phenyl derivatives. Biotechnol. Bioprocess Eng. 25: 149–163.
Jeon, H., S. Sarak, S. H. Lee, H. S. Bea, M. Patil, G. H. Kim, B. G. Kim, J. I. Won, and H. Yun (2018) Characterization of ELP-fused ω-transaminase and its application for the biosynthesis of β-amino acid. Biotechnol. Bioprocess Eng. 23: 481–489.
Cho, Y. J., J. H. Park, G. Y. Chung, and H. S. Shin (2020) Facile identification and isolation of protease using SDS-PAGE and zymography. Biotechnol. Bioprocess Eng. 25: 164–169.
Nadarajan, S. P., K. Deepankumar, J. H. Seo, and H. Yun (2017) Evaluating the role of puckering and fluorine atom in stability and folding of fluoroproline containing proteins. Biotechnol. Bioprocess Eng. 22: 504–511.
Mathew, S., S. P. Nadarajan, T. Chung, H. H. Park, and H. Yun (2016) Biochemical characterization of thermostable ω-transaminase from Sphaerobacter thermophilus and its application for producing aromatic β- and γ-amino acids. Enzyme Microb. Technol. 87–88: 52–60.
Oh, H. D., M. S. Cho, J. S. Kim, M. S. Kim, C. H. Kim, and J. Y. Kang (2020) Identification and characterization of a cocoon degradable enzyme from the isolated strain Bacillus subtilis Bs5C. Biotechnol. Bioprocess Eng. 25: 442–449.
Cantarel, B. L., H. G. Morrison, and W. Pearson (2006) Exploring the relationship between sequence similarity and accurate phylogenetic trees. Mol. Biol. Evol. 23: 2090–2100.
Qin, S., P. Wang, S. Huang, S. Liu, G. Wang, L. Wang, M. Sun, and X. Wang (2015) An efficient method for the production of cyclohexylamine from cyclohexanone and ammonia over Cu-Cr-La/γ-Al2O3. J. Korean Chem. Soc. 59: 493–498.
Patil, M. D., S. Yoon, H. Jeon, T. P. Khobragade, S. Sarak, A. D. Pagar, Y. Won, and H. Yun (2019) Kinetic resolution of racemic amines to enantiopure (S)-amines by a biocatalytic cascade employing amine dehydrogenase and alanine dehydrogenase. Catalysts. 9: 600.
Jakoblinnert, A. and D. Rother (2014) A two-step biocatalytic cascade in micro-aqueous medium: Using whole cells to obtain high concentrations of a vicinal diol. Green Chem. 16: 3472–3482.
Sheldon, R. A. and D. Brady (2018) The limits to biocatalysis: Pushing the envelope. Chem. Commun. 54: 6088–6104.
Wachtmeister, J. and D. Rother (2016) Recent advances in whole cell biocatalysis techniques bridging from investigative to industrial scale. Curr. Opin. Biotechnol. 42: 169–177.
Molnár, Z., E. Farkas, Á. Lakó, B. Erdélyi, W. Kroutil, B. G. Vértessy, C. Paizs, and L. Poppe (2019) Immobilized whole-cell transaminase biocatalysts for continuous-flow kinetic resolution of amines. Catalysts. 9: 438.
Gomm, A. and E. O’Reilly (2018) Transaminases for chiral amine synthesis. Curr. Opin. Chem. Biol. 43: 106–112.
Ruggieri, F., L. M. van Langen, D. T. Logan, B. Walse, and P. Berglund (2018) Transaminase-catalyzed racemization with potential for dynamic kinetic resolutions. ChemCatChem. 10: 5012–5018.
Costa, B. Z., J. L. Galman, I. Slabu, S. P. France, A. J. Marsaioli, and N. J. Turner (2018) Synthesis of 2,5-disubstituted pyrrolidine alkaloids via a one-pot cascade using transaminase and reductive aminase biocatalysts. ChemCatChem. 10: 4733–4738.
Slabu, I., J. L. Galman, R. C. Lloyd, and N. J. Turner (2017) Discovery, engineering, and synthetic application of transaminase biocatalysts. ACS Catal. 7: 8263–8284.
Shin, G., S. Mathew, M. Shon, B. G. Kim, and H. Yun (2013) One-pot one-step deracemization of amines using m-transaminases. Chem. Commun. 49: 8629–8631.
Patil, M. D., G. Grogan, and H. Yun (2018) Biocatalyzed C-C bond formation for the production of alkaloids. ChemCatChem. 10: 4783–4804.
Kelly, S. A., S. Pohle, S. Wharry, S. Mix, C. C. R. Allen, T. S. Moody, and B. F. Gilmore (2018) Application of ω-transaminases in the pharmaceutical industry. Chem. Rev. 118: 349–367.
Ferrandi, E. E. and D. Monti (2018) Amine transaminases in chiral amines synthesis: recent advances and challenges. World J. Microbiol. Biotechnol. 34: 13.
Kim, H., E. Kim, I. Lee, B. Bae, M. Park, and H. Nam (2020) Artificial intelligence in drug discovery: A comprehensive review of data-driven and machine learning approaches. Biotechnol. Bioprocess Eng. 25: 895–930.
Guex, N. and M. C. Peitsch (1997) SWISS-MODEL and the Swiss-PdbViewer: An environment for comparative protein modeling. Electrophoresis. 18: 2714–2723.
Acknowledgements
This research was supported by the Basic Science Research Program of the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT, and future Planning (2020R1A2C2009806) and supported by Konkuk University Researcher Fund in 2019.
The authors declare no conflict of interest.
Neither ethical approval nor informed consent was required for this study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lee, S., Jeon, H., Giri, P. et al. The Reductive Amination of Carbonyl Compounds Using Native Amine Dehydrogenase from Laribacter hongkongensis. Biotechnol Bioproc E 26, 384–391 (2021). https://doi.org/10.1007/s12257-021-0113-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12257-021-0113-2