Abstract
Human prion disease, also known as transmissible spongiform encephalopathy (TSEs), is caused by the conformational conversion of the normal cellular prion protein (PrPC) into the scrapie form (PrPSc). Pathogenic point mutations of prion proteins typically facilitate conformational conversion and lead to inherited prion diseases. A previous study has demonstrated that the pathogenic G131V mutation of human prion protein (HuPrP) brings in Gerstmann–Sträussler–Scheinker syndrome. However, the three-dimensional structure and dynamic features of the HuPrP(G131V) mutant remain unclear. It is expected that the determination of these structural bases will be beneficial to the pathogenic mechanistic understanding of G131V-related prion diseases. Here, we performed 1H, 15N, 13C backbone and side-chain resonance assignments of the G131V mutant of HuPrP(91–231) by using heteronuclear multi-dimensional NMR spectroscopy, and predicted the secondary structural elements and order parameters of the protein based on the assigned backbone chemical shifts. Our work lays the necessary foundation for further structural determination, dynamics characterization, and intermolecular interaction assay for the G131V mutant.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Biological context
Prion disease, the notorious transmissible spongiform encephalopathy (TSEs), is a kind of fatal neurodegenerative illness related to the central nervous system (Prusiner 1998). Four kinds of prion diseases have been identified in humans including fatal familial insomnia (FFI), Kuru, Gerstmann–Sträussler–Scheinker syndrome (GSS), and Creutzfeldt-Jakob disease (CJD) (Tee et al. 2018). These diseases result from the conformational conversion of the normal cellular prion protein (PrPC) into the scrapie isoform prion protein (PrPSc) (Prusiner 1982; Rayman and Kandel 2017). Interestingly, PrPC is mainly α-helix-rich, whereas PrPSc is β-sheet-rich (Pan et al. 1993).
The wild-type human prion protein (WT HuPrP) is composed of 230 amino acid residues, including an N-terminal random coil (residues 23–124) and a C-terminal globular domain (residues 125–231) associated with many pathogenic and protective mutations (Beck et al. 2010; Giachin et al. 2013). Pathogenic point mutations of human prion proteins were reported to accelerate conformational conversions and cause inherited prion diseases (Beck et al. 2010). For the M129 RPNP allele of HuPrP, the G127V mutation causes complete resistance to prion diseases (Mead et al. 2009; Asante et al. 2015), while the G131V mutation leads to the Gerstmann–Sträussler–Scheinker (GSS) disease (Panegyres et al. 2001).
The three-dimensional (3D) structure of the C-terminal domain of WT HuPrP(91–231) in solution has been determined by using heteronuclear multi-dimensional NMR spectroscopy, which is comprised of three α-helices (α1: 144–154, α2: 173–194, α3: 200–228), a short β-sheet (β1: 128–131, β2: 161–164) and a disulfide bridge (Cys179–Cys214) (Zahn et al. 2000). The 3D structure of WT HuPrP shows that the residue G131 is located in the β1-strand. Previously, we determined the 3D structure of the disease-resistant HuPrP(G127V) mutant spanning 91–231 in solution with NMR techniques,and indicated that HuPrP(G127V) consists of the α1, α2 and α3 helices and a stretch-strand (SS) pattern with two fragments (SS1: 128–131, SS2: 161–164) rather than the β-sheet contained in the WT protein (Zheng et al. 2018). Our previous work revealed that HuPrP(G127V) prevents formations of stable β-sheets and dimers through performing molecular dynamics simulations (Zheng et al. 2018). Notably, G131 is located in the SS1 fragment. Moreover, we carried out NMR dynamics analyses of HuPrP(G127V) and WT HuPrP, indicating that Gly131 underwent significant conformational exchange only in HuPrP(G127V) (Zheng et al. 2018). Furthermore, we found that the G131V mutation alleviates the fibrillization but promotes the oligomerization of HuPrP(91–231), suggesting that the oligomerization might play a vital role in the pathogenic mechanisms of the G131V mutant. We also exhibited that the flexible N-terminal fragment in both HuPrP(G127V) and WT HuPrP decreases fibrillization tendencies but increases oligomerization tendencies (Zhang et al. 2019).
Additionally, Santini et al. performed molecular dynamics simulations and showed that the G131V mutation enhances the flexibility of residues participating in the two-step process for prion propagation at neutral pH (Santini et al. 2003). Furthermore, we observed profound differences in 2D 1H-15N HSQC spectra between HuPrPC(G131V) and WT HuPrPC proteins, which implied that the G131 mutation induced significant conformational changes in HuPrP. Expectedly, the conformational distinctions between the two proteins potentially contribute to the pathogenesis of G131V-related prion diseases. However, the conformational distinctions remain unclear due to the lack of three-dimensional (3D) structure of HuPrP(G131V) until now.
In the present study, we performed 1H, 13C, 15N backbone and side-chain resonance assignments of the G131V mutant of HuPrP(91–231) spanning 91–231 by using heteronuclear multi-dimensional NMR spectroscopy, and predicted the secondary structures and order parameters of the protein based on the backbone resonance chemical shifts with the TALOS-N program. Our results lay the basis for further structural determination and NMR dynamics characterization as well as intermolecular interaction assay for the pathogenic HuPrP(G131V) mutant.
Methods and experiments
Plasmid construction, protein expression and purification
A recombinant of the pET30a plasmid bearing the DNA of the WT HuPrP (residues 91–231 with the G131M129 genotype) was used as a template plasmid (Wen et al. 2010a, b). A single point mutant on this plasmid containing a valine substitution of glycine 131 (G131V) was constructed via site-directed mutagenesis PCR and transformed into the E. coli BL21(DE3) strain. The uniformly 13C/15N-labeled proteins were overexpressed at 37 °C in the M9 medium, to which both 1 g/L 15NH4Cl and 3 g/L 13C6-glucose were added. The expression of HuPrP(G131V) protein was induced at OD600nm = 0.8–1.2 using 1 mM isopropyl-β-D-thiogalactopyranoside for 4–5 h at 37 °C. Cells were harvested by centrifugation and later resuspended in lysis buffer (20 mM Tris-HCl, 150 mM NaCl, 2 mM EDTA, 0.1% Triton X-100, pH 7.4), followed by the addition of PMSF (20 µg/mL) with subsequent sonication on ice for 35 min. After centrifugation at 11,000 rpm for 40 min at 4 °C to pellet down the cell debris, the sediment was dissolved in buffer B1 (20 mM Tris-HCl, 150 mM NaCl, 0.5% Triton X-100, pH 7.4), stirred for 1 h at 4 °C, and centrifuged at 11,000 rpm for 30 min at 4 °C. This step was repeated sequentially in buffer B2 (20 mM Tris-HCl, 2 M NaCl, pH 7.4) and buffer B3 (20 mM Tris-HCl, 150 mM NaCl, 2 M Urea, pH 7.4). To denature the protein, the resulting pellet was dissolved in buffer C (10 mM Tris-HCl, 100 mM Na2HPO4, 6 M GdnHCl, 0.5% Triton X-100, 10 mM β-mercaptoethanol, pH 8.0). Then, the denatured protein was stepwise dialyzed and refolded against buffer D (10 mM Tris-HCl, 100 mM Na2HPO4, pH 8.0) in a concentration gradient. The refolded HuPrPC(G131V) protein was further purified by size exclusion chromatography (SEC) with Superdex 75 column attached to an AKTA fast protein liquid chromatography system (GE Healthcare, Wisconsin, USA) pre-equilibrated with buffer F (20 mM NaAc, 0.02% NaN3, pH 4.5). Finally, the protein solutions were concentrated to approximately 0.65 mM in NMR buffer (20 mM NaAc, 0.02% NaN3, 10% D2O, pH 4.5).
NMR spectroscopy
To perform backbone and side-chain resonance assignments of the HuPrPC(G131V) protein, a suite of heteronuclear 2D/3D NMR spectra were recorded at 298 K on a Bruker Avance III 850-MHz spectrometer with a 1H/13C/15N triple-resonance cryogenic probe (TCI). The 2D NMR spectra included 1H-13C HSQC and 1H-15N HSQC. The 3D NMR spectra covered HNCACB, HNCA, HN(CA)CO, CBCA(CO)NH, HN(CO)CA, HCCH-TOCSY, HNCO, HBHA(CO)NH, H(CCCO)NH, CC(CO)NH, CCH-TOCSY, and 15N-edited TOCSY-HSQC using non-uniform sampling (NUS) with a sampling rate of 25%. Furthermore, both 3D 15N-edited NOSEY-HSQC and 13C-edited NOESY-HSQC spectra were recorded using traditional uniform sampling with a mixing time of 100 ms.
All NMR spectra were acquired with the Bruker TopSpin 3.6.2 software, processed with the NMRPipe software (Delaglio et al. 1995) and analyzed with the Sparky software (Lee et al. 2015). All the 3D spectra recorded with non-uniform sampling were reconstructed and processed by using the SMILE package of NMRPipe (Ying et al. 2017). Both the secondary structures and order parameters of HuPrPC(G131V) were predicted based on the assigned backbone chemical shifts using the TALOS-N program (Shen and Bax 2013).
Assignment and data deposition
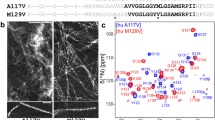
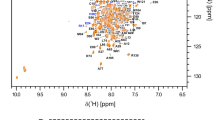
The recombinantly expressed HuPrPC(G131V) protein consists of 141 amino acid residues (residues 91–231) with 5 proline residues (102, 105, 137, 158 and 165). Superimposition of the 2D 1H-15N HSQC spectra of HuPrPC(G131V) and WT HuPrPC indicates that the G131V mutation induced significant structural changes in the protein (Fig. 1). Backbone and side-chain resonances of the G131V mutant were assigned based on the recorded heteronuclear 2D/3D NMR spectra (Fig. 2). In total, 89.7% of the 1HN and 15N resonances of all non-proline residues were unambiguously assigned. In detail, 93.6% of 13Cα (132 of 141), 92.2% of 13Cβ (119 of 129), 92.2% of 13C(O) (130 of 141), 93.9% of 1Hα (155 of 165), and 80.68% of side-chain 1H resonances have been assigned. Some residues (Met129-Ser132, Val161, Tyr162, Asp167-Asn171, Phe175, Ile182 and Gln217) had no available resonance assignments for backbone amide groups. Among these residues, however, Ser132, Tyr162, Asn171, Phe175, Ile182 and Gln217 had available resonance assignments for 13Cα, 13Cβ, 13 C(O), 1Hα atoms.
The solution structure of WT HuPrP(91–231) (Zheng et al. 2018; PDB: 5YJ5) contains two short β-strands (β1: 128–131, β2: 161–164). Backbone amide resonances of M129, I130, and V131 were visible in the HSQC spectrum of the WT protein, but invisible in that of the G131V mutant. The spectral differences suggested that the G131V mutation induced significant conformational changes in the β-sheet of the prion protein. The superimposed 1H-15N HSQC spectra of the G131V mutant and WT protein spanning 91–231 illustrate that the G131V mutation induced peak broadening for residues including Ala133, Ile215, Tyr218 and Arg220, and even peak disappearing for four residues (Met129, Ile130, Val161 and Gln217). The assigned chemical shifts of the backbone and side-chain atoms in HuPrPC(G131V) have been deposited in BioMagResBank (http://www.bmrb.wisc.edu/) under the access number 50,544.
Using the TALOS-N server (https://spin.niddk.nih.gov/bax/software/TALOS-N/), we predicted the secondary structure elements of HuPrPC(G131V) spanning 91–231 based on the assigned chemical shifts of the backbone HN, Hα, Cα, Cβ, C(O) and N (Fig. 3a). The prediction results suggested that the solution structure of the G131V mutant contained three α-helices (α1: 144–153, α2: 172–192, α3: 200–226) (Fig. 3a). The predicted helices in HuPrPC(G131V) were consistent with the experimentally determined helices in WT HuPrPC(144–154, 173–194, 200–228) (Zahn et al. 2000). Residues 161–164 were predicted to be a β-strand. Note that residues 128–131 form a β-strand comprising a β-sheet with another β-strand (residues 161–164) in WT HuPrP(91–231). In HuPrP(G131V), however, residues 128–131 were not predicted to be a β-strand due to unavailable resonance assignments of backbone amide groups for M129-S132 as described above. Whether the residues 161–164 could form a β-strand in HuPrP(G131V) need to be clarified by the 3D structure of HuPrP(G131V) determined in near future.
Secondary structures and order parameters of HuPrPC(G131V) spanning 91–231 predicted from the assigned backbone chemical shifts. a Secondary structural elements of HuPrP(G131V) were predicted based on backbone chemical shifts by using the TALOS-N program with a propensity cutoff of 0.5. b Random coil index (RCI) order parameters (S2) were calculated from secondary chemical shifts for individual residues by TALOS-N. A cutoff of 0.7 was used for distinguishing rigid and flexible backbone atoms
Apart from the prediction on this fragment, the prediction of secondary structures was generally reliable. Additionally, we used the TALOS-N program to predict random coil index (RCI) order parameters (S2) for the residues with assigned backbone chemical shifts (Fig. 3b). As is known, residues with S2 value < 0.7 are generally located in flexible fragments, while those with S2 value > 0.7 are typically associated with the rigid conformation. Thus, we used a cutoff of 0.7 to distinguish rigid and flexible backbone atoms. Similar to WT HuPrP, the fragment Gln91-Gly127 and the C-terminus showed evidence of conformational flexibility. Notably, the predicted β1 exhibited high S2 values comparable to the three α-helices. As the G131V mutation might affect the conformational stability of the β-sheet, the large S2 values of the fragment 161–164 could also result from the stabilization of their side chains by spatially neighboring residues in the helices. As an illustration, the fragment 161–164 in WT HuPrP(91–231) is stabilized by not only the hydrogen bonds with residues 128–131, but also hydrophobic interactions with non-polar residues like Phe175, Val210 and Met213 (PDB: 5YJ5). To conclude, the differences in the secondary structures between HuPrP(G131V) and WT HuPrP might contribute to the pathogenesis of G131V-related prion diseases.
In summary, we have performed 1H, 13C, 15N backbone and side-chain resonance assignments of HuPrPC(G131V) spanning 91–231 and predicted the secondary structural elements and order parameters of the prion protein. The G131V mutation significantly changes the conformation of the human prion protein. This work lays the basis for the further structural and dynamics studies of HuPrPC(G131V), and may be beneficial to an understanding of molecular mechanisms underlying the conformational conversion associated with the pathogenic G131V mutation.
References
Asante EA et al (2015) A naturally occurring variant of the human prion protein completely prevents prion disease. Nature 522(7557):478–478
Beck JA et al (2010) PRNP allelic series from 19 years of prion protein gene sequencing at the MRC prion unit. Hum Mutat 31(7):E1551–E1563
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6(3):277–293
Giachin G, Biljan I, Ilc G, Plavec J, Legname G (2013) Probing early misfolding events in prion protein mutants by NMR spectroscopy. Molecules 18(8):9451–9476
Lee W, Tonelli M, Markley JL (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31(8):1325–1327
Mead S et al (2009) A novel protective prion protein variant that colocalizes with kuru exposure. New Engl J Med 361(21):2056–2065
Pan KM et al (1993) Conversion of alpha-helices into beta-sheets features in the formation of the scrapie prion proteins. Proc Natl Acad Sci USA 90(23):10962–10966
Panegyres PK et al (2001) A new PRNP mutation (G131V) associated with Gerstmann-Straussler-Scheinker disease. Arch Neurol 58(11):1899–1902
Prusiner SB (1982) Novel proteinaceous infectious particles cause scrapie. Science 216(4542):136–144
Prusiner SB (1998) Prions. Proc Natl Acad Sci USA 95(23):13363–13383
Rayman JB, Kandel ER (2017) Functional prions in the brain. Cold Spring Harb Perspect Biol 9(1):a023671
Santini S, Claude JB, Audic S, Derreumaux P (2003) Impact of the tail and mutations G131V and M129V on prion protein flexibility. Proteins Struct Funct Bioinform 51(2):258–265
Shen Y, Bax A (2013) Protein backbone and sidechain torsion angles predicted from NMR chemical shifts using artificial neural networks. J Biomol NMR 56:227–241. https://doi.org/10.1007/s10858-013-9741-y
Tee BL, Longoria Ibarrola EM, Geschwind MD (2018) Prion diseases. Neurol Clin 36(4):865–897
Wen Y et al (2010a) Solution structure and dynamics of the I214V mutant of the rabbit prion protein. PLoS ONE 5(10):e13273
Wen Y et al (2010b) Unique structural characteristics of the rabbit prion protein. J Biol Chem 285(41):31682–31693
Ying JF, Delaglio F, Torchia DA, Bax A (2017) Sparse multidimensional iterative lineshape-enhanced (SMILE) reconstruction of both non-uniformly sampled and conventional NMR data. J Biomol Nmr 68(2):101–118
Zahn R et al (2000) NMR solution structure of the human prion protein. Proc Natl Acad Sci USA 97(1):145–150
Zhang M et al (2019) Biophysical characterization of oligomerization and fibrillization of the G131V pathogenic mutant of human prion protein. Acta Biochim Biophys Sin (Shanghai) 51(12):1223–1232
Zheng Z et al (2018) Structural basis for the complete resistance of the human prion protein mutant G127V to prion disease. Sci Rep 8(1):13211
Acknowledgements
This work was supported by grants from the Natural Science Foundation of China (No. 31670741), the National Key Research and Development Project of China (No. 2016YFA0500600) and the Foundation for Innovative Research Groups of the National Natural Science Foundation of China (No. 21521004).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhang, Q., Zhang, H., Zheng, F. et al. 1H, 13C, 15N backbone and side‐chain resonance assignments of the pathogenic G131V mutant of human prion protein (91–231). Biomol NMR Assign 15, 311–316 (2021). https://doi.org/10.1007/s12104-021-10022-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-021-10022-x