Abstract
Background
Although uncommon, medullary thyroid carcinoma (MTC) accounts for a significant proportion of thyroid cancer deaths. Recent studies have validated the two-tier International Medullary Thyroid Carcinoma Grading System (IMTCGS) to predict clinical outcomes. A 5% Ki67 proliferative index (Ki67PI) cut-off separates low-grade from high-grade MTC. In this study, we compared digital image analysis (DIA) to manual counting (MC) for determining the Ki67PI in a MTC cohort, and explored the challenges encountered.
Methods
Available slides from 85 MTCs were reviewed by two pathologists. The Ki67PI was documented by immunohistochemistry for each case, scanned with the Aperio® slide scanner at 40× magnification, and quantified using the QuPath® DIA platform. The same hotspots were screenshot, printed in color, and blindly counted. For each case, over 500 MTC cells were counted. Each MTC was graded using IMTCGS criteria.
Results
In our MTC cohort (n = 85), 84.7 and 15.3% were low- and high-grade with the IMTCGS. In the entire cohort, QuPath® DIA performed well (R2 = 0.9891) but appeared to undercall compared to MC. QuPath® performed better in high-grade cases (R2 = 0.99) compared to low-grade cases (R2 = 0.7071). Overall, Ki67PI determined with either MC or DIA did not affect IMTCGS grade. Encountered DIA challenges include optimizing cell detection, overlapping nuclei, and tissue artifacts. Encountered MC challenges include background staining, morphologic overlap with normal elements, and counting time.
Conclusion
Our study highlights the utility of DIA in quantifying Ki67PI for MTC and can serve as an adjunct for grading in conjunction with the other criteria of mitotic activity and necrosis.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
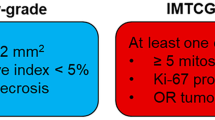
Medullary thyroid carcinoma (MTC) is a rare malignancy that arises from the calcitonin-secreting parafollicular C-cells of the thyroid [1]. While MTC represents < 10% (2%) of all thyroid malignancies, it represents 8% of thyroid cancer-related deaths [2, 3]. Investigations into this disproportionately high mortality rate have yielded three histologic features conferring worse survival: elevated Ki67 proliferative index (Ki67PI), elevated mitotic index, and presence of necrosis [4, 5]. These findings culminated into the International Medullary Thyroid Carcinoma Grading System (IMTCGS), a two-tiered grading system separating MTC into low-grade and high-grade [6]. The presence of mitotic index ≥ 5 per 2 mm2, tumor necrosis, or Ki67PI ≥ 5% confers a designation of high-grade MTC. It is important to emphasize that although the current IMTCGS uses binary cut-offs of ≥ 5% and ≥ 5 mitoses per 2 mm2 for Ki67PI and mitotic activity, respectively, both are continuous variables with worsening outcomes as each variable increases; hence, it is currently recommended to provide precise counts for both variables in the pathology report.
The standard for calculating Ki67PI, set by the World Health Organization (WHO), is by counting at least 500–2000 cells in areas with the highest density of Ki67 staining or “hotspots” [7]. This practice is well-established for neuroendocrine tumors of the gastropancreatic system and was applied during the validation of the IMTCGS. Three widely used methods for measuring the Ki67PI include “eye-balling,” manual cell counting (MC), and automated cell counting using digital image analysis (DIA). “Eye-balling” estimates the Ki67PI by scanning a slide section of the tumor at low power (× 10 objective). This method is quick, but consistent accuracy is difficult at Ki67PI < 5% with lower rates of interobserver agreement [8]. Manual cell counting is performed in real-time at the microscope or using a printed/camera image of tumor “hot spots.” This practice, while highly accurate and with high interobserver agreement, takes the longest to perform, making it less appealing in a busy practice setting [8, 9].
Automated cell counting using DIA software has grown in recent years with concordant and reproducible results in accurately measuring Ki67PI in neuroendocrine tumors [10]. The primary limitations of DIA software cited are operator dependency and cost [8,9,10]. QuPath® is an open source and free DIA software developed as a user-friendly and accessible solution to whole-image analysis, by Peter Bankhead and others [11]. The QuPath® cell detection algorithm is being evaluated and optimized by users to perform automated cell counting for measuring Ki67PI in both neuroendocrine and breast carcinomas [12,13,14,15,16].
In our study, we compared MC and automated DIA, using the Qupath® cell detection algorithm, in measuring the Ki67PI in our institutional MTC cohort. We also examined the performance of both counting methods after stratification of MTCs with the IMTCGS.
Materials and Methods
Our cohort included 85 primary MTC resections between 2000 and 2021 were retrieved from the pathology database at Emory University Hospital, Emory University Hospital Midtown, and Saint Joseph Hospital (all in Atlanta, GA), following approval from the institutional review board (IRB #00004034, K.V). Sections (5 micron) from formalin-fixed, paraffin-embedded tissue blocks were evaluated for the Ki67PI by immunohistochemistry (MIB-1 clone, 1:80 dilution, Agilent Dako) using the Leica Bond Max III (Leica Microsystems, Bannockburn, Illinois). Specimens are deparaffinized and antigen retrieved on the instrument. All slides are incubated with the primary antibody for 15 min, with post primary polymer for 8 min, blocked with 3% hydrogen peroxide for 5 min, 3,3-diaminobenzidine (DAB, brown chromogen) for 10 min, and hematoxylin as counterstain for 5 min. These incubations were performed at room temperature and sections were washed with Tris-buffered saline (Bond wash solution). Cover-slipping was performed using the Tissue-Tek SCA (Sakura Finetek USA, Inc, Torrance, CA) coverslipper. Available H&E slides from all cases were reviewed by two pathologists (D.J.L and K.V.) and histologic features were recorded. Slides were scanned with the Aperio® CS2 slide scanner (Leica Biosystems, San Diego, CA, USA) at 40× magnification and stored as.svs files. Areas with maximal-appearing Ki67 immunostain (“hotspots”) were selected with a uniform square box for all cases and quantified using the QuPath® image analysis platform (https://qupath.github.io/) by one primary operator (K.V.). Default QuPath® settings were applied with the following exceptions: nuclear area size cut-off of 27 determined by trial and error to minimize detection of normal elements, and slight variations in the DAB 1 + detection threshold between 0.1 and 0.2 to maximize detection of Ki67 positive nuclei. The same hotspots used for the QuPath quantification were screenshot, printed in color, and after a washout period of several weeks to minimize bias, were blindly counted by at least one pathologist (K.V., D.J.L, Q.S., and K.M.) and a pathology resident (D.B.B). Any degree of Ki67 staining in tumor nuclei was considered positive on MC. For each case, > 500 MTC cells were counted in the outlined hotspots by at least one of the two methods. Each MTC was graded with the IMTCGS (low/high) based on prior established criteria [6]. A random subset of low-grade (n = 16) and high-grade MTC cases (n = 8) were counted under timed conditions by D.B.B and K.V. Statistical analysis was performed using SPSS v27.0 (IBM, New York, NY) using the related-samples Wilcoxon signed rank test for non-parametric distributions.
Results
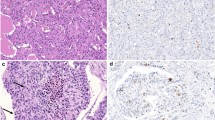
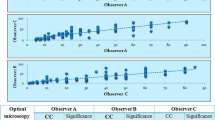
In our MTC cohort (n = 85), 84.7% were low-grade (n = 72) while 15.3% were high-grade (n = 13) by the IMTCGS (Table 1). The Ki67PI for the entire cohort were on average higher for cases measured by MC (median = 0.61%, mean = 1.92%, range = 0–58.37%) compared to DIA (median = 0.40%, mean = 1.59%, range = 0–51.17%). This was also true when evaluating low-grade cases (MC median = 0.47%, MC mean = 0.66%, range = 0–3.14%; DIA median = 0.30%, DIA mean = 0.50%, range 0–3.4%) and high-grade cases (MC median = 2.96%, MC mean = 8.94%, range = 0.64–58.37%; DIA median = 4.90%, DIA mean = 7.63%, range = 0.37–51.17%). A negative difference (i.e., DIA underestimation), positive difference (i.e., DIA overestimation), and a tie between MC and DIA was observed in 56.4% (48/85 cases), 25.9% (22/85), 17.6% (15/85) of the entire MTC cohort, respectively. While the difference between MC and DIA Ki67PI measurements was statistically significant for the entire MTC cohort and the low-grade subset (p < 0.001 for both), it was not statistically significant for the high-grade subset (p = 0.101). Representative examples of a MTC with a high and low Ki67PI are shown in Fig. 1. Compared to MC, QuPath® DIA performed well when considering the entire cohort (R2 = 0.9891). Upon further stratification of MTC cases with IMTCGS, QuPath® correlated better with MC in high-grade cases (R2 = 0.99) compared to low-grade cases (R2 = 0.7071). A comparison of MC to DIA for each case as well as with further stratification by histologic grade was performed with results displayed in Fig. 2.Overall, histologic grade by the IMTCGS was not affected by the method (MC or DIA) of measuring Ki67 proliferation indices for all cases (Table 1). Based on Ki67PI alone, all cases that met the high-grade threshold of ≥ 5% by MC were mirrored by DIA. Similarly, all low-grade cases demonstrated a Ki67PI < 5% with both MC and DIA.
Encountered challenges for QuPath® DIA included optimization of positive cell detection parameters for each case (albeit with minimal changes), nuclear segmentation difficulties due to overlap, and suboptimal focus from tissue folding due to sectioning, pigment, amyloid, or calcification artifact leading to false positive calls (Fig. 3A–D). Perceived challenges for MC of Ki67 positive nuclei included non-specific/weak background Ki67 staining, distinguishing endothelial cells from spindled tumor cells, and time for counting. In a subset of timed MC cases (n = 16 low-grade cases and 8 high-grade cases), manual counting took 8 min and 39 s on average (range 4–32 min 46 s, Supplemental Table 1), compared to < 10 s for DIA (for all cases).
Representative examples of challenges encountered with digital image analysis using QuPath® including (A) incomplete capture of positive cells (indicated by black arrows) requiring optimization of detection parameters, (B) nuclear segmentation difficulty (an example indicated by black arrow), and false positive detection due to pigment or tissue folding/sectioning artifacts (C and D)
Discussion
The use of artificial intelligence as an adjunct to daily clinical practice continues to increase in both surgical and cytopathology. Perhaps the most notable success has been in the quantification of the Ki67PI, an accurate count of which can be important for appropriate grading and prognostication, especially in neuroendocrine tumors [10, 16]. In our study in MTC, we observed that DIA performed comparatively well to MC in measuring Ki67PI for grading of MTC with no major changes in histologic grade based on method of counting. Overall, DIA performed well in identifying Ki67 positive cells compared to MC but appeared to underestimate the Ki67PI when considering the entire cohort. Furthermore, QuPath® DIA performed better in high-grade cases compared to low-grade cases when compared to MC. Our close correlation between MC and DIA for calculating Ki67PI was similar to those of previous studies evaluating DIA in pancreatic neuroendocrine neoplasms and breast tissue [10, 16, 17].
Standardization of DIA-based Ki67 counting in MTCs requires consideration of several pre-analytic and analytic variables, including, but not limited to, tissue processing, staining, selection of DIA platform, hotspot size selection, and optimization of cell detection [10, 15, 18]. All our cases were fixed in formalin, but time to fixation and processing times may have varied as these were cases as part of a routine clinical workflow over two decades, which could impact antigen detection between cases. Sectioning artifacts were a perceived challenge, particularly in heavily calcified cases where obtaining uniform tissue sections is difficult due to shearing. Tissue folding may also cause overlapping nuclei making it difficult for the software to identify individual nuclei. Limitations noted in the evaluation of DIA for pancreatic neuroendocrine included inability to distinguish non-tumor cells from contaminants in QuPath® without teaching the software through test cases [10]. Overstaining with hematoxylin could also impact DAB detection and differences in immunostaining protocols between laboratories could impact Ki67PI values [19]. Finally, the results could be impacted by the antibody clone used. The Ki67 antibody clone used in this study was the MIB-1 clone. Owens et al. found an overestimation of Ki67PI with three non-MIB-1 clones in the context of pancreatic neuroendocrine tumors [15]. Non-specific staining/weak staining of background cells may cause challenges for both DIA and the pathologist performing the manual count.
We did not explore the complexities of whole slide scan imaging in this study; however, the quality of the tissue section, artifacts, or suboptimal focusing can impact the quality of the image used for cell counting [20]. In our study, we used the same slide scanner and scanning parameters for all our cases to minimize this impact. Pai et al. captured 10 × images for their Ki67 counting using a microscope camera and stored them as.jpg files, which are considerably smaller compared to.svs scanned files (kilobytes for.jpg files compared to gigabytes for.svs files) [16]; however, it is difficult to ascertain if the resolution achieved from a 40 × scan would enable improved performance of the system and should be explored in future studies.
Several DIA platforms are available for automated cell counting. We chose to focus on QuPath® for this study due to its free availability and prior publications documenting its success. In the context of breast cancer, Acs et al. demonstrated that QuPath® performed equally well in Ki67 nuclei detection compared to two other DIA platforms—HALO® and Quantcenter®. Moreover, QuPath showed the lowest intra-DIA platform variability in that study [13]. That being said, a comparative study examining the performance of the various systems for Ki67 counting in MTCs will be prudent.
The primary difference between MC and DIA was an underestimation of the Ki67 count by DIA. Of interest, Acs et al. noted that QuPath® underestimated Ki67 counts compared to two other DIA platforms, although it did not significantly impact its overall performance [13]. In our study, the Ki67 positive nuclei detected by DIA was more often lower than that detected by MC (Fig. 3A). The cell detection parameters were optimized for each case, requiring slight adjustments, to ensure that the software had the maximal capability of identifying Ki67 positive nuclei while minimizing background noise, which could introduce systemic bias. Another challenge that might explain the observed overestimation or underestimation of DIA compared to MC includes overlapping nuclei, which may be considered as a single nucleus by the software (i.e., nuclear segmentation difficulties, Fig. 3B). In an optimization study by Pai et al. using QuPath®, the group noted that ~ 15% of their tumor images required changes in the settings to achieve an optimal Ki67 count, with some images requiring more than one adjustment [16]. Tissue artifacts also led to false positive detection (Fig. 3C and D) and requires assessment of the Ki67PI away from these areas to maximize accuracy, if feasible. A rapid quality check of the image would be useful to ensure that the software is appropriately capturing the cells of interest. That being said, despite these encountered challenges, there was no grading impact when using either DIA or MC in our study. Moreover, if this technology is applied to routine clinical practice, one would anticipate performing similar slight adjustments in the detection parameters to ensure that the software is rendering the most accurate possible count.
Another challenge we noted when optimizing cell detection parameters was the morphologic variation and overlap in medullary thyroid carcinoma with other normal elements. MTC can demonstrate diverse morphologies ranging from epithelioid, spindle, and pleomorphic/bizarre nuclei [1]. The spindle morphology will be challenging as it can overlap with endothelial cells. As the cell size and nuclear size of MTC tumor cells may not be necessarily consistent between each case, detection thresholds may need to be adjusted to ensure that lymphocytes and endothelial cells are excluded from the count.
The hotspot size used for the DIA and MC is also important to consider. In Owens et al., increasing the hotspot size from 500 to 2000 cells resulted in a decrease in Ki67PI, possibly representing dilutional effect, but this led to a stronger correlation with MC [15]. Similar findings have also been reported in Volynskaya et al. [21]. The hotspot shape may also be an important factor to consider [22]. For this study, we relied on the WHO recommendation of assessing at least 500–1000 cells for Ki67 counting, acknowledging the caveat that this minimum cell quantity was established in the context of pancreatic neuroendocrine tumors [7]. Owens et al. also raised concern for the potential over counting of Ki67PI by automated methods, but we did not observe this in our study [15]. The hotspot shape may also be an important factor to consider [22]. For our study, we relied on a uniform box encompassing the hotspot, which was consistent across all cases to minimize variations due to the hotspot shape. Nevertheless, additional studies exploring the optimal hotspot size and shape in a variety of tumors, including MTC, will be needed in the future.
Ultimately, these pre-analytic and analytic factors require careful consideration in the validation/calibration process. Nevertheless, a notable perceived advantage was the quicker time to count Ki67 using DIA compared to MC, which would be an incentive to eventually implement this technology into daily practice. In our study, a timed MC count took an average of 8 min and 39 s compared to < 10 s with the DIA count. This is consistent with prior studies demonstrating that MC can take 6 min on average with up to 55 min for some individual cases, whereas DIA can perform a similar analysis in seconds [21]. In addition, there was no significant difference in performance between DIA and MC when considering the entire cohort, low-grade, or high-grade cases. Finally, using the Ki67PI from either DIA or MC provided the same IMTCGS grade in our cohort; as such, we do not anticipate a significant impact on outcome stratification with either method. Hence, DIA could be used as an adjunct for quantifying Ki67.
There are several limitations recognized in this study. First, it is retrospective with a small cohort size. Second, we focused primarily on QuPath® which was open source and relatively easy to use, but it would be worthwhile to compare its performance to other DIA platforms for Ki67 counting in MTCs. Third, one primary pathologist performed the DIA analysis on the cohort, which as discussed previously could introduce bias, and intra-operator variability was not evaluated in this study; nevertheless, based on the study from Acs et al., intra-DIA reproducibility was excellent when using QuPath® for Ki67 quantification of breast cancer cases among four operating pathologists [13]. Finally, a machine-learning algorithmic approach with a training set was not employed in this study. Rather, slight manual parameter adjustments were made to maximize detection of the Ki67-positive nuclei by QuPath®. Future studies with a machine-learning approach in larger multi-institutional studies may enable standardization of detection parameters. Despite these limitations, we were able to use QuPath® with relative ease to quickly quantify Ki67 labeling without significant impact on the ultimate histologic grade.
It is worthwhile recognizing that one parameter alone does not determine the grade of the MTC. All MTC cases requires careful assessment of all three IMTCGS parameters—Ki67PI, necrosis, and mitoses. In our cohort, 13 MTCs met at least one of the three IMTCGS criteria for high-grade, most often necrosis (85%), followed by mitotic activity ≥ 5 per 2 mm2 (54%), and finally Ki67PI ≥ 5% (46%) and finally mitotic activity. The biologic expectation for a high-grade tumor is for increased cell turnover (indicated by necrosis) with expected increased proliferative activity (as judged by Ki67PI and mitotic cut-offs). For instance, four high-grade MTC cases with necrosis demonstrated a Ki67PI and mitotic cut-off that did not meet the cut-offs for high-grade. One potential explanation for the mismatch in parameters is tumor sampling and it is plausible that a stronger hot spot of proliferative activity may not have been represented on the sections available for review. Second, differences in the molecular landscape between these cohort subsets may lead to differences in the biologic behavior, but requires additional study. Finally, as mentioned previously, it is also important to recognize that the established Ki67PI and mitotic activity cut-offs of ≥ 5% and ≥ 5 mitoses/2 mm2 on continuous variables may themselves not truly be reflective of the underlying biologic behavior. As demonstrated in the initial study by Xu et al., when either continuous variable increased, outcomes worsened, and that a 5% cut-off was established primarily for ease of stratifying larger population cohorts [6]. Thus, even if either continuous variable fails to achieve the binary cut-off, the combination of parameters taken together may still confer a more aggressive biologic behavior to the MTC. Thus, reporting precise values for both continuous variables, Ki67PI and mitotic activity is key in the pathology report. Additional study will be needed to further refine the defining criteria to accurately predict a MTC with high-grade behavior.
Our study adds to the collective literature evaluating DIA, specifically QuPath®, for quicker and as accurate analysis of Ki67PI in malignancies similar to MC. To our knowledge, this is the first study exploring its use in Ki67PI in MTCs stratified by the IMTCGS. Furthermore, with the recent implementation of the IMTCGS, this technology will likely continue to be validated in other MTC cohorts. In summary, DIA using the open-source software QuPath® performed as well as MC in measuring the Ki67PI parameter in the IMTCGS for grading MTCs, and our study supports its continued exploration for clinical practice. Ultimately, Ki67PI represents one of the three parameters and regardless of methodology used should be used in conjunction with other histologic findings to determine the final MTC grade.
Data Availability (Data Transparency)
All the data are included in the manuscript.
Code Availability (Software Application or Custom Code)
QuPath® is a freely available, open-source software and can be accessed at the following link: https://qupath.github.io/.
References
R.Y. O, Gill AJ, Tallini G, M. V, M. B, Fuchs TL. Medullary thyroid carcinoma. WHO classification of tumours. Endocrine and neuroendocrine tumours. 5th ed; 2022.
Lim H, Devesa SS, Sosa JA, Check D, Kitahara CM (2017) Trends in thyroid cancer incidence and mortality in the United States, 1974–2013. JAMA 317:1338–1348
Pereira M, Williams VL, Hallanger Johnson J, Valderrabano P (2020) Thyroid cancer incidence trends in the United States: association with changes in professional guideline recommendations. Thyroid 30:1132–1140
Alzumaili B, Xu B, Spanheimer PM et al (2020) Grading of medullary thyroid carcinoma on the basis of tumor necrosis and high mitotic rate is an independent predictor of poor outcome. Mod Pathol 33:1690–1701
Fuchs TL, Nassour AJ, Glover A et al (2020) A proposed grading scheme for medullary thyroid carcinoma based on proliferative activity (Ki-67 and mitotic count) and coagulative necrosis. Am J Surg Pathol 44:1419–1428
Xu B, Fuchs TL, Ahmadi S et al (2022) International medullary thyroid carcinoma grading system: a validated grading system for medullary thyroid carcinoma. J Clin Oncol 40:96–104
Klimstra DS, G. K, S. L, G. R. Classification of neuroendocrine neoplasms of the digestive system. WHO classifcation of tumours. Digestive system tumours; 2019.
Tang LH, Gonen M, Hedvat C, Modlin IM, Klimstra DS (2012) Objective quantification of the Ki67 proliferative index in neuroendocrine tumors of the gastroenteropancreatic system: a comparison of digital image analysis with manual methods. Am J Surg Pathol 36:1761–1770
Reid MD, Bagci P, Ohike N et al (2015) Calculation of the Ki67 index in pancreatic neuroendocrine tumors: a comparative analysis of four counting methodologies. Mod Pathol 28:686–694
Luchini C, Pantanowitz L, Adsay V et al (2022) Ki-67 assessment of pancreatic neuroendocrine neoplasms: systematic review and meta-analysis of manual vs. digital pathology scoring. Mod Pathol 35:712–720
Bankhead P, Loughrey MB, Fernandez JA et al (2017) QuPath: open source software for digital pathology image analysis. Sci Rep 7:16878
Acs B, Leung SCY, Kidwell KM et al (2022) Systematically higher Ki67 scores on core biopsy samples compared to corresponding resection specimen in breast cancer: a multi-operator and multi-institutional study. Mod Pathol 35:1362–1369
Acs B, Pelekanou V, Bai Y et al (2019) Ki67 reproducibility using digital image analysis: an inter-platform and inter-operator study. Lab Invest 99:107–117
Aung TN, Acs B, Warrell J et al (2021) A new tool for technical standardization of the Ki67 immunohistochemical assay. Mod Pathol 34:1261–1270
Owens R, Gilmore E, Bingham V et al (2020) Comparison of different anti-Ki67 antibody clones and hot-spot sizes for assessing proliferative index and grading in pancreatic neuroendocrine tumours using manual and image analysis. Histopathology 77:646–658
Pai R, Karki S, Agarwal R, Sieber S, Barasch S (2022) Optimal settings and clinical validation for automated Ki67 calculation in neuroendocrine tumors with open source informatics (QuPath). J Pathol Inform 13:100141
Skjervold AH, Pettersen HS, Valla M, Opdahl S, Bofin AM (2022) Visual and digital assessment of Ki-67 in breast cancer tissue—a comparison of methods. Diagn Pathol 17:45
Kim JY, Hong SM (2016) Recent updates on neuroendocrine tumors from the gastrointestinal and pancreatobiliary tracts. Arch Pathol Lab Med 140:437–448
Blank A, Wehweck L, Marinoni I et al (2015) Interlaboratory variability of MIB1 staining in well-differentiated pancreatic neuroendocrine tumors. Virchows Arch 467:543–550
Basak K, Ozyoruk KB, Demir D (2023) Whole slide images in artificial intelligence applications in digital pathology: challenges and pitfalls. Turk Patoloji Derg. https://doi.org/10.5146/tjpath.2023.01601
Volynskaya Z, Mete O, Pakbaz S, Al-Ghamdi D, Asa SL (2019) Ki67 quantitative interpretation: insights using image analysis. J Pathol Inform 10:8
Khan Niazi MK, Yearsley MM, Zhou X, Frankel WL, Gurcan MN (2014) Perceptual clustering for automatic hotspot detection from Ki-67-stained neuroendocrine tumour images. J Microsc 256:213–225
Funding
Research reported in this publication was supported in part by the Cancer Tissue and Pathology shared resource of Winship Cancer Institute of Emory University and NIH/NCI under award number P30CA138292. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
KV was involved in the study design. DBB and KV wrote the initial manuscript draft and created the figures. DBB, KV, DJL, KM, QS were involved in the data collection and provided feedback on the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no conflicts of interest or competing interests.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to Participate
Due to the retrospective nature of this study, formal consent is not required. This study was approved by the institutional review board at Emory University (IRB #00004034, K.V).
Consent for Publication (Include Appropriate Statements)
Not applicable for this retrospective study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Behrman, D.B., Lubin, D.J., Magliocca, K. et al. Exploration of Digital Image Analysis for Ki67 Quantification in the Grading of Medullary Thyroid Carcinoma: A Pilot Study with 85 Cases. Head and Neck Pathol 17, 638–646 (2023). https://doi.org/10.1007/s12105-023-01564-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12105-023-01564-8