Abstract
Soils play an important role in the ecosystem of karstic landscapes both as a buffer zone and as a source of acidity to belowground water. Although the microbiota of karstic soils is known to have a great effect on karstification processes, the activity and composition of these communities are largely unknown. This study gives a comparative analysis of soil microbial profiles from different parts of a doline located at Aggtelek, Hungary. The aim was to reveal the relationships between the vegetation type and genetic fingerprints and substrate utilisation (multi-SIR) profiles of the soil microbiota. Soil samples were collected in early and late springs along a transect in a doline covered with different types of vegetation. Genetic fingerprints of bacterial communities were examined by denaturing gradient gel electrophoresis (DGGE) based on the 16S rRNA gene, along with multi-SIR profiles of the microbial communities measured by the MicroResp method using 15 different carbon sources. Genetic fingerprinting indicated that vegetation cover had a strong effect on the composition of soil bacterial communities. Procrustean analysis showed only a weak connection between DGGE and multi-SIR profiles, probably due to the high functional redundancy of the communities. Seasonality had a significant effect on substrate usage, which can be an important factor to consider in future studies.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Karstic landscapes provide many ecosystem services, such as the production of drinking water for about 25% of the global population (Ford and Williams 2007). Also, because of their special microclimatic effects, karst dolines provide refugee to many vascular plants (Bátori et al. 2014). In the past few centuries, however, human activities had great impact on karstic ecosystems by influencing both karst forming processes and ecosystem services (Móga et al. 2013; Gutiérrez et al. 2014).
The microbial ecology of subsurface waters and caves, and the role of microbes in direct weathering of carbonate rocks and formation of minerals are intensively studied fields (Barton and Northup 2007; Lian et al. 2008; Baskar et al. 2016). Besides these, individual indicator organisms in karstic waters are also studied during the monitoring of human effects on karstic ecosystems (Mulec et al. 2012). The microbial ecology of epikarstic soils, however, is rarely included in the karst researches, despite the crucial role of soil microbiota playing in karstification processes.
The soil layer, when present, plays an important role as a buffer zone of water perturbations, as in most areas, the precipitation filtrates through the soil before reaching the subsurface aquifers (Williams 2008). Epikarstic soils are also hypothesised to have a huge impact on larger scale karst formation as the main source of acidity in subsurface waters (Williams 2008; Phillips 2016).
In Hungary, a few previous studies examined the relationship between karstic processes and soil microbiota. Bárány and Mezősi (1977) showed that in the case of dolines covered with grasslands, the number of culturable microbes in the upper 5 cm soil layer is mainly determined by the temperature, while in deeper layers, it is more related to soil moisture. That said, the number of microbes in different parts of a doline can be greatly influenced by the exposure, especially north vs south of the sampling site. It was also shown that microbial strains from terra rossa type soil types can have higher limestone corrosion abilities, than microbes from rendzinas (Darabos 1999).
Soil pH was recently shown to be the main factor in shaping the composition of belowground bacterial communities of karstic areas (Yun et al. 2016). Different vegetation types, however, are also known to correspond with belowground microbial populations (Hooper et al. 2000), mainly through the composition of plant litter and root exudates (Berg and Smalla 2009). Besides this, Kevei and Zámbó (1985) showed that the density of aerobic culturable bacteria is higher in dolines covered with forests, possibly because of more stable microclimatic – soil moisture and temperature – conditions.
In another study, soil microbial communities from two different Hungarian karst areas were compared by substrate induced respiration (SIR) and genetic fingerprinting by denaturing gradient gel electrophoresis (DGGE) (Knáb et al. 2012). The results showed that the structure of soil bacterial communities were clearly distinct in different sampling sites; however, microbial respiration rates only slightly differed in the top soil but more influenced by the soil depth.
Beside the widespread study of soil microbial community composition, in the last decade, the research of catabolic processes using fingerprinting methods also became a widely used in the assessment of soil functioning and in soil monitoring (Wakelin et al. 2013; Nazaries et al. 2015; Creamer et al. 2016). However, these methods have been rarely applied to epikarstic soils.
Therefore, this study aimed to reveal differences in the DGGE and multi-SIR profiles of soil microbial communities from a typical karst doline in the Aggtelek National Park, Hungary. The relationship between soil properties, vegetation cover and microbiota was investigated, as well as possible correlations between genetic and catabolic fingerprints of the communities. Seasonal changes of the community fingerprints were also assessed.
Materials and methods
Study site and sampling
The doline is located in the karst of Aggtelek National Park, near Lake Vörös (48.4715° N, 20.5426° E, 326 m above sea level, Fig. 1). As a result of reforestation, the southern part and the bottom of the doline is now covered by forest dominated by Acer campestre and Carpinus betulus without significant understory. On the northern slope, the forest is gradually transitioned into diverse grassland which is maintained by mowing and chopping of shrubs, resulting in a mosaic forest-steppe vegetation (see Online Resource 1–5 for photos of the vegetation at the sampling sites). The soils in the doline were identified according to the Hungarian Soil Classification System by Kiss (2012) as brown forest soil with clay illuviation at southern slope, red clay rendzina at northern slope and slope sediment soil at the bottom of the doline, all three developed on red clay. The former two can be classified in the WRB system as leptic Luvisols (clayic, chromic) and the soils from the doline bottom as Regosols.
The samplings were performed at the early (April) and late (June) spring period. Five sampling sites were set in north-south direction, with two sampling sites on the southern slope, one at the bottom of the doline and further two on the northern slope (Fig. 1). Triplicate soil samples from each site were collected by spade from the 0–10 cm layer of 1 m × 1 m quadrates (15 samples altogether). Samples were stored in PE zip-lock bags at 4 °C until the physical-chemical and catabolic experiments. Approximately 10 g soil of each sample was immediately put in sterile tubes and was stored at − 20 °C for molecular analysis.
Soil physical and chemical properties
Soil water content was determined by drying 15 g of each sample at 105 °C. pH-KCl and pH-H2O were determined from 1:2.5 suspensions of soil:KCl and soil:H2O, respectively. Total organic carbon (Corg) content, total salt content (%) from electrical conductivity, and soil texture were also determined. The above measurements were done by the Hungarian standard methodology (Buzás 1988, 1993).
DNA extraction and PCR-DGGE
Total soil community DNA was extracted from approximately 0.25 g soil of each sample using PowerSoil DNA Isolation Kit (MO BIO Laboratories Inc., Carlsbad, CA, USA). The V1 region of the bacterial 16S rRNA gene was amplified by two consecutive PCR (Polymerase Chain Reaction), using first the primers 27F (Lane 1991) and 1401R (Nübel et al. 1996) and a GC-clamp 27F-GC and 519R (Turner et al. 1999) primers thereafter. The PCR mixtures contained 2 μL of purified genomic DNA, 0.2 mM of each deoxynucleotide, 2 mM MgCl2, 1 U LC Taq DNA Polymerase (Fermentas, Vilnius, Lithuania), 1× PCR Buffer (Fermentas, Vilnius, Lithuania) and 0.325 μM of the primers in a final volume of 50 μL. Temperature profile of both PCRs included an initial denaturation at 95 °C for 5 min, followed by 32 cycles (denaturation at 94 °C for 30 s, annealing at 52 °C for 30 s, extension at 72 °C for 1 min) and a final extension at 72 °C for 10 min. The DNA content of the samples was checked by electrophoresis in 1% agarose gel after each step.
DGGE was carried out using an INGENY phorU gel electrophoresis apparatus (Ingeny International BV, Goes, Netherlands), at 60 °C and a charge of 120 V for 14 h in 7% polyacrylamide gel containing 40 to 60% gradient of denaturants (100% was defined as 40% formamide and 7 M urea). The gel was stained by ethidium bromide for 20 min, then the patterns were visualized by an UV transilluminator. The patterns were digitalised for later analysis by taking a photograph.
Soil multi-SIR profiles
Measurements were done with the MicroResp system (Campbell et al. 2003), using the protocol provided by the manufacturer (MSC Ltd., Aberdeen, UK). Soil samples were filled into deep-well microplates (one sample/plate) and covered by Parafilm. They were then put into a desiccator together with a wet towel and sodalime for a 5-day preincubation at room temperature. For the measurements, 15 carbon sources (D-glucose, trehalose, D-galactose, L-arabinose, D-fructose, citric acid, malic acid, Na-succinate, L-arginine, L-alanine, L-leucine, L-lysine, L-glutamine, L-glutamic acid and 3,4-dihydroxybenzoic acid) were used in 6 repetitions on each plate, with distilled water as control. After substrate addition, a 5 h incubation period at 25.2 °C was applied; then, substrate utilization patterns were measured by a microplate reader (Anthos 2010, Biochrom, Cambridge, UK) as absorbance changes of the gel in the detector plates. Absorbance values measured at 570 nm were converted into %CO2 values using the equation given by the manufacturer (%CO2 = A + B / (1 + D × Ai), where A, B and D are constants and Ai are the individual absorbance values after incubation). Detector plates were calibrated on different CO2 concentrations using a gas chromatograph (Fisons GC 8000) prior to the experiment to determine the values of the A, B and D constants. %CO2 values were then converted into CO2 production rates (μgCO2−C × g soil−1 × h−1) using the calculation given by the manufacturer.
Statistical analysis
DGGE patterns were analysed using the TotalLab 120 software (TotalLab LTD., UK). Bands were detected automatically after subtracting the background level with the rolling ball method. Bands were grouped automatically based on their position in the lanes. Automatic detection and grouping were then manually checked for errors, e.g. impurities in the gel detected as bands. Similarity matrices were created on the basis of presence or absence of bands, then dendrograms were generated using the UPGMA method. Raw presence-absence data was then exported for further analysis with the R 3.5.0 (R Core Team 2018). Bands that had zero variance along the samples were removed from the data sets, then principal components analysis (PCA) was applied on the centred data matrix, using the prcomp function of the vegan package (Oksanen et al. 2018). Visualization of the ordinations was done using the ggbiplot package (Vu 2011) with custom modifications for visuals. The ordinations from the two seasons were compared based on the first two principal components, determined by the broken stick method. For the comparison of the two season, symmetric Procrustes test (Peres-Neto and Jackson 2001) was applied using the protest function, with 999 permutations to test the significance of the statistic.
Data from MicroResp were analysed using MS Excel and R. Raw data were processed as described in the MicroResp protocol to determine CO2 production rates (μg CO2-C × g soil−1 × h−1) for each well. Respiration data were standardized by the average respiration rate for each plate. PCA was carried out on the data from the two seasons separately; then, Procrustes test was applied similarly as before, to detect significant differences between the two sampling times. For testing the differences between locations, PERMANOVA was applied with different a priori sample groups, using Bray-Curtis index and 9999 permutations (Anderson 2001).
To determine how well the genetic and catabolic fingerprints correlate, the ordinations from DGGE and MicroResp data were also compared by Procrustes tests.
Results
Soil physical and chemical properties
Soils were slightly acidic, with high organic C content (Table 1.). Soil textures varied between silty loam, silty clay loam and silty clay on different parts of the doline. While the samples from the two slopes had similar texture, the doline bottom had much higher proportion (over 71%) of silt and more variability in the pH. It can be because soils from the slopes were redepositioned by rainfall erosion, resulting in a mixed sediment at the doline bottom which is a common phenomenon (Kiss 2012).
DGGE profiles of bacterial communities
Dendrograms based on the DGGE profiles were similar in the two sampling times but some differences occurred (Fig. 2a and b). From the DB1 sample in late spring, only low amounts of DNA could be extracted thus it was placed in outsider position on the dendrogram. The DB samples clustered with the northern slope of the doline (NSUP and NSL) in the early spring, and with the southern slope (SSUP and SSL) in the late spring. Sites with different vegetation cover—forest, open forest, and grassland—had clearly distinct bacterial communities in both samplings.
The PCA of the data confirmed that the microbial community profiles were different in the five parts of doline, and showed that doline bottom has higher variance, probably because of the slope sediment nature of this area. Temporal differences can also be observed in the relative position of the samples (Online Resource 6 a and b). However, Procrustes test (Fig. 3) indicated that despite the different clustering of the doline bottom samples, the DGGE profiles of the microbial communities was quite similar in the two sampling times (r = 0.855, p = 0.001).
Multi-SIR profiles
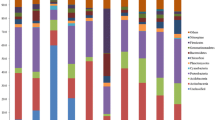
The average respiration rates of the samples were similar in the two sampling times but showed a high variability between parallel samples. The PCA could not reveal differences among the substrate usage patterns of the microbial communities in the late spring but it could separate some of the sample groups in the early spring (Fig. 4a and b). Procrustes test showed that the catabolic fingerprints were very different in the two sampling times (r2 = 0.272, p = 0.581). The PCA of the combined dataset from the two sampling times (Online Resource 4) showed that the main difference was along PC 1, which had strong correlation (|r| > 0.6) with three of the organic substrates (malic acid, lysine and leucine), among which malic acid had the highest loading value (0.94). Early spring samples had higher respiration rates for malic acid (1.32 ± 0.52 μg CO2-C × g soil−1 × h−1 for late spring, vs. 5.5 ± 1.33 μg CO2-C × g soil−1 × h−1 for early spring).
Results of principal components analysis of MicroResp fingerprints from early (a) and late (b) spring period. Ellipses represent the 95% confidence intervals of the groups. Arrows of the biplot correspond to the contribution of the original substrates to the variance represented by the principal components
Although not clearly separated by PCA, further analysis of the samples by PERMANOVA revealed that in the early spring, samples from different parts of the doline had distinct multi-SIR profiles. This difference was the most pronounced (p = 0.0004, pseudo-F = 3.96, all pairwise p-values below 0.05) when the samples were classified into three groups by their geographical location as northern slope, doline bottom, and southern slope. In the late spring, however, no statistically significant difference was found between samples.
Comparisons of MicroResp and DGGE fingerprints differed in the two sampling times. While in the early spring period there was a moderate Procrustean correlation between the two fingerprints (r2 = 0.605, p = 0.004), there was no significant correlation (r2 = 0.187, p = 0.848) between the two fingerprints in the late spring period.
Discussion
Soil microbial density and diversity are known to be influenced by many factors. It was shown recently that in a karstic ecosystem, soil pH is the main parameter influencing the microbial communities (Yun et al. 2016). Although the method used in the present study is not suitable for detailed and quantitative analysis of community structure, and hence a direct comparison with the results of Yun et al. (2016) is not possible, DGGE is an excellent tool for comparative analysis of DGGE profiles (Nielsen et al. 2013), and is often used to indicate changes in the bacterial community structure after soil treatment or different land uses (e.g. Stagniari et al. 2014; Orlewska et al. 2018a, b). In our study, pH was not different significantly in different parts of doline—slightly acidic through the whole transect—still, there were significant differences in the genetic fingerprints of bacterial communities.
It is also known that vegetation can have a strong influence on soil microbial communities (Hooper et al. 2000; Haichar et al. 2014; Khlifa et al. 2017). Our results are similar to those of Li et al. (2014), who found in that different soil microbial communities developed at different successional stages of a karstic. In our study, areas with different vegetation type had clearly different bacterial communities. Plant communities can also affect soil microbes through changes in microclimatic relations. It was previously that areas covered by forests had much higher bacterial counts than those covered by grassland in karst dolines, because of more stable water conditions (Kevei and Zámbó 1985). Zhang et al. (2006) also found that in karstic areas, DGGE fingerprints, basal respiration and SIR rates better reflected the changes in plant coverage and vegetation type than the soil biochemical properties which is in good agreement with our results.
In the case of catabolic activities, however, we found that the relationship between vegetation and multi-SIR profiles was much weaker. In the late spring period, we could not indicate any differences in the substrate usage of microbial communities. In the early spring period, catabolic fingerprints were different on the two slopes and the bottom of the doline, but unlike in the case of DGGE profiles, no further distinction were possible e.g. between shrubs and grassland vegetation. The northern slope was slightly drier in this season; therefore, it is possible that the slightly different soil moisture played a role in the better separation of the samples, because it can have an effect on microbial communities (Kevei and Zámbó 1985).
There were, however, significant differences between the two sampling times, especially in the ability of microbial communities to utilise malic acid. It is already known that microbial communities are affected by root exudates (Haichar et al. 2014), which can possibly result in changes in catabolic fingerprints. Also, plant roots are known to actively exudate malate—along with citrate—to increase phosphate availability in the rhizosphere (Jones 1998), and also to recruit beneficial bacteria from the soil by malic acid exudation (Berendsen et al. 2012). Although drawing conclusions regarding root exudates is beyond the scope of this study, it is possible that the observed differences of catabolic fingerprints are a result of vegetation changes during the seasons. Nevertheless, seasonality clearly has a great effect on soil multi-SIR profiles. This effect is surprisingly rarely addressed in the literature, even in areas other than karst research, but it should definitely be taken into consideration in future studies.
We found that the correlation between genetic and catabolic fingerprints was weak in the early spring, and not significant in the late spring. This, together with the high number of bands in DGGE fingerprints, suggests that karstic soils can have very high microbial diversity and that it can result in significant redundancy of soil functional—in this case, catabolic—diversity (Wertz et al. 2006; Nielsen et al. 2011). Because of this, changes in bacterial community structure might be masked by functional redundancy which might limit the use of MicroResp method in itself for soil monitoring. Our results are in accordance with the study of Zhu et al. (2012), who examined a karstic area with five different vegetation succession stages, and found that DGGE was more sensitive than catabolic fingerprinting.
Conclusions
Our study showed that vegetation differences can have a significant effect on the genetic fingerprint of soil microbial communities in a karst doline. However, these differences are not necessarily reflected in the catabolic activity and diversity because of the high functional redundancy of these soils. Our results also confirmed that seasonality can have a significant effect on the multi-SIR profiles of microbial communities. These effects can have a great influence on the results of soil monitoring and assessment of soil functions and should be taken into consideration in further studies.
Data availability
Not applicable.
References
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Austral Ecol 26:32–46
Bárány I, Mezősi G (1977) Interrelations of some factors of karst corrosion in a Bükk doline. Acta Univ Szeged Acta Geogr 17:133–140
Barton HA, Northup DE (2007) Geomicrobiology in cave environments: past, current and future perspectives. J Cave Karst Stud 69:163–178
Baskar S, Routh J, Baskar R, Kumar A, Miettinen H, Itävaara M (2016) Evidences for microbial precipitation of calcite in speleothems from Krem Syndai in Jaintia Hills, Meghalaya, India. Geomicrobiol J 33:906–933. https://doi.org/10.1080/01490451.2015.1127447
Bátori Z, Csiky J, Farkas T, Vojtkó A, Erdős L, Kovács D, Wirth T, Körmöczi L, Vojtkó A (2014) The conservation value of karst dolines for vascular plants in woodland habitats of Hungary: refugia and climate change. Int J Speleol 43:15–26. https://doi.org/10.5038/1827-806X.43.1.2
Berendsen RL, Pieterse CMJ, Bakker PAHM (2012) The rhizosphere microbiome and plant health. Trends Plant Sci 17:478–486. https://doi.org/10.1016/j.tplants.2012.04.001
Berg G, Smalla K (2009) Plant species and soil type cooperatively shape the structure and function of microbial communities in the rhizosphere. FEMS Microbiol Ecol 68:1–13. https://doi.org/10.1111/j.1574-6941.2009.00654.x
Buzás I (1988) Manual for soil and agrochemical analyses 2: Physico-chemical and chemical analysis of soils. (in Hungarian). Mezőgazda Kiadó, Budapest
Buzás I (1993) Manual for soil and agrochemical analyses 1: physical, water management and mineral analysis of soils. (in Hungarian). INDA 4321 Kiadó, Budapest
Campbell CD, Chapman SJ, Cameron CM, Davidson MS, Potts JM (2003) A rapid microtiter plate method to measure carbon dioxide evolved from carbon substrate amendments so as to determine the physiological profiles of soil microbial communities by using whole soil. Appl Environ Microbiol 69:3593–3599. https://doi.org/10.1128/AEM.69.6.3593
Creamer RE, Stone D, Berry P, Kuiper I (2016) Measuring respiration profiles of soil microbial communities across Europe using MicroResp™ method. Appl Soil Ecol 97:36–43. https://doi.org/10.1016/j.apsoil.2015.08.004
Darabos G (1999) The role of soil microrganisms in karst corrosion. Acta Univ Szeged Acta Geogr TOMUS XXXVII:54–59
Ford D, Williams P (2007) Karst hydrogeology and geomorphology. Wiley, West Sussex
Gutiérrez F, Parise M, De Waele J, Jourde H (2014) A review on natural and human-induced geohazards and impacts in karst. Earth-Sci Rev 138:61–88. https://doi.org/10.1016/j.earscirev.2014.08.002
Haichar FZ e, Santaella C, Heulin T, Achouak W (2014) Root exudates mediated interactions belowground. Soil Biol Biochem 77:69–80. https://doi.org/10.1016/j.soilbio.2014.06.017
Hooper DU, Bignell DE, Brown VK et al (2000) Interactions between aboveground and belowground biodiversity in terrestrial ecosystems: patterns, mechanisms, and feedbacks. Bioscience 50:1049–1061
Jones DL (1998) Organic acids in the rhizosphere–a critical review. Plant Soil 205:25–44. https://doi.org/10.1023/A:1004356007312
Kevei I, Zámbó L (1985) Study of the relationship between bacteria activity in karstic soils and corrosion. Ann Univ Sci Budapestinensis Rolando Eötvös Nomin Sect Geogr 20–21:325–334
Khlifa R, Paquette A, Messier C, Reich PB, Munson AD (2017) Do temperate tree species diversity and identity influence soil microbial community function and composition? Ecol Evol 7:7965–7974. https://doi.org/10.1002/ece3.3313
Kiss K (2012) Analysis of red clay soils of the Aggtelek karst (in the catchment area of Béke cave). Karstdevelopment 17:89–103
Knáb M, Szili-Kovács T, Kiss K, Palatinszky M, Márialigeti K, Móga J, Borsodi A (2012) Comparison of soil microbial communities from two distinct karst areas in Hungary. Acta Microbiol Immunol Hung 59:91–105. https://doi.org/10.1556/AMicr.59.2012.1.10
Lane DJ (1991) 16S/23S rRNA Sequencing. In: Nucleic acid techniques in bacterial systematics. Wiley, Chichester, pp 115–175
Li L, Wang D, Liu X, Zhang B, Liu Y, Xie T, du Y, Pan G (2014) Soil organic carbon fractions and microbial community and functions under changes in vegetation: a case of vegetation succession in karst forest. Environ Earth Sci 71:3727–3735. https://doi.org/10.1007/s12665-013-2767-3
Lian B, Chen Y, Zhu L, Yang R (2008) Effect of microbial weathering on carbonate rocks. Earth Sci Front 15:90–99. https://doi.org/10.1016/S1872-5791(09)60009-9
Móga J, Kiss K, Szabó M et al (2013) Hazards and landscape changes (degradations) on Hungarian karst mountains due to natural and human effects. J Mt Sci 10:16–28. https://doi.org/10.1007/s11629-013-2400-7
Mulec J, Krištůfek V, Chroňáková A (2012) Monitoring of microbial indicator groups in caves through the use of RIDA®COUNT kits. Acta Carsol 41. https://doi.org/10.3986/ac.v41i2-3.565
Nazaries L, Tottey W, Robinson L, Khachane A, al-Soud WA, Sørensen S, Singh BK (2015) Shifts in the microbial community structure explain the response of soil respiration to land-use change but not to climate warming. Soil Biol Biochem 89:123–134. https://doi.org/10.1016/j.soilbio.2015.06.027
Nielsen UN, Ayres E, Wall DH, Bardgett RD (2011) Soil biodiversity and carbon cycling: a review and synthesis of studies examining diversity-function relationships. Eur J Soil Sci 62:105–116. https://doi.org/10.1111/j.1365-2389.2010.01314.x
Nielsen JW, Jordan FL, Maier RM (2013) Analysis of artifacts suggests DGGE should not be used for quantitative diversity analysis. J Microbiol Methods 92:256–263. https://doi.org/10.1016/j.mimet.2012.12.021
Nübel U, Engelen B, Felske A, Snaidr J, Wieshuber A, Amann RI, Ludwig W, Backhaus H (1996) Sequence heterogeneities of genes encoding 16S rRNAs in Paenibacillus polymyxa detected by temperature gradient gel electrophoresis. J Bacteriol 178:5636–5643
Oksanen J, Blanchet FG, Friendly M, et al (2018) Vegan: community ecology package. Version 2.5-2URL https://cran.r-project.org/web/packages/vegan/index.html. Accessed 17 May 2018
Orlewska K, Piotrowska-Seget Z, Cycoń M (2018a) Use of the PCR-DGGE method for the analysis of the bacterial community structure in soil treated with the cephalosporin antibiotic cefuroxime and/or inoculated with a multidrug-resistant Pseudomonas putida strain MC1. Front Microbiol 9:1384. https://doi.org/10.3389/fmicb.2018.01387
Orlewska K, Piotrowska-Seget Z, Bratosiewicz-Wąsik J, Cycoń M (2018b) Characterization of bacterial diversity in soil contaminated with the macrolide antibiotic erythromycin and/or inoculated with a multidrug-resistant Raoultella sp. strain using the PCR-DGGE approach. Appl Soil Ecol 126:57–64. https://doi.org/10.1016/j.apsoil.2018.02.019
Peres-Neto PR, Jackson DA (2001) How well do multivariate data sets match? The advantages of a procrustean superimposition approach over the Mantel test. Oecologia 129:169–178. https://doi.org/10.1007/s004420100720
Phillips JD (2016) Biogeomorphology and contingent ecosystem engineering in karst landscapes. Prog Phys Geogr 40:503–526. https://doi.org/10.1177/0309133315624641
R Core Team (2018) A language and environment for statistical computing. Version 3.5.0. R Foundation for Statistical Computing, Vienna URL https://www.R-project.org/
Stagniari F, Perpetuini G, Tofala R, Campanelli G, Leteo F, Della Vella U, Schirone M, Suzzi G, Pisante M (2014) Long-term impact of farm management and crops on soil microorganisms assessed by combined DGGE and PLFA analyses. Front Microbiol 5:644. https://doi.org/10.3389/fmicb.2014.00644
Turner S, Pryer KM, Miao VPW, Palmer JD (1999) Investigating deep phylogenetic relationships among cyanobacteria and plastids by small subunit rRNA sequence analysis. J Eukaryot Microbiol 46:327–338. https://doi.org/10.1111/j.1550-7408.1999.tb04612.x
Vu VQ (2011) ggbiplot: a ggplot2 based biplot. Version 0.55URL http://github.com/vqv/ggbiplot
Wakelin S, Lombi E, Donner E, MacDonald L, Black A, O’Callaghan M (2013) Application of MicroResp™ for soil ecotoxicology. Environ Pollut 179:177–184. https://doi.org/10.1016/j.envpol.2013.04.010
Wertz S, Degrange V, Prosser JI, Poly F, Commeaux C, Freitag T, Guillaumaud N, Roux XL (2006) Maintenance of soil functioning following erosion of microbial diversity. Environ Microbiol 8:2162–2169. https://doi.org/10.1111/j.1462-2920.2006.01098.x
Williams PW (2008) The role of the epikarst in karst and cave hydrogeology: a review. Int J Speleol 37:1–10
Yun Y, Wang H, Man B, Xiang X, Zhou J, Qiu X, Duan Y, Engel AS (2016) The relationship between pH and bacterial communities in a single karst ecosystem and its implication for soil acidification. Front Microbiol 7:1955. https://doi.org/10.3389/fmicb.2016.01955
Zhang P, Li L, Pan G, Ren J (2006) Soil quality changes in land degradation as indicated by soil chemical, biochemical and microbiological properties in a karst area of southwest Guizhou, China. Environ Geol 51:609–619. https://doi.org/10.1007/s00254-006-0356-4
Zhu H, He X, Wang K, Su Y, Wu J (2012) Interactions of vegetation succession, soil bio-chemical properties and microbial communities in a karst ecosystem. Eur J Soil Biol 51:1–7. https://doi.org/10.1016/j.ejsobi.2012.03.003
Acknowledgements
We thank Klaudia Kiss for her help in sampling and providing information about soil types of the field.
Code availability
Not applicable.
Funding
Open access funding provided by ELKH Centre for Agricultural Research. This research was supported by the Hungarian Scientific Research Fund (OTKA, Grants No. T79135).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Permissions
Permission for conducting research in the area of Aggtelek National Park was issued by the Northern Hungarian Inspectorate for Environmental Protection, Natural Protection and Water (15590-10/2010, 2010.09.27.)
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
Vegetation of the SSUP sampling sites on the southern slope (early spring). (JPG 3717 kb)
ESM 2
Vegetation of the SSL sampling sites on the southern slope (early spring). (JPG 3855 kb)
ESM 3
Vegetation of the DB sampling sites on the doline bottom (early spring). (JPG 3734 kb)
ESM 4
Vegetation of the NSL sampling sites (shrubs) on the northern slope from above (late spring). Due to dense vegetation we could not take informative photos on the site itself. (JPG 4577 kb)
ESM 5
Vegetation of the NSUP (grassland) sampling sites on the northern slope (late spring). (JPG 4684 kb)
ESM 6
Results of Principal Components Analysis of DGGE fingerprints from early (a) and late (b) spring period. Ellipses represent the 95% confidence intervals of the groups. (PPTX 67 kb)
ESM 7
Principal components analysis of the MicroResp data for the early spring (group of point on the right), and late spring (group pf points on the left) samples. Contribution of the original substrates to the variance represented by the principal components are shown as arrows of the biplot. (PPTX 56 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mucsi, M., Krett, G., Szili-Kovács, T. et al. Denaturing gradient gel electrophoresis and multi-SIR profiles of soil microbial communities from a karst doline at Aggtelek National Park, Hungary. Folia Microbiol 66, 107–114 (2021). https://doi.org/10.1007/s12223-020-00828-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-020-00828-y