Summary
Molecular profiling of circulating tumor DNA (ctDNA) to guide treatment decisions has found its way into routine management of patients with advanced cancer. This represents a pivotal advancement in precision oncology, offering a non-invasive and fast-tracked method to detecting clinically relevant biomarkers. With the backing of international oncology guidelines, ctDNA analysis is now a standard approach to consider in molecular diagnostics. Despite the promise of ctDNA in refining treatment strategies through the detection of genomic alterations and treatment-relevant biomarkers with high concordance to tissue biopsies, challenges persist. These include the interpretation of discordances due to tumor heterogeneity, sampling biases, and technical limitations, alongside the differentiation of tumor-derived mutations from clonal hematopoiesis. The current consensus supports the utility of comprehensive genomic profiling (CGP) panels for a broad spectrum of actionable targets, while acknowledging the limitations and advocating for a balanced application of “tissue-first” and “plasma-first” approaches tailored to individual patient scenarios. The essential role of molecular tumor boards (MTBs) is in navigating the complexities of ctDNA data interpretation, thereby ensuring the effective incorporation of liquid biopsy into personalized cancer treatment regimens.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
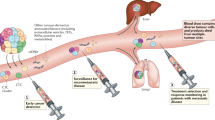
The integration of liquid biopsy into clinical practice marks a significant advancement in the realm of precision oncology, signaling a shift towards more refined and patient-centric approaches in routine cancer care. Clinicians are increasingly performing molecular profiling on circulating tumor DNA (ctDNA) obtained from plasma to non-invasively monitor residual disease [1], guide treatment options [2,3,4], or to match patients to suitable clinical trials [5], efforts which are underscored by the issuance of comprehensive implementation guidelines from international oncology and molecular pathology consortia such as ASCO/CAP [6, 7], ESMO [8], NCCN [9, 10], and IASLC. Certain indications now even exist that support the “plasma first” use case, meaning that traditional tissue biopsies may be bypassed in favor of a minimally invasive testing strategy. One example would be the testing for ESR1 mutations in plasma at endocrine resistance in breast cancer to guide addition of selective estrogen receptor degraders. While ctDNA-based assays are redefining clinical pathways, it can be difficult to sift through the growing literature base and clinical trial evidence [11], impeding the adoption of liquid biopsy in everyday cancer management. In this short review, we focus on the advanced cancer setting and offer brief, high-level summaries of the current guidelines, molecular profiling strategies and everyday challenges for incorporating liquid biopsy into real-world precision oncology approaches.
Concordance between alterations in tumor tissue and ctDNA
Many studies have assessed the concordance of genomic alterations detected in major driver genes and treatment-relevant biomarkers between plasma and tissue and have found consistent sensitivities, ranging approximately between 70–90% across various solid tumors [5, 12,13,14]. Discordance is mostly related to tumor heterogeneity and temporal dynamics, sampling biases and technical limitations. Tumors are often heterogeneous and the portion of the tumor sampled for tissue analysis may not fully represent the entire genetic landscape of the tumor, whereas ctDNA represents a mixture of DNA shed from various tumor regions. Moreover, tumor genomes evolve over time due to therapeutic pressure and clonal selection. The ctDNA shed into the bloodstream reflects the most recent state of the tumor, whereas tissue samples may have been obtained at an earlier stage of the disease and may not capture these changes. On the other hand, mutations derived from the hematopoietic system, which can lead to clonal expansions, can be picked up in cfDNA. Clonal hematopoiesis-related mutations can introduce specificity concerns in ctDNA analysis. Without careful validation and discrimination strategies, there is a risk of misinterpreting clonal hematopoiesis-related mutations as tumor-derived mutations, leading to false-positive results.
Taken together, tissue and ctDNA analysis each have strengths and limitations, and their findings may be complementary rather than identical. However, a robust detection agreement among key driver events as well as a comparable number of targetable alterations between ctDNA and tissue profiling [15,16,17] has established the viability of ctDNA-based assays as a substitute for tissue-based testing.
Genomic profiling of ctDNA in patients with advanced cancer for treatment selection
Selecting the right testing approach
Currently, the standard clinical application of genomic profiling of ctDNA is for treatment selection in the advanced cancer setting, meaning detecting alterations that can be matched to targeted therapies or identifying alterations that would be a contraindication for a particular therapy. As with tissue analysis, several scenarios may warrant the limited analysis of a single gene [18,19,20] or using a cancer hotspot panel ([4, 21]; Table 1). However, the general direction of the treatment selection setting is moving toward employing larger comprehensive genomic profiling (CGP) panels to maximize the detection of therapeutic targets and to harvest the additional, valuable information such as mutations, copy numbers, fusions, tumor fraction in plasma (Fig. 1; [22, 23]). In fact, several ctDNA-based CGP tests have already received FDA approval/clearance for select indications (Table 2). Putting algorithmically estimated levels of plasma tumor fraction (TF) into context with the detected variant allele frequencies (VAF) of detected mutations can help with the interpretation of detected mutations and indicate whether they are derived from the germline, from subclones or even from the hematopoietic system (Fig. 2). For CGP panels, TF is usually estimated based on tumor aneuploidy measured as deviations in coverage across the genome. When TFs are below 5–10%, such an estimate is no longer informative, while SNVs can still reliably be detected down to 0.1% [16]. If the panel is large enough, also complex biomarkers such as microsatellite instability (MSI) status and blood tumor mutational burden (bTMB), which have significant relevance for immunotherapies, can be inferred [24]. The added insight that can only be obtained from CGP approaches is critical to downstream interpretation of ctDNA results and provides the best comprehensive overview of the patient’s sample (Table 1).
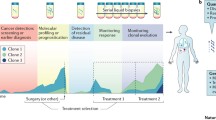
Single biomarker testing vs. CGP of DNA. Outside of the academic hospital setting, many clinics still predominantly employ single marker testing in their cancer diagnostic workflows. This routine approach is often cancer type-specific and it is able to identify alterations from a pre-specified gene of interest, but the technology overlooks potential existing mutations in other genes (left). Comprehensive genomic profiling (CGP) is gaining traction as a pan-cancer approach to detecting all four classes of alterations across hundreds of clinically relevant cancer-associated genes (right). CGP extends beyond the limited hotspot mutations and includes insertions and deletions, copy number alterations and fusions all from a single sample and test. In addition, large gene panels can also measure complex biomarkers, such as tumor fraction (TF), microsatellite instability (MSI) and blood tumor mutational burden (bTMB). The goal of CGP via NGS is to maximize the detection of therapeutically relevant and targetable genomic alterations that can be used to direct selection of suitable individualized treatment options
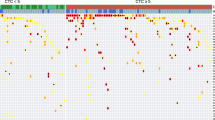
Potential outcomes of liquid profiling using CGP. a Samples with low tumor fraction (TF). Mutations in the range of a variant allele frequency (VAF) of 40–60% indicate germline origin. Pathogenic MSH2 mutation is a clear indication for genetic counseling regarding Lynch syndrome. b Sample with intermediate tumor fractions. While mutations in the range of the estimated TF are likely to be tumor-derived, higher VAFs are more likely associated with germline variants. Variants with substantially lower VAFs than the estimated TF can be of subclonal or hematopoietic origin. c Samples with a high TF. VAFs in the range of the estimated TF cannot distinguish between tumor-derived and germline variants. Germline testing and genetic counseling is indicated
Selecting the right analyte: tissue first, plasma first, or both in parallel?
An increasing body of evidence supporting the usage, advantages and limitations of plasma-based CGP for guiding treatment decisions has been instrumental in updating clinical guidelines for routine ctDNA testing. This has led to terminologies and concepts such as “tissue-first”, “plasma-first” or matched tissue and liquid profiling, indicating in which clinical scenario it makes sense to perform initial testing on biopsy material, cfDNA from plasma, or both, respectively. Because each approach has its obvious advantages and disadvantages, it is difficult to design a universal testing strategy for patients with advanced cancer and, in the past, recommendations have often conflicted among authors of clinical guidelines [10, 25,26,27,28] Recently, several of these professional multidisciplinary expert panels have reconvened to provide a general but thorough framework for assay and analyte selection for advanced cancer genotyping which are, for reasons of brevity, summarized here [8, 9]. Generally, it is recognized that ctDNA assays have demonstrated utility in the identification of actionable alterations to inform targeted treatment and may be used routinely for the management of advanced cancer patients, but the assay limitations must be considered. The “tissue-first” approach remains the gold standard for the majority of patients, especially as ctDNA assays are often limited in detecting important events of therapeutic relevance, such as fusions and SCNAs. As such, most guidelines stress the importance of the “tissue-first” approach and generally recommend “plasma-first” for most tumor types when tissue material is unavailable or inadequate [8]. However, a “plasma-first” approach may be performed when a quicker turnaround time is critical for a clinical decision, like in aggressive tumors such as advanced NSCLC [8]. Additionally, there are several scenarios in which ctDNA testing is preferred to standard tissue profiling, such as for the detection of ESR1 mutations in breast cancer and the detection of resistance-related mutations in NSCLC patients who previously received tyrosine kinase inhibitor (TKI) therapy [8]. However, negative ctDNA results can have different interpretations depending on the clinical context and the specific characteristics of the patient and the tumor. While it may suggest the absence of detectable tumor DNA in the bloodstream or a favorable response to treatment, it does not definitively rule out the presence of a mutations, in particular when the sample harbors low tumor content. Therefore, expert guidelines advise reflex tumor testing when non-informative results are obtained. There is also accumulating evidence that performing molecular profiling on both tissue and plasma in parallel significantly enhances the detection of actionable alterations, thus increasing the chance of being able to match patients to targeted therapies [4, 15]. In addition, joint tissue and liquid testing provides valuable complementary, technical and biological information that enables a more holistic interpretation and evaluation of the molecular results.
Interpretation of liquid biopsy data poses challenges for integration into routine clinical care
Much emphasis is put on the challenges of the technical implementation of liquid biopsy in the clinic. However, the complexity of downstream interpretation of molecular testing results from ctDNA is often greatly underestimated (Table 3). The increasing broad coverage and high accuracy of liquid CGP panels has propelled the potential of this technology for guiding treatment decisions, but it comes with everyday challenges in interpreting genomic variance from hundreds of genes and alteration types. This information can only be processed accurately, efficiently and with high confidence within the framework of a clinical team that covers diverse medical disciplines, a construct referred to as the molecular tumor board (MTB) [29,30,31]. MTBs bring together specialists in oncology, pathology, genetics, molecular biology, bioinformatics, patient care and clinical trials. They collaborate to tailor personalized treatment choices for patients, taking into account genomic alterations within their tumors and other relevant clinical factors and data. MTBs may also decide which patients to test, which analytes to assess and which molecular assays to employ. This helps provide clinicians with essential diagnostic, prognostic and actionable insights, enabling them to integrate molecular findings into optimized and individualized care plans for their patients. In routine MTB settings, results from ctDNA testing pose several interpretation challenges that require careful discussion among panel members before reaching treatment or further diagnostic workup recommendations.
Concluding remarks
The accumulating evidence supporting the use of liquid biopsy in routine oncology has paved the way for regulatory approval of several ctDNA-based tests, which has in turn driven an increase in clinical adoption in the advanced disease setting. Currently, one of the main challenges for those starting with liquid biopsy is determining which test best aligns with the clinical question at hand, how many biomarkers should be tested, whether to partner with an academic laboratory or outsource testing to an industry provider, and how to convert NGS readouts into evidence-based treatment decisions, particularly when the results are not as straightforward as with standard tissue testing. While the introduction of CGP has improved the probability of detecting a biomarker-based indication, the additional information provided in liquid biopsy medical reports has posed interpretation challenges that interfere with streamlined treatment decision-making, thus necessitating the interdisciplinary collaboration among medical professionals in the form of an MTB.
References
Semenkovich NP, Szymanski JJ, Earland N, Chauhan PS, Pellini B, Chaudhuri AA. Genomic approaches to cancer and minimal residual disease detection using circulating tumor DNA. J Immunother Cancer. 2023; 01;11(6):e006284.
Brannon RA, Jayakumaran G, Diosdado M, Patel J, Razumova A, Hu Y, et al. Enhanced specificity of clinical high-sensitivity tumor mutation profiling in cell-free DNA via paired normal sequencing using MSK-ACCESS. Nat Commun. 2021; 18;12(1):3770–5.
Sicklick JK, Kato S, Okamura R, Schwaederle M, Hahn ME, Williams CB, et al. Molecular profiling of cancer patients enables personalized combination therapy: the I‑PREDICT study. Nat Med. 2019; 01;25(5):744–50.
Riedl JM, Hasenleithner SO, Pregartner G, Scheipner L, Posch F, Groller K, et al. Profiling of circulating tumor DNA and tumor tissue for treatment selection in patients with advanced and refractory carcinoma: a prospective, two-stage phase II Individualized Cancer Treatment trial. Ther Adv Med Oncol. 2021; 27(13):1758835920987658.
Rothwell DG, Ayub M, Cook N, Thistlethwaite F, Carter L, Dean E, et al. Utility of ctDNA to support patient selection for early phase clinical trials: the TARGET study. Nat Med. 2019; 01;25(5):738–43.
Merker JD, Oxnard GR, Compton C, Diehn M, Hurley P, Lazar AJ, et al. Circulating Tumor DNA Analysis in Patients With Cancer: American Society of Clinical Oncology and College of American Pathologists Joint Review. J Clin Oncol. 2018; 01;36(16):1631–41.
Lockwood CM, Borsu L, Cankovic M, Earle JSL, Gocke CD, Hameed M, et al. Recommendations for Cell-Free DNA Assay Validations: A Joint Consensus Recommendation of the Association for Molecular Pathology and College of American Pathologists. J Mol Diagn. 2023; 01;25(12):876–97.
Pascual J, Attard G, Bidard F, Curigliano G, De Mattos-Arruda L, Diehn M, et al. ESMO recommendations on the use of circulating tumour DNA assays for patients with cancer: a report from the ESMO Precision Medicine Working Group. Ann Oncol. 2022; 01;33(8):750–68.
Ettinger DS, Wood DE, Aisner DL, Akerley W, Bauman JR, Bharat A, et al. Non-Small Cell Lung Cancer, Version 3.2022, NCCN Clinical Practice Guidelines in Oncology. J Natl Compr Canc Netw. 2022; 01;20(5):497–530.
Gradishar WJ, Anderson BO, Abraham J, Aft R, Agnese D, Allison KH, et al. Breast Cancer, Version 3.2020, NCCN Clinical Practice Guidelines in Oncology. J Natl Compr Canc Netw. 2020; 01;18(4):452–78.
Hasenleithner SO, Speicher MR. A clinician’s handbook for using ctDNA throughout the patient journey. Mol Cancer. 2022; 21;21(1):81–7.
Adalsteinsson VA, Ha G, Freeman SS, Choudhury AD, Stover DG, Parsons HA, et al. Scalable whole-exome sequencing of cell-free DNA reveals high concordance with metastatic tumors. Nat Commun. 2017 06;8(1):1324‑y.
Odegaard JI, Vincent JJ, Mortimer S, Vowles JV, Ulrich BC, Banks KC, et al. Validation of a Plasma-Based Comprehensive Cancer Genotyping Assay Utilizing Orthogonal Tissue- and Plasma-Based Methodologies. Clin Cancer Res. 2018 01;24(15):3539–49.
Park S, Olsen S, Ku BM, Lee M, Jung H, Sun J, et al. High concordance of actionable genomic alterations identified between circulating tumor DNA-based and tissue-based next-generation sequencing testing in advanced non-small cell lung cancer: The Korean Lung Liquid Versus Invasive Biopsy Program. Cancer. 2021 15;127(16):3019–28.
Iams WT, Mackay M, Ben-Shachar R, Drews J, Manghnani K, Hockenberry AJ, et al. Concurrent Tissue and Circulating Tumor DNA Molecular Profiling to Detect Guideline-Based Targeted Mutations in a Multicancer Cohort. JAMA Netw Open. 2024 02;7(1):e2351700.
Husain H, Pavlick DC, Fendler BJ, Madison RW, Decker B, Gjoerup O, et al. Tumor Fraction Correlates With Detection of Actionable Variants Across 23,000 Circulating Tumor DNA. Samples JCO Precis Oncol. 2022;01(6):e2200261.
Zhang Y, Yao Y, Xu Y, Li L, Gong Y, Zhang K, et al. Pan-cancer circulating tumor DNA detection in over 10,000 Chinese patients. Nat Commun. 2021; 12(1):11–8.
Osumi H, Takashima A, Ooki A, Yoshinari Y, Wakatsuki T, Hirano H, et al. A multi-institutional observational study evaluating the incidence and the clinicopathological characteristics of NeoRAS wild-type metastatic colorectal cancer. Transl Oncol. 2023; 01(35):101718.
Burstein HJ, DeMichele A, Somerfield MR, Henry NL. Biomarker Testing and Endocrine and Targeted Therapy in Metastatic Breast Cancer Expert Panels. Testing for ESR1 Mutations to Guide Therapy for Hormone Receptor-Positive, Human Epidermal Growth Factor Receptor 2‑Negative Metastatic Breast Cancer: ASCO Guideline Rapid Recommendation Update. J Clin Oncol. 2023; 20;41(18):3423–5. Jun.
Suppan C, Graf R, Jahn S, Zhou Q, Klocker EV, Bartsch R, et al. Sensitive and robust liquid biopsy-based detection of PIK3CA mutations in hormone-receptor-positive metastatic breast cancer patients. Br J Cancer. 2022; 126(3):456–63.
Rothe F, Laes J, Lambrechts D, Smeets D, Vincent D, Maetens M, et al. Plasma circulating tumor DNA as an alternative to metastatic biopsies for mutational analysis in breast cancer. Ann Oncol. 2014; 25(10):1959–65.
Zehir A, Benayed R, Shah RH, Syed A, Middha S, Kim HR, et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat Med. 2017; 01;23(6):703–13.
Reitsma M, Fox J, Borre PV, Cavanaugh M, Chudnovsky Y, Erlich RL, et al. Effect of a Collaboration Between a Health Plan, Oncology Practice, and Comprehensive Genomic Profiling Company from the Payer Perspective. J Manag Care Spec Pharm. 2019; 01;25(5):601–11.
Kim ES, Velcheti V, Mekhail T, Yun C, Shagan SM, Hu S, et al. Blood-based tumor mutational burden as a biomarker for atezolizumab in non-small cell lung cancer: the phase 2 B‑F1RST trial. Nat Med. 2022; 01;28(5):939–45.
Cardoso F, Paluch-Shimon S, Senkus E, Curigliano G, Aapro MS, Andre F, et al. 5th ESO-ESMO international consensus guidelines for advanced breast cancer (ABC 5). Ann Oncol. 2020; 31(12):1623–49.
Planchard D, Popat S, Kerr K, Novello S, Smit EF, Faivre-Finn C, et al. Metastatic non-small cell lung cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol. 2018; 29(Suppl 4):iv192–237.
Kalemkerian GP, Narula N, Kennedy EB, Biermann WA, Donington J, Leighl NB, et al. Molecular Testing Guideline for the Selection of Patients With Lung Cancer for Treatment With Targeted Tyrosine Kinase Inhibitors: American Society of Clinical Oncology Endorsement of the College of American Pathologists/International Association for the Study of Lung Cancer/Association for Molecular Pathology Clin Practice Guideline Update. J Clin Oncol. 2018; 20;36(9):911–9.
Burstein HJ, Somerfield MR, Barton DL, Dorris A, Fallowfield LJ, Jain D, et al. Endocrine Treatment and Targeted Therapy for Hormone Receptor-Positive, Human Epidermal Growth Factor Receptor 2‑Negative Metastatic Breast Cancer: ASCO Guideline Update. J Clin Oncol. 2021; 39(35):3959–77.
Wahida A, Buschhorn L, Frohling S, Jost PJ, Schneeweiss A, Lichter P, et al. The coming decade in precision oncology: six riddles. Nat Rev Cancer. 2023; 23(1):43–54.
Tsimberidou AM, Kahle M, Vo HH, Baysal MA, Johnson A, Meric-Bernstam F. Molecular tumour boards—current and future considerations for precision oncology. Nat Rev Clin Oncol. 2023; 20(12):843–63.
Koopman B, Groen HJM, Ligtenberg MJL, Grunberg K, Monkhorst K, de Langen AJ, et al. Multicenter Comparison of Molecular Tumor Boards in The Netherlands: Definition, Composition, Methods, and Targeted Therapy Recommendations. Oncologist. 2021; 26(8):e1347–58.
https://www.fda.gov/medical-devices/in-vitro-diagnostics/list-cleared-or-approved-companion-diagnostic-devices-in-vitro-and-imaging-tools, accessed April 14th, 2024
Funding
Open access funding provided by Medical University of Graz.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
S.O. Hasenleithner and E. Heitzer declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hasenleithner, S.O., Heitzer, E. Liquid profiling for patients with advanced cancer is ready for clinical integration. memo (2024). https://doi.org/10.1007/s12254-024-00978-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12254-024-00978-6