Abstract
It is possible and risky for fur animal carcasses to be mixed into meat products, which is a potential danger for meat quality safety and human health. Therefore, meat validation of quality and constituents is crucial. A variety of methods have been developed to identify muscle tissues of different species. However, most of these methods have the disadvantages of poor repeatability, complex operation and low efficiency, and cannot simultaneously detect multiple species of muscle tissue. The purpose of this study was to construct a multiplex PCR protocol to detect the samples of mink, fox, and raccoon dog. In this study, the specific primers of mink, fox, and raccoon dog were designed according to the variable region sequence of mitochondrial cytochrome b (Cytb) gene. The primers showed good specificity and 50 ℃ was determined as the optimal annealing temperature. The lowest concentration of DNA template of mink, fox, or raccoon dog that could be determined simultaneously by a single tube was 1 pg/µL. Clinical tissue samples detect analysis test results showed that this method could identify whether the tissue samples of three fur animals were mixed from the muscles of chickens, ducks, dogs, cattle, sheep, pigs and rabbits in one PCR reaction simultaneously. In conclusion, the scheme exhibited the advantages of convenient operation, low cost, strong species specificity, high sensitivity, good stability, and repeatability. The systematic optimized inspection process can be applied to meat detection to ensure veterinary public health safety, which has important scientific significance, production, public health, and safety significance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mink, fox, and raccoon dog are important economic animals for fur. Their high-quality fur is used for fur processing and eventually becomes popular high-grade fur clothing. Like Denmark, Finland, the USA, and Canada, China is also an important fur animal breeding country, raw material fur producing country, fur processing country, and fur garment producing, exporting, and consuming country in the world (Li et al. 2020; Yang et al. 2017; Liu 2022; Wang 2022). Fur is the main product of fur-based economic animals. The carcasses left after these animals are slaughtered and skinned become by-products. Carcass can be processed into meat and bone meal for animal feed (Xu et al. 2016). However, it may also be mixed with human food, which is a potential hidden danger to food quality safety and veterinary public health. Therefore, meat validation of quality and constituents is paramount. It is necessary to explore and build a rapid, practical, simple, and specific detection technology for the muscle tissue of mink, fox, and raccoon dog to distinguish their muscle tissue from the muscle tissue of conventional livestock and poultry, so as to promote the healthy development of the industry and ensure the food quality and safety.
To date, a variety of DNA-based analytical methods have been developed to identify tissues of different species (Shahrooz et al. 2016). DNA detection methods based on polymerase chain reaction (PCR) for meat analysis are also increasing (Fajardo et al. 2010). For example, PCR (Karabasanavar et al. 2013), polymerase chain reaction-restriction fragment length polymorphism (PCR–RFLP) (Girish et al. 2005), random amplified polymorphic DNA marker PCR (RAPD-PCR) (Martinez and Yman 1998), and PCR-Southern blotting hybridization (Mutalib et al. 2015) have been used for the identification of meat products. However, these methods have the disadvantages of poor repeatability, complex operation, and low efficiency, and cannot simultaneously detect multiple species of muscle tissue. Nowadays, real-time fluorescent quantitative PCR and digital PCR have become common identification methods because of their fast and sensitive characteristics (Qamar et al. 2020; Xu et al. 2018). However, compared with conventional PCR, its high reagent and equipment costs limit its application.
At the same time, people pay more and more attention to the food quality and safety testing program which is fast, accurate, easy to operate, economic, and can detect many and complete samples. Therefore, it is very necessary to establish a reliable multiplex PCR method to quickly and accurately identify meat products. It can not only detect multiple species samples simultaneously through a reaction system (Matsunaga et al. 1999), but also, compared with real-time fluorescent quantitative PCR, multiplex PCR only requires conventional PCR primers without other modifications, which can save detection time and reduce detection costs (Izadpanah et al. 2018a, b).
The cytochrome b gene (Cytb gene) is a gene sequence in the mitochondrial DNA (mtDNA) (Yuttamol et al. 2018). Cytb gene has a sustainable or conserved region at the species level. The relatively moderate evolution speed of the Cytb gene in the mtDNA genome can provide phylogenetic and genetic evolution information. The sequence variability of Cytb gene makes it suitable for differential identification between species or genera, which has been used in many studies of phylogenetic relationships within mammals (Castresana 2001; Yuttamol et al. 2018; Suyadi et al. 2022). In this study, the specific primers of mink, fox, and raccoon dog were designed according to the variable region sequence of mitochondrial cytochrome b (Cytb) gene. By optimizing the multiplex PCR reaction system and amplification conditions, the multiplex PCR detection scheme was constructed. The multiplex PCR detection scheme can realize single tube PCR reaction and detect the samples of mink, fox, and raccoon dog from the mixed muscle tissue samples of conventional livestock and poultry, mink, fox, and raccoon dog, which will provide reference for food quality safety and veterinary public health.
Materials and Methods
Materials
Carcass samples of mink (black mink, white mink, iron gray mink), fox (white fox, blue fox), and raccoon dog (Wusuli raccoon dog, white raccoon dog) were given by Shandong Zhucheng fur animal research institute. Chicken, duck, beef, and pork samples were purchased from a supermarket in Tai'an, Shandong Province; dog meat was purchased from a dog meat shop in Tai'an, Shandong Province, and mutton and rabbit meat were preserved by our laboratory.
Primers
To establish a multiple PCR analysis protocol, sequence analysis was performed using the biological software Meglin (Bioinformatics Software DNASTAR) based on the mitochondrial Cytb gene sequences of minks, foxes, and raccoons in GenBank. And three pairs of specific upstream and downstream primers were designed using Primer Premier 6.0 (Products for Genomics and Mass Spectrometry premierbiosoft.com), and then performed PCR validation one by one. The detailed process is as follows: Based on the published Cytb gene sequences of mink (GenBank: KF990329.1), fox (GenBank: DQ498127.1), and raccoon dog (GenBank: JX099889.1) by NCBI, specific primers were designed based on the alignment results of the Cytb gene sequences of mink, fox, and raccoon dog (Fig. 1). The primers and the respective amplification lengths are shown in Table 1. At the same time, a pair of universal primers that can amplify vertebrates, with a size of 1200 bp, were designed. Upstream of universal primer (5′-3′) is GTAGTCATATGCTTGTCTC, and downstream of universal primer (5′-3′) is CTTCCGTCAATTCCTTTAAG. All the primers used in current study were synthesized by Beijing Liuhe Huada Gene Co., Ltd.
DNA Extraction
Using TIANamp Blood/Cell/Tissue DNA Kit (Tiangen, Beijing, China) according to the Kit instructions, genomic DNA was extracted from the carcass muscle of mink, fox, and raccoon dog, as well as the tissue samples of chicken, duck, dog, beef, mutton, pork, and rabbit muscle. The extracted muscle tissue samples’ DNA was detected by nucleic acid analyzer (DS-11 Series Spectrophotometer, Purchased from DeNovix Company). The extracted DNA was stored at − 20 °C.
PCR Reaction Verification of Universal Primers
Using universal primers, the DNA of muscle tissue samples from mink, fox and raccoon dog, chicken, duck, dog, cow, sheep, pig, and rabbit was PCR reacted to verify whether the DNA of muscle tissue samples could meet the requirements of PCR amplification reaction in vertebrates.
The PCR reaction were carried out in 50 µL reaction mixtures containing 25 µL of 2 × Taq Master Mix, 5 µL for each of the muscle tissue samples’ DNA templates, and 2 µL for each of the upstream and downstream universal primer (10 µmol/L). Finally, the volume of the reaction mixtures was filled up to 50 µL with sterilized double distilled water. The amplification conditions consisted of a pre-denaturation at 94 °C for 1 min 30 s, 30 cycles of denaturation at 94 °C for 50 s, annealing at 55 °C for 1 min, extension at 72 °C for 1 min, and final extension for 5 min at 72 °C. PCR products were electrophoresed on 2% agarose gel and the results were observed on the gel imager (Azure c150, Purchased from Azure Biosystems).
Single PCR Amplification Reaction and Enzyme Digestion Verification of Amplification Products
Using specific primers, the DNA of muscle tissue samples from mink, fox, and raccoon dog was PCR reacted. The PCR reaction were carried out in 25 µL reaction mixtures containing 12.5 µL of 2 × Taq Master Mix, 2 µL for each of the muscle tissue samples DNA templates, and 1 µL for each of the upstream and downstream universal primer (10 µmol/L). Finally, the volume of the reaction mixtures was filled up to 25 µL with sterilized double distilled water. The amplification conditions consisted of a pre-denaturation at 94 °C for 1 min 30 s, 30 cycles of denaturation at 94 °C for 50 s, annealing at 55 °C for 1 min, extension at 72 °C for 50 s, and final extension for 5 min at 72 °C.
The amplified products of mink, fox, and raccoon dog were digested and verified with BamH I, ApoI and PflMI endonucleases (Dalian TaKaRa Co., Ltd), respectively. The enzyme digestion validation reaction system contains 2 µL of 10 × buffer, 1 µL of restriction enzyme, and 10 µL of PCR product. The enzyme digestion validation reaction condition is 37 °C for 30 min. PCR products and enzyme digestion products were electrophoresed by 2% agarose gel and observed on the gel imager.
Primer Cross-Reaction Test
The purpose of this test is to determine whether there is cross-reaction between primers. The specific primers of mink, fox, and raccoon dog were mixed, and the single DNA template and the mixed DNA template were amplified by PCR, respectively, to verify whether there was cross-reaction between the primers. The reaction conditions of primer cross test were the same as those of single PCR amplification. The total volume of the reaction system remains the same; however, the number of primers needs to be changed, and the number of ddH2O should be reduced accordingly.
Multiplex PCR Amplification Reaction and Annealing Temperature Optimization
After verified by primer cross test, ensure that there is no cross-reaction between primers. Subsequently, three specific primers were mixed, and the DNA templates of mink, fox, and raccoon dog muscle samples were amplified by PCR. At the same time, the annealing temperature was optimized. Five temperature gradients of 50 °C, 52 °C, 54 °C, 56 °C, and 58 °C were set to determine the optimal annealing temperature.
Specificity Test of Multiplex PCR
Mixing the specific primers of three fur animals, and then the DNA of muscle tissue samples from mink, fox and raccoon dog, chicken, duck, dog, cow, sheep, pig, and rabbit were used as templates in the optimized reaction system for multiplex PCR amplification to detect the specificity of primers. In the specificity test, ddH2O was used as the template for the negative control.
Sensitivity Test of Multiplex PCR
The DNA template was serially diluted to 1 ng/µL, 100 pg/µL, 10 pg/µL, 1 pg/µL, and 0.1 pg/µL by tenfold gradient with sterile double distilled water, and then amplified by the optimized single PCR reaction system and the optimized multiplex PCR reaction system. Finally, the PCR products were electrophoresis to detect the sensitivity of each primer.
Verification Test of Mixed Samples
Mix the muscle tissue samples of different conventional livestock and poultry, and take one of the similar species. Every four kinds of conventional livestock and poultry muscles are mixed to form a combination. In this way, a total of six kinds of conventional livestock and poultry mixed samples are formed: (1) dog, beef, chicken, and rabbit; (2) dog, beef, duck, and rabbit; (3) dog, mutton, chicken, and rabbit; (4) dog, mutton, duck, and rabbit; (5) dog, pork, chicken, and rabbit; and (6) dog, pork, duck, and rabbit. The sample addition amount of each animal is 20 mg, and the total mass of the conventional livestock and poultry mixed samples of each combination is 80 mg.
On this basis, 20 mg of mink, fox, and raccoon dog muscle tissue samples or ddH2O is separately added to each combined mixed sample. In this way, a total of 24 mixed sample combinations are formed, and the total mass of each mixed sample is 100 mg. The mixing methods of different animal muscle samples and their corresponding combination numbers are shown in Table 2.
The mixed samples of each group were fully mixed, the genomic DNA of each group was extracted, and the DNA template of each group was detected and verified by the established multiple PCR method.
Results and Analysis
Amplification Results of Universal Primers
The results of universal primer amplification experiment showed that 1200 bp target gene band could be amplified from every DNA template extracted from muscle tissue samples of different animals (Fig. 2), and the size of target gene fragments was consistent with the expected results. The results showed that the DNA quality of the tissue samples extracted in this study met the test requirements, and the amplification results of the target gene were stable, which ensures the subsequent multiple PCR scheme optimization test.
Results of Single PCR Amplification and Enzyme Digestion Verification of Amplified Products
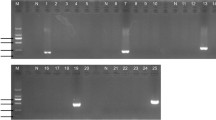
The muscle tissue DNA of mink, fox, and raccoon dog was amplified by PCR using their specific primers, and the amplified products were confirmed by enzyme digestion. The results showed that the fragment size of PCR amplification products of mink samples was 180 bp and 112 bp, respectively, after enzyme digestion. The fragment size of PCR amplification product of raccoon dog sample is 411 bp and 150 bp, respectively. The fragment size of PCR amplification product of fox sample is 159 bp and 700 bp, respectively. These indicated that the results of PCR amplification and enzyme digestion were in line with expectations (Fig. 3).
Results of single PCR amplification and enzyme digestion verification of amplified products. Note: lane M, D2000 DNA marker; lane 1, black mink; lane 2, bands after BamH I digestion; lane 3, white mink; lane 4, bands after BamH I digestion; lane 5, iron gray mink; lane 6, bands after BamH I digestion; lane 7, white raccoon dog; lane 8, bands after PflMI digestion; lane 9, Wusuli raccoon dog; lane 10, bands after PflMI digestion; lane 11, white fox; lane 12, bands after ApoI digestion; lane 13, blue fox; lane 14, bands after ApoI digestion
Results of Primer Cross-Reaction Test
The specific primers of mink, fox, and raccoon dog were mixed, and the single DNA template and the mixed DNA template were amplified by PCR, respectively, to verify whether there was cross-reaction between the primers. The results showed that there was no cross-reaction between the primers (Fig. 4), which ensured the normal implementation of the multiplex PCR detection scheme.
Results of primer cross reaction test. Note: Lane M, D2000 DNA marker. (1) Primer of each lane. Lanes 1–4, mink and raccoon dog; lane 5–8, mink and fox; lane 9–12, raccoon dog and fox. (2) DNA template of each lane. Lane 1, mink; lane 2, raccoon dog; lane 3, mink, and raccoon dog; lane 4, ddH2O. Lane 5, mink; lane 6, fox; lane 7, mink and fox; lane 8, ddH2O; lane 9, raccoon dog; lane 10, fox; lane 11, raccoon dog and fox; lane 12, ddH2O
Results of Multiplex PCR Amplification Reaction and Annealing Temperature Optimization
Three specific primers were mixed, and the DNA templates of mink, fox, and raccoon dog muscle samples were amplified by multiplex PCR. The results showed that multiplex PCR could amplify the corresponding species-specific target band, which was in line with the expected results (Fig. 5). In addition, 50 ℃ was determined as the optimal annealing temperature (Fig. 6).
Results of Specificity and Sensitivity Test of Multiplex PCR
Results of specificity test of multiplex PCR showed that the DNA of mink, fox, and raccoon dog tissue samples showed their respective specific target bands, while the DNA of other animals and the blank control did not amplify the target bands, indicating that the primers showed good specificity (Fig. 7). The results of sensitivity test showed that the lowest concentration of DNA template of mink, fox, and raccoon dog that could be determined simultaneously by a single tube was 1 pg/µL (Fig. 8).
Results of Validation Test Results of Mixed Samples
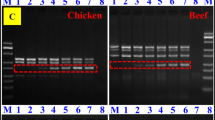
The verification experiment results of mixed samples showed that as long as the DNA template of mixed samples was mixed with mink tissues, 292 bp-specific bands could be amplified. As long as mixed sample DNA template of fox tissue is incorporated, 859 bp-specific bands can be amplified. As long as mixed sample DNA template of raccoon dog tissue is incorporated, 561 bp-specific bands can be amplified (Fig. 9).
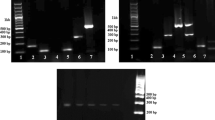
Results of validation test results of mixed samples. Note: Lane M, D2000 DNA marker. A Lane 1, dog, beef, chicken and rabbit; lane 2, dog, beef, chicken, rabbit, and mink; lane 3, dog, beef, chicken, rabbit, and fox; lane 4, dog, beef, chicken, rabbit, and raccoon dog. B Lane 5, dog, beef, duck, and rabbit; lane 6, dog, beef, duck, rabbit, and mink; lane 7, dog, beef, duck, rabbit, and fox; lane 8, dog, beef, duck, rabbit, and raccoon dog. C Lane 9, dog, mutton, chicken, and rabbit; lane 10, dog, mutton, chicken, rabbit, and mink; lane 11, dog, mutton, chicken, rabbit, and fox; lane 12, dog, mutton, chicken, rabbit, and raccoon dog. D Lane 13, dog, mutton, duck, and rabbit; lane 14, dog, mutton, duck, rabbit, and mink; lane 15, dog, mutton, duck, rabbit, and fox; lane 16, dog, mutton, duck, rabbit, and raccoon dog. E Lane 17, dog, pork, chicken, and rabbit; lane 18, dog, pork, chicken, rabbit, and mink; lane 19, dog, pork, chicken, rabbit, and fox; lane 20, dog, pork, chicken, rabbit, and raccoon dog. F Lane 21, dog, pork, duck, and rabbit; lane 22, dog, pork, duck, rabbit, and mink; lane 23, dog, pork, duck, rabbit, and fox; lane 24, dog, pork, duck, rabbit, and raccoon dog
Discussion
After decades of development, PCR technology has become an important scientific research means in modern molecular biology and other fields, and has played an important role in the field of differential diagnosis. However, with the rapid development of biological detection technology, the needs of users have also developed from specificity and sensitivity to the consideration of simple operation, cost saving, and wide detection range. Multiplex PCR diagnostic technology is a PCR reaction in which two or more pairs of specific primers are added to the same PCR reaction system to simultaneously amplify multiple target fragments in a short time in the same reaction system. In order to establish a multiplex PCR reaction system, it is necessary to optimize the reaction conditions. Cyt b gene is an important part of mitochondrial DNA (Shamsuddin 2017; Suyadi et al. 2022). The evolution speed of Cyt b gene is moderate, and Cyt b gene shows diversity among genera, species, and even varieties. Therefore, the phylogenetic information and genetic evolution level analysis provided by Cyt b gene can be used to analyze the differences between species or genera (Castresana 2001; Afshari et al. 2021; Yuttamol et al. 2018).
In this study, mitochondrial DNA Cyt b gene was selected, and specific primers for Cyt b gene of three fur animals were designed. In this study, single PCR amplification and product digestion validation test were carried out to ensure that there would be no false positive. Then the primer cross test was carried out to ensure the smooth progress of the multiplex PCR detection reaction. Finally, a series of optimizations were carried out on the reaction conditions and reaction system of multiplex PCR, and a multiplex PCR detection scheme was successfully constructed ultimately, so that the samples of mink, fox, and raccoon dog could be identified simultaneously in one reaction. The sensitivity of this method could reach 1 pg/µL which was much higher than that of other PCR detection methods (Izadpanah et al. 2018a, b; Junan et al. 2017; Qin et al. 2019; He et al. 2012; Zhao et al. 2015). The sensitivity of these studies is generally in the range of 16–6250 pg/µL. By analyzing and comparing the specificity of the detection methods, the detection scheme constructed in this study can detect the meat of order varieties from 10 species of meat, which is much higher than other studies. Most of their studies distinguish and identify 4 to 7 kinds of meat (Izadpanah et al. 2018a, b; Qin et al. 2019; He et al. 2012). The results of this study reveal that the detection method showed good specificity and high sensitivity. The above results indicate that if this detection technology is applied to meat detection practice, the efficiency and accuracy of inspection can be greatly improved.
The detection scheme constructed in this study was used to detect and analyze clinical tissue samples. The results showed that even for samples of different varieties of the same species, this detection method could also amplify its corresponding expected target bands. In addition, even for mixed samples of different species, this method could also detect whether the tissue samples of three fur animals were mixed, and the detection results could clearly show the specific target bands of three fur animals. The above results suggest that if this multiplex PCR scheme is applied to the meat detection on the market, it will show better precision, accuracy, practicality, reliability, and stability than other detection methods. However, the main limitation of multiplex PCR is the increased complexity and potential for primer-dimer formation when multiple primer pairs are used in the same reaction. This can lead to non-specific amplification and interfere with the accuracy of the results. Additionally, as you increase the number of targets in a multiplex reaction, it can become more challenging to optimize conditions for all the primers simultaneously. It can be seen that multiple PCR detection schemes also have certain limitations, which may reduce their application scope.
In summary, a systematic optimized multiplex PCR detection scheme constructed in this study could identify minks, fox, and raccoon dogs’ meat from the muscles of chickens, ducks, dogs, cattle, sheep, pigs, and rabbits in one PCR reaction simultaneously. In addition, this detection technique also showed the advantages of convenient operation, low cost, strong specificity, and high sensitivity, which can be regarded as a convenient, time-saving, fast, and accurate meat detection technique. Thus, it can be seen that the detection scheme can be applied to meat product detection and other fields, which has important scientific, clinical, production significance, public health, and safety significance.
Availability of Data and Materials
All data generated or analyzed during this study are included in this published article and its supplementary information files. The datasets are also available from the corresponding author on reasonable request.
References
Afshari A, Tavassoli M, Esmaelnejad B, Habibi GH, Esmaeilnia K (2021) Molecular characterization and phylogenetic analysis of pathogenic Theileria spp. isolated from cattle and sheep based on cytochrome b gene in Iran. Arch Razi Inst 76:243–252. https://doi.org/10.22092/ari.2020.341390.1431
Castresana J (2001) Cytochrome b phylogeny and the taxonomy of great apes and mammals. Mol Biol Evol 18:465–471. https://doi.org/10.1093/oxfordjournals.molbev.a003825
Fajardo V, Gonzalez I, Rojas M, Garcia T, Martin R (2010) A review of current PCR - based methodologies for the authentication of meats from game animal species. Trends Food Sci Technol 21:408–421. https://doi.org/10.1016/ji.tifs.2010.06.002
Girish PS, Anjaneyulu AS, Viswas KN, Shivakumar BM, Anand M, Patel M, Sharma B (2005) Meat species identification by polymerase chain reaction-restriction fragment length polymorphism (PCR - RFLP) of mitochondrial 12S rRNA gene. Meat Sci 70:107–112. https://doi.org/10.1016/j.meatsci.2004.12.004
He W, Zhang C, Yang J, Huang M, Yang J (2012) A quick multiplex PCR method for the identification of four meat ingredients in food products. Scientia Agricultura Sinica 45:1873–1880. https://doi.org/10.3864/j.issn.0578-1752.2012.09.024. ((In Chinese))
Izadpanah M, Mohebali N, Gorji ZE (2018a) Simple and fast multiplex PCR method for detection of species origin in meat products. J Food Sci Technol 55:698–703. https://doi.org/10.1007/s13197-017-2980-2
Izadpanah M, Mohebali N, Elyasi GZ, Farzaneh P, Vakhshiteh F, Shahzadeh Fazeli SA (2018b) Simple and fast multiplex PCR method for detection of species origin in meat products. J Food Sci Technol 55:698–703. https://doi.org/10.1007/s13197-017-2980-2
Junan R, Tingting D, Wensheng H (2017) A digital PCR method for identifying and quantifying adulteration of meat species in raw and processed food. PLoS ONE 12:e0173567. https://doi.org/10.1371/journal.pone.0173567
Karabasanavar NS, Singh SP, Kumar D, Shebannavar SN (2013) Development and application of highly specific PCR for detection of chicken (Gallus gallus) meat adulteration. Eur Food Res Technol 236:129–134. https://doi.org/10.1007/s00217-012-1868-7
Li P, Zhang D, Li H, Pang J, Guo H, Qiu J (2020) Establishment and application of a multiplex PCR for simultaneously detecting Escherichia coli, Salmonella, Klebsiella pneumoniae and Staphylococcus aureus in minks. Frontiers in Veterinary Science 7(eCollection 2020):588173. https://doi.org/10.3389/fvets.2020.588173
Liu Z (2022) Analysis on the situation of fur animal industry in 2021 and prospects for 2022. Animal Agriculture 4:32–36 ((In Chinese) Website of the thesis: http://202.194.143.28/rwt/CNKI/https/NNYHGLUDN3WXTLUPMW4A/kcms/detail/detail.aspx?dbcode=CJFQ&dbname=CJFDLAST2022&filename=SNCM202204005&v=MjczMTBQTXE0OUZZWVI4ZVgxTHV4WVM3RGgxVDNxVHJXTTFGckNVUjdpZVorUnRGeXZoVXI3Tk5pUElZN0c0SE4=)
Martinez I, Yman IM (1998) Species identification in meat products by RAPD analysis. Food Res Int 31:459–466. https://doi.org/10.1016/S0963-9969(99)00013-7
Matsunaga T, Chikuni K, Tanabe R, Muroyab S, Shibataa K, Yamadaa J, Shinmuraa Y (1999) A quick and simple method for the identification of meat species and meat products by PCR assay. Meat Sci 51:143–148. https://doi.org/10.1016/S0309-1740(98)00112-0
Mutalib SA, Muin NM, Abdullah A, Muin M, Abdullah A, Hassan O, Mustapha WAW, Sani NA, Maskat MY (2015) Sensitivity of polymerase chain reaction (PCR) - southern hybridization and conventional PCR analysis for Halal authentication of gelatin capsules. LWT Food Sci Technol 63:714–719. https://doi.org/10.1016/j.lwt.2015.03.006
Qamar Z, Alawami M, Mokhtar NFK, Nhari RMHR, Hanish I (2020) Current analytical methods for porcine identification in meat and meat products. Food Chem 324:126664. https://doi.org/10.1016/j.foodchem.2020.126664
Qin P, Qu W, Xu J, Qiao D, Yao L, Xue F, Chen W (2019) A sensitive multiplex PCR protocol for simultaneous detection of chicken, duck, and pork in beef samples. J Food Sci Technol 56:1266–1274. https://doi.org/10.1007/s13197-019-03591-2
Shahrooz R, Nurhidayatullaili MJ, Wageeh AY, Wan JB (2016) Identification of meat origin in food products - a review. Food Control 68:379–390. https://doi.org/10.1016/j.foodcont.2016.04.013
Shamsuddin M (2017) Preface: special issue on “genetic diversity of small ruminants in Asia.” Small Rumin Res 148:1. https://doi.org/10.1016/j.smallrumres.2017.02.017
Suyadi S, Murtika IL, Susilorini TE, Septian WA, Saputra F, Furqon A (2022) Genetic diversity of goats in East Java through analysis of the cytochrome B and cytochrome oxidase I genes in mitochondrial DNA. Small Rumin Res 209:106659. https://doi.org/10.1016/j.smallrumres.2022.106659
Wang D (2022) Statistical report on the number of mink, fox and raccoon dog taking skins in China in 2021. Westleather 44:2–4 (In Chinese) Website of the thesis: http://202.194.143.28/rwt/CNKI/https/NNYHGLUDN3WXTLUPMW4A/kcms/detail/detail.aspx?dbcode=CJFQ&dbname=CJFDLAST2022&filename=XBPG202204002&v=MTAzNjRSN2llWitSdEZ5dmhVTC9MUFMvYmFiRzRITlBNcTQ5RlpvUjhlWDFMdXhZUzdEaDFUM3FUcldNMUZyQ1U=)
Xu M, Liang Y, Liu Z (2016) Application status of meat and bone meal and prospect of processing fur animal carcass into meat and bone meal. Heilongjiang Anim Sci Vit Med 10:209–211 (https://doi.org/10.13881/j.cnki.hljxmsy.2016.1925 (In Chinese))
Xu R, Wei S, Zhou G, Ren J, Liu Z, Tang S, Pck C, Wu X (2018) Multiplex TaqMan locked nucleic acid real-time PCR for the differential identification of various meat and meat products. Meat Sci 137:41–46. https://doi.org/10.1016/j.meatsci.2017.11.003
Yang ML, Wang FY, Jiang DH, Ma ZF, Cui K (2017) Investigation on the operation of special economic animal industry in Shandong Province, China. Shangdong J Anim Sci Vet Med 8:47–49. https://doi.org/10.3969/j.issn.1007-1733.2017.08.035. ((In Chinese))
Yuttamol M, Worawidh W, Akira A, Manakorn S (2018) The novel primers for mammal species identification-based mitochondrial cytochrome b sequence implication for reserved wild animals in Thailand and endangered mammal species in Southeast Asia. Mitochondrial DNA A DNA Mapp Seq Anal 29:62–72. https://doi.org/10.1080/24701394.2016.1238902
Zhao X, Wang Y, Lan Q, Liu N, Chen R, Zhu Z, Liu O (2015) Real-time PCR detection of sheep derived materials in meat product. Sci Technol Food Industry 36:299–302+ 208 ((In Chinese))
Funding
This work was mainly supported by Major Science and Technology Innovation Project in Shandong Province (2023CXGC010704), Shandong Province Modern Agricultural Technology System Innovation Team Program (SDAIT-21–10), the Earmarked Fund for the Protection for the State Forest and Wild Animals (No. 2130211), and Funds of Shandong “Double Tops” program.
Author information
Authors and Affiliations
Contributions
Huijun Guo conceived and designed the experiments. Yao Pan, Xuemin Wang and Wenjie Ma are mainly responsible for experimental implementation. Hongmei Li, Jianhua Qiu and Yuxin Sun is mainly responsible for sample collection and helped to do some experiments. Jianhua Qiu, Xuemin Wang and Yao Pan wrote the manuscript. All authors have read and approved the final manuscript
Corresponding authors
Ethics declarations
Conflict of Interest
Yao Pan declares that he has no conflict of interest. Xuemin Wang declares that she has no conflict of interest. Wenjie Ma declares that he has no conflict of interest. Hongmei Li declares that she has no conflict of interest. Yuxin Sun declares that he has no conflict of interest. Jianhua Qiu declares that he has no conflict of interest. Huijun Guo declares that he has no conflict of interest.
Ethics Approval
Ethical approval was not required for the study because the sampling process did not harm the animals.
Competing Interest
The authors declare no competing interests
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pan, Y., Wang, X., Ma, W. et al. Establishment and Application of Multiplex PCR Scheme for Simultaneously Distinguishing Muscle Tissues of Mink, Fox, and Raccoon Dog from Conventional Livestock and Poultry. Food Anal. Methods 17, 226–235 (2024). https://doi.org/10.1007/s12161-023-02560-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12161-023-02560-y