Abstract
Purpose
To investigate the role of PRDX2 in esophageal carcinoma (ESCA).
Methods
The expression of PRDX2 was detected in ESCA tissues. And PRDX2 expression in two ESCA cell lines was knocked down. Cell proliferation, metastasis and invasion were detected in these cells.
Results
Here, we found that PRDX2 expression was significantly increased in ESCA tissues and was associated with a poor prognosis in ESCA patients. In addition, PRDX2 expression was significantly associated with pathological grading, infiltration degree and 5-year survival time in ESCA patients. Next, we knocked down PRDX2 expression by PRDX2-shRNA transfection in two ESCA cell lines, Eca-109 and TE-1. Proliferation analysis indicated that in vitro PRDX2 knockdown decreased growth and clone formation of ESCA cells. Scratch and transwell assays indicated that cell migration and invasion were significantly inhibited by PRDX2 knockdown. In addition, PRDX2 knockdown inhibited cell cycle of ESCA cells and down-regulated Cyclin D1-CDK4/6. Moreover, PRDX2 knockdown regulated proteins involved in mitochondrial-dependent apoptosis, including increased Bax and Caspase9/3 and decreased Bcl2. Mechanism investigation indicated that PRDX2 knockdown led to inactivation of Wnt/β-catenin and AKT pathways.
Conclusions
Our data suggest that PRDX2 may function as an oncogene in the development of ESCA via regulating Wnt/β-catenin and AKT pathways. Our study fills a gap in the understanding of the role of PRDX2 in ESCA and provides a potential target for ESCA treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Esophageal carcinoma (ESCA) is a malignant tumor of the digestive system, which ranks the eighth among the most prevalent cancer and the sixth among the causes of cancer deaths worldwide [1]. Esophageal squamous cell carcinoma accounts for 90% of the major histological types of ESCA [1]. The 5-year survival rate of patients with ESCA is only 15–20% due to the difficulty in early diagnosis and recurrence after treatment [2, 3]. Therefore, it is of great importance to identify new diagnostic markers and therapeutic targets.
Reactive oxygen species (ROS) are mainly produced by the highly reduced respiratory chain of mitochondria [4]. ROS can impair DNA, proteins and lipids and cause TNF-alpha-related cell death [4, 5]. Peroxiredoxin 2 (PRDX2) is a cysteine-dependent cellular antioxidant enzyme that can eliminate ROS [6]. PRDX2 is widely distributed in various tissues and cells. PRDX2 has many functions, including protecting intracellular DNA, proteins and lipids from damages induced by ROS [7] and regulating signalling pathways involved in proliferation, differentiation and survival [8]. In addition, it is also reported that PRDX2 is abnormally expressed in a variety of tumor tissues and closely related to cancer development [9,10,11]. However, the expression and effect (oncogenic or tumor-suppressor effect) of PRDX2 depend on the types of tumor. For example, PRDX2 expression is increased in colorectal cancer (CRC) and cervical cancer, and negatively correlates with the prognosis of CRC patients [12, 13]. Knockdown of PRDX2 induces growth inhibition in CRC by inactivating Wnt/β-catenin pathway [14]. PRDX2 protects hepatocellular carcinoma cells SMMC-7721 against oxidative stress [9]. However, PRDX2 plays a tumor suppressor role in some other tumors. For example, PRDX2 is down-regulated in pancreatic cancer, melanoma and gastric cancer and positively correlates with the prognosis of gastric cancer patients [15,16,17]. Restoring the expression of PRDX2 can inhibit colony formation and migration of gastric cancer [17]. Genomic analysis of histone H3 acetylation patterns in acute myelocytic leukemia (AML) showed that PRDX2 was an epigenetically silenced tumor suppressor gene [18]. However, the function of PRDX2 in ESCA has not been studied.
In this study, we first investigated the expression and prognostic value of PRDX2 in ESCA patients. Then, the roles of PRDX2 in ESCA and the regulatory mechanisms were determined. Our study provided new sights into the therapeutic application of PRDX2 in ESCA.
Materials and methods
Tissue chip
The tissue sections were obtained from Shandong Provincial Hospital affiliated to Shandong University from January 2010 to January 2012. This work has been approved by the ethical committees at Shandong Provincial Hospital affiliated to Shandong University. All patients provided written informed consent.
Immunohistochemistry
ESCA and para-carcinoma tissues (normal esophageal mucosa tissues) were cut into 4-μm tissue section on SuperFrost (Braunschweig, Germany). The sections were then dewaxed, rehydrated and antigen retrieved using microwave. Endogenous peroxidase activity was inactivated by 0.5% H2O2. Then, the sections were blocked for 1 h at room temperature and incubated with primary antibodies. 3,3′-diaminobenzidine tetrahydrochloride (DAB) developer was used for slide development. After counterstaining with hematoxylin, dehydration with gradient ethanol, and mounting with neutral resins, tissue sections were observed under a microscope (Leica Microsystems, Germany).
The images were taken in 5 random fields under magnification 40×. The staining intensity was evaluated as follows: “0” (no staining), “1” (weakly positive), “2” (moderately positive), and “3” (strongly positive). The percentage of positively stained cells was scored as: 0 = 0%, 1 = 1–25%, 2 = 26–50%, 3 = 51–75%, and 4 = 76–100%. The evaluation was performed by two independent pathologists. Low expression was identified when calculation of the score was less than 6; otherwise, they were defined as high expression.
Cell culture and transfection
ESCA cell lines, Eca-109 and TE-1, were purchased from the Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences. Cell lines were cultured in RPMI 1640 medium (Sigma, Milan, IT) with 10% FBS (Gibco, Euroclone, Milan, IT) at 37 °C in 5% CO2.
For transfection, ESCA cells were seeded into a 6-well plate. When the confluence reached 80%, cells were transfected with the PRDX2 shRNA (shPRDX2) or shNC plasmid (Santa Cruz Biotechnology, TX, USA) using Lipofectamine 6000 (Beyotime Institute of Biotechnology, Shanghai, China).
qRT-PCR
Total RNA of cells was isolated using TRIzol reagent (Invitrogen) according to the manufacturer’s instructions. The concentration and purity of RNA were measured at 260/280 nm using ultraviolet spectrophotometer. cDNA was synthesized through a RT reaction using the Primer-Script one step RT-PCR kit (TaKaRa). The mRNA expression of PRDX2 was determined with SYBR (Applied Biosystems, USA) on 7500 RealTime PCR System (Applied Biosystems, USA) with β-actin as an internal control. The gene expression data were analyzed using the 2−ΔΔCt method. The experiment was performed in triplicate.
CCK8 assay
After transfection for 24 h, cells were digested and re-seeded into a 96-well plate at 1000 per well. Every 24 h, 10-μl CCK8 solution was added to each well and incubated for 2 h. OD values (450 nm) were measured on microplate reader (BioTek, VT, United States).
Clone formation assay
ESCA cells transfected with shPRDX2 or shNC were seeded in a culture dish and incubated at 37 °C for 14 days. Then, cells were fixed with 1-ml 4% paraformaldehyde for 30 min and stained with crystal violet for 30 min. Cell clones were counted under a microscope (Leica Microsystems, Germany) and pictured.
Scratch assay
After 24 h of transfection and when the confluence reached 100%, a 200-μl pipette tip was used to scratch a straight line on cell monolayer. The cells were washed to remove the detached cells, Then, serum-free medium was added and cells were cultured at 37 °C for 24 h. The pictures were taken at 0 h and 24 h. The wound closure rate was calculated as: (wound closure area/initial wound area) × 100%.
Transwell assay
For invasion, 100 μl of Matrigel (BD Biosciences Discovery Labware, Bedford, MA) was added to the upper chamber of a transwell (BD Biosciences Discovery Labware, Bedford, MA) and incubated at 37 °C for 4–6 h. After the gel was formed, 500 μl of complete medium was added to the lower chamber. ESCA cells transfected with shPRDX2 or shNC were suspended in 500 μl serum-free medium and transferred to transwell inserts. After 24 h of culture, the residual cells on the surface of membrane were removed. Then, the cells were fixed with 4% paraformaldehyde and stained with 0.1% crystal violet. The invaded cells were photographed under a microscope (Leica Microsystems, Germany) and counted in five random view fields.
The migration experiment procedure was similar to that of the invasion experiment except that no Matrigel was added.
Cell cycle analysis
After 48 h of transfection, cells were collected and fixed in 500 μl of 70% ethanol at − 20 °C for two days. After washing with PBS for 3 times, cells were incubated with pre-prepared 500 μl of PI/RNaseA solution for 30–60 min before detection by a flow cytometer (FACS Calibur; Becton Dickinson, San Jose, CA, USA). The results were analyzed by Flowjo software (Tree Star Corp, Ashland, OR).
Western blot
After 48 h of transfection, total proteins were extracted using RIPA lysate (added with 1% protease inhibitor). Protein concentration was determined by BCA method. Protein samples (20 μg) were separated by SDS-PAGE and transferred to a PVDF membrane. After blocking with 5% non-fat milk for 1 h, the membrane was incubated with primary antibodies overnight at 4 °C. After washing, the membrane was incubated with the secondary antibodies for 1 h at room temperature and then visualized using ECL development system. The density of protein bands was determined by Image J software (Software Inquiry, Quebec, Canada). The primary antibodies against Cyclin D1 (Cat# 60,186–1-Ig, 1:10,000),, GAPDH (Cat# 60,004–1-Ig, 1:10,000), Bcl2 (Cat# 12,789–1-AP, 1:1000), Bax (Cat# 60,267–1-Ig, 1:1000), Caspase 9 (Cat# 10,380–1-AP, 1:300) and Caspase 3 (Cat# 19,677–1-AP, 1:10,000), AKT (Cat# 60,203–2-Ig, 1:5000), p-AKT (Cat# 66,444–1-Ig, 1:2000), p70S6K (Cat# 66,638–1-Ig, 1:3000) were purchased from ProteinTech (Rosemont, IL, USA). The primary antibodies against CDK4 (Cat# 12,790, 1:1000), CDK6 (Cat# 13,331, 1:1000), Wnt3a (Cat# 2391, 1:10,000), β-catenin (Cat# 8480, 1:10,000), E-cadherin (Cat# 14,472, 1:10,000), N-cadherin (Cat# 99,377, 1:1000), Vimentin (Cat# 5741, 1:10,000), mTOR (Cat# 2972, 1:1000), Snail1 (Cat# 3879, 1:1000), Snail2 (Cat# 9585, 1:1000), p-mTOR (Cat# 5536, 1:1000) were purchased from Cell Signaling Technology (Danvers, MA, USA). The secondary antibodies were purchased from Pierce Biotechnology (Rockford, IL).
Statistical analysis
GraphPad Prism 7 software was used for data analysis. All experiments were performed for 3 times. Difference between two groups was evaluated by Student’s t-test. P < 0.05 represented significant difference.
Results
PRDX2 is remarkably up-regulated in ESCA tissues and may be used as a prognostic marker in ESCA
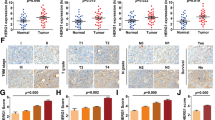
We first detected the endogenous level of PRDX2 in ESCA tissues and para-carcinoma tissues. As shown in Fig. 1a, PRDX2 was highly expressed in ESCA tissues compared with para-carcinoma tissues. Statistically, the high expression rate of PRDX2 in ESCA tissues was significantly higher than that of normal para-carcinoma tissues (66/74 vs 20/68, P < 0.05, Table 1). In addition, Fig. 1b showed that ESCA patients with high PDRX2 expression had a worse prognosis than those with low PRDX expression. The correlation between PRDX2 expression and clinical pathological features of ESCA patients was analyzed and listed in Table 2. The mRNA expression level of PRDX2 was significantly associated with pathological grading, infiltration degree and 5-year survival time (P < 0.05). These data suggest that PRDX2 might be involved in ESCA progression and used as a prognostic marker in gastric cancer.
PRDX2 was up-regulated in ESCA tissues and represented a poor prognosis. a Endogenous levels of prdx2 in ESCA tissues and para-carcinoma tissues was detected by IHC analysis. b The correlation between PRDX2 expression and patients’ survival. The mRNA expression of PRDX2 in c Eca-109 and d TE-1 cells was detected using qRT-PCR after shPRDX2 transfection. All experiments were performed at 3 times. *P < 0.05
PRDX2 knockdown inhibits cell proliferation of ESCA
To investigate the function of PRDX2 in ESCA, we knocked down PRDX2 expression by shRNA-PRDX2. The PRDX2 expression was significantly suppressed by shPRDX2 compared with NC group in Eca-109 and TE-1 cells (Fig. 1c, d). The sh1-PRDX2 for Eca-109 and sh2-PRDX2 for TE-1 cells achieved the optimal interference efficiency and were, thus, used for the following experiments.
We further detected cell proliferation of ESCA cells using CCK8 assay. Cell viability of ESCA cells were significantly inhibited by shPRDX2 at 72 h in both in Eca-109 and TE-1 cells (Fig. 2a, b) (P < 0.05). Then, colony formation ability was detected. As shown in Fig. 2c, d, Eca-109 and TE-1 cells transfected with shPRDX2 formed much less clones compared with NC group. Quantification analysis further showed that clone number was significantly decreased in shPRDX2 group, compared to NC group (P < 0.05). The above data indicate that PRDX2 knockdown induces growth inhibition in ESCA cells.
PRDX2 knockdown inhibits cell mobility of ESCA
To investigate whether PRDX2 plays a role in cell movements of ESCA, we performed a scratch assay. As shown in Fig. 3a, b, photographs of shPRDX2 group indicated a lower migrating rate into the cell-free region compared to NC group in Eca-109 and TE-1 cells. Quantification analysis indicated that the percentage of wound closure was decreased from 30.17 to 10.57% in Eca-109 cells and 35.77 to 17.9% in TE-1 cells (P < 0.05). What is more, the role of PRDX2 in migration and invasion of ESCA cells was detected through transwell assay. As shown in Fig. 4, PRDX2 knockdown significantly inhibited migration and invasion of both Eca-109 and TE-1 cells (P < 0.05). Taken together, silence of PRDX2 led to a significant down-regulation of cell migration and invasion in ESCA cells.
PRDX2 knockdown induces cell cycle arrest and activates mitochondrial-dependent apoptosis.
To investigate whether PRDX2 induces growth inhibition of ESCA through regulating cell cycle, we performed a flow cytometry. As shown in Fig. 5, cell cycle of Eca-109 and TE-1 was arrested at S phase by shPRDX2 compared with NC group. To determine the regulating mechanisms of PRDX2 in ESCA proliferation and survival, we detected the key signaling proteins by Western blot. As shown in Fig. 6a, PRDX2 knockdown significantly down-regulated Cyclin D1 and CDK4/6 in Eca-109 and TE-1 cells. Taken together, PRDX2 silencing induced cell cycle arrest by regulating the expression of cell cycle check point proteins.
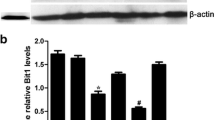
Knockdown of PRDX2 down-regulated Cyclin D1-CDK4/6 and activated mitochondrial-dependent pathway in ESCA cells. a Western blot analysis of cell cycle related proteins, Cyclin D1 and CDK4/6; b western blot analysis of mitochondrial-dependent pathway including Bcl2, Bax, Caspase9 and Caspase3. The density of protein bands was determined by Image J software. All experiments were performed at 3 times. *P < 0.05
Mitochondrial-dependent apoptosis mainly included Bcl2, Bax, Caspase9 and Caspase3. The expressions of these proteins were measured with Western blot. Figure 6b showed that pro-apoptotic protein Bax, Caspase9 and Caspase3 were significantly up-regulated and anti-apoptotic protein Bcl2 was significantly down-regulated by knockdown of PRDX2 in both Eca-109 and TE-1 cells. The results indicate that proteins involved in mitochondrial-dependent apoptosis are activated when knocking down PRDX2, suggesting the increase in apoptosis.
PRDX2 knockdown induced inhibition of ESCA is mediated by inactivation of Wnt/β-catenin and AKT pathways
Wnt/β-catenin pathway is involved in tumor epithelial–mesenchymal transition (EMT) and metastasis [19, 20]. Based on the above results that PRDX2 knockdown inhibited cell migration and invasion, we further examined whether this effect is mediated by Wnt/β-catenin pathway. As shown in Fig. 7, PRDX2 knockdown significantly decreased Wnt3a, β-catenin, N-cadherin, Vimentin, Snail1 and Snail2; while increased E-cadherin in Eca-109 and TE-1 cells, indicating an inactivated status of Wnt/β-catenin pathway.
Knockdown of PRDX2 inactivated Wnt/β-catenin pathway in ESCA cells. a Members of Wnt/β-catenin pathway including Wnt3a, β-catenin, E-cadherin, N-cadherin, Vimentin, Snail1 and Snail2 were analyzed by western blot; b, c The density of protein bands was determined by Image J software. All experiments were performed at 3 times. *P < 0.05
The protein kinase B (AKT) pathway is involved in many pathological processes of tumors, including proliferation, survival, cell cycle, etc. [21, 22]. The analysis of AKT pathway indicated that PRDX2 knockdown significantly down-regulated phosphorylation level of AKT and mTOR and decreased the expression of p70S6K (Fig. 8). The results showed that AKT pathway was inactivated when knocking down PRDX2. Taken together, PRDX2 knockdown may inhibit proliferation and metastasis of esophageal cancer cells by inactivating Wnt/β-catenin and AKT pathways.
Discussion
ESCA, with high aggressiveness and lethality, continues to be one of the notable global issues that threaten human health. However, there are few biomarkers for the early diagnosis and clinical treatment of ESCA. Here, for the first time, we identified that the antioxidant enzyme PRDX2 promoted the progression of in ESCA cell lines (Eca-109 and TE-1).
Previous studies have found that PRDX2 is up-regulated in many tumor types such as cervical cancer and nasopharyngeal carcinoma [20, 21], and regulates many biological features of tumors. Shiota et al. found that PRDX2 promoted cell cycle progression in prostate cancer by regulating androgen receptor (AR) activity [23]. PRDX2 knockdown induced growth inhibition in breast cancer metastatic cells by regulating ROS level [22]. Zhang et al. reported that PRDX2 promoted angiopoiesis in colon cancer through activation of VEGFR2 [24]. In addition, PRDX2 was also a target of miR-122a, which could inhibit proliferation and promote apoptosis in hepatocellular carcinoma (HCC) [25]. These results suggest that PRDX2 is a crucial functional protein involved in tumor progression. Here, we found that PRDX2 expression was significantly increased in ESCA tissues and predicted a bad prognosis in ESCA patients. PRDX2 expression also correlated with pathological grading, infiltration degree and 5-year survival time in ESCA patients. We knocked down PRDX2 in ESSC cell lines, Eca-109 and TE1, using RNA interference and found that cell proliferation and clone formation ability were significantly inhibited. And, PRDX2 knockdown significantly inhibited migration and invasion of both Eca-109 and TE-1 cells. In addition, PRDX2 knockdown also induced cell cycle arrest by decreasing the expression of Cyclin D1-CDK4/6. Cyclin D1 forms a complex with CDK4/6 to regulate G1/S transition in cell cycle [26]. Assessment of apoptosis-related proteins indicated that PRDX2 knockdown decreased Bcl2/Bax ratio and increased Caspase 9 and Caspase 3. During apoptosis, Bax is transferred to the mitochondrial outer membrane and forms membrane channels that mediate release of cytochrome C, which in turn causes apoptosis through Caspae9/3 cascade [27]. Bcl2 inhibits apoptosis mainly through competitive combination with Bax [27]. The above data suggest that PRDX2 functions as an oncogene in the progression of ESCA.
We further investigated relevant mechanisms underlying the oncogenic role of PRDX2 in ESCA. Previous studies have shown that PRDX2 is involved in regulating various signaling pathways in tumors, including Wnt, Hedgehog, PI3K/AKT, and AR signaling pathways [14, 28]. These signaling pathways mediate cell proliferation, migration, maintenance of stem cell phenotypes and other physiological functions of tumor cells. Here, the key node proteins in Wnt/β-catenin and AKT signaling pathways were analyzed by western blot. It is widely known that Wnt/β-catenin and AKT pathways regulate numerous processes in tumor development, including stimulating tumor cell proliferation, EMT and apoptosis [29,30,31]. Here, we found that PRDX2 knockdown down-regulated Wnt3a, β-catenin, N-cadherin, Vimentin, Snail1 and Snail2 but increased E-cadherin expression, indicating an inactivated status of Wnt/β-catenin pathway. Assessment of AKT pathway indicated that p-AKT, p-mTOR and p70S6K were significantly down-regulated by knockdown of PRDX2. These data suggest that Wnt/β-catenin and AKT pathways are significantly suppressed by silencing of PRDX2.
In conclusion, PRDX2 knockdown by RNA interference inhibited proliferation and mobility of ESCA cells by down-regulating Wnt/β-catenin and AKT signaling pathways. Our study will provide a promising target for clinical treatment of ESCA.
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65(2):87–108.
Kim T, Grobmyer SR, Smith R, Ben-David K, Ang D, Vogel SB, et al. Esophageal cancer–the five year survivors. J Surg Oncol. 2011;103(2):179–83.
Cavallin F, Scarpa M, Cagol M, Alfieri R, Ruol A, ChiarionSileni V, et al. Time to diagnosis in esophageal cancer: a cohort study. Acta Oncol. 2018;57(9):1179–84.
Bouzigues CI, Nguyen TL, Ramodiharilafy R, Claeson A, Tharaux PL, Alexandrou A. Regulation of the ROS response dynamics and organization to PDGF motile stimuli revealed by single nanoparticle imaging. Chem Biol. 2014;21(5):647–56.
Feng J, Fu Z, Guo J, Lu W, Wen K, Chen W, et al. Overexpression of peroxiredoxin 2 inhibits TGF-beta 1-induced epithelial-mesenchymal transition and cell migration in colorectal cancer. Molecular medicine reports. 2014;10(2):867–73.
Brown JD, Day AM, Taylor SR, Tomalin LE, Morgan BA, Veal EA. A peroxiredoxin promotes H2O2 signaling and oxidative stress resistance by oxidizing a thioredoxin family protein. Cell Rep. 2013;5(5):1425–35.
Yim MB, Chae HZ, Rhee SG, Chock PB, Stadtman ER. On the protective mechanism of the thiol-specific antioxidant enzyme against the oxidative damage of biomacromolecules. J Biol Chem. 1994;269(3):1621–6.
Fang J, Nakamura T, Cho D-H, Gu Z, Lipton SA. S-nitrosylation of peroxiredoxin 2 promotes oxidative stress-induced neuronal cell death in Parkinson's disease. Proc Natl Acad Sci USA. 2007;104(47):18742–7.
Zhou S, Han Q, Wang R, Li X, Wang Q, Wang H, et al. PRDX2 protects hepatocellular carcinoma SMMC-7721 cells from oxidative stress. Oncol Lett. 2016;12(3):2217–21.
Kinnula VL, Lehtonen S, Sormunen R, Kaarteenaho-Wiik R, Kang SW, Rhee SG, et al. Overexpression of peroxiredoxins I, II, III, V, and VI in malignant mesothelioma. J Pathol. 2002;196(3):316–23.
Lu W, Fu Z, Wang H, Feng J, Wei J, Guo J. Peroxiredoxin 2 is upregulated in colorectal cancer and contributes to colorectal cancer cells' survival by protecting cells from oxidative stress. Mol Cellular biochemistry. 2014;387(1–2):261–70.
Peng L, Wang R, Shang J, Xiong Y, Fu Z. Peroxiredoxin 2 is associated with colorectal cancer progression and poor survival of patients. Oncotarget. 2017;8(9):15057–700.
Kontostathi G, Zoidakis J, Makridakis M, Lygirou V, Mermelekas G, Papadopoulos T, et al. Cervical cancer cell line secretome highlights the roles of transforming growth factor-beta-induced protein ig-h3, peroxiredoxin-2, and NRF2 on cervical carcinogenesis. Biomed Res Int. 2017;2017:4180703.
Lu W, Fu Z, Wang H, Feng J, Wei J, Guo J. Peroxiredoxin 2 knockdown by RNA interference inhibits the growth of colorectal cancer cells by downregulating Wnt/beta-catenin signaling. Cancer Lett. 2014;343(2):190–9.
Cui L, Li F, Zhao Q, Li Z. Screening and verification of differentially expressed proteins from pancreatic cancer tissue. Chin J Chem. 2010;28:884–90.
Furuta J, Nobeyama Y, Umebayashi Y, Otsuka F, Kikuchi K, Ushijima T. Silencing of peroxiredoxin 2 and aberrant methylation of 33 CpG islands in putative promoter regions in human malignant melanomas. Cancer Res. 2006;66(12):6080–6.
Hong SH, Min C, Jun Y, Lee DJ, Kim SH, Park JH, et al. Silencing of peroxiredoxin II by promoter methylation is necessary for the survival and migration of gastric cancer cells. Exp Mol Med. 2018;50(2):e443.
Agrawal-Singh S, Isken F, Agelopoulos K, Klein H-U, Thoennissen NH, Koehler G, et al. Genome-wide analysis of histone H3 acetylation patterns in AML identifies PRDX2 as an epigenetically silenced tumor suppressor gene. Blood. 2012;119(10):2346–57.
Carnero A, Blancoaparicio C, Renner O, Link W, Leal JF. The PTEN/PI3K/AKT signalling pathway in cancer, therapeutic implications. Curr Cancer Drug Targets. 2008;8(3):187.
Lin L-H, Xu Y-W, Huang L-S, Hong C-Q, Zhai T-T, Liao L-D, et al. Serum proteomic-based analysis identifying autoantibodies against PRDX2 and PRDX3 as potential diagnostic biomarkers in nasopharyngeal carcinoma. Clin Proteomics. 2017;14:6.
Nikoshkov A, Broliden K, Attarha S, Sviatoha V, Hellstrom A-C, Mints M, et al. Expression pattern of the PRDX25 RAB1A, RAB1B, RAB5A and RAB25 genes in normal and cancer cervical tissues. Int J Oncol. 2015;46(1):107–12.
Stresing V, Baltziskueta E, Rubio N, Blanco J, Arriba MC, Valls J, et al. Peroxiredoxin 2 specifically regulates the oxidative and metabolic stress response of human metastatic breast cancer cells in lungs. Oncogene. 2013;32(6):724–35.
Shiota M, Izumi H, Miyamoto N, Onitsuka T, Kashiwagi E, Kidani A, et al. Ets regulates peroxiredoxin1 and 5 expressions through their interaction with the high-mobility group protein B1. Cancer Sci. 2008;99(10):1950–9.
Zhang S, Fu Z, Wei J, Guo J, Liu M, Du K. Peroxiredoxin 2 is involved in vasculogenic mimicry formation by targeting VEGFR2 activation in colorectal cancer. Med Oncol. 2015;32(1):414.
Diao S, Zhang J-F, Wang H, He M-I, Lin MC, Chen Y, et al. Proteomic identification of microRNA-122a target proteins in hepatocellular carcinoma. Proteomics. 2010;10(20):3723–31.
Jiang Q-Q, Liu B, Yuan T. MicroRNA-16 inhibits bladder cancer proliferation by targeting cyclin D1. Asian Pac J Cancer Prev. 2013;14(7):4127–30.
Guidolin D, Agnati LF, Tortorella C, Marcoli M, Maura G, Albertin G, et al. Neuroglobin as a regulator of mitochondrial-dependent apoptosis: a bioinformatics analysis. Int J Mol Med. 2014;33(1):111–6.
Wang R, Wei J, Zhang S, Wu X, Guo J, Liu M, et al. Peroxiredoxin 2 is essential for maintaining cancer stem cell-like phenotype through activation of Hedgehog signaling pathway in colon cancer. Oncotarget. 2016;7(52):86816–28.
Fatima I, El-Ayachi I, Taotao L, Lillo MA, Krutilina RI, Seagroves TN, et al. The natural compound Jatrophone interferes with Wnt/β-catenin signaling and inhibits proliferation and EMT in human triple-negative breast cancer. PLoS ONE. 2017;12(12):e0189864.
Martini M, De Santis MC, Braccini L, Gulluni F, Hirsch E. PI3K/AKT signaling pathway and cancer: an updated review. Ann Med. 2014;46(6):372–83.
He R, Zhang FH, Shen N. LncRNA FEZF1-AS1 enhances epithelial–mesenchymal transition (EMT) through suppressing E-cadherin and regulating WNT pathway in non-small cell lung cancer (NSCLC). Biomed Pharmacother. 2017;95:331–8.
Acknowledgements
This study was funded by National Natural Science Foundation of China (Foundation Number: 81502508 and 81500496).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by ALF, XH, XM, ZC, QL, WS, HD, JZ and ZY. The first draft of the manuscript was written by ALF and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This work has been approved by the ethical committees at Shandong Provincial Hospital affiliated to Shandong University.
Informed consent
All patients provided written informed consent.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Feng, A.L., Han, X., Meng, X. et al. PRDX2 plays an oncogenic role in esophageal squamous cell carcinoma via Wnt/β-catenin and AKT pathways. Clin Transl Oncol 22, 1838–1848 (2020). https://doi.org/10.1007/s12094-020-02323-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-020-02323-9