Abstract
In the present study, we studied downstream signals of BCR-ABL with regard to Src family kinases and YAP, a transcription cofactor and an effector of the Hippo pathway. We first checked the phosphorylation status of YAP and found that it was constitutively phosphorylated at tyrosine 357 in CML-derived cell lines (TCC-S and K562) but not in AML-derived cell lines (HL-60 and KG-1a). Treatment with imatinib or RK-20449 inhibited cell growth and decreased tyrosine phosphorylation of YAP in both CML lines. Expression of Survivin or Cyclin D1 was decreased in TCC-S, but not in either HL-60 or KG-1a. Furthermore, we established BCR-ABL stable transfectant and control empty vector transfectant from TF-1, a factor-dependent human erythroleukemia cell line, to verify our results obtained with CML cell lines. YAP was phosphorylated at Y357 constitutively in BCR-ABL stable transfectant but not in control transfectant, and treatment with imatinib or RK-20449, a Src family kinase-specific inhibitor, inhibited cell growth, YAP tyrosine phosphorylation, and expression of Cyclin D1 in BCR-ABL stable transfectant. These results suggest that BCR-ABL induces tyrosine phosphorylation of YAP presumably through Src family kinases, which results in expression of Survivin and Cyclin D leading to leukemogenesis in CML cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chronic myeloid leukemia (CML) is caused by the fusion protein BCR-ABL generated by a reciprocal chromosomal translocation t(9;22) (q34;q11) [1]. To date, several downstream signaling pathways have been reported to underlie the leukemogenesis of CML such as the JAK/STAT pathway, the PI3 K/AKT pathway, and the Grb2/MAPK pathway [2]. For example, the JAK/STAT pathway plays an important role in cellular transformation, because conditional knockout of STAT5 suppressed the cell growth in BCR-ABL-induced CML mouse [3]. Inhibition of PI3K signaling suppresses the growth of CML cells, suggesting that PI3K/AKT is involved in BCR-ABL-mediated leukemogenesis [4]. BCR-ABL has been reported to induce Grb2-mediated MAPK activation, which results in transformation of primary human hematopoietic cells [5]. Furthermore, the Src family kinases (SFKs), especially Hck, Lyn, and Fyn, have been suggested to be involved in BCR-ABL-induced transformation [6]. Although these studies have revealed important aspects of the downstream signals of BCR-ABL, the detailed molecular mechanism of CML has not been thoroughly elucidated.
Concerning the SFKs involvement, the tyrosine phosphatase SHP2 that is known to be required for BCR-ABL-mediated hematopoietic cell transformation [5] plays a key role in their activation. Namely, BCR-ABL, via its phosphorylated Tyr177, recruits the adapter GRB2/GRB2-associated binding protein 2 (GAB2) complex and SHP2. Then activated SHP2 dephosphorylates PAG1 and paxillin, allowing SFKs to be activated through dissociation from CSK [7].
Yes-associated protein (YAP) is a transcriptional cofactor that functions as an effector of the Hippo pathway which regulates cell growth and survival. In the classical Hippo pathway, YAP phosphorylated at serine 127 (S127) by LATS1/2 is bound to 14-3-3 and prevented from nuclear translocation [8]. Apart from this serine/threonine phosphorylation, YAP undergoes phosphorylation at several tyrosine residues by various kinases to be activated [9]. SFKs can phosphorylate and activate YAP, which has been demonstrated in some tumors [10] [11]. Among the several possible phosphorylated tyrosine residues, the phosphorylation at Y357 (p-Y357) has been demonstrated to be the most important for tumorigenesis [12]. Therefore, it is possible that BCR-ABL directly or indirectly phosphorylates YAP through SFKs, and thus activated YAP is translocated into the nucleus and together with TEAD induces the expression of genes necessary for cell growth and survival. With regard to this point, it should be noted that cell cycle regulator protein Cyclin D1 [13] and anti-apoptotic factor Survivin [14], have been reported to be YAP dependent and highly expressed in CML cells [15].
In the present study, we investigated the effects of imatinib and an SFK-specific inhibitor RK-20449 [16] on viable cell number, YAP p-Y357 and expression of Survivin as well as Cyclin D1 in CML-derived cell lines in comparison with AML-derived cell lines. Furthermore, we established BCR-ABL stable transfectants and the control lines derived from TF-1, a factor-dependent human erythroleukemia cell line, to verify our results obtained with CML-derived cell lines.
Materials and methods
Cell lines and culture
CML-derived BCR-ABL-positive cell lines, TCC-S [17] and K562, and AML-derived BCR-ABL-negative cell lines, HL-60 and KG-1a, were cultured in RPIM1640 medium supplemented with 10% fetal bovine serum (FBS) and antibiotics. Erythroleukemia-derived TF-1 [18] was cultured in the same medium in the presence of 1 ng/ml recombinant human GM-CSF.
Plasmids and antibodies
BCR-ABL cDNA was subcloned from MSCV-BCR-ABL-IRES-GFP (a kind gift of Dr. Jian) into pcDNA3.1.
The following antibodies were used in Western blotting: anti-YAP1 (Abnova, Taipei City, Taiwan), anti- Phospho-YAP (Y357) (Abcam, Cambridge, UK), anti-c-Abl (Cell Signaling Technology, Inc., Danvers, MA), anti-GAPDH (Cell Signaling Technology), HRP-linked anti-mouse IgG (Cell Signaling Technology).
Cell proliferation assay
Cells were cultured at 2 × 105/ml in 96-well flat-bottomed plates in the absence or presence of serially diluted imatinib mesylate (Wako Pure Chemical, Osaka, Japan) or an SFK-specific inhibitor RK-20449 (Cayman Chemical, Ann Arbor, MI) for 48 h at 37 °C in a CO2 incubator. Then viable cells were counted using a microscope by trypan blue exclusion.
Western blot analysis
Expression of YAP and its tyrosine phosphorylation was visualized by SDS-PAGE and Western blotting. In brief, cells were harvested and lysed in lysis buffer with Phosphatase Inhibitor Cocktail (Nakalai Tesque, Kyoto, Japan), Protease Inhibitor Cocktail EDTA free (Nakalai Tesque), and phenylmethylsulfonyl fluoride, and applied to SDA-PAGE. After electrophoresis, samples were transferred to Immobilon-P Transfer Membrane (Millipore, Bedford, MA). Sample bands were visualized by chemiluminescence using Amersham Imager 600 (GE Healthcare Life Science, Marlborough, MA). In some experiments, densitometric analysis was done with the data files using Image J program and the ratio of YAP/GVHD was calculated for each sample.
Quantitative PCR
RNA was isolated with Sepasol-RNA (Nakalai Tesque) and used as template for reverse transcription reaction with RevatraAce (Toyobo, Osaka, Japan) to make cDNA. Diluted cDNA was mixed with Fast SYBR Green Master Mix (Thermo Fisher Science, Waltham) and quantitative PCR (qPCR) was done with StepOnePlus Real-time PCR System (Thermo Fisher) using GAPDH as an internal control. The primers used are as follows: human Survivin forward TGAACTTCAGGTGGATGAGGAGA, reverse GTCTAATCACACAGCAGTGGCAA, human Cyclin D1 forward AACTACCTGGACCGCTTCTT, reverse CCACTTGAGCTTGTTCACCA, human GAPDH forward GTCAGCCGCATCTTCTTTTG, reverse GCGCCCAATACGACCAAATC.
Establishment of BCR-ABL stable transfectant lines
TF-1 cells were transfected with linearized pcDNA3.1-BCR-ABL or empty vector by electroporation using Neon® Transfection System (Thermo Fisher Scientific, Waltham, MA), and after 48 h, cells were subjected to selection with 800 μg/ml G418. After selection, stable transfectant clones were isolated by limiting dilution and checked for expression of BCR-ABL by Western blotting.
Results
CML cells are sensitive not only to imatinib but also to SFKs inhibitor
We first confirmed the sensitivity of CML cell lines, TCC-S and K562, as well as BCR-ABL-negative AML cell lines, HL-60 and KG-1a, to imatinib. Cells were cultured at 2 × 105/ml in 96-well plates in the absence or presence of serially diluted imatinib for 48 h at 37 °C in a CO2 incubator. Then viable cells were counted using a microscope by trypan blue exclusion. As shown in Fig. 1a, TCC-S was sensitive to relatively low concentrations of imatinib while K562 was less sensitive compared to TCC-S but responded to high concentrations of imatinib. Neither of the AML cell lines was sensitive to imatinib.
Effect of imatinib and RK-20449 on cell proliferation of TCC-S, K562, HL-60, and KG-1a. Cells were cultured in the absence or presence of serial dilutions of imatinib (a) or RK-20449 (b) in flat-bottomed 96-well plates for 48 h. Viable cells were counted by trypan blue dye exclusion. Bars indicate S.E. of the mean (n = 3)
Next, we checked the sensitivity of these cell lines to an SFK-specific inhibitor RK-20449 in a similar way. As shown in Fig. 1b, TCC-S was sensitive to RK-20449, while K562 and KG-1a needed higher concentrations of RK-20449. HL-60 was hardly sensitive to RK-20449.
YAP is constitutively phosphorylated at Y357 in CML cell lines
To explore the possible involvement of YAP in the downstream signaling of BCR-ABL, we examined the phosphorylation of YAP at tyrosine 357 (p-Y357) in these cell lines. Western blot analysis using anti-YAP p-Y357 Ab indicated that YAP of both CML cell lines was phosphorylated at Y357 constitutively while such a band was scarcely detected in either of AML cell lines. Treatment with imatinib decreased YAP p-Y357 in TCC-S while much higher concentrations of imatinib were required to decrease YAP p-Y357 in K562, which was compatible to its low sensitivity to imatinib in cell proliferation assay (Fig. 2a). These data suggest that BCR-ABL induces YAP p-Y357 directly or indirectly in CML cells, enabling them to proliferate. In the case of K562, it is possible that another tyrosine phosphorylation site on YAP or some signal transduction targets of SFKs other than YAP may be involved in proliferation in this particular cell line.
Phosphorylation of YAP at Y357 in TCC-S, K562, HL-60, and KG-1a. After culture with serial dilutions of imatinib (a) or RK-20449 (b) for 12 h, cell lysates were analyzed by Western blotting using anti-YAP p-Y357 (Abcam, Cambridge, UK), anti-YAP (Abnova, Taipei City, Taiwan), anti-c-ABL (CST, Danvers, MA), and anti-GAPDH (CST) antibodies
We next examined the effect of RK-20449 on YAP p-Y357 in these cell lines. Treatment with RK-20449 decreased YAP p-Y357 in both CML cell lines (Fig. 2b), suggesting that SFKs are involved in tyrosine phosphorylation of YAP in CML cells. Since this reagent is highly specific for SFKs (IC50 less than 40 nM) and has ~ 100 times less inhibitory effect on c-ABL than imatinib [16], the inhibition of YAP p-Y357 was likely to be a direct effect on SFKs rather than BCR-ABL.
In these Western blots, we measured and compared the relative expression levels of YAP by densitometry. The YAP/GAPDH ratios in the four cell lines were as follows, HL-60 0.87, KG-1a 0.71, TCC-S 0.79, and K562 1.91, indicating that K562 but not TCC-S expressed higher levels of YAP than AML cell lines.
BCR-ABL and SFKs are involved in YAP-dependent gene expression
Then the next question was whether BCR-ABL or SFKs are involved in YAP-dependent gene expression. So, we examined the effects of imatinib and RK-20449 on expressions of Survivin and Cyclin D1 mRNA which are both YAP dependent and closely associated with CML leukemogenesis. As shown in Fig. 3, treatment with imatinib decreased the expression of Survivin as well as Cyclin D1 in TCC-S but not in the AML cell lines. As expected, K562 was less sensitive to imatinib in terms of Survivin and Cyclin D1 expression and needed much higher concentrations of imatinib.
Expressions of Survivin and Cyclin D1 mRNA in HL-60, KG-1a, TCC-S, and K562. After culture with serial dilutions of imatinib (a, c) or RK-20449 (b, d) for 24 h, total RNA was isolated and expression of Survivin (a, b) or Cyclin D1 (c, d) mRNA was measured by qPCR. The primers used in these experiments were as follows: Survivin forward TGAACTTCAGGTGGATGAGGAGA, Survivin reverse GTCTAATCACACAGCAGTGGCAA, Cyclin D1 forward AACTACCTGGACCGCTTCTT, Cyclin D1 reverse CCACTTGAGCTTGTTCACCA, GAPDH forward GTCAGCCGCATCTTCTTTTG, GAPDH reverse GCGCCCAATACGACCAAATC. Bars indicate S.E. of the mean (n = 3)
Treatment with RK-20449 also decreased the expression of Survivin and Cyclin D1 in TCC-S, while it had no or only weak effect in K562 and the AML cell lines. These results suggest that certain SFKs may be involved in the signaling pathway from BCR-ABL to Survivin and Cyclin D1 expression at least in TCC-S. With regard to the results with K562, it is possible that some of SFKs or other tyrosine kinases might be activated independent of BCR-ABL and play roles in induction of Survivin and Cyclin D. It is likely that TCC-S is more dependent on the signals of BCR-ABL than K562 which seems to have various genetic abnormalities other than BCR-ABL that generate signals for cell growth and survival.
Establishment of stable transfectants of BCR-ABL from TF-1
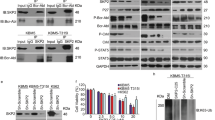
In abovementioned experiments, we used two AML cell lines, KG-1a and HL-60, as controls for CML cell lines. However, they have distinct genetic abnormalities from CML cell lines, and therefore are not considered to be appropriate controls. Therefore, we established BCR-ABL stable transfectant lines from TF-1, an erythroleukemia-derived factor-dependent cell line [18]. TF-1 cells were transfected with pcDNA3.1-BCR-ABL or empty vector by electroporation using Neon® Transfection System (Thermo Fisher Scientific, Waltham, MA) and subjected to G418 selection. Several BCR-ABL stable transfectant lines were isolated which turned out to be very similar to each other and able to grow without GM-CSF. Here, we show the data of Western blot using anti-c-ABL Ab with a representative stable line of BCR-ABL (TF-1-BCR-ABL) and a control line of empty vector (TF-1-EV) (Fig. 4a).
Establishment of TF-1-BCR-ABL and its characterization. Expression of BCR-ABL protein in the TF-1-derived stable transfectant in Western blotting using anti-c-Abl Ab (CST) (a). Stable transfectant TF-1-BCR-ABL was sensitive to imatinib (b) as well as RK-20449 (c). YAP was constitutively phosphorylated at Y357 in TF-1-BCR-ABL and this phosphorylation was inhibited by either imatinib or RK-20449 (d). Expression of Survivin mRNA in TF-1-BCR-ABL was not inhibited by imatinib or RK-20449 (e), while expression of Cyclin D1 mRNA was inhibited by these reagents (f). Bars indicate S.E. of the mean (n = 3). As to Fig. 4e, relative amount of Survivin of each sample as the ratio to GAPDH instead of comparison with time 0 was as follows: ΔCt (Survivin Ct—GAPDH Ct) of TF-EV with GM-CSF, TF-EV without GM-CSF, TF-BCR-ABL were 4.13, 3.60, and 3.78, respectively
Stable BCR-ABL transfectant is sensitive to both imatinib and RK-20449
Then we examined the sensitivity of the transfectants to imatinib as well as RK-20449 by viable cell counting. As shown in Fig. 4b and c, GM-SCF independent growth of TF-1-BCR-ABL was sensitive to both reagents, while GM-CSF-dependent growth of TF-1-EV was not, indicating that signals of BCR-ABL and certain SFKs are crucial for the autonomous growth of TF-1-BCR-ABL.
Next, we examined whether the tyrosine phosphorylation of YAP that we detected in CML cell lines could be detected in the stable transfectants. As shown in Fig. 4d, YAP was constitutively phosphorylated at Y357 in TF-1-BCR-ABL but not TF-1-EV even in the presence of GM-CSF, and this phosphorylation was inhibited by treatment with imatinib or RK-20449 in a dose-dependent manner, indicating that BCR-ABL and SFKs are involved in tyrosine phosphorylation of YAP at Y357.
Unexpectedly, expression of Survivin was inhibited neither by imatinib nor RK-20449 in TF-1-BCR-ABL. With regard to this, we noticed that parental TF-1 with or without GM-CSF expressed high levels of Survivin, which was suggested by relative amounts of Survivin at time 0: △Ct (Survivin Ct—GAPDH Ct) of TF-EV with GM-CSF, TF-EV without GM-CSF, and TF-BCR-ABL were 4.13, 3.60, and 3.78, respectively. In addition, original TF-1 has been reported to express high levels of Survivin that is dependent on STAT3 [19]. Therefore, it is clear that this cell line was not suitable for analysis of BCR-ABL-induced expression of Survivin. On the other hand, expression of Cyclin D1 in TF-1-BCR-ABL was inhibited by imatinib as well as RK-20499, suggesting that BCR-ABL and SFKs are involved in autonomous cell growth of this transfectant.
Discussion
In the present study, we investigated the role of YAP in the leukemogenesis of CML with regard to its tyrosine phosphorylation. We found that YAP was constitutively phosphorylated at Y357 in CML cell lines but not in AML cell lines. Not only imatinib but also RK-20449 inhibited this phosphorylation and at the same time expression of Survivin and Cyclin D1, both YAP-dependent genes. Experiments with TF-1 derived stable transfectants indicated that YAP was phosphorylated at Y357 in TF-1-BCR-ABL but not in TF-1-EV, and again this phosphorylation and expression of Cyclin D1 were inhibited by imatinib or RK-20449. These results suggest that BCR-ABL induces these gene expressions through the activation of SFKs and subsequent tyrosine phosphorylation of YAP.
Accumulating evidence has indicated that YAP plays a critical role in progression of various cancers [20,21,22]. YAP promotes malignant progression through the expression of Survivin [21] and Cyclin D1 [22]. Nuclear translocation and activation of YAP are not only induced by dephosphorylation at S127 but also by tyrosine phosphorylation at Y357 [23, 12]. Upon DNA damage, c-Abl phosphorylates YAP at Y357 and induces its association with p73 resulting in cellular apoptosis [24]. It has not been determined, however, whether BCR-ABL directly phosphorylates YAP at Y357 without DNA damage. Our results suggest that BCR-ABL induces YAP p-Y357 directly or indirectly via SFKs, leading to YAP-dependent gene expression for cell growth and cell survival.
As to the involvement of SFKs in pathogenesis of CML, Hu et al. reported that Lyn, Hck, and Fgr are required for BCR-ABL-induced B ALL but not CML because retrovirus-transduced marrow from mice lacking all three Src kinases efficiently induced CML but not B-ALL in recipients. In addition, the kinase inhibitor CGP76030 impaired the proliferation of B-lymphoid cells expressing Bcr-Abl in vitro and prolonged survival of mice with B-ALL but not CML [25]. However, in our view, this report did not demonstrate that any SFKs are not required for CML, because in their knockout mice models, BCR-ABL might use some SFKs other than Lyn, Hck, and Fgr to cause CML due to the redundancy among SFKs. Furthermore, CGP76030 is now known to be not specific for SFKs.
A recent paper has suggested that YAP/TAZ is dispensable for normal as well as pathological hematopoiesis, using an established orthotopic mouse model of AML, based on the retroviral transduction of hematopoietic progenitor cells with the MLL-AF9 oncogene [26]. It should be noted that their claim is that YAP/TAZ is dispensable in the particular case of MLL-AF9-caused leukemia and may not be applicable to other types of leukemia. Their results should not be generalized to other hematopoietic malignancies.
In fact, Li et al. have reported that inhibition of YAP suppresses CML cell proliferation and enhances the efficacy of imatinib in vitro and in vivo [27]. They showed that YAP is upregulated in samples from CML patients and CML cell lines and silencing of YAP inhibits the proliferation of CML cells. In the present study, we observed that YAP expression was upregulated in K562 but not TCC-S compared to two AML cell lines, suggesting that their claim is not always adequate in CML. Instead, we demonstrated for the first time that YAP is constitutively phosphorylated in CML cells and treatment with imatinib or RK-20449 inhibited this phosphorylation and expression of Survivin or Cyclin D1 coincidentally at least in one cell line.
Taken together, our results suggest that BCR-ABL phosphorylates and activates YAP directly or indirectly through phosphorylation at Y357 leading to overexpression of YAP-dependent Survivin and Cyclin D1. We have not determined which SFK is involved in the downstream signaling of BCR-ABL. Our preliminary overexpression experiments in HEK293T have indicated that some of SFKs can phosphorylate YAP at Y357 without BCR-ABL. However, it remains to be determined which SFK is activated by BCR-ABL and practically involved in YAP phosphorylation resulting in leukemogenesis. Further studies are required to specify the relevant SFKs and disclose the downstream signaling from activated YAP.
References
Inoue A, Kobayashi CI, Shinohara H, Miyamoto K, Yamauchi N, Yuda J, et al. Chronic myeloid leukemia stem cells and molecular target therapies for overcoming resistance and disease persistence. Int J Hematol. 2018;108:365–70.
Chereda B, Melo JV. Natural course and biology of CML. Ann Hematol. 2015;94:107–21.
Hoelbl A, Schuster C, Kovacic B, Zhu B, Wickre M, Hoelzl MA, et al. Stat5 is indispensable for the maintenance of bcr/abl -positive leukaemia: stat5 in leukaemia maintenance. EMBO Mol Med. 2010;2:98–110.
Klejman A, Rushen L, Morrione A, Slupianek A, Skorski T. Phosphatidylinositol-3 kinase inhibitors enhance the anti-leukemia effect of STI571. Oncogene. 2002;21:5868–76.
Sattler M, Mohi MG, Pride YB, Quinnan LR, Malouf NA, Podar K, et al. Critical role for Gab2 in transformation by BCR/ABL. Cancer Cell. 2002;1:479–92.
Wilson MB, Schreiner SJ, Choi H-J, Kamens J, Smithgall TE. Selective pyrrolo-pyrimidine inhibitors reveal a necessary role for Src family kinases in Bcr-Abl signal transduction and oncogenesis. Oncogene. 2002;21:8075–88.
Rubbi L, Titz B, Brown L, Galvan E, Komisopoulou E, Chen SS, et al. Global phosphoproteomics reveals crosstalk between Bcr-Abl and negative feedback mechanisms controlling src signaling. Sci Signal. 2011;4:ra18.
Zhao B, Tumaneng K, Guan K-L. The Hippo pathway in organ size control, tissue regeneration and stem cell self-renewal. Nat Cell Biol. 2011;13:877–83.
Taniguchi K, Wu L-W, Grivennikov SI, de Jong PR, Lian I, Yu F-X, et al. A gp130–Src–YAP module links inflammation to epithelial regeneration. Nature. 2015;519:57–62.
Li P, Silvis MR, Honaker Y, Lien W-H, Arron ST, Vasioukhin V. αE-catenin inhibits a Src–YAP1 oncogenic module that couples tyrosine kinases and the effector of Hippo signaling pathway. Genes Dev. 2016;30:798–811.
Taniguchi K, Moroishi T, de Jong PR, Krawczyk M, Grebbin BM, Luo H, et al. YAP–IL-6ST autoregulatory loop activated on APC loss controls colonic tumorigenesis. Proc Natl Acad Sci. 2017;114:1643–8.
Sugihara T, Werneburg NW, Hernandez MC, Yang L, Kabashima A, Hirsova P, et al. YAP tyrosine phosphorylation and nuclear localization in cholangiocarcinoma cells are regulated by LCK and independent of LATS activity. Mol Cancer Res. 2018;16:1556–67.
Afar DE, McLaughlin J, Sherr CJ, Witte ON, Roussel MF. Signaling by ABL oncogenes through cyclin D1. Proc Natl Acad Sci. 1995;92:9540–4.
Wang Z, Sampath J, Fukuda S, Pelus LM. Disruption of the Inhibitor of apoptosis protein survivin sensitizes Bcr-abl–Positive cells to STI571-induced apoptosis. Cancer Res. 2005;65:8224–32.
Rangatia J, Bonnet D. Transient or long-term silencing of BCR-ABL alone induces cell cycle and proliferation arrest, apoptosis and differentiation. Leukemia. 2006;20:68–76.
Saito Y, Yuki H, Kuratani M, Hashizume Y, Takagi S, Honma T, et al. A pyrrolo-pyrimidine derivative targets human primary aml stem cells in vivo. Sci Transl Med. 2013;5:181ra52.
Kano Y. In vitro cytotoxic effects of a tyrosine kinase inhibitor STI571 in combination with commonly used antileukemic agents. Blood. 2001;97:1999–2007.
Kitamura T, Tange T, Terasawa T, Chiba S, Kuwaki T, Miyagawa K, et al. Establishment and characterization of a unique human cell line that proliferates dependently on GM-CSF, IL-3, or erythropoietin. J Cell Physiol. 1989;140:323–34.
Gu L, Chiang K-Y, Zhu N, Findley HW, Zhou M. Contribution of STAT3 to the activation of survivin by GM-CSF in CD34 + cell lines. Exp Hematol. 2007;35:957–66.
Muramatsu T, Imoto I, Matsui T, Kozaki K, Haruki S, Sudol M, et al. YAP is a candidate oncogene for esophageal squamous cell carcinoma. Carcinogenesis. 2011;32:389–98.
Zhang W, Gao Y, Li F, Tong X, Ren Y, Han X, et al. YAP promotes malignant progression of Lkb1 -deficient lung adenocarcinoma through downstream regulation of survivin. Cancer Res. 2015;75:4450–7.
Mizuno T, Murakami H, Fujii M, Ishiguro F, Tanaka I, Kondo Y, et al. YAP induces malignant mesothelioma cell proliferation by upregulating transcription of cell cycle-promoting genes. Oncogene. 2012;31:5117–22.
Vlahov N, Scrace S, Soto MS, Grawenda AM, Bradley L, Pankova D, et al. Alternate RASSF1 transcripts control src activity, E-cadherin contacts, and YAP-mediated invasion. Curr Biol. 2015;25:3019–34.
Levy D, Adamovich Y, Reuven N, Shaul Y. Yap1 phosphorylation by c-Abl Is a critical step in selective activation of proapoptotic genes in response to DNA damage. Mol Cell. 2008;29:350–61.
Hu Y, Liu Y, Pelletier S, Buchdunger E, Warmuth M, Fabbro D, et al. Requirement of Src kinases Lyn, Hck and Fgr for BCR-ABL1-induced B-lymphoblastic leukemia but not chronic myeloid leukemia. Nat Genet. 2004;36:453–61.
Donato E, Biagioni F, Bisso A, Caganova M, Amati B, Campaner S. YAP and TAZ are dispensable for physiological and malignant haematopoiesis. Leukemia. 2018;32:2037–40.
Li H, Huang Z, Gao M, Huang N, Luo Z, Shen H, et al. Inhibition of YAP suppresses CML cell proliferation and enhances efficacy of imatinib in vitro and in vivo. J Exp Clin Cancer Res. 2016. https://doi.org/10.1186/s13046-016-0414-z.
Acknowledgements
We thank Dr. Xiaoyan Jiang (BC Cancer Agency—Terry Fox Laboratory) for MSCV-BCR-ABL-IRES-GFP, Dr. Yusuke Furukawa (Jichi University) for TCC-S, and Dr. Toshio Kitamura (University of Tokyo) for TF-1. We also thank our laboratory members, M.Sc. Konomi Maeda, M.Sc. Haruka Oshima, and Dr. Ryotaro Nishi for their collaborations.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
We declare that we have no competing interests in this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
About this article
Cite this article
Moriyama, K., Hori, T. BCR-ABL induces tyrosine phosphorylation of YAP leading to expression of Survivin and Cyclin D1 in chronic myeloid leukemia cells. Int J Hematol 110, 591–598 (2019). https://doi.org/10.1007/s12185-019-02726-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12185-019-02726-7