Abstract
The virulence factors, antibiotic resistance patterns, and the associated genetic elements have been investigated in Staphylococcus species. A total of 100 strains has been isolated from clinical samples in the Microbiology Laboratory of Hesperia Hospital, Modena, Italy, and identified as Staphylococcus aureus (65), Staphylococcus epidermidis (24), Staphylococcus hominis (3), Staphylococcus saprophyticus (3), and Staphylococcus warneri (5). All the strains were analyzed to determine phenotypic and genotypic characters, notably the virulence factors, the antibiotics susceptibility, and the genetic determinants. The highest percentage of resistance in Staphylococcus spp. was found for erythromycin and benzylpenicillin (87% and 85%, respectively). All S. aureus, two S. epidermidis (8.3%), and one S. saprophyticus (33.3%) strains were resistant to oxacillin. The methicillin resistance gene (mecA) was detected by polymerase chain reaction (PCR) amplification in 65 S. aureus strains and in 3 coagulase-negative staphylococci (CoNS) (8.6%). With regard to the virulence characteristics, all the S. aureus were positive to all virulence tests, except for slime test. Among the CoNS isolates, 19 (79.1%) S. epidermidis and one (33.3%) S. saprophyticus strains resulted positive for the slime test only. The results obtained are useful for a more in-depth understanding of the function and contribution of S. aureus and CoNS antibiotic resistance and virulence factors to staphylococcal infections. In particular, the production of slime is very important for CoNS, a virulence factor frequently found in infections caused by these strains. Further investigations on the genetic relatedness among strains of different sources will be useful for epidemiological and monitoring purposes and will enable us to develop new strategies to counteract the diffusion of methicillin-resistant S. aureus (MRSA) and CoNS strains not only in clinical field, but also in other related environments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Staphylococcus species are frequently isolated from clinical specimens. These bacteria are widespread in the environment and are commensal inhabitants of the skin, mucous membranes, and other body sites in humans and animals. Staphylococcus aureus is the leading causative agent of a broad variety of illnesses, ranging from minor infections of the skin to severe diseases like pneumonia and bacteremia, and it is also responsible for hospital-acquired infections (HAIs) with surgical wounds and indwelling medical device infections (Tong et al. 2015).
Treatment of S. aureus infections is complicated by antibiotic resistance, and methicillin-resistant S. aureus (MRSA) and multidrug-resistant S. aureus (MDRSA) cause more severe infections in hospitalized patients, being endowed with high pathogenic potential, e.g., carrying gene coding for virulence factors and antimicrobial resistance. Resistance to methicillin and related antibiotics is attributed to mecA gene expression, which alters penicillin-binding protein (PBP-2) to PBP-2a, resulting in loss of target affinity (Zhan and Zhu 2018).
The mecA gene is part of a mec complex found on mobile genetic elements called the Staphylococcal Cassette Chromosome mec (SCCmec). Different and specific types of structural organization of SCCmec elements have been found in S. aureus of humans and animals (Jiang et al. 2019) and in methicillin-resistant coagulase-negative staphylococci (CoNS). To differentiate between MRSA and CoNS positive for mecA, the fem genes (femA and femB) were used, which encode proteins that influence the level of methicillin resistance of S. aureus. Moreover, there is an increasing interest in other virulence determinants that S. aureus produces and how they impact disease. Different virulence factors encoded by diverse genes play a major role during pathogenesis. Such factors include Panton–Valentine Leucocidin toxins (PVL) (encoded by lukS/F-PV and lukE/D genes), capable to damage erythrocytes and membranes of host defense cells (Shallcross et al. 2013), fibronectin-binding proteins (fnbA and fnbB) involved in tissue invasion, exfoliative toxins (ETs, etb and eta genes), arginine catabolic mobile element (ACME, arcA gene), β-hemolysin (Hlb, hlb gene) (Burlak et al. 2007), toxic shock syndrome toxin-1 (TSST-1, tst gene), accessory gene regulator (Agr, agr gene), and α-hemolysin (Hla, hla gene) (Heilmann et al. 2019).
Lastly, in recent years, despite their commensal status, CoNS have also been implicated as a cause of infections. CoNS cover a large and continuously expanding group of bacteria, with more than 50 species described so far, currently distributed into 41 main species, divided into more than 20 subspecies (Becker et al. 2020). CoNS infections represent an emerging public health problem, largely linked to the demographic aging of people, which creates older, multimorbid, and immunocompromised patients. In nosocomial environments, CoNS are transmitted mainly by medical and/or nursing procedures, followed by colonization and growth on indwelling device’s surface. The degree of pathogenicity expressed by the members of this group is the result of a series of factors such as the development of different host defense strategies and the presence of strain-specific virulence characteristics. Some of the key virulence factors associated with CoNS are (1) adhesion factors, CoNS have surface proteins and adhesins that enable them to adhere to several different surfaces including abiotic (polyethylene, stain steel, rubber, and glass) or biotic surfaces (living tissue or abiotic surfaces covered with proteins) and medical devices. These adhesins help establish initial infections and facilitate biofilm formation. (2) For biofilm formation, CoNS are known for their ability to form biofilms on various surfaces, including medical devices like catheters and prosthetic implants. Biofilms are complex structures made up of bacterial cells encased in a matrix of extracellular polymeric substances (EPS), which protect them from the host’s immune response and make them resistant to antibiotics. This is a significant virulence factor, as it allows CoNS to persist and cause chronic infections. (3) For toxin production, while CoNS typically produce fewer toxins than S. aureus, some strains can release toxins like hemolysins and exotoxins that contribute to their virulence. (4) For immune evasion mechanisms, CoNS can employ strategies to evade the host immune system, such as resisting phagocytosis and interfering with immune responses. (5) For intracellular persistence, some CoNS species can invade and persist within host cells, enabling them to evade immune responses and treatment. (6) For iron uptake mechanisms, CoNS may have iron acquisition systems that allow them to scavenge iron, an essential nutrient, from the host environment, promoting bacterial growth and survival. (7) For antibiotic resistance, many CoNS strains have developed antibiotic resistance, making them challenging to treat with conventional antibiotics. Methicillin-resistant coagulase-negative staphylococci (MR-CoNS) have become a clinical concern, as they can cause infections that are resistant to multiple antibiotics (França et al. 2021).
The virulence capacity of CoNS, associated with pathogenic processes, increases above all in the presence of other risk factors, such as immunosuppression, long-term hospitalization, or the use of medical devices (catheters, joint prostheses, and others) (Argemi et al. 2019). The most frequently involved species are Staphylococcus epidermidis, Staphylococcus haemolyticus, Staphylococcus saprophyticus, Staphylococcus capitis, and Staphylococcus lugdunensis (Cadieux et al. 2014). S. epidermidis, from the CoNS group, is the most frequently isolated of each group, and for this reason, it is the main studied CoNS species. S. epidermidis is a saprophyte that is part of the normal mucosa and skin microflora and emerged, together with S. aureus, as a frequent etiologic agent of infections associated with catheters and other indwelling medical devices. In humans, most of the isolates correspond to CoNS represented by S. epidermidis, considered to be the most abundant species that lives on the skin, and is also frequently encountered in the hospital environment, where healthcare workers may have a role in the transmission of S. epidermidis to patients (Cherifi et al. 2014). Some isolates have become increasingly concerning, as S. lugdunensis, a pathogenic bacterium with a high virulence feature responsible for skin infections (García-Malinis et al. 2021), highly acute, and destructive events of infective endocarditis with high mortality rates (Argemi et al. 2018). It is important to note that the specific virulence factors and the degree of virulence can vary among different CoNS species and strains belonging to this heterogeneous group, that includes both non-pathogenic and facultative pathogenic species, with distinct virulence potential levels (Rosenstein and Götz 2013). Thus, factors leading to staphylococcal infections are both host- and bacterial cell-dependent, and the fate of infection is influenced by bacterial characteristics (such as bacterial tolerance and resistance to antimicrobials or the production of different virulence factors that promote bacterial adherence and/or invasion) and host defense mechanisms (Larsson and Flach 2022). All these characteristics favor the survival and persistence of bacteria in hostile environments and lead to difficult-to-treat infections with long and sometimes severe evolution (Preda et al. 2021).
This study aims to investigate both coagulase-positive and coagulase-negative Staphylococcus strains isolated from clinical samples, evaluating the main virulence factors and antibiotic resistances.
Materials and methods
Bacterial strain identifications and antimicrobial susceptibility testing
One hundred Staphylococcus strains were isolated in the Microbiology Laboratory of Hesperia Hospital, Modena, Italy. Sites of isolation included nasal and superficial wound swabs and urine and blood specimens. Each specimen was subcultured onto Mannitol Salt Agar (MSA, Oxoid S.p.A, Milan, Italy) and incubated at 37 °C for 24 h. The identification of species and antimicrobial susceptibility testing were performed using the Vitek 2 system and AST-GP 580 card (bioMérieux Florence, Italy) according to the manufacturer’s instructions. Briefly, three to five colonies of Staphylococcus spp., with a concentration of approximately 0.5 McFarland, were inoculated into a sterile 0.45% NaCl solution. Then, the solution was loaded with the card into the Vitek 2 system and incubated for 5–8 h. Antibiotics tested included clindamycin, daptomycin, erythromycin, gentamicin, ciprofloxacin, levofloxacin, linezolid, mupirocin, oxacillin, benzylpenicillin, rifampicin, tetracycline, trimethoprim/sulfoxide, tigecycline, and glycopeptides (teicoplanin, vancomycin). VITEK® 2, an automated ID/AST analyzer, uses the Advanced Expert System (AES) that integrates EUCAST Expert and breakpoint documents (European Committee on Antimicrobial Susceptibility Testing EUCAST 2023). AES also determines the phenotype of isolates by comparing their antibiogram with MIC distributions of resistant and wild-type organisms stored in the AES database, which utilizes a MIC distribution concept very similar to the EUCAST wild-type MIC distributions.
Virulence factor determination
Hemolysin production
For the hemolysis test, the isolate was transferred onto Petri plates containing Tryptic Soy Agar (TSA, bioMérieux, Florence, Italy) supplemented with 7% horse blood and incubated at 37 °C for 24 h. The β-hemolytic reaction leads to complete lysis of the erythrocyte cells with the appearance of a clear halo around the colony, while the α-hemolytic reaction involves the partial lysis of the erythrocyte cells (Iseppi et al. 2020). Streptococcus pneumoniae ATCC 49619 was used as a positive control for α-hemolysis and S. aureus ATCC 25923 for β-hemolysis.
Lipase and lecithinase production
Lipolytic activity was determined on plates containing 2% agar base, 1% peptone, 1% yeast extract, 0.1% CaCl2, and 2% Tween 80 (Jessen et al. 1959). Lecithinase production was studied in Baird–Parker agar (bioMérieux, Florence, Italy) (Matos et al. 1995). In both tests, a positive result was indicated by the formation of an opaque halo around the colonies after incubation at 37 °C for 48 h.
DNAse and thermonuclease activity
Nuclease (DNAse) and thermonuclease (TNAse) were determined by the metachromatic toluidine blue O agar diffusion-DNA technique, according to Lachica et al. (1971). For both tests, S. aureus ATCC 25923 and S. epidermidis ATCC 12228 were used as positive and negative controls, respectively.
Hyaluronidase activity
The presence of hyaluronidase was evaluated on Brain Heart Infusion agar (BHI, bioMérieux, Florence, Italy) containing hyaluronic acid (0.4 mg/mL) (Makris et al. 2004). After incubation at 37 °C for 24 h, plates were covered with cetylpyridinium chloride, and a positive result was indicated by the formation of a transparent halo around the colonies.
Slime production
Slime production was evaluated by Congo red agar method. The medium was prepared with 37 g/L Brain Heart Infusion broth (bioMérieux, Florence, Italy), 50 g/L sucrose, 10 g/L agar, and 0.8 g/L Congo red (Baldassarri et al. 1993). Plates were incubated at 37 °C for 24 h and then incubated at room temperature for 12 h. A positive result was indicated by black colonies on the surface.
Detection of virulence genes
Bacterial DNA was extracted with the DNeasy tissue kit (Qiagen, Milan, Italy) as specified by the manufacturer and using lysostaphin (100 μg/mL; Sigma, Italy) to achieve bacterial lysis. Then, genomic DNA was used in PCR amplification using primers targeting the following genes staphylococcal enterotoxins (sea), toxic shock syndrome toxin (tst), alpha/beta-hemolysins (hla and hlb), fibronectin-binding proteins A and B (fnbA and fnbB), adhesins map/eap (map/eap), slime production (icaA and icaD), and methicillin resistance (mecA). Primer sequences and PCR methods were carried out as already described in other investigation (Booth et al. 2001; Campbell et al. 2008; Choi et al. 2003; Metwally et al. 2017; Nashev et al. 2004). The primer sequence, amplification conditions, and product length are presented in Table 1. PCR was performed in a DNA thermal cycler (Applied Biosystems PCR system2700). The reaction was done in a 25 µl volume containing the above-mentioned primers (1 mM each) together with 150 ng of the extracted DNA; 100 mM of each of dATP, dCTP, dGTP, and dTTP; 1 U of Taq DNA polymerase; and 10 mM PCR buffer (pH 9.0). The magnesium concentration in the mixture was 3 mM for each gene. After amplification, 10 µl of the PCR mixture was analyzed by agarose gel electrophoresis (2% agarose in Tris-borate-EDTA).
Results
Bacterial strain identification and antimicrobial susceptibility testing
The 100 Staphylococcus strains used in the study were identified as S. aureus (65), S. epidermidis (24), S. hominis (3), S. saprophyticus (3), and S. warneri (5) strains. Table 2 shows the resistance of the strains to 16 antimicrobials, tested by the Vitek 2 system (card AST-GP 580, bioMérieux Florence, Italy).
All S. aureus (100%), 2 (8.3%) S. epidermidis, and 1 (33.3%) S. saprophyticus strains were resistant to oxacillin. The results also show the resistance of all S. aureus and S. saprophyticus to erythromycin and benzylpenicillin and in a smaller percentage for S. epidermidis (resistance to erythromycin and benzylpenicillin of 79.1% and 70.3%, respectively). For the remaining drugs, resistance ranging from 4.1 (clindamycin for CoNS) to 100% ciprofloxacin for S. saprophyticus) emerged from this study. Both S. hominis and S. warneri species resulted sensitive to the antibiotics used in the test. The remaining staphylococcal species were susceptible to daptomycin, gentamicin, linezolid, rifampicin, trimethoprim/sulfoxide, tigecycline, teicoplanin, and vancomycin.
Virulence factor determination
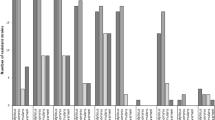
Regarding the virulence characteristics (Fig. 1 and Table 3), the 65 S. aureus strains were positive to all virulence tests, of which 87.6% presented positivity to both hemolysins and lipase production, 100% to lecithinase and DNase, while 92.3% and 69.2% showed positivity to thermonuclease and to hyaluronidase, respectively. One S. aureus only was positive to the slime test (7.6%), simultaneously to a β-hemolytic profile and lipase.
About the CoNS isolates, all were negative for virulence characteristics, except 19 (79.1%) S. epidermidis and only one (33.3%) S. saprophyticus strains that resulted positive to the slime test.
Detection of virulence genes
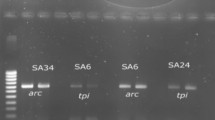
Our results on the antibiotic resistance show that all the 100% of S. aureus and 41.6% of CoNS (S. epidermidis and S. saprophyticus) clinical isolates are resistant to oxacillin. The molecular analysis (Table 4) showed that in these strains, the mecA gene was found in all S. aureus, in 2 S. epidermidis, and 1 of S. saprophyticus strains. Regarding the other genes, 30 S. aureus strains (46.1%) harbored the hla gene, 36 (53.8%) the hlb gene, 16 (24.6%) the sea gene, 15 (23.7%) the tst gene, 49 (75.3%) the fnbA and fnbB genes, and 1 (1.5%) the map/eap gene, while neither the icaA nor the icaD genes were found. All PCR reactions were negative for CoNS, apart for 20 S. epidermidis, in which icaA and icaD genes, coding for the slime production, were present (83,3%), and for 3 S. saprophyticus strains, in which icaA and icaD genes were found in one (33,3%) and two (66,6%) strains, respectively (Fig. 2a–c).
Example of agarose gel electrophoresis of amplicons of different genes obtained by PCR of samples from a S. aureus lanes, M marker (234–2176 pb), 1–2 hla with size of 534 bp 3–4 hla 833 bp; b S. aureus lanes, M markers (10–1000 pb), 1–3. tst gene350bp; c lanes, M markers (234–2176 pb), 1–2 fnbA (1278 bp), 3–4 fnbB (812): d S. epidermidis M markers (50–1500 pb), 1–6 icaA (188pb, 7–10 icaD gene (198 pb); e S. aureus lane M marker (100–1500pb) mecA gene (314pb)
Discussion
In recent years, the most common infections in humans have been caused by several different Staphylococcus spp. Staphylococcus aureus is the major pathogen in healthcare settings (Taylor and Unakal 2023; Lakhundi and Zhang 2018). S. aureus can synthesize some enzymes, and antibiotic resistance can increase its pathogenicity (Ahmad-Mansour et al. 2021). Methicillin-resistant S. aureus (MRSA) may display a multidrug-resistant pattern, not only to penicillin but also to others antimicrobial classes including macrolides, fluoroquinolones, aminoglycosides, tetracyclines, and lincosamides (Algammal et al. 2020). Coagulase-negative staphylococci (CoNS) are a group of microorganisms that are increasingly implicated as a cause of significant infections (Michalik et al. 2020) and tend to increase their resistance to antimicrobial agents, especially methicillin and aminoglycosides (Petrillo et al. 2021). In the present study, all S. aureus were resistant to oxacillin, benzylpenicillin, and erythromycin; furthermore, many strains displayed resistance to clindamycin, ciprofloxacin, tetraciclin, and levofloxacin. Also, many CoNS clinical isolates (S. epidermidis and S. saprophyticus) exhibited resistance to oxacillin (41.5%), erythromycin (62.9%), and benzylpenicillin (57.1%). All isolates were susceptible to daptomycin, gentamicin, linezolid, rifampicin, trimethoprim/sulfoxide, tigecycline, teicoplanin, and vancomycin, in accordance with Marincola et al. (2021). All S. aureus strains showed at least one virulence factors tested, while some CoNS strains resulted in positive only for slime production. Slime is a virulence factor related to biofilm formation, frequently involved in pathogenesis in CoNS strains isolated from bacterial keratitis (Fey and Olson 2010; Nayak and Satpathy 2000). In the present investigation, all S. aureus exhibited methicillin resistance gene (mecA), and some strains also displayed staphylococcal enterotoxins (sea), toxic shock syndrome toxin (tst), alpha/beta-hemolysins (hla and hlb), fibronectin-binding proteins (fnbA and fnbB), and adhesins (map/eap). Concerning CoNS strains, 19 S. epidermidis (79.1%) and one S. saprophyticus (33.3%) were slime producers and detected as icaA and icaD positive by PCR analysis (83.3% for S. epidermidis, 33.3% and 66.6% for icaA and icaD, respectively in S. saprophyticus). Also, other authors reported that the percentage of slime producing clinically isolated CoNS strains can vary from 20 to 89% (Preda et al. 2021; Fredheim et al. 2009), a result consistent with those obtained in the present study. The production of slime and the presence of ica genes appear to be involved in the production of biofilm by the microorganism which is thus more resistant to detergents or drugs (Sharma et al. 2023). Recent studies show that the most frequently identified genes in Staphylococcus spp. were hla, hlb, fnbA, fnbB, tst, and map/eap (Khan et al. 2021; Khodabux et al. 2023), as emerged in the present investigation also. These genes encoding specific virulence traits are involved in the pathogenesis of staphylococcal infections. Fibronectin-binding proteins (fnbA and fnbB genes) and the extracellular adhesion protein map/eap help S. aureus adhere to epithelial cells leading to chronic infections. The first step in the process of bacterial infection is the adherence of bacteria to human epithelial cells; this property has been used to define the pathogenicity of an infecting agent. Fibronectin-binding proteins and their corresponding fnbA and fnbB genes have been proposed to be some of the major ligands on the staphylococcal cell surface that help S. aureus adhere to epithelial cells (Khan et al. 2021). The presence of both genes in S. aureus provides strong adherence properties and heightens pathogenicity. In our study, we observed that the 53.8% of S. aureus contained the hla gene and 24% the hlb gene.
The tst gene encodes toxic shock syndrome toxin, an exoprotein that impairs the immunological response of cells, ultimately causing cell death. The Tst gene was detected in 23.7% S. aureus isolates in our study. The percentage was close to these studies (24.5%) (Costa et al. 2019) (28.8%) (Zhao et al. 2019). Other studies recorded highly prevalent Tst gene (72.2%) in MRSA isolates from blood (Peck et al. 2009), whereas in another study, Tst genes were nondetected (Motallebi et al. 2016).
Lastly, the hla gene encodes α-hemolysin, while beta-hemolysin, encoded by the hlb gene, plays an important role in skin and lung infections, respectively.
Conclusions
In this study, staphylococcal isolates recovered from clinical specimens exhibited antibiotic resistance and significant virulence factors, in both MRSA and CoNS strains. All S. aureus and some CoNS strains were resistant to oxacillin, erythromycin, and benzylpenicillin. Concerning virulence characteristics, all S. aureus strains were positive for at least one virulence factors tested. Conversely, only some CoNS isolates were positive exclusively for the slime test, a virulence factor mediated by the ica operon which plays an important role in the pathogenesis of infections. Indeed, the presence of ica genes may be a predictive indicator of virulence for staphylococcal infections. These findings could be useful for a deeper understanding of the function and impact of staphylococcal virulence factors in human infections and for developing new strategies to counteract the spread of both MRSA and CoNS strains.
Data availability
The data presented in this study are available on request from the corresponding authors.
References
Ahmad-Mansour N, Loubet P, Pouget C, Dunyach-Remy C, Sotto A, Lavigne JP, Molle V (2021) Staphylococcus aureus toxins: an update on their pathogenic properties and potential treatments. Toxins (Basel) 13:677. https://doi.org/10.3390/toxins13100677
Algammal AM, Hetta HF, Elkelish A, Alkhalifah DHH, Hozzein WN, Batiha GE, El Nahhas N, Mabrok MA (2020) Methicillin-resistant Staphylococcus aureus (MRSA): one health perspective approach to the bacterium epidemiology, virulence factors, antibiotic-resistance, and zoonotic impact. Infect Drug Resist 13:3255–3265. https://doi.org/10.2147/IDR.S272733
Argemi X, Matelska D, Ginalski K, Riegel P, Hansmann Y, Bloom J, Pestel-Caron M, Dahyot S, Lebeurre J, Prévost G (2018) Comparative genomic analysis of Staphylococcus lugdunensis shows a closed pan-genome and multiple barriers to horizontal gene transfer. BMC Genomics 19:621. https://doi.org/10.1186/s12864-018-4978-1
Argemi X, Hansmann Y, Prola K, Prevost G (2019) Coagulase-negative staphylococci pathogenomics. Int J Mol Sci 20:1215. https://doi.org/10.3390/ijms20051215
Baldassarri L, Simpson WA, Donelli G, Christensen GD (1993) Variable fixation of staphylococcal slime by different histochemical fixatives. Eur J Clin Microbiol Infect Dis 12:866–868. https://doi.org/10.1007/BF02000411
Becker K, Both A, Weißelberg S, Heilmann C, Rohde H (2020) Emergence of coagulase-negative staphylococci. Expert Rev Anti Infect Ther 18:349–366. https://doi.org/10.1080/14787210.2020.1730813
Booth MC, Pence LM, Mahasreshti P, Callegan MC, Gilmore MS (2001) Clonal associations among Staphylococcus aureus isolates from various sites of infection. Infect Immun 69:345–352. https://doi.org/10.1128/IAI.69.1.345-352.2001.Erratum.In:InfectImmun69:1976
Burlak C, Hammer CH, Robinson MA, Whitney AR, McGavin MJ, Kreiswirth BN, Deleo FR (2007) Global analysis of community-associated methicillin-resistant Staphylococcus aureus exoproteins reveals molecules produced in vitro and during infection. Cell Microbiol 9:1172–1190. https://doi.org/10.1111/j.1462-5822.2006.00858.x
Cadieux B, Vijayakumaran V, Bernards MA, McGavin MJ, Heinrichs DE (2014) Role of lipase from community-associated methicillin-resistant Staphylococcus aureus strain USA300 in hydrolyzing triglycerides into growth-inhibitory free fatty acids. J Bacteriol 196:4044–4056. https://doi.org/10.1128/JB.02044-14
Campbell SJ, Deshmukh HS, Nelson CL, Bae IG, Stryjewski ME, Federspiel JJ, Tonthat GT, Rude TH, Barriere SL, Corey R, Fowler VG Jr (2008) Genotypic characteristics of Staphylococcus aureus isolates from a multinational trial of complicated skin and skin structure infections. J Clin Microbiol 46:678–684. https://doi.org/10.1128/JCM.01822-07
Cherifi S, Byl B, Deplano A, Nagant C, Nonhoff C, Denis O, Hallin M (2014) Genetic characteristics and antimicrobial resistance of Staphylococcus epidermidis isolates from patients with catheter-related bloodstream infections and from colonized healthcare workers in a Belgian hospital. Ann Clin Microbiol Antimicrob 4:13–20. https://doi.org/10.1186/1476-0711-13-20
Choi SM, Kim SH, Kim HJ, Lee DG, Choi JH, Yoo JH, Kang JH, Shin WS, Kang MW (2003) Multiplex PCR for the detection of genes encoding aminoglycoside modifying enzymes and methicillin resistance among Staphylococcus species. J Korean Med Sci 18:631–636. https://doi.org/10.3346/jkms.2003.18.5.631
Costa SS, Sobkowiak B, Parreira R, Edgeworth JD, Viveiros M, Clark TG, Couto I (2019) Genetic diversity of norA, coding for a main efflux pump of Staphylococcus aureus. Front Genet 9:710. https://doi.org/10.3389/fgene.2018.00710
European Committee on Antimicrobial Susceptibility Testing EUCAST (2023) Breakpoint tables for interpretation of MICs and zone diameters version 13.1. http://www.eucast.org/clinical_breakpoints
Fey PD, Olson ME (2010) Current concepts in biofilm formation of Staphylococcus epidermidis. Future Microbiol 5:917–933. https://doi.org/10.2217/fmb.10.56
França A, Gaio V, Lopes N, Melo LDR (2021) Virulence Factors in Coagulase-Negative Staphylococci Pathogens 10:170. https://doi.org/10.3390/pathogens10020170
Fredheim EG, Klingenberg C, Rohde H, Frankenberger S, Gaustad P, Flaegstad T, Sollid JE (2009) Biofilm formation by Staphylococcus haemolyticus. J Clin Microbiol 47:1172–1180. https://doi.org/10.1128/JCM.01891-08
García-Malinis AJ, Milagro A, Torres Sopena L, Gilaberte Y (2021) Staphylococcus lugdunensis skin infection: report of 16 cases. Actas Dermosifiliogr (Engl Ed) 112:261–265. English, Spanish. https://doi.org/10.1016/j.ad.2019.05.017
Heilmann C, Ziebuhr W, Becker K (2019) Are coagulase-negative staphylococci virulent? Clin Microbiol Infect 25:1071–1080. https://doi.org/10.1016/j.cmi.2018.11.012
Iseppi R, Sabia C, Bondi M, Mariani M, Messi P (2020) Virulence factors, drug resistance and biofilm formation in Pseudomonas species isolated from healthcare water systems. Curr Microbiol 77:1737–1745. https://doi.org/10.1007/s00284-020-01990-9
Jessen O, Faber V, Rosendal K, Eriksen KR (1959) Some properties of Staphylococcus aureus, possibly related to pathogenicity. Part 1. A study of 446 strains from different types of human infection. Acta Pathol Microbiol Scand 47:316–326
Jiang N, Li J, Feßler AT, Wang Y, Schwarz S, Wu C (2019) Novel pseudo-staphylococcal cassette chromosome mec element (φSCCmecT55) in MRSA ST9. J Antimicrob Chemother 74:819–820. https://doi.org/10.1093/jac/dky457
Khan S, Marasa BS, Sung K, Nawaz M (2021) Genotypic characterization of clinical isolates of Staphylococcus aureus from Pakistan. Pathogens 10:918. https://doi.org/10.3390/pathogens10080918
Khodabux RMJ, Mariappan S, Sekar U (2023) Spectrum of virulence factors in clinical isolates of Staphylococcus aureus and prevalence of SCCmec types in methicillin-resistant Staphylococcus aureus in a tertiary care center. J Lab Physicians 15:450–461. https://doi.org/10.1055/s-0043-1764483
Lachica RV, Genigeorgis C, Hoeprich PD (1971) Metachromatic agar-diffusion methods for detecting staphylococcal nuclease activity. Appl Microbiol 21:585–587. https://doi.org/10.1128/am.21.4.585-587.1971
Lakhundi S, Zhang K (2018) Methicillin-resistant Staphylococcus aureus: molecular characterization, evolution, and epidemiology. Clin Microbiol Rev 31:e00020-e118. https://doi.org/10.1128/CMR.00020-18
Larsson DGJ, Flach CF (2022) Antibiotic resistance in the environment. Nat Rev Microbiol 20:257–269. https://doi.org/10.1038/s41579-021-00649-x
Makris G, Wright JD, Ingham E, Holland KT (2004) The hyaluronate lyase of Staphylococcus aureus - a virulence factor? Microbiology (reading) 150:2005–2013. https://doi.org/10.1099/mic.0.26942-0
Marincola G, Liong O, Schoen C, Abouelfetouh A, Hamdy A, Wencker FDR, Marciniak T, Becker K, Köck R, Ziebuhr W (2021) Antimicrobial resistance profiles of coagulase-negative staphylococci in community-based healthy individuals in Germany. Front Public Health 9:684456. https://doi.org/10.3389/fpubh.2021.684456
Matos JE, Harmon RJ, Langlois BE (1995) Lecithinase reaction of Staphylococcus aureus strains of different origin on Baird-Parker medium. Lett Appl Microbiol 21:334–335. https://doi.org/10.1111/j.1472-765X.1995.tb01073
Metwally LA, Hashem AA, ElAzab SZ, El Islam HN, Mansour MK (2017) Biofilm production and antibiotic resistance of Staphylococcus epidermidis in catheter related bloodstream infections. Egypt J Med Microbiol 26:73–80. https://doi.org/10.12816/0046254
Michalik M, Samet A, Podbielska-Kubera A, Savini V, Międzobrodzki J, Kosecka-Strojek M (2020) Coagulase-negative staphylococci (CoNS) as a significant etiological factor of laryngological infections: a review. Ann Clin Microbiol Antimicrob 19:26. https://doi.org/10.1186/s12941-020-00367-x
Motallebi M, Jabalameli F, Asadollahi K, Taherikalani M, Emaneini M (2016) Spreading of genes encoding enterotoxins, haemolysins, adhesin and biofilm among methicillin resistant Staphylococcus aureus strains with staphylococcal cassette chromosome mec type III. A isolated from burn patients. Microb Pathog 97:34–37
Nashev D, Toshkova K, Salasia SIO, Hassan AA, Lämmler C, Zschöck M (2004) Distribution of virulence genes of Staphylococcus aureus isolated from stable nasal carriers. FEMS Microbiol Lett 233:45–52. https://doi.org/10.1016/j.femsle.2004.01.032
Nayak N, Satpathy G (2000) Slime production as a virulence factor in Staphylococcus epidermidis isolated from bacterial keratitis. Indian J Med Res 111:6–10
Peck KR, Baek JY, Song JH, Ko KS (2009) Comparison of genotypes and enterotoxin genes between Staphylococcus aureus isolates from blood and nasal colonizers in a Korean hospital. J Korean Med Sci 24:585–591. https://doi.org/10.3346/jkms.2009.24.4.585
Petrillo F, Pignataro D, Di Lella FM, Reibaldi M, Fallico M, Castellino N, Parisi G, Trotta MC, D’Amico M, Santella B, Folliero V, Della Rocca MT, Rinaldi M, Franci G, Avitabile T, Galdiero M, Boccia G (2021) Antimicrobial susceptibility patterns and resistance trends of Staphylococcus aureus and coagulase-negative staphylococci strains isolated from ocular infections. Antibiotics (Basel) 10:527. https://doi.org/10.3390/antibiotics10050527
Preda M, Mihai MM, Popa LI, Dițu LM, Holban AM, Manolescu LSC, Popa GL, Muntean AA, Gheorghe I, Chifiriuc CM, Popa MI (2021) Phenotypic and genotypic virulence features of staphylococcal strains isolated from difficult-to-treat skin and soft tissue infections. PLoS ONE 16:e0246478. https://doi.org/10.1371/journal.pone.0246478
Rosenstein R, Götz F (2013) What distinguishes highly pathogenic staphylococci from medium- and non-pathogenic? Curr Top Microbiol Immunol 358:33–89. https://doi.org/10.1007/82_2012_286
Shallcross LJ, Fragaszy E, Johnson AM, Hayward AC (2013) The role of the panton-valentine leucocidin toxin in staphylococcal disease: a systematic review and meta-analysis. Lancet Infect Dis 13:43–54. https://doi.org/10.1016/S1473-3099(12)70238-4
Sharma S, Mohler J, Mahajan SD, Schwartz SA, Bruggemann L, Aalinkeel R (2023) Microbial biofilm: a review on formation, infection, antibiotic resistance, control measures, and innovative treatment. Microorganisms 11:1614. https://doi.org/10.3390/microorganisms11061614
Taylor TA, Unakal CG (2023) Staphylococcus aureus infection. In: StatPearls [Internet]. StatPearls Publishing, Treasure Island (FL). https://www.ncbi.nlm.nih.gov/books/NBK441868/. PMID: 28722898
Tong SY, Davis JS, Eichenberger E, Holland TL, Fowler VG Jr (2015) Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin Microbiol Rev 28:603–661. https://doi.org/10.1128/CMR.00134-14
Zhan XY, Zhu QY (2018) Evolution of methicillin-resistant Staphylococcus aureus: evidence of positive selection in a penicillin-binding protein (PBP) 2a coding gene mecA. Infect GeNet Evol 59:16–22. https://doi.org/10.1016/j.meegid.2018.01.018
Zhao H, Xu S, Yang H, He C, Xu X, Hu F, Shu W, Gong F, Zhang C, Liu Q (2019) Molecular typing and variations in amount of tst gene expression of TSST-1-producing clinical Staphylococcus aureus isolates. Front Microbiol 10:1388. https://doi.org/10.3389/fmicb.2019.01388
Author information
Authors and Affiliations
Contributions
Conceptualization, C.S.; methodology, N.P, R.I., and C.C.; validation, P.M., C.S., and A.D.C.; formal analysis, N.P, C.C., and R.I.; investigation, N.P, C.C., and R.I.; resources, C.S., A.D.C., and P.M.; data curation, S.G., C.C., and R.I.; writing—original draft preparation, C.S., R.I., A.D.C., and P.M.; writing—review and editing, A.D.C., C.C., and S.G.; visualization, C.S., R.I., A.D.C., and P.M.; supervision, C.S.; project administration, P.M. and C.S.; funding acquisition, P.M., C.S., and A.D.C. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pedretti, N., Iseppi, R., Condò, C. et al. Characterization of virulence factors and antimicrobial resistance in Staphylococcus spp. isolated from clinical samples. Folia Microbiol 69, 1043–1052 (2024). https://doi.org/10.1007/s12223-024-01148-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-024-01148-1