Abstract
Severe oral mucositis occurs frequently in patients receiving hematopoietic stem cell transplantation (HCT). Oral mucosal bacteria can be associated with progression of oral mucositis, and systemic infection may occur via ulcerative oral mucositis. However, little information is available regarding the oral microbiota after HCT. Here, PCR-denaturing gradient gel electrophoresis (DGGE) was performed to characterize the oral mucosal microbiota, which can be affected by antibiotics, before and after HCT. Sixty reduced-intensity HCT patients were enrolled. Three patients with the least antibiotic use (quinolone prophylaxis and/or β-lactam monotherapy group) and three patients with the most antibiotic use (β-lactam-glycopeptide combination therapy group) were selected. Bacterial DNA samples obtained from the oral mucosa before and after HCT were subjected to PCR-DGGE. The trajectory of oral mucositis was evaluated. The oral mucosal microbiota in the β-lactam-glycopeptide combination therapy group was different from that in the quinolone prophylaxis and/or β-lactam monotherapy group, and Staphylococcus spp. and Enterococcus spp. were identified. Lautropia mirabilis was dominant in one patient. Ulcerative oral mucositis was observed only in the β-lactam-glycopeptide combination therapy group. In conclusion, especially with the use of strong antibiotics, such as glycopeptides, the oral mucosal microbiota differed completely from that under normal conditions and consisted of Staphylococcus spp., Enterococcus spp., and unexpectedly L. mirabilis. The normal oral microbiota consists not only of bacteria, but these unexpected bacteria could be involved in the pathophysiology as well as systemic infection via oral mucositis. Our results can be used as the basis for future studies in larger patient populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oral mucositis occurs in approximately 80% of patients receiving high-dose chemotherapy as conditioning for hematopoietic stem cell transplantation (HCT) (Vera-Llonch et al. 2007). Severe mucositis is associated with not only intolerable pain but also the possible risk of systemic bacteremia. Oral mucosal bacteria may play an important role in the progression of oral mucositis, as well as systemic infection via oral ulcerative mucositis. However, limited information is available regarding the oral microbiota after HCT. Streptococci are common components of the oral mucosal flora, and therefore, streptococcal infection is generally suspected to be of oral origin. However, we have encountered a case in which the gingiva in a patient undergoing leukemia treatment acted as a site of proliferation and a reservoir for multidrug-resistant Stenotrophomonas maltophilia (Soga et al. 2008). We showed that bacterial substitution of mainly coagulase-negative staphylococci (CoNS) for streptococci occurred frequently on the oral buccal mucosa after HCT by surveillance culture of oral mucosa just before and after HCT and discussed the importance of considering the presence of oral mucositis in cases of CoNS infection after HCT (Soga et al. 2011). The oral mucosa can be a reservoir of unexpected bacteria because of bacterial substitution due to the use of many antibiotics. Indeed, empirical antibacterial management has reduced infection-related mortality (Bow 2005), while prophylactic administration of fluoroquinolone was reported to induce fluoroquinolone-resistant Escherichia coli (Bucaneve et al. 2005; Kern et al. 2005), Pseudomonas aeruginosa (Bucaneve et al. 2005), Clostridium difficile (Muto et al. 2005; Pepin et al. 2005), etc. Appropriate empirical antibacterial management is important, while another option would be to not use antibiotic prophylaxis for supportive care in HCT.
Many reports regarding bacteria from clinical laboratories were limited to the genus level both because of examination capacity and clinical necessity. This strategy has limitations to reveal the whole oral mucosal microbiota. Denaturing gradient gel electrophoresis (DGGE) introduced by Fisher and Lerman (1983) and Myers et al. (1987) for mutation analysis or detection of gene polymorphisms has been widely used in microbial ecology (Muyzer et al. 1993). As sequence-specific separation of 16S rDNA amplicons of the same length and further sequencing or hybridization analysis are possible, DGGE has become a powerful tool to examine bacterial diversity in various natural habitats (Muyzer and Smalla 1998), such as marine, lake, and soil environments. This method is often employed in medical fields for examination of polymicrobial communities in humans (Fujimoto et al. 2003; Muyzer and Smalla 1998; Schabereiter-Gurtner et al. 2001).
The severity of clinically evident mucosal damage increases and peaks between 6 and 12 days post-HCT, with resolution of uncomplicated mucositis occurring over the subsequent 7–10 days (Kolbinson et al. 1988; Tardieu et al. 1996). Recently, we reported that the severity of oral mucositis was reduced and the peak day of oral mucositis was delayed in reduced-intensity stem cell transplantation (RIST) patients compared to those receiving conventional HCT, but almost all patients healed within 2–3 weeks after HCT (Takahashi et al. 2010). Therefore, examining the oral microbiota after 1–3 weeks of HCT can contribute to determination of bacteria related to the pathophysiology of oral mucositis and potential bacteria associated with infection via oral mucositis.
This study was performed to determine the changes in the oral mucosal microbiota before and after HCT, which would be affected by antibiotic use, using the DGGE method, and to provide information on the microbiota associated with progression of oral mucositis as well as systemic infection via oral ulcerative mucositis.
Materials and methods
Subjects
A total of 60 consecutive patients (2014–2015) who received HCT at Okayama University Hospital because of hematological malignancies were enrolled in this study. Oral mucosal bacterial sample collection was performed in all subjects, as described below. As DGGE analysis is highly labor intensive, a total of six subjects were chosen to evaluate typical changes in the oral mucosal microbiota after HCT, which would be affected by antibiotic use. Three patients with the least antibiotic use (quinolone prophylaxis and/or β-lactam monotherapy group) and three patients with the most antibiotic use (β-lactam-glycopeptide combination therapy group) were selected. The groups are summarized in Table 1, and the antibiotics used are summarized in Fig. 1. All subjects provided informed consent, and the study was approved by the Ethics Committee of Okayama University Graduate School of Medicine, Dentistry, and Pharmaceutical Sciences (Approval No. 902).
Summary of antibiotics used and the timing of sample collection. Antibiotics used in each patients and the timing of sample collection are shown. LVFX, levofloxacin; SMX/TMP, sulfamethoxazole/trimethoprim; TEIC, teicoplanin; MEPM, meropenem; VCM, vancomycin; CLDM, clindamycin; CFPM, cefepime; DAP, daptomycin. The top three subjects are defined as the quinolone prophylaxis and/or β-lactam monotherapy group, and the bottom three subjects are defined as the β-lactam-glycopeptide combination therapy group. The timings of mucosal bacteria sample collection are also shown by arrows (before HCT: white arrows; after HCT: black arrows). Double lines show the days of engraftment
HCT conditioning regimens
All patients received RIST. In the majority of cases, reduced-intensity conditioning was performed with a fludarabine-based regimen associated with busulfan, melphalan, or cyclophosphamide, as shown in Table 1.
General infection control
All patients were isolated in a room equipped with a laminar airflow system. Patients received azole drugs or micafungin for prophylaxis against fungal infection. Prophylaxis was also given against herpes virus infection with acyclovir. Neutropenic fever was managed according to the guidelines of Hughes et al. (2002). Briefly, empirical antibiotic therapy was administered promptly in all neutropenic patients at the onset of fever and in afebrile patients who were neutropenic but had signs or symptoms compatible with infection. The details of the antibiotics administered in each patient are shown in Fig. 1. G-CSF (lenograstim 5 μg/kg/day or filgrastim 300 μg/m2) was given intravenously for 60 min starting on day 1 or 5 and was continued until the absolute neutrophil count exceeded 500/μL.
Oral management
All subjects were referred to dentists, and necessary dental treatment was completed before HCT. All subjects received instruction regarding self-management of oral hygiene; tooth brushing after every meal and before going to bed and oral rinsing with normal saline solution every 3 h during the day were also indicated. In cases in which the patient’s condition was poor, nurses, dental hygienists, and dentists performed these oral managements. We usually perform surveillance culture of oral mucosa once per week, and antifungal rinses are indicated in cases with the detection of fungi. However, no fungi were detected in these patients, and therefore, no antimicrobial rinses were used.
Assessment of oral mucositis
The severity of oral mucositis in patients undergoing HCT was evaluated every day according to the National Cancer Institute Common Terminology Criteria for Adverse Events (NCI-CTCAE) version 3.0 (http://ctep.cancer.gov/protocolDevelopment/electronic_applications/docs/ctcaev3.pdf, accessed on 31st Dec 2016). The criteria for oral mucositis (clinical exam) were as follows:
-
Grade 1: Erythema of the mucosa
-
Grade 2: Patchy ulcerations or pseudomembranes
-
Grade 3: Confluent ulcerations or pseudomembranes; bleeding in response to minor trauma
-
Grade 4: Tissue necrosis; significant spontaneous bleeding; life-threatening consequences
-
Grade 5: Death
As the clinical examination criteria for oral mucositis are no longer included and only functional/symptomatic criteria are available in the newest version of NCI-CTCAE (4.0), version 3.0 was used to evaluate oral mucositis itself. Assessments were performed as part of daily nursing by nurses under the instruction of dentists and dental hygienists, and the consistency of assessments was checked during the rounds of dentists and dental hygienists at least once per week.
Laboratory culture analysis to identify microorganisms from the oral mucosa and blood
We usually perform surveillance culture from the oral mucosa four times (days − 7 to − 1; days 0 to + 6; days + 7 to + 13; days + 14 to + 20) in all patients. Microbial samples were obtained about 2 h after lunch by swabbing from the whole surface of the buccal mucosa regardless of whether mucositis was observed. Culture and identification of microorganisms were performed at the Central Clinical Laboratory of Okayama University Hospital. Microbial samples from mucosal swabs were plated onto Vitalmedia series-sheep blood agar plates (Kyokuto Pharmaceutical Industrial Co., Ltd., Tokyo, Japan) and cultured under aerobic conditions at 37 °C. Identification of colonies thus obtained was performed using matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS) (MALDI Biotyper; Bruker, Billerica, MA) according to the manufacturer’s instructions.
Blood culture was performed when bloodstream infection was suspected using the automated VITEK system (SYSMEX; bioMérieux, Marcy l’Etoile, France), and the obtained colonies were identified by MALDI-TOF MS (MALDI Biotyper; Bruker) according to the manufacturer’s instructions.
Oral mucosal bacterial sample collection and DNA extraction
Bacterial samples were obtained by gently wiping the buccal mucosa with sterilized cotton swabs before (around 1 week) and after (around 2–3 weeks) HCT. Samples were suspended in 400 μL of PBS (Invitrogen, Carlsbad, CA). Bacterial DNA was extracted using a QIAamp® DNA Mini Kit (QIAGEN, Venlo, The Netherlands).
Polymerase chain reaction denaturing gradient gel electrophoresis analysis
Polymerase chain reaction denaturing gradient gel electrophoresis (PCR-DGGE) fingerprinting analysis of 16S rRNA genes was carried out according to Muyzer et al. (1993) using the primers GC-341f and 534r with Platinum® PCR SuperMix (Invitrogen Ltd., Glasgow, UK). For PCR, we employed a touchdown protocol according to Nishijima et al. (2010) with minor modifications; 2 min of Taq DNA polymerase activation at 94 °C; 20 cycles of denaturation at 94 °C for 15 s, annealing at decreasing temperature from 65 to 56 °C (1 °C decrease for every two cycles) for 15 s, and extension at 68 °C for 30 s; followed by 15 cycles of 94 °C for 15 s, at 55 °C for 15 s, and at 68 °C for 30 s using GeneAmp PCR System 9600 (Applied Biosystems, Foster City, CA). For DGGE analyses, the amplicons and DGGE marker I (Nippon Gene, Tokyo, Japan) were electrophoresed using a D-code DGGE complete system (Bio-Rad, Hercules, CA) operated at 60 °C for 12 h at 100 V in a linear 25–65% denaturing agent gradient (100% denaturing agent consisted of 7 mol/L urea and 40% deionized formamide) with 8% polyacrylamide gels (polyacrylamide gel, ratio of acrylamide HG (Wako Pure Chemical Industries, Osaka, Japan) to bisacrylamide (Wako Pure Chemical Industries), 37.5:1). After DGGE, the gels were soaked for 30 min in SYBR Green I nucleic acid gel stain (1:10000 dilution; Lonza, Rockland, ME) and photographed on a UV transilluminator with a CCD camera.
Counting of DGGE bands
All gels were scanned at 300 dpi. The number of DGGE bands was calculated from the densitometric curves of the scanned DGGE profiles with the CLIQS ver1.1 software (TotalLab Ltd., Newcastle upon Tyne, UK) using the detect band function with the default settings.
Sequence analysis of 16S rRNA gene fragments and homology search
For sequence determination of DGGE bands, the bands were excised from the gel, and the purified DNA was analyzed using an ABI PRISM 3100xl Genetic Analyzer (Applied Biosystems) with a BigDye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems) and the primers for the DGGE band sequencing kit for analysis of the bacterial v3 region (DS-0001) (TechnoSuruga Laboratory Co., Ltd., Shizuoka, Japan). The obtained DNA sequences were subjected to National Center for Biotechnology Information (NCBI)-BLAST against the 16S ribosomal RNA sequence (Bacteria and Archaea) database (https://blast.ncbi.nlm.nih.gov/Blast.cgi accessed on Dec 31, 2016) to identify sequences with the greatest similarity.
Results
DGGE profiles of amplified 16S rDNA from mucosal bacterial samples
PCR-DGGE band patterns representing oral mucosal microbiota before and after HCT are shown in Fig. 2. Bands at the same location before and after HCT samples were observed in the quinolone prophylaxis and/or β-lactam monotherapy group (especially in A1 and A3), while bands were seen at completely different positions between before and after HCT samples in the β-lactam-glycopeptide combination therapy group.
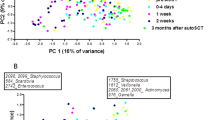
DGGE profiles of amplified 16S rDNA from mucosal bacterial samples of HCT patients. DGGE profiles of amplified 16S rDNA from mucosal bacterial samples of HCT patients are shown. a Quinolone prophylaxis and/or β-lactam monotherapy group. b β-Lactam-glycopeptide combination therapy group. bfr, before HCT sample; aft, after HCT sample. Intense bands (a a–ao, b a–z) were sequenced, and bacterial species identified from the sequence data are shown in Table 2. M, marker
Number of DGGE bands and their changes before and after HCT
The number of band on DGGE decreased markedly after HCT in the β-lactam-glycopeptide combination therapy group: before HCT, 6–12 (median 12); after HCT, 4–6 (median 4). The change in number of bands in the quinolone prophylaxis and/or β-lactam monotherapy group was slight: before HCT, 17–25 (median 20); after HCT, 17–22 (median 21). The number of bands after HCT in the β-lactam-glycopeptide combination therapy group was decreased compared to that in the quinolone prophylaxis and/or β-lactam monotherapy group.
Bacteria composing the oral mucosal microbiota before and after HCT
The bacterial species identified from the major bands on DGGE are shown in Table 2. Band sequence analysis of 16S rRNA gene fragments and homology search were performed as described in the previous section, and in cases where the BLAST search result was 100% identity with multiple bacterial species, all are described in this table. All identified bacteria had > 99% identity.
As shown in Table 2, in the quinolone prophylaxis and/or β-lactam monotherapy group, Streptococcus spp. were frequently identified in samples from both before and after HCT, which was correlated with the results of surveillance oral mucosal culture, as shown in Table 3. A number of bacteria considered to be components of the normal oral microbiota, such as Gemella spp., Veillonella spp., Rothia spp., and Actinomyces spp., were also identified although they were not detected by surveillance culture. Bacteria not detected by surveillance culture were revealed by sequencing of PCR-DGGE bands.
Almost all bacteria identified in the quinolone prophylaxis and/or β-lactam monotherapy group could be explained as normal oral microbiota, while bacteria that are not usually components of the normal oral microbiota, such as Staphylococcus spp. and Enterococcus spp., were identified after HCT in the β-lactam-glycopeptide combination therapy group, which was correlated with the results of surveillance oral mucosal culture as shown in Table 3. Interestingly, Lautropia mirabilis, which is rare, was newly identified only recently (Gerner-Smidt et al. 1994) and was not detected by the culture method, was dominant in one patient.
Trajectory of oral mucositis
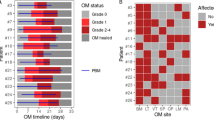
The trajectories of oral mucositis in each patient are shown in Fig. 3. There were no cases in which oral mucositis was suspected to be caused by acute graft versus host disease, and all were induced by the conditioning regimen for HCT. Oral mucositis in all patients in the quinolone prophylaxis and/or β-lactam monotherapy group (A1–A3) was less than grade 1 or zero (limited to redness), while all patients in the β-lactam-glycopeptide combination therapy group (B1–B3) reached grade 2 (ulcerative oral mucositis) around days 7–19. Blood culture was performed when ulcerative mucositis appeared only once for patient B2 on day 7, and Streptococcus agalactiae was identified. This species was also detected in oral surveillance culture performed on day 5.
Trajectory of oral mucositis. The trajectories of oral mucositis in all patients are shown. Oral mucositis of all patients in the quinolone prophylaxis and/or β-lactam monotherapy group (A1–A3) was < grade 1 or zero (limited to redness), while that in all patients in the β-lactam-glycopeptide combination therapy group (B1–B3) reached grade 2 (ulcerative oral mucositis) around days 7–19
Discussion
The oral mucosal microbiota changed rapidly after HCT and was completely different from that before HCT in the β-lactam-glycopeptide combination therapy group. Many antibiotics are used clinically to prevent infections in highly compromised patients, and therefore, such changes in the microbiota can occur in many cases of HCT. The observed decrease in number of bands in the β-lactam-glycopeptide combination therapy group indicated antibiotic selection, and bacteria corresponding to the bands after HCT could be strongly resistant to antibiotics. Bacteria that are not normal components of the oral microbiota, such as Staphylococcus spp., Enterococcus spp., and Lautropia mirabilis, were shown to be involved in the pathophysiology of oral mucositis after HCT (Table 2). Interestingly, all cases of ulcerative mucositis (> grade 2) occurred in the β-lactam-glycopeptide combination therapy group, and the normal oral microbiota disappeared. Our observations suggested that unusual microbiota could lead to progression of oral mucositis. Furthermore, positive blood culture for S. agalactiae was also detected in oral surveillance when ulcerative mucositis appeared in one case, and therefore, infection via oral mucositis was suspected.
While we reported previously that bacterial substitution of mainly coagulase-negative staphylococci (CoNS) for streptococci occurred frequently on the oral buccal mucosa after HCT based on surveillance culture of oral mucosa just before and after HCT (Soga et al. 2011), Enterococcus spp. were identified after HCT in the β-lactam-glycopeptide combination therapy group. Enterococci are part of the normal human microbial flora and are common inhabitants of the human gastrointestinal (GI) tract. Although these species normally constitute a small proportion of the gut microbiota (Eckburg et al. 2005), an important first step toward nosocomial enterococcal infection seems to be increased density of colonization of the GI tract (Arias and Murray 2012). The present study showed that oral mucosal microbiota after HCT with antibiotic treatment could be similar to the conditions in the GI tract and that oral mucositis could be associated with enterococcal bacteremia.
Unexpectedly, L. mirabilis was detected in one subject as one of the major bacteria composing the oral microflora after HCT. L. mirabilis, a motile gram-negative coccus characterized in 1994, has been isolated from oral and pulmonary sites (Gerner-Smidt et al. 1994). Rossmann et al. reported a significant association of L. mirabilis isolation with human immunodeficiency virus (HIV) infection in 1998 (Rossmann et al. 1998). Recent microarray analyses have detected L. mirabilis in the subgingival microbiota of periodontally healthy subjects (Colombo et al. 2012; Colombo et al. 2009), and it was shown to be most abundant in the subgingival microbiota of healthy children (Shaddox et al. 2012). There have been few studies regarding the pathogenicity of L. mirabilis, and this is the first report of L. mirabilis as a dominant species in the oral mucosa. In patient B2, only two bands were clear after HCT, with the clearest being L. mirabilis. Unexpected bacteria, such as L. mirabilis, could be involved in the pathophysiology of oral mucositis after HCT. Furthermore, oral mucositis after HCT could be a route of infection for unusual bacteria, such as L. mirabilis.
The oral mucosa after HCT is a reservoir of unusual bacteria, which may be involved in the pathophysiology of oral mucositis, and systemic infection may occur via ulcerative oral mucositis. The pathogenic effects of bacteria composing the mucosal microbiota and bacterium-epithelial interactions may play important roles in the progression of oral mucositis and be involved in systemic infection via ulcerative oral mucositis. Especially, strong antibiotic therapy causes remarkable selection of microbiota in the oral mucosa, and therefore, candidate bacteria that should be studied with regard to their roles in oral mucositis could appear. Clinically, we would like to emphasize the importance of intensive oral care focusing on the oral mucosa in patients undergoing HCT.
DGGE is a powerful tool to examine bacterial diversity (Muyzer and Smalla 1998), but it is highly labor intensive and difficult to perform. The small number of subjects (limited to six subjects) represented a limitation of this study. However, it was unavoidable with use of the DGGE method, while we observed clear changes in the oral mucosal microbiota in the β-lactam-glycopeptide combination therapy group before and after HCT and showed that Staphylococcus spp., Enterococcus spp., and unexpectedly L. mirabilis were candidates for future studies regarding management of oral mucositis and infection via oral mucositis after HCT. Molecular technology is rapidly advancing, and newly developed techniques will provide further clear information regarding changes in the oral mucosal microbiota. The results presented here can be used as a basis for future studies.
Whether there is an association between these microorganisms and the development and/or presence of ulcerative mucositis could be determined in future studies in a larger patient population and with more sampling points. Other factors, such as the level and duration of neutropenia, and local oral factors, such as changes in the quantity and composition of saliva, may also influence alterations in the microbiome. In addition, in vitro studies using an experimental mucositis model could yield further insights.
In conclusion, we demonstrated changes in the oral mucosal microbiota before and after HCT by the DGGE method, which were affected by the use of antibiotics. Especially with the use of strong antibiotics, such as glycopeptides, the oral mucosal microbiota differed completely from that seen under normal conditions and consisted of Staphylococcus spp., Enterococcus spp., and unexpectedly L. mirabilis. The normal oral microbiota consists not only of bacteria but these unexpected bacteria could also be involved in the pathophysiology of oral mucositis as well as systemic infection via oral mucositis. Our results can be used as the basis for future studies in larger patient populations.
References
Arias CA, Murray BE (2012) The rise of the Enterococcus: beyond vancomycin resistance. Nat Rev Microbiol 10:266–278. https://doi.org/10.1038/nrmicro2761
Bow EJ (2005) Management of the febrile neutropenic cancer patient: lessons from 40 years of study. Clin Microbiol Infect: Off Publ Eur Soc Clin Microbiol Infect Dis 11(Suppl 5):24–29. https://doi.org/10.1111/j.1469-0691.2005.01240.x
Bucaneve G, Micozzi A, Menichetti F, Martino P, Dionisi MS, Martinelli G, Allione B, D'Antonio D, Buelli M, Nosari AM, Cilloni D, Zuffa E, Cantaffa R, Specchia G, Amadori S, Fabbiano F, Deliliers GL, Lauria F, Foà R, del Favero A, Gruppo Italiano Malattie Ematologiche dell'Adulto (GIMEMA) Infection Program (2005) Levofloxacin to prevent bacterial infection in patients with cancer and neutropenia. N Engl J Med 353:977–987. https://doi.org/10.1056/NEJMoa044097
Colombo AP et al (2009) Comparisons of subgingival microbial profiles of refractory periodontitis, severe periodontitis, and periodontal health using the human oral microbe identification microarray. J Periodontol 80:1421–1432. https://doi.org/10.1902/jop.2009.090185
Colombo AP et al (2012) Impact of periodontal therapy on the subgingival microbiota of severe periodontitis: comparison between good responders and individuals with refractory periodontitis using the human oral microbe identification microarray. J Periodontol 83:1279–1287. https://doi.org/10.1902/jop.2012.110566
Eckburg PB, Bik EM, Bernstein CN, Purdom E, Dethlefsen L, Sargent M, Gill SR, Nelson KE, Relman DA (2005) Diversity of the human intestinal microbial flora. Science 308:1635–1638. https://doi.org/10.1126/science.1110591
Fischer SG, Lerman LS (1983) DNA fragments differing by single base-pair substitutions are separated in denaturing gradient gels: correspondence with melting theory. Proc Natl Acad Sci U S A 80:1579–1583
Fujimoto C, Maeda H, Kokeguchi S, Takashiba S, Nishimura F, Arai H, Fukui K, Murayama Y (2003) Application of denaturing gradient gel electrophoresis (DGGE) to the analysis of microbial communities of subgingival plaque. J Periodontal Res 38:440–445
Gerner-Smidt P et al (1994) Lautropia mirabilis gen. nov., sp. nov., a gram-negative motile coccus with unusual morphology isolated from the human mouth. Microbiology 140(Pt 7):1787–1797. https://doi.org/10.1099/13500872-140-7-1787
Hughes WT et al (2002) 2002 guidelines for the use of antimicrobial agents in neutropenic patients with cancer. Clin Infect Dis: Off Publ Infect Dis Soc Am 34:730–751. https://doi.org/10.1086/339215
Kern WV, Klose K, Jellen-Ritter AS, Oethinger M, Bohnert J, Kern P, Reuter S, Baum H, Marre R (2005) Fluoroquinolone resistance of Escherichia coli at a cancer center: epidemiologic evolution and effects of discontinuing prophylactic fluoroquinolone use in neutropenic patients with leukemia. Eur J Clin Microbiol Infect Dis 24:111–118. https://doi.org/10.1007/s10096-005-1278-x
Kolbinson DA, Schubert MM, Flournoy N, Truelove EL (1988) Early oral changes following bone marrow transplantation. Oral Surg, Oral Med, Oral Pathol 66:130–138
Muto CA, Pokrywka M, Shutt K, Mendelsohn AB, Nouri K, Posey K, Roberts T, Croyle K, Krystoflak S, Patel-Brown S, Pasculle AW, Paterson DL, Saul M, Harrison LH (2005) A large outbreak of Clostridium difficile-associated disease with an unexpected proportion of deaths and colectomies at a teaching hospital following increased fluoroquinolone use. Infect Control Hosp Epidemiol 26:273–280. https://doi.org/10.1086/502539
Muyzer G, Smalla K (1998) Application of denaturing gradient gel electrophoresis (DGGE) and temperature gradient gel electrophoresis (TGGE) in microbial ecology. Antonie Van Leeuwenhoek 73:127–141
Muyzer G, de Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
Myers RM, Maniatis T, Lerman LS (1987) Detection and localization of single base changes by denaturing gradient gel electrophoresis. Methods Enzymol 155:501–527
Nishijima M, Lindsay DJ, Hata J, Nakamura A, Kasai H, Ise Y, Fisher CR, Fujiwara Y, Kawato M, Maruyama T (2010) Association of thioautotrophic bacteria with deep-sea sponges. Mar Biotechnol (NY) 12:253–260. https://doi.org/10.1007/s10126-009-9253-7
Pepin J, Saheb N, Coulombe MA, Alary ME, Corriveau MP, Authier S, Leblanc M, Rivard G, Bettez M, Primeau V, Nguyen M, Jacob CE, Lanthier L (2005) Emergence of fluoroquinolones as the predominant risk factor for Clostridium difficile-associated diarrhea: a cohort study during an epidemic in Quebec. Clin Infect Dis: Off Publ Infect Dis Soc Am 41:1254–1260. https://doi.org/10.1086/496986
Rossmann SN, Wilson PH, Hicks J, Carter B, Cron SG, Simon C, Flaitz CM, Demmler GJ, Shearer WT, Kline MW (1998) Isolation of Lautropia mirabilis from oral cavities of human immunodeficiency virus-infected children. J Clin Microbiol 36:1756–1760
Schabereiter-Gurtner C et al (2001) 16S rDNA-based identification of bacteria from conjunctival swabs by PCR and DGGE fingerprinting. Invest Ophthalmol Vis Sci 42:1164–1171
Shaddox LM, Huang H, Lin T, Hou W, Harrison PL, Aukhil I, Walker CB, Klepac-Ceraj V, Paster BJ (2012) Microbiological characterization in children with aggressive periodontitis. J Dent Res 91:927–933. https://doi.org/10.1177/0022034512456039
Soga Y, Saito T, Nishimura F, Ishimaru F, Mineshiba J, Mineshiba F, Takaya H, Sato H, Kudo C, Kokeguchi S, Fujii N, Tanimoto M, Takashiba S (2008) Appearance of multidrug-resistant opportunistic bacteria on the gingiva during leukemia treatment. J Periodontol 79:181–186. https://doi.org/10.1902/jop.2008.070205
Soga Y, Maeda Y, Ishimaru F, Tanimoto M, Maeda H, Nishimura F, Takashiba S (2011) Bacterial substitution of coagulase-negative staphylococci for streptococci on the oral mucosa after hematopoietic cell transplantation. Support Care Cancer: Off J Multinational Assoc Support Care Cancer 19:995–1000. https://doi.org/10.1007/s00520-010-0923-9
Takahashi K, Soga Y, Murayama Y, Udagawa M, Nishimoto H, Sugiura Y, Maeda Y, Tanimoto M, Takashiba S (2010) Oral mucositis in patients receiving reduced-intensity regimens for allogeneic hematopoietic cell transplantation: comparison with conventional regimen. Support Care Cancer: Off J Multinational Assoc Support Care Cancer 18:115–119. https://doi.org/10.1007/s00520-009-0637-z
Tardieu C, Cowen D, Thirion X, Franquin JC (1996) Quantitative scale of oral mucositis associated with autologous bone marrow transplantation. Eur J Cancer. Part B Oral Oncol 32B:381–387
Vera-Llonch M, Oster G, Ford CM, Lu J, Sonis S (2007) Oral mucositis and outcomes of allogeneic hematopoietic stem-cell transplantation in patients with hematologic malignancies. Support Care Cancer: Off J Multinational Assoc Support Care Cancer 15:491–496. https://doi.org/10.1007/s00520-006-0176-9
Acknowledgments
PCR-DGGE analysis and sequence analysis were performed by a professional technical analysis service (Techno Suruga Lab Service, Shizuoka, Japan). We are grateful to Ms. Junko Tomita and Mr. Shunsuke Takahashi, technicians of Techno Suruga Laboratory Co., Ltd., for performing these analyses. We are also grateful to Ms. Motoko Nose, medical technologist of Clinical Laboratory, Central Clinical Department, Okayama University Hospital, Okayama, Japan, for the advice.
Funding
This study was partly supported by a Grant-in-Aid from JSPS KAKENHI Grant Number JP26462881 (to Y.S.), a Grant-in-Aid for the COE projects by MEXT, Japan, entitled “Center of excellence for molecular and gene targeting therapies with micro-dose molecular imaging modalities” (to Y.S.), and a Grant-in-Aid for the COE projects by Ministry of Health, Labour and Welfare, Japan, entitled “Designated regional hub hospital promoting hematopoietic stem cell transplantation” (to M.M.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
All subjects provided informed consent, and the study was approved by the Ethics Committee of Okayama University Graduate School of Medicine, Dentistry, and Pharmaceutical Sciences (Approval No. 902).
Conflict of interest
The authors declare that they have no conflicts of interest.
Rights and permissions
About this article
Cite this article
Muro, M., Soga, Y., Higuchi, T. et al. Unusual oral mucosal microbiota after hematopoietic cell transplantation with glycopeptide antibiotics: potential association with pathophysiology of oral mucositis. Folia Microbiol 63, 587–597 (2018). https://doi.org/10.1007/s12223-018-0596-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-018-0596-1