Abstract
In order to identify potential specific gene networks of Hepatitis C virus (HCV) related hepatocellular carcinoma (HCC), weighted gene co-expression network analysis (WGCNA) was performed, which may provide an insight into the potential mechanism of the HCC development. HCV-related HCC and normal sample data were downloaded from GEO, T test of limma package was used to screen different expression genes (DEGs); KEGG pathway was used to analyze related biochemical pathways, and WGCNA was used to construct clustering trees and screen hub genes in the HCC-specific modules. A total of 1151 DEGs were authenticated between the HCC and normal liver tissue samples, including 433 upregulated and 718 downregulated genes. Among these genes, three specific modules of HCC were constructed, including Tan, Yellow and Cyan, but only Yellow module had a significant enrichment score in substance combination module with three hub genes: SLA2547, EFNA4 and MME. Although Tan and Cyan separately had four and three hub genes, but the bio-functions of them did not have significant enrichment scores (score < 2). SLA2547, EFNA4 and MME may play important roles in the substance combination of HCV-related HCC, so studying the function of this gene network may provide us a deeper understanding of HCV-related HCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatocellular carcinoma (HCC) is one of the most common malignant cancers and the third leading cause of death induced by cancer worldwide [1]. Generally, HCC develops in patients suffering from chronic liver diseases, such as hepatitis, steatosis, cirrhosis. Hepatitis C virus (HCV) is one of the causes of chronic liver diseases. HCV is a positive single-stranded RNA virus and belongs to the Flaviviridae family [2]. Although it has identified for many years that the development of HCC induced by HCV contains four sections including: hepatitis, fibrosis, cirrhosis and HCC, the molecular mechanism of this procedure still remains unclear [3].
HCV-related HCC is a highly heterogeneous and comprehensive tumor with several alterations of genes and molecular pathways. For example, the expressions of insulin growth factor 2 and aldo-keto reductase family 1 member B10 as well as mRNA-binding protein 3 are significantly upregulated in HCV-related HCC [4, 5]. The crosstalks among inflammation, lipid metabolism and epigenetic alteration are important parts of the hottest topics in hepatocellular carcinogenesis. Inflammation is the first response of HCV infection and may run through all of the pathologic processes, Wnt signal pathway [6], P53 pathway and NF-κB pathway are all involved in the inflammatory responses [7]. HCV infection and replication are the original causes and accomplice of the development of HCV-related HCC. As a previous review reported that HCV virus hijacks the host cell lipid metabolism to facilitate their infections and replications by upregulating lipid synthesis and downregulating secretion and catabolism in infected hepatocytes [8]. It has been clearly reported that fatty acid synthase (FAS) and sterol-regulatory element binding proteins (SREBPs) are significantly upregulated during HCV infection [1]. DNA methylation is a new research topic of the development of HCC, it has been clarified that the promoters of RASSF1 (Ras association domain-containing protein 1), DOK1 (Docking protein 1) and CHRNA3 (Cholinergic receptor nicotinic alpha-3 subunit) were significantly methylated in HCV-related HCC [9].
Microarray data have been used for the investigation of the molecular mechanism underlying HCV-related HCC for a period time. For example, in order to clarify the association of IL28B (Interleukin 28B) and HCC recurrence, a series of human HCV-related HCC samples was collected and detected and the final microarray data were submitted under the dataset of GSE41804 [10]. According to this dataset, Hodo et al. have documented that patients with Il28B rs8099917 TT genotype showed higher recurrence than patients with TG/GG genotype after curative therapy. Besides, based on this microarray dataset, there also some other researches have been done, including the studies on HCC carbon metabolism [11], cancer invasion and metastasis [12], gene expressions profiles of different lesions by Gaussian model [13], and liver cell dedifferentiation [14]. However, the difference of gene co-expression network analysis between HCV-related HCC and normal tissues has been not yet investigated.
In order to further uncover the potential molecular mechanisms of Il28B rs8099917 TT genotype involved HCV-related HCC, microarray data in GSE41804 [10] was used to screen differentially expressed genes (DEGs) between TT genotype HCV-related tumor tissues and normal controls. Kyoto encyclopedia of genes and genomes (KEGG) was used to identified the biochemical pathways of DEGs, then the weighted gene co-expression network analysis (WGCNA) was used to screen hub factors of the potential specific genes co-expressed in HCV-related HCC, and finally used Gene ontology (GO) to identify the bio-functions of the selected genes. The results may provide novel information for the screening of target genes and diagnostic biomarkers for HCV-related HCC.
Materials and Methods
Data of Hepatocellular Carcinoma Gene Expression Chips
Expressional profile of GSE41804 [10] was downloaded from Gene Expression Omnibus (GEO, http://www.ncbi.nlm.nih.gov/geo/), and the data of tissue samples with Il28B rs8099917 TT genotype were produced from the platform of GPL570 Affymetrix Human Genome U133 Plus 2.0 Array to analyze the difference between HCV-related HCC and normal liver tissues. Finally, a total of 20 TT samples, including 10 HCV-related HCC samples and 10 non-carcinoma liver tissue samples were included.
Data Preprocessing and Identification of DEGs
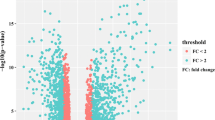
Firstly, robust multi-array average (RMA) method in the Affy package in R [15] was used to preprocess the downloaded chip data, including background correction, data normalization, and calculation of expression values. Subsequently, DEGs between the HCC and normal tissue samples were identified using T-test in the Linear Models for Microarray Data (limma) package in R [16], and thresholds were set as |log fold change (FC)| ≥ 0.58 and P-value ≤0.05.
Functional Enrichment Analysis of DEGs
KEGG Pathway Database [17] is a relatively common and comprehensive database in different biochemical pathway studies. In this study, the DAVID [18] online tool was selected to perform the KEGG pathway enrichment analysis for DEGs. Only the pathway terms with P-value <0.05 were considered significant.
Construction of Clustering Tree for WGCNA
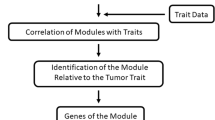
WGCNA arithmetic [19] is a typical systemic biological arithmetic for the construction of gene co-expression network, and this arithmetic is based on high-throughput chip data of mRNA and is widely used in biomedical area all over the world. For WGCNA, it is assumed that all the gene networks obey scale-free and define the adjacent function of gene co-expression related matrix and gene-network formation to calculate discrepancy coefficient of different nodes, and the hierarchical clustering tree is constructed by the computed results. For hierarchical clustering tree, each line represents a single gene and each branch represents the co-expression gene, so the different branches can obviously reflect the different gene modules, different co-expression of genes in these modules and the levels of co-expression genes of different modules. Except these advantages for searching co-expression modules, this network analysis is also able to help us to focus on the genes whose fold changes are not obvious (0.58 ≤ |logFC| ≤ 1) but have potential to affect the development of HCV-related HCC.
Here, WGCNA package in R was used to construct clustering trees for both normal liver and HCV-related HCC samples, and then continually used to compare and screen the specific modules for HCV-related HCC for the further study.
Construction of WGCNA Network and Screening for the hub Genes
R package of WGCNA was used to construct networks for the selected modules. Due to too much genes in modules, according to the connectivity degree of node, the top 30 hub genes were firstly selected, and about 5 genes were then chosen from the 30 genes based on their topological overlaps. Finally, GO functional enrichment analysis was performed via DAVID online tool to select gene modules and analyze these gene functions in molecular level.
Results
Screening of DEGs
Based on the criteria of |logFC| ≥ 0.58 and P-value ≤0.05, a total of 1151 DEGs were identified in the HCV-related HCC samples compared with the normal samples, including 433 up-regulated genes and 718 down-regulated genes.
KEGG Pathway Enrichment Analysis
With the criterion of 0.05 in the KEGG pathway enrichment analysis, the significant enriched biochemistry pathways of the upregulated genes, such as LAMA4, MDM4, COL4A2, COL4A1 and CDKN2B, were mainly concentrated in ECM-receptor interaction, Cell cycle, Pathway in cancer, P53 signaling pathway and small cell lung cancer (Table 1). Meanwhile, with the same cut-off criteria, the significant enriched biochemistry pathways of the downregulated genes, such as ACADM, HMGCS2, CXCL14, ACADS, and CXCL2, were mainly concentrated in cytokine-cytokine receptor interaction, peroxisome proliferator-activated receptor (PPAR) signaling pathway, complement and coagulation cascades, tryptophan metabolism, metabolism of xenobiotic by cytochrome P450 and drug metabolism pathways and so on (Table 2).
Analysis of WGCNA Clustering Trees
Based on the WGCNA, the systematic clustering were generated (Fig. 1). The gene modules were signified by different colors and the grey module indicated the genes that cannot be merged. For the HCC tissue clustering tree (Fig. 1a), with the soft-threshold of 7, 14 gene modules were identified, and the gene number of 14 modules ranged from 31 (Cyan) to 219 (Turquoise). For the normal tissue clustering tree (Fig. 1b), with the soft-threshold of 6, 8 gene modules were identified, and the numbers of genes in these modules ranged from 43 (Pink) to 372 (Turquoise).
The cluster dendrogram and color display of co-expression network modules for differentially expressed genes in hepatitis C virus related hepatocellular carcinoma samples (a) and normal samples (b) Each short vertical line represents a gene; each branch represents the co-expression genes in each cluster module; and the area of each colors represents the number size of each cluster module
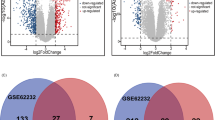
Furthermore, repeatability analysis was performed for the modules of two clustering trees (Fig. 2), it clearly showed that genes only in Grey module of normal tissue were not overlapped with modules of HCV-related HCC, but the genes in Tan, Yellow and Cyan modules in HCV–related HCC were not overlapped with any other modules in normal tissues. This result may indicate that only modules Tan, Yellow and Cyan modules specifically belonged to HCV-related HCC, and genes in these 3 modules were not overlapped with genes expressed in normal tissue clustering trees.
Comparisons between HCV related HCC and normal liver modules. Vertical axis represents HCV-related HCC modules, and the horizontal axis represents normal tissue modules; the depth of red color is positively related to coincidence depth of crossed modules, and the numbers in each rectangle represents the number of overlapping genes
Analysis of WGCNA Network
To further explore the biological functions of these included genes, WGCNA network analysis map was constructed by combining distinctive modules of HCV-related HCC. In the Tan network, with the cut-off criterion of 0.05 and node connection >10, a total of 4 hub genes in the network were identified, including two upregulated genes EPS8L3 and WNK4 and two downregulated genes GCKR and SDC4 (Fig. 3a). In the Yellow network, with the cut-off criterion of 0.1 and node connection >10, a total of 3 hub genes in the network were identified, including two upregulated genes SLA25A47, EFNA4 and one downregulated gene MME (Fig. 3b). Additionally, in the Cyan network, with the cut-off criterion of 0.03 and node connection >10, a total of 3 downregulated genes hub genes: DHX32, SERTAD1 and STBD1 were obtained in the network (Fig. 3c).
Furthermore, in order to assess whether these genes share specific biological function features, GO enrichment analysis was performed for the genes of these 3 specific modules. Only Yellow module had a significant enrichment effect (Enrichment score > 2, Table 3) and its function was mainly concentrated on the extracellular matrix integrated, polysaccharide combination, heparin-binding etc., so this module was named substance combination module.
Discussion
Based on the analysis of HCV-related microarray data, a total of 1151 DEGs were identified including 433 upregulated and 718 downregulated genes. Based on these genes, Tan, Yellow and Cyan were three HCV-related HCC-specific-modules. Based on the functional enrichment analysis by the GO method, the result showed that only module Yellow had a significant enrichment score, so we named it as substance combination module.
Yellow module was the unique module which had a meaningful enrichment score in the GO enrichment analysis. SLC25A47, EFNA4 and MME were the hub genes in this module. SLC25A47 (Solute carrier family 25 member 47) is related to the development of liver disease [20]. A recent study has reported that SLC25A47 contributes to the hepatitis-associated inflammation [20] and is upregulated in livers of the mice with the deficiency of Rictor, which plays a vital role in mammalian target of rapamycin complex 2 (mTORC2) signaling pathway of HCC development [21]. Given that the expression of SLC25A47 was also increased in HCV-related HCC, this gene might play a vital role in the development of HCV-related HCC. MME, encoding membrane metallo-endopeptidase, is an indicator of cancers, especially in leukemia. The bio-functions of MME are mainly concentrated on cell-cell signaling and proteolysis [22]. It has previously reported that the CpG islands of MME were hyper-methylated in HCC compared with pre-neoplastic tissues and normal tissues [22], which is consistent with the result of this study that MME was downregulated in HCC tissues compared with the normal controls. In addition, the research of Tong et al. also has indicated that MEM also has a decrease in the focal segmental glomerulosclerosis which is caused by podocyte injury [23]. EFNA4 encodes ephrin A4, which belongs to the family of receptor tyrosine kinases which has two subclasses: A (A1-A5) and B (B1-B3) [24]. Members of epherin A subclass are anchored to the membrane via glycosylphosphatidylinositol linkage to mediate signaling about growth, migration and invasion in cancer [24]. As it previously reported, EFNA4 is found to be upregulated in lung cancer compared with normal lung tissues [25], which is consistent with the result in this study. In a clinical research, ephrin A4 was also found to be a potential risk factor for chemotherapy and prognosis of osteosarcoma patients [26]. Based on the bio-functions of the hub genes in Yellow module, we suggested when liver was infected by HCV, inflammation induced by the immune response will lead to a upregulated expression of SLC25A47, which promoted hepatitis to develop into HCC; on the same time, hyper-methylation would downregulate the expression of MME and this results may impair the hydrolysis of MME protein to maintain the disordered increasing of liver cells so that can trigger EFNA4 to promote the migration and invasion of infiltrating tumor cells.
Furthermore, in this study, Tan and Cyan were another two HCC-specific modules. There were four hub genes including two upregulated genes WNK4 and EPS8L3, and two downregulated genes GCKR and SDC4 in Tan module. The downregulated GCKR is the coding gene of glucokinase regulator, which is the key enzyme for de novo lipogenesis from glucose [27]. Another downregulated gene SDC4 is an important sensor of the extracellular matrix in wound contraction, fibrosis, regulation of motility [28]. WNK4 was one of the upregulated genes in the module Tan. It has reported that WNK can affect the HCV entry via modulating the phosphorylation of claudins [29]. Despite there was no direct evidence to prove that these four genes participated in HCC occurrence, the bio-functions of them also could clearly show that they may play vital roles in the development and metabolism of HCC. EPS8L3 is encoding gene of a protein which is a substrate of the epidermal growth factor receptor, but the function of this protein is still unclear. Three downregulated hub genes (DHX32, SERTAD1 and STBD1) were included in the module Cycan. DHX32 is originally identified as an RNA helicase, and the function of this gene is largely unknown. A recent research has reported that DHX32 promotes proliferation, migration and invasion of colorectal cancer cells with an upregulated expression [30], but this result was not according to the result of HCC. The conflict may suggest that DHX32 may be differentially expressed in different tumor types. SERTAD1 was mainly reported being upregulated with cell proliferation. STBD1 is the coding gene of starch-binding domain-containing protein 1 which can fix glycogen to autophagic membrane to help lysosomal degradation [31]. In addition, it has also reported that STDB1 plays a vital role in the transportation of glycogen to lysosomes in liver [32]. It is worthy to point out that the expression levels of DHX32 and SERTAD1 in the above studies are contrary to their expressions in this study. Whether they had any other mechanism involved should be concerned in the future.
In this study, WGCNA was used to analyze the expression of mRNA between HCV-related HCC and normal tissues in order to obtain some specific genes and modules of HCC which were not identified in known pathways and studies. Among the selected genes, for some genes, only names and variations in expression have been reported, and the functions of some genes in HCV-related HCC are still unknown. Performing researches on these selected genes and modules may allow us to have a further understanding of the development of HCC and provide some potential drug targets in targeted therapy. Meanwhile, in order to screen more potential remarkable genes, we also set a wider threshold (|logFC| ≥ 0.58 and P-value ≤0.05) than general (|logFC| ≥ 1 and P-value ≤0.05) selected criteria. However, this study still has some limitations. The clinical information was not included in the analyses. Sufficient clinical data not only can eliminate the influence of gender and age but also can provide a specific analysis for certain symptom, but there is only scarce data in GEO (https://www.ncbi.nlm.nih.gov/geo/), so a larger clinical information should be collected to have a better analysis in the future. In addition, the absence of experimental verification was also another deficiency of this study.
In conclusion, a total of 10 hub genes were identified in three specific modules in HCV-related HCC tissues compared with normal tissues. Among three modules, Yellow module had significant enrichment score, the hub genes SLC25A47, EFNA4 and MME in this module are new–found to be related to the development of HCV-related HCC. Studies on these genes may not only enable us to have a further understanding of the HCC development, but also can provide potential treatment targets for HCC therapy.
References
Jin B, Wang W, Du G, Huang GZ, Han LT, Tang ZY et al (2015) Identifying hub genes and dysregulated pathways in hepatocellular carcinoma. Eur Rev Med Pharmacol Sci 19:592–601
Luna J, Scheel TH, Danino T, Shaw K, Mele A, Fak J et al (2015) Hepatitis C virus RNA functionally sequesters miR-122. Cell 160:1099–1110
Castello G, Scala S, Palmieri G, Curley SA, Izzo F (2010) HCV-related hepatocellular carcinoma: from chronic inflammation to cancer. Clin Immunol 134:237–250
Petrizzo A, Caruso FP, Tagliamonte M, Tornesello ML, Ceccarelli M, Costa V et al (2016) Identification and validation of HCC-specific Gene transcriptional signature for tumor antigen discovery. Sci Rep 6:29258
Kakehashi A, Ishii N, Sugihara E, Min G, Saya H, Wanibuchi H (2016) CD44 variant 9 is a potential biomarker of tumor initiating cells predicting survival outcome in hepatitis C virus-positive patients with resected hepatocellular carcinoma. Cancer Sci 107:609–618
Boj S, Vanes J, Huch M, Li VW, José A, Hatzis P et al (2012) Diabetes risk gene and Wnt effector Tcf7l2/TCF4 controls hepatic response to perinatal and adult metabolic demand. Cell 151:1595–1607
Park SH, Rehermann B (2014) Immune responses to HCV and other hepatitis viruses. Immunity 40:13–24
Syed GH, Amako Y, Siddiqui A (2009) Hepatitis C virus hijacks host lipid metabolism. Trends Endocrinol Metab 21:33–40
Lambert MP, Paliwal A, Vaissière T, Chemin I, Zoulim F, Tommasino M et al (2011) Aberrant DNA methylation distinguishes hepatocellular carcinoma associated with HBV and HCV infection and alcohol intake. J Hepatol 54:705–715
Hodo Y, Honda M, Tanaka A, Nomura Y, Arai K, Yamashita T et al (2013) Association of interleukin-28B genotype and hepatocellular carcinoma recurrence in patients with chronic hepatitis C. Clin Cancer Res 19:1827–1837
Zhang J, Baddoo M, Han C, Strong MJ, Cvitanovic J, Moroz K et al (2016) Gene network analysis reveals a novel 22-gene signature of carbon metabolism in hepatocellular carcinoma. Oncotarget 7:49232–49245
Xu Y, Cui J, Puett D (2014) Understanding cancer invasion and metastasis. Springer, New York
Ueda T, Honda M, Horimoto K, Aburatani S, Saito S, Yamashita T et al (2013) Gene expression profiling of hepatitis B- and hepatitis C-related hepatocellular carcinoma using graphical Gaussian modeling ☆. Genomics 101:238–248
† SC, † PSS, Lee J, Park J, Shin EC, Choi C (2015) Prolonged silencing of diacylglycerol acyltransferase-1 induces a dedifferentiated phenotype in human liver cells. J Cell Mol Med 20:38–47
Irizarry RA, Hobbs B, Collin F, Beazerbarclay YD, Antonellis KJ, Scherf U et al (2003) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4:249–264
Diboun I, Wernisch L, Orengo CA, Koltzenburg M (2006) Microarray analysis after RNA amplification can detect pronounced differences in gene expression using limma. BMC Genomics 7:1–14
Wixon J, Kell D (2000) The Kyoto encyclopedia of genes and genomes--KEGG. Yeast 17:48–55
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature Protocol 4:44–57
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinf 9:1–13
Bushkofsky JR, Maguire M, Larsen MC, Fong YH, Jefcoate CR (2016) Cyp1b1 affects external control of mouse hepatocytes, fatty acid homeostasis and signaling involving HNF4α and PPARα. Arch Biochem Biophys 597:30–47
Lamming DW, Demirkan G, Boylan JM, Mihaylova MM, Peng T, Ferreira J et al (2014) Hepatic signaling by the mechanistic target of rapamycin complex 2 (mTORC2). Faseb Journal Official Publication of the Federation of American Societies for Experimental Biology 28:300–315
Archer KJ, Mas VR, Maluf DG, Fisher RA (2010) High-throughput assessment of CpG site methylation for distinguishing between HCV-cirrhosis and HCV-associated hepatocellular carcinoma. Molecular Genetics & Genomics Mgg 283:341–349
Tong J, Xie J, Ren H, Liu J, Zhang W, Wei C et al (2015) Comparison of glomerular Transcriptome profiles of adult-onset steroid sensitive focal segmental Glomerulosclerosis and minimal change disease. PLoS One 10:e0140453
Arvanitis DN, Davy A (1900) Regulation and misregulation of Eph/ephrin expression. Cell Adhes Migr 6:131–137
Saintigny P, Peng S, Zhang L, Sen B, Wistuba II, Lippman SM et al (2012) Global evaluation of Eph receptors and Ephrins in lung adenocarcinomas identifies EphA4 as an inhibitor of cell migration and invasion. Mol Cancer Ther 11:2021–2032
Abdou AG, Abd el-Wahed MM, Asaad NY, Samaka RM, Abdallaha R (2010) Ephrin A4 expression in osteosarcoma, impact on prognosis, and patient outcome. Indian J Cancer 47:46–52
Hernaez R (2011) Genetic factors associated with the presence and progression of nonalcoholic fatty liver disease: a narrative review. Gastroenterol Hepatol 35:32–41
Sanchez-Antolín G, Almohalla-Alvarez C, Bueno P, Almansa R, Iglesias V, Rico L et al (2015) Evidence of active pro-fibrotic response in blood of patients with cirrhosis. PLoS One 10:e0137128
English DP, Santin AD (2012) Claudins overexpression in ovarian cancer: potential targets for Clostridium perfringens enterotoxin (CPE) based diagnosis and therapy. Int J Mol Sci 14:10412–10437
Lin H, Liu W, Fang Z, Liang X, Li J, Bai Y et al (2015) Overexpression of DHX32 contributes to the growth and metastasis of colorectal cancer. Sci Rep 5:9247
Kim KH, Lee MS (2014) Autophagy - a key player in cellular and body metabolism. Nat Rev Endocrinol 10:322–337
Sun T, Yi H, Yang C, Kishnani PS, Sun B (2016) Starch binding domain-containing protein 1 plays a dominant role in glycogen transport to lysosomes in liver. J Biol Chem 291:16479–16484
Acknowledgements
This work is funded by grants from Outstanding Leaders Training Program of Pudong Health Bureau of Shanghai (PWRL2016-01), three-year plan of action of traditional Chinese medicine in Shanghai (ZY3-RCPY-1-1001 and ZY3-JSFC-1-1011), Science and Technology Commission of Pudong New Area Shanghai (PKJ2014-Y37), Prof. Jian-jie Chen Studio (Shanghai Legendary Medical Practitioner of Traditional Chinese Medicine, ZYSNXD-CC-MZY003).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Cheng, Y., Ping, J. & Chen, J. Identification of Potential Gene Network Associated with HCV-Related Hepatocellular Carcinoma Using Microarray Analysis. Pathol. Oncol. Res. 24, 507–514 (2018). https://doi.org/10.1007/s12253-017-0273-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12253-017-0273-8