Abstract
Background
Colorectal cancer has a high incidence and mortality rate due to a low rate of early diagnosis. Therefore, efficient diagnostic methods are urgently needed.
Purpose
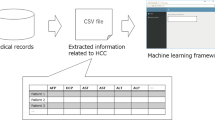
This study assesses the diagnostic effectiveness of Carbohydrate Antigen 19–9 (CA19-9), Carcinoembryonic Antigen (CEA), Alpha-fetoprotein (AFP), and Cancer Antigen 125 (CA125) serum tumor markers for colorectal cancer (CRC) and investigates a machine learning-based diagnostic model incorporating these markers with blood biochemical indices for improved CRC detection.
Method
Between January 2019 and December 2021, data from 800 CRC patients and 697 controls were collected; 52 patients and 63 controls attending the same hospital in 2022 were collected as an external validation set. Markers’ effectiveness was analyzed individually and collectively, using metrics like ROC curve AUC and F1 score. Variables chosen through backward regression, including demographics and blood tests, were tested on six machine learning models using these metrics.
Result
In the case group, the levels of CEA, CA199, and CA125 were found to be higher than those in the control group. Combining these with a fourth serum marker significantly improved predictive efficacy over using any single marker alone, achieving an Area Under the Curve (AUC) value of 0.801. Using stepwise regression (backward), 17 variables were meticulously selected for evaluation in six machine learning models. Among these models, the Gradient Boosting Machine (GBM) emerged as the top performer in the training set, test set, and external validation set, boasting an AUC value of over 0.9, indicating its superior predictive power.
Conclusion
Machine learning models integrating tumor markers and blood indices offer superior CRC diagnostic accuracy, potentially enhancing clinical practice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colorectal cancer is the third most common cancer and the second leading cause of cancer-related deaths globally, accounting for approximately 10% of all cancer cases and deaths [1]. In 2018, the number of new cases and deaths from colorectal cancer worldwide was 1.9 million and 540,000, respectively, and it is projected that its global burden will increase by 60% by 2030, with the number of new cases and deaths reaching 2.2 million and 1.1 million, respectively [2, 3]. Statistically, the 5 year relative survival rate for colorectal cancer cases in the United States can be as high as 90% when detected at an early stage and only 14% when distant metastases are present [4]. Therefore, early screening and identification of colorectal cancer patients is important for reducing colorectal cancer mortality and prolonging the quality and duration of patient survival. Colonoscopy is currently the gold standard for the diagnosis of colorectal cancer, but its shortcomings such as invasiveness, complex bowel preparation, certain risks, and high cost limit its use in large-scale population screening [5, 6]. Fecal noninvasive testing and serum tumor marker testing are currently commonly used noninvasive testing methods for CRC in the clinic [7, 8], and compared with feces, people’s compliance with blood sample collection is better in practical application [9]. Serum tumor markers are mainly abnormally elevated when tumor-related genes are expressed or when the body recognizes tumors, so they have important value in both the diagnosis and prognosis of tumors and are widely used in clinical practice [10]. In recent years, with the rapid development of information technology, more and more researchers have applied artificial intelligence technology to disease diagnosis and prognosis prediction and achieved good results [11,12,13]. This research undertakes a thorough examination of the diagnostic efficacy of four serum tumor markers (CA199, CEA, AFP, CA125), which are prevalently utilized in the clinical detection of colorectal cancer (CRC), evaluating their performance both singularly and synergistically. Building on this, the study endeavors to craft a machine learning-based diagnostic model for CRC. This advanced model will integrate the aforementioned tumor markers with clinical blood biochemistry indicators that are readily obtainable from patients. By conducting a comparative analysis between this innovative model and the traditional tumor marker approaches, the study aims to uncover a more efficacious diagnostic route for CRC. This endeavor is set to contribute significantly toward the enhancement of early screening procedures and the development of more targeted management strategies for CRC. Ultimately, the study's objective is to identify and deploy a dependable model capable of enabling the early detection and timely therapeutic intervention for individuals diagnosed with CRC, thereby potentially improving patient outcomes.
Materials and methods
Participants
Internal validation set: Case group: retrospective collection of colorectal cancer patients who attended the Department of Anorectal and Gastroenterology of the First Affiliated Hospital of Dalian Medical University from January 2019 to December 2021. Inclusion criteria: 1. patients diagnosed with primary colorectal cancer by pathologic findings; 2. aged between 18 and 85 years old; 3. none of them had undergone surgery and radiotherapy for related diseases; 4. all of them were tested for four serum markers; 5. consent was obtained from the patients and their family members and an informed consent form was signed. Exclusion criteria: 1. pregnant and lactating women; 2. accompanied by acute and critical illnesses or organ failure; 3. serious deficiencies in blood counts and serum markers in the medical records; and 4. 800 cases were finally included.
Control group: retrospective collection of people who visited the health management center of the First Hospital of Dalian Medical University during the same period. Inclusion criteria: 1. people who underwent colonoscopy and serum markers; 2. aged between 18 and 85 years old; 3. consent from the person and family members and signing of the informed consent. Exclusion criteria: 1. patients diagnosed with colorectal cancer by colonoscopy; 2. pregnant and lactating women; 3. those with serious deficiencies in blood routine and serum markers in their medical records; and 4. 697 controls were finally included.
External validation set: Case group: retrospective collection of colorectal cancer patients who attended the Department of Anus and Intestines of the First Hospital of Dalian Medical University in 2022.
Control group: retrospective collection of people who visited the health management center of the First Hospital of Dalian Medical University during the same period. The inclusion and exclusion criteria of the external validation set were the same as those of the internal validation set.The study was approved by the Ethics Committee of the First Hospital of Dalian Medical University.
Data collection
The study collected comprehensive data encompassing both demographic and health-related metrics. Demographic information included gender and age. Health-related data comprised a wide range of measures:
Basic health indicators: Body Mass Index (BMI), Systolic Blood Pressure (SBP), and Diastolic Blood Pressure (DBP).
Blood composition analysis:
Complete blood count: White Blood Cell (WBC), Neutrophil (NEUT), Lymphocyte (LYMPH), Red Blood Cell (RBC), Hemoglobin (Hb), Hematocrit (HCT).
Blood cell metrics: Red Cell Distribution Width—Standard Deviation (RDW-SD), Red Cell Distribution Width—Coefficient of Variation (RDW-CV), Platelet (PLT), Platelet Distribution Width (PDW), Mean Platelet Volume (MPV), Platelet Large Cell Ratio (P-LCR).
Kidney function tests: Creatinine (Cr) and Uric Acid (UA).
Liver function tests: Glutamic Acid (Glu), Alanine Aminotransferase (ALT), Aspartate Aminotransferase (AST), Albumin (ALB), Gamma-glutamyl Transferase (GGT), Total Bilirubin (TBIL), Direct Bilirubin (DBIL).
Cancer markers: Carbohydrate Antigen 19–9 (CA19-9), Carcinoembryonic Antigen (CEA), Alpha-fetoprotein (AFP), Cancer Antigen 125 (CA125).
This detailed collection of data aims to provide a holistic view of the participants’ health, facilitating a nuanced analysis of the relationship between these variables and colorectal cancer.
Sample collection and testing methods
Fasting venous blood of 2 ~ 4 ml was collected, and the serum was separated by centrifugation at 3000 r/min for 10 min and stored at − 20 ℃ in the refrigerator for examination. Blood routine items were detected by Japanese sysmexXN-10 instrument; blood glucose, liver function and kidney function were detected by Hitachi 7600–210 automatic biochemical analyzer; four serum markers were detected by electrochemiluminescence method, and the instrument was Myeri CL-6000i. AFP (< 20 ng/ml), CA125 (< 35 U/ml), beyond the above range is considered positive.
Data cleaning
The outliers in the data were assigned as NA, and BMI was classified into categorical variables (< 18.5, 18.5–24.9, ≥ 25) according to WHO standards. Missing values were interpolated using the “MissForest” R software, which was proposed by Stekhoven in 2012 and is an iterative interpolation method based on random forests, which essentially treats missing value interpolation as a prediction problem and can simultaneously deal with mixed data consisting of both categorical and continuous variables, and is superior to the K-nearest-neighbors, MICE package-based chain interpolation, and other interpolation methods [14, 15]. For the interpolated data, multicollinearity test was performed using the “performance” R software to determine whether multicollinearity exists among independent variables by variance inflation factor (VIF), and the variables with VIF < 5 were included in the study [16].
Model construction
In this study, several indicators, including area under the ROC curve AUC, accuracy, sensitivity, specificity, precision, and F1 score, were used to assess the efficacy of four serum tumor markers, CA199, CEA, AFP, and CA125, for diagnosing colorectal cancer alone and in combination. In general, we believe that the higher the AUC value, the better the differentiation of the model, and when the AUC ≥ 0.9, the model performs well; when the AUC is between 0.8 and 0.9, the model performs well; when the AUC is between 0.7 and 0.8, the model performance is fair; and when the AUC < 0.7, the model performance is poor [17, 18]. Then, the variables were screened using the stepwise regression (backward) method, which was implemented by the stepAIC function in the “MASS” R package, which was based on the AIC (Akaikei Information Criterion), in which all independent variables were firstly put into the model, and then the insignificant variables were gradually eliminated, so that the fewest independent variables were obtained, which resulted in the lowest AIC value and the best model performance [19]. The screened variables were then used to construct the machine learning model. In this study, six machine learning algorithms, logistic regression (LR), support vector machine (SVM), gradient boosting machine (GBM), plain Bayes (NB), artificial neural network (ANN), and random forest (RF), were selected to construct the model, and the dataset was randomly divided into the training set and the test set according to the ratio of 7:3, and the tenfold cross-validation was used to internally validate the model. The predictive performance of the models was evaluated using several indicators, including area under the ROC curve AUC, accuracy, sensitivity, specificity, precision, and F1 score, and compared with the diagnostic performance of tumor markers, and the better performing model was entered into the external validation set for validation.. The machine learning models in this study were all constructed by the “caret” package in R4.3.2, and the ROC curves were plotted by the “pROC” package.
Statistical analysis
This study used R4.3.2 to process and analyze the data, and the measurement information was expressed as mean ± standard deviation (x ± s), and t test was used for comparison between groups; the count information was expressed as percentage (%), and χ2 test was used for comparison between groups. All statistical tests were two-sided, and the differences were considered statistically significant at P < 0.05.
Results
Baseline information
The flow chart for this study is shown in Fig. 1. As shown in Table 1, 800 CRC patients and 697 non-patients were included in this study. Among the demographic variables, the differences in age and BMI between the two groups were statistically significant; although there was no difference in gender, the proportion of males was significantly higher than that of females in both groups. Among the four serum markers, three of them, CA199, CEA, and CA125, were significantly associated with CRC, and their mean values were significantly higher in the case group than in the control group. Among the rest of the biochemical markers, all of them were significantly correlated with CRC, except for RDW-CV, PLT, PDW, Cr, AST, and GGT, the differences of which were not statistically significant between the two groups.
Test of covariance
Multiple covariance test was performed on the independent variables and it was concluded that the VIF values of all the independent variables were less than 5 (Fig. 2) and there was no multiple covariance among the variables, so all the variables could be included in the study normally and no need to be excluded.
Serum marker model
As shown in Fig. 3, when the four serum markers were utilized to predict CRC alone, only the area under the ROC curve of CEA (AUC = 0.79) had an AUC of 0.7 or more, and the model performance was fair, followed by CA199 (AUC = 0.643), and the two indicators of CA125 (AUC = 0.592) and AFP (AUC = 0.553) had a poorer ability to predict CRC. The combination of four serum markers to predict CRC was better than the use of a single tumor marker when predicting, the AUC value reached more than 0.8, in Table 2, the AUC value, accuracy, specificity and precision of the model combining the four tumor markers were higher than that of the four single-prediction models.
Machine learning models
The backward stepwise regression method was applied to screen out 17 variables (Table 3), which were incorporated into six machine learning models. The ROC curves and model evaluation metrics of the training set are shown in Fig. 4A and Table 4, respectively. In the training set, except for RF, which showed overfitting, the AUC values of all the models reached more than 0.8, among which, the three models, GBM, SVM, and LR, with AUC values of 0.9 or more, had excellent prediction performance. GBM had the highest AUC value in the training set, reaching 0.945, with the best model performance; SVM was the second highest, at 0.936; NB had relatively poor performance, at 0.865.The ROC curves and model evaluation metrics for the test set are shown in Fig. 4B and Table 5. In the test set, the AUC values of GBM and RF were both 0.931, but the accuracy, sensitivity, and F1 score of GBM were higher than those of RF, so GBM was the best model for diagnosing CRC.RF had the highest specificity and accuracy of the six models, and its prediction performance was only second to that of RF.GBM was the best model for diagnosing CRC. highest among the six models, and the predictive performance was second only to GBM. The AUC values of the two models, ANN and NB, still did not reach 0.9, and the predictive performance was relatively poor. The ROC curves and model evaluation indexes of the external validation set are shown in Fig. 3C and Table 6.In the external validation set, the AUC value of GBM is still the highest among all the models, except that the AUC value of RF is slightly lower than that of GBM, but the remaining evaluation indexes, such as accuracy and specificity, are all the highest among all the models. In conclusion, after internal and external validation, the diagnostic ability of GBM and RF for CRC is more prominent and has certain extrapolation ability. Among the six machine learning models, the variables in the top five in terms of variable importance are CEA and ALB, followed by age, which is in the first place in terms of importance contribution in all five models except ANN; DBIL and HCT are in the top order of contribution in several models, and they are important variables in the prediction of CRC, as shown in Supplementary Fig. 1.
Discussion
Colorectal cancer, as one of the most common malignant tumors, is known for its high morbidity and mortality, which brings a huge burden to patients and society [20, 21]. Early screening and diagnosis are of profound significance in reducing the morbidity and mortality of colorectal cancer. As a noninvasive, economical and conveniently sampled test, serum tumor marker assay is now commonly used in the clinic for screening and diagnosis of various types of tumors and prognostic assessment [22, 23]. However, an increasing number of studies have found that tumor markers have low sensitivity or specificity when utilized for cancer diagnosis [22, 24], and thus their ability to serve as an independent screening tool for malignant tumors remains to be considered. Carcinoembryonic antigen (CEA) is the most widely used tumor marker for colorectal cancer that has been identified, and was first demonstrated in human colorectal adenocarcinoma by Gold and Freedman in 1965 [25].ASCO recommends that CEA be used as an important factor in performing postoperative surveillance and prognostic evaluation of colorectal cancer but should not be used in the early screening diagnosis of CRC, due to its diagnostic sensitivity is low and can lead to excessive occurrence of false positives [26]. This is consistent with our findings that although CEA was the most well differentiated tumor marker for independent diagnosis of CRC in this study, with an AUC value close to 0.8, its sensitivity was only 0.589, the lowest among the four tumor markers. In addition, CEA is not only found in CRC patients, but also in esophageal, gastric, and breast cancers, which can also cause elevated serum CEA levels [27], and in non-cancerous diseases such as hepatitis and pancreatitis [28]. Therefore, CEA is often used in conjunction with other tumor markers or as an adjunct to diagnosis rather than as an independent diagnostic tool.
In addition to carcinoembryonic antigen (CEA), commonly used tumor markers for colorectal cancer include carbohydrate antigen CA19-9 and CA125 [29, 30]. Despite the disadvantages of low sensitivity for early identification and inability to effectively differentiate from benign diseases, CA199 is still the only tumor marker designated by the FDA for monitoring pancreatic ductal carcinoma in clinical practice [31]; whereas CA125 is mainly used for screening and monitoring of patients with ovarian cancer [32], and CA199 and CA125 also have certain clinical diagnosis and prognosis of CRC. Previous studies have demonstrated [33, 34] that the efficacy of CEA in the diagnosis of CRC is superior to that of CA125 and CA199, and our study came to a similar conclusion that the AUC value, accuracy, and specificity of CEA were higher than those of other tumor markers. Therefore, both are not significant when used alone for CRC diagnosis and are usually used as a complement to CEA or in combination with other tumor markers. Alpha-fetoprotein (AFP) is a glycoprotein produced by the fetal liver and yolk sac. In healthy adults, levels of AFP are usually low. However, it is significantly elevated in the serum of patients with hepatocellular carcinoma, and therefore, in clinical practice, AFP is mainly used as an important tumor marker in the diagnosis and prognostic assessment of hepatocellular carcinoma. In addition to this, AFP has also been used to monitor other types of cancer such as gastric and colorectal cancers [35]. In this study, AFP differed from the other three tumor markers in that its value did not differ significantly between the case group and the control group, and its AUC value for predicting CRC alone was only 0.553, which is a poor predictive performance and is only used in combination with other tumor markers for the diagnosis or monitoring of CRC.
With the development of AI technology, more and more researchers are integrating it into the practice of disease diagnosis and treatment. Our study constructed six machine learning models using common demographic and laboratory indicators in the clinic, and compared their diagnostic efficacy by AUC value, accuracy, sensitivity, specificity, etc., and finally selected the best-performing gradient boosting machine model. In addition to this, we also analyzed the variable importance of each of the six machine learning models. In the best performing GBM model, the top five variables in terms of importance were Age, ALB, CEA, HCT, Hb. In our study, age was the most important risk factor for colorectal cancer and the incidence increased with age. This is in agreement with the conclusion reached by USPSTF [36] and since most of the new cases were above 45 years of age, 45 years was set as the age node for which colorectal cancer screening is recommended.ALB is synthesized mainly by the liver and is an important protein in plasma and is often used clinically as a measure of the nutritional and health status of patients [37]. In a study by Heys et al. [38], pre-treatment serum albumin concentration could be used as an independent prognostic indicator for colorectal cancer, suggesting that we can include albumin in the screening of prognostic and screening markers for colorectal cancer. In particular, ALB was found to be of high importance in our study, only after age. This may be related to the effect of tumor burden on the status of the body, and attention to changes in ALB in cancer screening is important for the diagnosis of colorectal cancer. Colorectal cancer patients are often associated with the development of anemia [39], and both HCT and Hb are diagnostic markers of anemia, and a decrease suggests a risk of anemia [40, 41]. Ben et al. [42] combined HCT and Hb with several other laboratory indicators, respectively, to construct a prediction model for sporadic colorectal cancer, and after linear correction, the AUC value of the HCT model reached 0.76, and the Hb model had an AUC value of 0.80, indicating that both had good predictive value for colorectal cancer.
Our study has several limitations. First, this study only applies external validation of the models with populations attending the same hospital at different times of the day, and lacks validation from different hospitals or districts, which may affect the extrapolation ability of the model to some extent. Second, the variables in this study were mainly laboratory indicators and did not incorporate information on lifestyle, dietary habits, and past medical history; in future, it is hoped that such variables can be further collected and incorporated into the model to improve the predictive ability of the model and expand the scope of model application.
Conclusion
In summary, compared with the traditional serological tumor markers, the machine learning model shows more excellent performance in colorectal cancer diagnosis, with the gradient boosting machine model as the best choice. When this model is applied to large-scale population screening, it can more accurately distinguish colorectal cancer patients from healthy people, provide doctors with reliable diagnostic basis, and provide important support for the rational allocation of medical resources. In addition, ALB, HCT and Hb, as very common and economical tests in clinical practice, show similar predictive efficacy to that of tumor markers in the prediction of CRC, which will be of great value in the prediction and prognosis of CRC in the future.
Data availability
The data from this study can be obtained from the corresponding author.
References
Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer statistics, 2022. CA Cancer J Clin. 2022;72(1):7–33.
Arnold M, Sierra MS, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global patterns and trends in colorectal cancer incidence and mortality. Gut. 2017;66(4):683–91.
Vabi BW, Gibbs JF, Parker GS. Implications of the growing incidence of global colorectal cancer. J Gastrointest Oncol [Internet]. 2021 Jul [Accessed 4 Feb 2024];12(Suppl 2). Available from: https://jgo.amegroups.org/article/view/43295
Siegel RL, Miller KD, Goding Sauer A, Fedewa SA, Butterly LF, Anderson JC, et al. Colorectal cancer statistics, 2020. CA Cancer J Clin. 2020;70(3):145–64.
Chan SCH, Liang JQ. Advances in tests for colorectal cancer screening and diagnosis. Expert Rev Mol Diagn. 2022;22(4):449–60.
Weiss JB, Cetel NS, Weiss DE. Colorectal cancer screening: colonoscopy has disadvantages. Cleve Clin J Med. 2019;86(12):774–6.
Werner S, Krause F, Rolny V, Strobl M, Morgenstern D, Datz C, et al. Evaluation of a 5-marker blood test for colorectal cancer early detection in a colorectal cancer screening setting. Clin Cancer Res. 2016;22(7):1725–33.
Luo H, Shen K, Li B, Li R, Wang Z, Xie Z. Clinical significance and diagnostic value of serum NSE, CEA, CA19-9, CA125 and CA242 levels in colorectal cancer. Oncol Lett. 2020;20(1):742–50.
Osborne J, Wilson C, Moore V, Gregory T, Flight I, Young G. Sample preference for colorectal cancer screening tests: blood or stool? [Internet]. 2012 [Accessed 5 Feb 2024]; Available from: https://digital.library.adelaide.edu.au/dspace/handle/2440/76944
Pang C, Ma Y, Shi W, Zi M, Chen J, Liang C, et al. Prognostic significance of serum tumor markers in various pathologic subtypes of gastric cancer. J Gastrointest Surg [Internet]. 2024 Feb 20 [Accessed 2 Apr 2024]; Available from: https://www.sciencedirect.com/science/article/pii/S1091255X24003317
Cao R, Yang F, Ma SC, Liu L, Zhao Y, Li Y, et al. Development and interpretation of a pathomics-based model for the prediction of microsatellite instability in colorectal cancer. Theranostics. 2020;10(24):11080–91.
Waljee AK, Weinheimer-Haus EM, Abubakar A, Ngugi AK, Siwo GH, Kwakye G, et al. Artificial intelligence and machine learning for early detection and diagnosis of colorectal cancer in sub-Saharan Africa. Gut. 2022;71(7):1259–65.
Lee C, Light A, Alaa A, Thurtle D, van der Schaar M, Gnanapragasam VJ. Application of a novel machine learning framework for predicting non-metastatic prostate cancer-specific mortality in men using the Surveillance, Epidemiology, and End Results (SEER) database. Lancet Digit Health. 2021;3(3):e158–65.
Stekhoven DJ, Bühlmann P. MissForest—non-parametric missing value imputation for mixed-type data. Bioinformatics. 2012;28(1):112–8.
You J, Ellis JL, Adams S, Sahar M, Jacobs M, Tulpan D. Comparison of imputation methods for missing production data of dairy cattle. Animal. 2023;17:100921.
Applied linear statistical models.djvu - Contents.pdf [Internet]. [Accessed 22 Jan 2024]. Available from: https://thuvienso.hoasen.edu.vn/v/web/viewer.html?file=/bitstream/handle/123456789/9564/Contents.pdf?sequence=1&isAllowed=y
Muller MP, Tomlinson G, Marrie TJ, Tang P, McGeer A, Low DE, et al. Can routine laboratory tests discriminate between severe acute respiratory syndrome and other causes of community-acquired pneumonia? Clin Infect Dis Off Publ Infect Dis Soc Am. 2005;40(8):1079–86.
Hanley JA. Receiver operating characteristic (ROC) methodology: the state of the art. Crit Rev Diagn Imaging. 1989;29(3):307–35.
Zhang Z. Variable selection with stepwise and best subset approaches. Ann Transl Med. 2016;4(7):136.
Xi Y, Xu P. Global colorectal cancer burden in 2020 and projections to 2040. Transl Oncol. 2021;14(10): 101174.
Keum N, Giovannucci E. Global burden of colorectal cancer: emerging trends, risk factors and prevention strategies. Nat Rev Gastroenterol Hepatol. 2019;16(12):713–32.
Clinical value of combining serum tumor marker detection with fecal occult blood testing in diagnosing colorectal cancer. J Physiol Pharmacol [Internet]. 2022 [Accessed 18 Feb 2024]; Available from: https://doi.org/10.26402/jpp.2022.3.12
Sisik A, Kaya M, Bas G, Basak F, Alimoglu O. CEA and CA 19–9 are still valuable markers for the prognosis of colorectal and gastric cancer patients. Asian Pac J Cancer Prev. 2013;14(7):4289–94.
Palmqvist R, Engarås B, Lindmark G, Hallmans G, Tavelin B, Nilsson O, et al. Prediagnostic levels of carcinoembryonic antigen and CA 242 in colorectal cancer: a matched case-control study. Dis Colon Rectum. 2003;46(11):1538.
Gold P, Freedman SO. Demonstration of tumor-specific antigens in human colonic carcinomata by immunological tolerance and absorption techniqueS. J Exp Med. 1965;121(3):439–62.
Locker GY, Hamilton S, Harris J, Jessup JM, Kemeny N, Macdonald JS, et al. ASCO 2006 update of recommendations for the use of tumor markers in gastrointestinal cancer. J Clin Oncol Off J Am Soc Clin Oncol. 2006;24(33):5313–27.
Filella X, Rodríguez-Garcia M, Fernández-Galán E. Clinical usefulness of circulating tumor markers. Clin Chem Lab Med CCLM. 2023;61(5):895–905.
Stojkovic Lalosevic M, Stankovic S, Stojkovic M, Markovic V, Dimitrijevic I, Lalosevic J, et al. Can preoperative CEA and CA19-9 serum concentrations suggest metastatic disease in colorectal cancer patients? Hell J Nucl Med. 2017;20(1):41–5.
Yamashita K, Watanabe M. Clinical significance of tumor markers and an emerging perspective on colorectal cancer. Cancer Sci. 2009;100(2):195–9.
Li L, Zhang L, Tian Y, Zhang T, Duan G, Liu Y, et al. Serum chemokine CXCL7 as a diagnostic biomarker for colorectal cancer. Front Oncol. 2019;9(9):921.
Chan A, Prassas I, Dimitromanolakis A, Brand RE, Serra S, Diamandis EP, et al. Validation of biomarkers that complement CA19.9 in detecting early pancreatic cancer. Clin Cancer Res. 2014;20(22):5787–95.
Felder M, Kapur A, Gonzalez-Bosquet J, Horibata S, Heintz J, Albrecht R, et al. MUC16 (CA125): tumor biomarker to cancer therapy, a work in progress. Mol Cancer. 2014;29(13):129.
Acharya A, Markar SR, Matar M, Ni M, Hanna GB. Use of tumor markers in gastrointestinal cancers: surgeon perceptions and cost-benefit trade-off analysis. Ann Surg Oncol. 2017;24(5):1165–73.
Cao H, Zhu L, Li L, Wang W, Niu X. Serum CA724 has no diagnostic value for gastrointestinal tumors. Clin Exp Med. 2023;23(6):2433–42.
Wang, et al. An integrated giant magnetoimpedance biosensor for detection of biomarker. Biosens Bioelectron. 2014;58:338–44.
US Preventive Services Task Force. Screening for colorectal cancer: us preventive services task force recommendation statement. JAMA. 2021;325(19):1965–77.
Walker HK, Hall WD, Hurst JW, editors. Clinical Methods: The History, Physical, and Laboratory Examinations [Internet]. 3rd ed. Boston: Butterworths; 1990 [Accessed 27 Mar 2024]. Available from: http://www.ncbi.nlm.nih.gov/books/NBK201/
Heys SD, Walker LG, Deehan DJ, Eremin OE. Serum albumin: a prognostic indicator in patients with colorectal cancer. J R Coll Surg Edinb. 1998;43(3):163–8.
Frazzoni L, Laterza L, Marca ML, Zagari RM, Radaelli F, Hassan C, et al. Clinical value of alarm features for colorectal cancer: a meta-analysis. Endoscopy. 2023;55(5):458–68.
Kiebach J, de Vries-ten HJ, van Duijnhoven FJB, Kok DE, van Lanen AS, Kouwenhoven EA, et al. Hematocrit is associated with cancer-related fatigue in colorectal cancer survivors: a longitudinal analysis. Cancer Epidemiol Biomarkers Prev. 2024;33(3):411–8.
Vieth JT, Lane DR. Anemia. Hematol Oncol Clin North Am. 2017;31(6):1045–60.
Boursi B, Mamtani R, Hwang WT, Haynes K, Yang YX. A risk prediction model for sporadic CRC based on routine lab results. Dig Dis Sci. 2016;61(7):2076–86.
Acknowledgements
Thanks to the reviewers and editors for their sincere comments.
Funding
This study was supported by Natural Science Foundation of Liaoning Province of China (2022-MS-320).
Author information
Authors and Affiliations
Contributions
R.X.: original manuscript preparation, methods, and data curation. H.C. and Q.Z.: manuscript review and editing. Additional study supervision: X.L and Z.H.. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Ethical approval
The study was approved by the Ethics Committee of the First Hospital of Dalian Medical University.
Informed consent
For this type of study, formal consent is not required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, R., Chi, H., Zhang, Q. et al. Enhancing the diagnostic accuracy of colorectal cancer through the integration of serum tumor markers and hematological indicators with machine learning algorithms. Clin Transl Oncol (2024). https://doi.org/10.1007/s12094-024-03564-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12094-024-03564-8