Abstract
Oxidation of methionine (Met) to Met sulfoxide (MetSO) is a frequently found reversible posttranslational modification. It has been presumed that the major functional role for oxidation-labile Met residues is to protect proteins/cells from oxidative stress. However, Met oxidation has been established as a key mechanism for direct regulation of a wide range of protein functions and cellular processes. Furthermore, recent reports suggest an interaction between Met oxidation and O-phosphorylation. Such interactions are a potentially direct interface between redox sensing and signaling, and cellular protein kinase/phosphatase-based signaling. Herein, we describe the current state of Met oxidation research, provide some mechanistic insight into crosstalk between these two major posttranslational modifications, and consider the evolutionary significance and regulatory potential of this crosstalk.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As an inevitable consequence of aerobic metabolism, cellular components sustain damage from reactive oxygen species (ROS). Proteins are major targets for oxidative modifications, which often cause loss of function that is manifested in a variety of pathophysiological conditions including aging (Curtis et al. 2012). Consequently, oxidative stress is a major determinant of cellular fate, and is involved with cellular malfunction, disease, and death. Proteins can be oxidatively modified by a number of mechanisms at a variety of amino acid residues (Møller et al. 2007). Oxidized proteins can be effectively repaired or recycled when there is a low level of oxidative damage; however, more extensive oxidative damage leads to protein aggregation and cellular dysfunction (Aiken et al. 2011). Carbonylation, for example, is an irreversible modification induced by severe oxidative stress that can affect several amino acid residues (e.g., Arg, Lys, Pro, and Thr). Protein carbonylation results in irreversible loss of function (Møller et al. 2011); although more recently, regulatory and signaling functions in cellular homeostasis have additionally been attributed to carbonylation (Curtis et al. 2012).
While most oxidative modifications are nonenzymatic and irreversible, two modifications—oxidation of Cys to cystine and Met to Met sulfoxide (MetSO)—are enzymatic and reversible. The roles of Cys-oxidation in protein folding and structural stability are manifold (Wedemeyer et al. 2000; Trivedi et al. 2009), and more recently, there has been an increased appreciation of the potential for Cys oxidation/reduction as a component of cellular signaling pathways (Balsera et al. 2014; García-Santamarina et al. 2014; Kim et al. 2014a, b). Furthermore, many enzymes have an active-site Cys residue (e.g., Waszczak et al. 2014). In contrast, there is no established catalytic role for Met, and the structural role of Met can often be replaced by any other hydrophobic residues (Ile, Leu, Phe, and Val; Levine et al. 1996; Rao et al. 2013). Thus, reversible Met oxidation has a separate, distinct, and more specific regulatory potential.
Met oxidation is common
There is a widespread anecdotal belief that Met oxidation in proteins is an artifact of sample preparation (Potgieter et al. 1997). However, the position of MetSO in proteins has a preference for neighboring polar residues indicating a nonrandom pattern (Ghesquière et al. 2011). Based on synthetic MetSO-containing peptides and structural modeling of the MetSO reductase A (MsrA) catalytic center, Ghesquière et al. (2011) concluded that sequence specificity of cellular reductases is the major determinant for the steady-state MetSO/Met. The occurrence of an Arg or Lys residue at the −1 position from MetSO allows the formation of an additional H-bond, which promotes binding of the client protein by MsrA and preferential reduction of MetSO to Met. Conversely, a negatively charged amino acid, such as Asp at the −1 position does not allow the formation of the stabilizing H-bond. This likely explains the strong preference for a negatively charged residue at the −1 position from MetSO observed by Ghesquière et al. (2011). Using 18O stable-isotope labeling to discriminate analytical artifacts, Liu et al. (2013) have verified the occurrence of bona fide, physiological MetSO in target proteins.
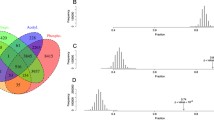
Recent reports suggest that Met oxidation in proteins can be extensive. For example, Ghesquière et al. (2011) identified 2626 instances of MetSO in 1655 proteins from Jurkat cells stressed with hydrogen peroxide. This does not, however, mean that Met oxidation is predominantly found in proteins from stressed cells. Salvato et al. (2014) identified 1373 MetSO sites in 447 proteins (∼40 % of identified proteome) as well as the occurrence of an MsrA and an MsrB in mitochondria from potato tubers. In these analyses, the plant material had been stored under standard commercial conditions which are considered to be “unstressed.” The proteins were denatured and resolved under reducing conditions, and immediately processed for in-gel digestion in the presence of DTT to minimized oxidative modifications. It is notable that mitochondria are the major site of cellular ROS production (Møller 2001; Brand 2010). Song et al. (2012) reported hundreds of oxidized Met residues within thousands of phosphopeptides identified from human liver.
Although most protein amino acid side-chains can be oxidized provided the treatment is sufficiently harsh, Met (and Cys) residues are readily oxidized by even quite mild oxidation agents such as hydrogen peroxide (Shacter 2000) in proteins, and MetSO production therefore often precedes more severe oxidative damages to other amino acid residues (Luo and Levine 2009). It has been proposed that Met residues act as antioxidants that can protect proteins and cells against oxidative stress. In this capacity, it has been termed a “molecular bodyguard” (Levine et al. 1996; Luo and Levine 2009). While surface-exposed residues are the most readily oxidized (Levine et al. 1996), most Met residues in proteins are susceptible to oxidation. Local protein backbone flexibility facilitates oxidation of buried Met residues (Xu et al. 2012). There is, of course, a limit to the capability of the molecular bodyguard and the accumulation of “excessive” MetSO residues has been implicated in aging (Stadtman et al. 2005).
Met oxidation is reversible
Oxidation of Met is reversible (Hoshi and Heinemann 2001), a critical attribute for participation in dynamic regulation of protein function. There are two structural forms or diastereomers of MetSO, S- and R- (Khor et al. 2010). The enzyme MsrA reduces both free and protein-associated S-forms of MetSO, whereas MsrB reduces only the protein-associated R-form. A third enzyme, fRmsr, reduces the free, nonprotein R-form of MetSO (Lee et al. 2009). The MsrA enzyme has been shown to regulate protein function in the context of the redox environment. MsrA-null mice are sensitive to oxidative stress and neurological disorders (Moskovitz 2005). If MetSO is not reduced by MsrA/B, further oxidation yields Met sulfone (MetSO2) and irreversibly damages proteins (Hoshi and Heinemann 2001).
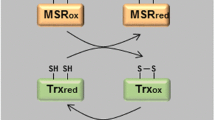
It is generally believed that the conversion of Met to MetSO is nonenzymatic, catalyzed by ROS. However, Lim et al. (2011) recently reported that MsrA, which reduces the S-epimer of MetSO, can also function as a Met oxidase in catalyzing the forward reaction of Met to S-epimer of MetSO (Fig. 1). The ability of the same protein to catalyze both forward and reverse reactions is not without precedent. For example, haloalkaliphilic bacterial arsenate reductase can also function as an oxidase (Richey et al. 2009). It is not yet clear if MsrA functions as an oxidase by a reversible covalent modification-induced conformational change and/or by interaction with a regulatory protein (Lim et al. 2011). Additionally, whether Met oxidase requires an oxidative context to catalyze the forward reaction is unknown.
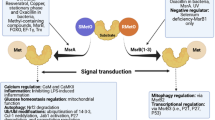
Relevance and context of crosstalk between Met oxidation and O-phosphorylation. a Reversible phosphorylation is unrestricted in the absence of Met oxidation. However, oxidation of Met to MetSO can alter the O-phosphorylation cycle. Crosstalk between the two PTM’s might involve disruption of recognition/binding by either protein kinases or phosphatases. b Structures of Met, MetSO, and MetSO2, and reactions mediating the interconversions. c The bases of crosstalk are manifold, and might depend upon relevance in/to a regulatory network, protein sequence and structure, position of Met and Ser/Thr/Tyr residues, nature of the flanking amino acid residues, functional and evolutionary conservation, and response to stress, hypoxic conditions, and physiological/developmental transitions
Met oxidation can directly regulate protein function
The underlying molecular mechanisms that explain changes in protein function directly caused by Met oxidation have been previously reviewed (e.g., Levine et al. 2000; Drazic and Winter 2014; Kim et al. 2014a), and will only be addressed briefly herein. Apart from a general antioxidant role, Met functions through an oxidation/reduction cycle (Levine et al. 1996, 2000; Stadtman et al. 2002). The roles of reversible Met-oxidation in controlling cellular functions have been previously reviewed (Levine et al. 2000; Drazic and Winter 2014; Kim et al. 2014a, b), and will not be further elaborated herein. Instead, our intent is to focus on cross-talk between reversible Met oxidation and reversible O-phosphorylation (phosphorylation of the -hydroxyl group of Ser/Thr/Tyr), including effects on kinase/phosphatase activities. It is, however, important to highlight an exceptional recent observation. Previously, the invariant sequence of events began with oxidation of Met to MetSO ultimately leading to inhibition/inactivation of downstream events. However, recently, an instance was reported where Met oxidation led to the activation of a protein function. The oxidation of multiple Met residues in the hypochlorite-responsive transcription factor (HypT) was found to stimulate the role that it plays in preventing hypochlorite-induced stress in Escherichia coli (Drazic et al. 2013). Based upon the results from structural modeling of HypT, and comparison with the redox active and structurally similar E. coli transcription factor OxyR, the authors speculated that oxidation-mediated conformational rearrangements in HypT alter contacts within the dimer/tetramer interface and facilitate binding to target DNA. Downstream effects of HypT involve downregulation of ion acquisition and maintenance of intracellular ion levels.
Crosstalk between Met oxidation and O-phosphorylation
While reversible Met oxidation is an effective mechanism to directly regulate many protein activities, interactions (or crosstalk) with other PTM has the potential for a far broader dynamic control of protein activities and cellular processes (Rao et al. 2013). An obvious candidate for such a crosstalk would be O-phosphorylation, the most well-known PTM-based signaling mechanism. Additionally, such crosstalk has the potential for coupling oxidative signals with mainstream regulation and signaling.
The interplay between O-phosphorylation and redox-dependent signaling based on Cys oxidation is well established (Chiarugi and Buricchi 2007). While both modifications are reversible, their effects on protein Tyr-kinase and phosphatase activities are often in opposition. Cys-oxidation frequently results in the activation of Tyr-kinase activity, but typically causes inhibition of Tyr-phosphatases. A similar interplay involving reversible Met oxidation has only recently come to light (Table 1). Kanayama et al. (2002) were perhaps the first to report a possible interaction between Met oxidation and Ser phosphorylation. They observed that Met45 of IκBα in Jurkat cells treated with taurine chloramine could be oxidized, and, when converted to MetSO, the protein was resistant to proteolysis. There are at least two possible explanations for this; firstly, structural obstruction by oxidized Met45 might prevent IκB-kinase (IKK) from approaching the IκBα phosphorylation sites (Ser32 and 36). Secondly, if IKK can still phosphorylate oxidized IκBα, it might then interfere with recognition/binding by the F-box protein that is required for ubiquitination of Lys21 and 22 of IκBα.
Oxidation of a pair of Met residues (281 and 282) in the regulatory domain of Ca2+/calmodulin (CaM)-dependent protein kinase II (CaMKII) has been shown to activate the enzyme in the absence of Ca2+/CaM (Erickson et al. 2008). Oxidation of the Met residues in essence substitutes for the Ca2+/CaM-dependent autophosphorylation of nearby Thr287 that is required for activation of CaMKII. A mouse mutant lacking msrA showed increased CaMKII oxidation and exacerbation of the effect of myocardial infarction. In a related study, Snijder et al. (2011) showed that oxidation of Met144 and 145 in CaM inhibits interaction with CaMKII. Although no phosphorylation is involved in this instance, Met oxidation affects the kinase recognition.
The transient receptor potential melastatin type 6 (TRPM6) protein is primarily found in kidney and colon cells, where it is involved with Mg2+ homeostasis (van der Wijst et al. 2014). This channel protein includes an α-kinase domain, and oxidation of Met1755 within this domain inhibited channel activity in response to H2O2 treatment (Cao et al. 2010). While the direct mechanism of this inhibition remains obscure, TRPM6 activity requires that the ATP-binding motif within the α-kinase domain be occupied. Co-expression of MsrB1 rescues TRPM6 activity, suggesting that reversible oxidation of Met1755 regulates ATP binding and this underlying control mechanism.
Oien et al. (2009, 2011) reported that Met oxidation inhibits the in vitro phosphorylation of yeast or mouse α-synuclein by casein kinase. Furthermore, msrA-null mutants showed lower degradation of α-synuclein protein. Two studies (Hardin et al. 2009; Miernyk et al. 2009) have specifically tested the effect of Met oxidation on the subsequent phosphorylation of nearby Ser residues. Using a multiple combinations of synthetic peptides and recombinant kinases, Hardin et al. (2009) found that Met oxidation strongly inhibits phosphorylation in vitro. They additionally found evidence for an in vivo effect of Met oxidation on the Ser534 phosphorylation of Arabidopsis thaliana leaf nitrate reductase (NR). Oxidation of the +4 Met residue, which serves as a hydrophobic recognition element for the kinase, markedly inhibited phosphorylation. In a related study, also using synthetic peptides and a recombinant kinase, Miernyk et al. (2009) observed that oxidation of the +1 Met of pyruvate dehydrogenase (PDH) strongly inhibited phosphorylation of the adjacent Ser residue.
The H2O2-based oxidation of Met406 of calcineurin, a Ca2+/CaM-activated Ser/Thr-phosphatase, was shown to inhibit CaM binding and thus phosphatase activation (Carruthers and Stemmer 2008). The inhibitory effect of oxidation could be reversed in vitro by treating the phosphatase with MsrA. Results from functional analysis of the Met406Leu mutation suggests that the more hydrophilic MetSO interferes with CaM binding/activation of calcineurin. Thus, Met oxidation has the potential to mediate Ca2+/CaM-based signaling.
There are currently relatively few explicit instances of crosstalk between Met oxidation and O-phosphorylation (Table 1). However, the common occurrence of Met adjacent to Ser/Thr/Tyr residues (Rao et al. 2013) and the frequent occurrence of MetSO (Ghesquière et al. 2011; Liu et al. 2013; Salvato et al. 2014) near O-phosphorylation-sites (Song et al. 2012) suggests that crosstalk between these two PTM’s is widespread.
Bases for crosstalk between Met oxidation and Ser/Thr/Tyr O-phosphorylation
Hardin et al. (2009) were the first to formally propose the idea of coupling oxidative signals with protein phosphorylation via Met oxidation. There are multiple potential mechanisms for interaction between Met oxidation and O-phosphorylation. First, as with NR and PDH (Hardin et al. 2009; Miernyk et al. 2009), oxidation of Met might directly inhibit O-phosphorylation of a nearby Ser/Thr/Tyr-residue by interfering with the kinase recognition/binding (Fig. 1). Second, oxidation of Met in a cohort protein (e.g., CaM) might inhibit recognition/binding of the kinase to the client protein (Snijder et al. 2011). Finally, there might be oxidation of a Met in the kinase sequence itself (for example, CaMKII) which affects activity or signaling efficiency (Cao et al. 2010; Erickson et al. 2008). One might additionally envision similar scenarios involving protein phosphatases (e.g., Carruthers and Stemmer 2008).
In most instances, Met oxidation has a negative effect on subsequent O-phosphorylation (Table 1), although the ultimate downstream readout can be positive. However, as is the case of HypT (Drazic et al. 2013), there are some events immediately downstream where Met oxidation has an immediate positive regulatory effect (e.g., Met oxidation allows Ca2+/CaM-independent activation of CaMKII; Erickson et al. 2008).
The oxidation of Met to MetSO reduces side-chain hydrophobicity, increases the capacity for H-bonding, and alters the spacing of the flanking amino acid residues. It is equally plausible to imagine that if there is an increase in side-chain polarity then the effects of substitution with a nonpolar side chain would be minor. While MetSO is less hydrophobic than Met, it has been reported (Chao et al. 1997) that in some instances, Met oxidation can actually increase the surface-hydrophobicity of a protein, presumably via conformational changes. At our present state of knowledge, it is difficult to predict the structural response upon oxidation of a given Met residue.
If Met oxidation results in a small and relatively subtle change in surface hydrophobicity, for example, then downstream affects might be the result of changes in recognition or binding by Msr, or of a kinase or phosphatase targeting a vicinal Ser/Thr/Tyr residue. However, there is also precedent for Met oxidation causing large-scale changes in protein structure (Griffiths and Cooney 2002; Wolschner et al. 2009; Pan et al. 2010; Marondedze et al. 2013). If this were the case, it might easily lead to exposure of a previously cryptic phosphorylation site or to hide a previously exposed site.
Biological insight into crosstalk between Met oxidation and O-phosphorylation
One physiological response to an adverse environment is avoidance. This can be in the form of encapsulation for microbes (O’Meara and Alspaugh 2012; Lemire et al. 2012) or the formation of propagules such as spores or seeds (McKenney et al. 2013; Nonogaki 2014). Typically, while in the avoiding or replicating state, metabolism is greatly reduced but not curtailed completely (Finch-Savage and Leubner-Metzger 2006), so there is ongoing ROS production (Tudzynski et al. 2012).
Germination of both seeds and spores involves complex signaling pathways (Finkelstein et al. 2008; Setlow 2014) including O-phosphorylation (Arc et al. 2011). It has been proposed (El-Maarouf-Bouteau et al. 2013) that germination takes place when ROS levels are sufficient for signaling but before ROS damage occurs (Dębska et al. 2013). This suggests the importance of crosstalk between Met oxidation and O-phosphorylation at the interface of dormancy, quiescence, and resumption of full biological activity.
Is Met oxidation relevant only in an oxygenic environment? Oxidatively modified proteins/peptides are excellent couriers for communicating the redox status within a cell or subcellular component (Møller et al. 2007; Møller and Sweetlove 2010), and reversible Met oxidation could provide an equally elegant mechanism to link oxidative signals with mainstream cellular signaling in an oxidatively assaultive environment. Reactive oxygen species might be useful for communication among cellular components even under hypoxic conditions. In fact, mitochondria and cells are known to produce ROS under conditions that are virtually anaerobic (Esposti and McLennan 1998). Thus, even in the latter context, there is an opportunity for crosstalk between Met oxidation and O-phosphorylation. The occurrence of MetSO reductase in obligate anaerobes such as the Actinomyces or Bacteroides adds credence to the possibility of PTM crosstalk even in a nonoxygenic environment.
Concluding remarks
The relative simplicity of Met belies the complexity of its structural biochemical functions. As a within-protein antioxidant, Met has been referred to as a “molecular bodyguard.” Under physiological conditions, Met can be readily and reversibly oxidized to MetSO, a reaction that often precedes oxidation of other amino acid residues. The Met redox cycle is dynamic, and has been shown to regulate protein function in a wide range of cellular contexts. Recently, there have been several reports that Met oxidation inhibits neighboring O-phosphorylation. This interaction or crosstalk can depend on several factors: position of the Met residue, structural and evolutionary context of the flanking amino acid sequences, and exposure to oxidative stress (Fig. 1). The reversible oxidation of Met might either directly or indirectly control kinase/phosphatase interactions with their client proteins, and provides an elegant mechanism for coupling the monitoring of ROS with mainstream cellular signaling. Given the importance and ubiquity of ROS and the necessity of cellular signaling, our appreciation for the scope of regulatory crosstalk between Met oxidation and O-phosphorylation can be expected to grow rapidly in the near future.
References
Aiken CT, Kaake RM, Wang X, Huang L (2011) Oxidative stress-mediated regulation of proteasome complexes. Mol Cell Proteomics 10:R110.006924. doi:10.1074/mcp.M110.006924
Arc E, Galland M, Cueff G, Godin B, Lounifi I, Job D, Rajjou L (2011) Reboot the system thanks to protein post-translational modifications and proteome diversity: how quiescent seeds restart their metabolism to prepare seedling establishment. Proteomics 11:1606–1618
Balsera M, Uberegui E, Schürmann P, Buchanan BB (2014) Evolutionary Development of Redox Regulation in Chloroplasts. Antioxid Redox Signal 21:1327–55
Brand MD (2010) The sites and topology of mitochondrial superoxide production. Exp Gerontol 45:466–472
Cao G, Lee KP, van der Wijst J, de Graaf M, van der Kemp A, Bindels RJ, Hoenderop JG (2010) Methionine sulfoxide reductase B1 (MsrB1) recovers TRPM6 channel activity during oxidative stress. J Biol Chem 285:26081–26087
Carruthers NJ, Stemmer PM (2008) Methionine oxidation in the calmodulin-binding domain of calcineurin disrupts calmodulin binding and calcineurin activation. Biochemistry 47:3085–3095
Chao CC, Ma YS, Stadtman ER (1997) Modification of protein surface hydrophobicity and methionine oxidation by oxidative systems. Proc Natl Acad Sci USA 94:2969–2974
Chiarugi P, Buricchi F (2007) Protein tyrosine phosphorylation and reversible oxidation: two cross-talking posttranslation modifications. Antioxid Redox Signal 9:1–24
Curtis JM, Hahn WS, Long EK, Burrill JS, Arriaga EA, Bernlohr DA (2012) Protein carbonylation and metabolic control systems. Trends Endocrinol Metab 23:399–406
Dębska K, Krasuska U, Budnicka K, Bogatek R, Gniazdowska A (2013) Dormancy removal of apple seeds by cold stratification is associated with fluctuation in H2O2, NO production and protein carbonylation level. J Plant Physiol 170:480–488
Drazic A, Winter J (2014) The physiological role of reversible methionine oxidation. Biochim Biophys Acta. doi:10.1016/j.bbapap.2014.01.001
Drazic A, Miura H, Peschek J, Le Y, Bach NC, Kriehuber T, Winter J (2013) Methionine oxidation activates a transcription factor in response to oxidative stress. Proc Natl Acad Sci USA 110:9493–9498
El-Maarouf-Bouteau H, Meimoun P, Job C, Job D, Bailly C (2013) Role of protein and mRNA oxidation in seed dormancy and germination. Front Plant Sci 4:77
Erickson JR, Joiner ML, Guan X, Kutschke W, Yang J, Oddis CV, Bartlett RK, Lowe JS, O’Donnell SE, Aykin-Burns N, Zimmerman MC, Zimmerman K, Ham AJ, Weiss RM, Spitz DR, Shea MA, Colbran RJ, Mohler PJ, Anderson ME (2008) A dynamic pathway for calcium-independent activation of CaMKII by methionine oxidation. Cell 133:462–474
Esposti MD, McLennan H (1998) Mitochondria and cells produce reactive oxygen species in virtual anaerobiosis: relevance to ceramide-induced apoptosis. FEBS Lett 430:338–342
Finch-Savage WE, Leubner-Metzger G (2006) Seed dormancy and the control of germination. New Phytol 171:501–523
Finkelstein R, Reeves W, Ariizumi T, Steber C (2008) Molecular aspects of seed dormancy. Annu Rev Plant Biol 59:387–415
García-Santamarina S, Boronat S, Hidalgo E (2014) Reversible cysteine oxidation in hydrogen peroxide sensing and signal transduction. Biochemistry 53:2560–2580
Ghesquière B, Jonckheere V, Colaert N, Van Durme J, Timmerman E, Goethals M, Schymkowitz J, Rousseau F, Vandekerckhove J, Gevaert K (2011) Redox proteomics of protein-bound methionine oxidation. Mol Cell Proteomics 10:M110.006866
Griffiths SW, Cooney CL (2002) Relationship between protein structure and methionine oxidation in recombinant human alpha 1-antitrypsin. Biochemistry 41:6245–6252
Hardin SC, Larue CT, Oh MH, Jain V, Huber SC (2009) Coupling oxidative signals to protein phosphorylation via methionine oxidation in Arabidopsis. Biochem J 422:305–312
Hoshi T, Heinemann S (2001) Regulation of cell function by methionine oxidation and reduction. J Physiol 531:1–11
Kanayama A, Inoue J, Sugita-Konishi Y, Shimizu M, Miyamoto Y (2002) Oxidation of IκBα at methionine 45 is one cause of taurine chloramine-induced inhibition of NF-κB activation. J Biol Chem 277:24049–24056
Khor HK, Jacoby ME, Squier TC, Chu GC, Chelius D (2010) Identification of methionine sulfoxide diastereomers in immunoglobulin gamma antibodies using methionine sulfoxide reductase enzymes. MAbs 2:299–308
Kim G, Weiss SJ, Levine RL (2014a) Methionine oxidation and reduction in proteins. Biochim Biophys Acta 1840:901–905
Kim HJ, Ha S, Lee HY, Lee KJ (2014b) ROSics: chemistry and proteomics of cysteine modifications in redox biology. Mass Spectrom Rev. doi:10.1002/mas.21430
Lee BC, Dikiy A, Kim HY, Gladyshev VN (2009) Functions and evolution of selenoprotein methionine sulfoxide reductases. Biochim Biophys Acta 1790:1471–1477
Leichert LI (2014) Thiol-based redox processes. Biochim Biophys Acta 1844:1333–1334
Lemire P, Houde M, Lecours MP, Fittipaldi N, Segura M (2012) Role of capsular polysaccharide in Group B Streptococcus interactions with dendritic cells. Microbes Infect 14:1064–1076
Levine RL, Mosoni L, Berlett BS, Stadtman ER (1996) Methionine residues as endogenous antioxidants in proteins. Proc Natl Acad Sci USA 93:15036–15040
Levine RL, Moskovitz J, Stadtman ER (2000) Oxidation of methionine in proteins: roles in antioxidant defense and cellular regulation. IUBMB Life 50:301–307
Lim JC, You Z, Kim G, Levine RL (2011) Methionine sulfoxide reductase A is a stereospecific methionine oxidase. Proc Natl Acad Sci U S A 108:10472–10477
Liu H, Ponniah G, Neill A, Patel R, Andrien B (2013) Accurate determination of protein methionine oxidation by stable isotope labeling and LC-MS analysis. Anal Chem 85:11705–11709
Luo S, Levine RL (2009) Methionine in proteins defends against oxidative stress. FASEB J 23:464–472
Marondedze C, Turek I, Parrott B, Thomas L, Jankovic B, Lilley KS, Gehring C (2013) Structural and functional characteristics of cGMP dependent methionine oxidation in Arabidopsis thaliana proteins. Cell Commun Signal 11:1
McKenney PT, Driks A, Eichenberger P (2013) The Bacillus subtilis endospore: assembly and functions of the multilayered coat. Nat Rev Microbiol 11:33–44
Miernyk JA, Johnston ML, Huber SC, Tovar-Méndez A, Hoyos E, Randall DD (2009) Oxidation of an adjacent methionine residue inhibits regulatory seryl-phosphorylation of pyruvate dehydrogenase. Proteomics Insights 2:15–22
Møller IM (2001) Plant mitochondria and oxidative stress. Electron transport, NADPH turnover and metabolism of reactive oxygen species. Annu Rev Plant Physiol Plant Mol Biol 52:561–591
Møller IM, Sweetlove LJ (2010) ROS signaling—specificity is required. Trends Plant Sci 15:370–374
Møller IM, Jensen PE, Hansson A (2007) Oxidative modifications to cellular components in plants. Annu Rev Plant Biol 58:459–481
Møller IM, Rogowska-Wrzesinska A, Rao RSP (2011) Protein carbonylation and metal-catalyzed protein oxidation in a cellular perspective. J Proteomics 74:2228–2242
Moskovitz J (2005) Methionine sulfoxide reductases: ubiquitous enzymes involved in antioxidant defense, protein regulation, and prevention of aging-associated diseases. Biochim Biophys Acta 1703:213–219
Nonogaki H (2014) Seed dormancy and germination-emerging mechanisms and new hypotheses. Front Plant Sci 5:233
Oien DB, Shinogle HE, Moore DS, Moskovitz J (2009) Clearance and phosphorylation of alpha-synuclein are inhibited in methionine sulfoxide reductase A null yeast cells. J Mol Neurosci 39:323–332
Oien DB, Carrasco GA, Moskovitz J (2011) Decreased phosphorylation and increased methionine oxidation of a-synuclein in the methionine sulfoxide reductase A knockout mouse. J Amino Acids 2011:721094
O’Meara TR, Alspaugh JA (2012) The Cryptococcus neoformans capsule: a sword and a shield. Clin Microbiol Rev 25:387–408
Pan Y, Brown L, Konermanna L (2010) Site-directed mutagenesis combined with oxidative methionine labeling for probing structural transitions of a membrane protein by mass spectrometry. J Am Soc Mass Spectrom 21:1947–1956
Potgieter HC, Ubbink JB, Bissbort S, Bester MJ, Spies JH, Vermaak WJ (1997) Spontaneous oxidation of methionine: effect on the quantification of plasma methionine levels. Anal Biochem 248:86–93
Rao RSP, Xu D, Thelen JJ, Miernyk JA (2013) Circles within circles: crosstalk between protein Ser/Thr/Tyr-phosphorylation and Met oxidation. BMC Bioinforma 14:S14
Richey C, Chovanec P, Hoeft SE, Oremland RS, Basu P, Stolz JF (2009) Respiratory arsenate reductase as a bidirectional enzyme. Biochem Biophys Res Commun 382:298–302
Salvato F, Havelund JF, Chen M, Rao RSP, Rogowska-Wrzesinska A, Jensen ON, Gang DR, Thelen JJ, Møller IM (2014) The potato tuber mitochondrial proteome. Plant Physiol 164:637–653
Setlow P (2014) Germination of spores of Bacillus species: what we know and do not know. J Bacteriol 196:1297–1305
Shacter E (2000) Quantification and significance of protein oxidation in biological samples. Drug Metab Rev 32:307–326
Snijder J, Rose RJ, Raijmakers R, Heck AJ (2011) Site-specific methionine oxidation in calmodulin affects structural integrity and interaction with Ca2+/calmodulin-dependent protein kinase II. J Struct Biol 174:187–195
Song C, Ye M, Liu Z, Cheng H, Jiang X, Han G, Songyang Z, Tan Y, Wang H, Ren J, Xue Y, Zou H (2012) Systematic analysis of protein phosphorylation networks from phosphoproteomic data. Mol Cell Proteomics 11:1070–1083
Stadtman ER, Moskovitz J, Berlett BS, Levine RL (2002) Cyclic oxidation and reduction of protein methionine residues is an important antioxidant mechanism. Mol Cell Biochem 234(235):3–9
Stadtman ER, van Remmen H, Richardson A, Wehr NB, Levine RL (2005) Methionine oxidation and aging. Biochim Biophys Acta 1703:135–140
Trivedi MV, Laurence JS, Siahaan TJ (2009) The role of thiols and disulfides on protein stability. Curr Protein Pept Sci 10:614–625
Tudzynski P, Heller J, Siegmund U (2012) Reactive oxygen species generation in fungal development and pathogenesis. Curr Opin Microbiol 15:653–659
van der Wijst J, Bindels RJ, Hoenderop JG (2014) Mg2+ homeostasis: the balancing act of TRPM6. Curr Opin Nephrol Hypertens 23:361–369
Waszczak C, Akter S, Eeckhout D, Persiau G, Wahni K, Bodra N, Van Molle I, De Smet B, Vertommen D, Gevaert K, De Jaeger G, Van Montagu M, Messens J, Van Breusegem F (2014) Sulfenome mining in Arabidopsis thaliana. Proc Natl Acad Sci USA 111:11545–11550
Wedemeyer WJ, Welker E, Narayan M, Scheraga HA (2000) Disulfide bonds and protein folding. Biochemistry 39:4207–4216
Wolschner C, Giese A, Kretzschmar H, Huber R, Moroder L, Budisa N (2009) Design of anti- and pro-aggregation variants to assess the effects of methionine oxidation in human prion protein. Proc Natl Acad Sci USA 106:7756–7761
Xu K, Uversky VN, Xue B (2012) Local flexibility facilitates oxidization of buried methionine residues. Protein Pept Lett 19:688–697
Acknowledgments
Research in the Thelen Lab is supported by a grant from the National Science Foundation Plant Genome Research Program Grant No. DBI-0604439 and research in the Møller lab was supported by a grant from the Danish Council for Independent Research-Natural Sciences.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rao, R.S.P., Møller, I.M., Thelen, J.J. et al. Convergent signaling pathways—interaction between methionine oxidation and serine/threonine/tyrosine O-phosphorylation. Cell Stress and Chaperones 20, 15–21 (2015). https://doi.org/10.1007/s12192-014-0544-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12192-014-0544-1