Abstract
In the present study, a total of 102 samples were collected from chickens of different flocks, died due to suspected colibacillosis. Bacteriological and PCR methods were applied to detect avian pathogenic Escherichia coli (APEC). Phenotypic antimicrobial resistance (AMR) was determined by disk diffusion method. Extended spectrum beta lactamases (ESBL) detection was carried out via PCR by targeting blaTEM, blaSHV, blaOXA, and blaCTX-M groups 1, 2, and 9. Genes of eight virulence factors and class I integrons were also detected by PCR using gene specific primers. Culture, microscopic, biochemical tests and PCR recognised 69/102 (67.64%) samples as E. coli. Phenotypic AST revealed higher resistance against fluoroquinolone antibiotics, i.e., enrofloxacin (72.46%), levofloxacin (69.56%) & ciprofloxacin (66.66%), followed by amoxyclav (63.77%) and tetracycline (59.42%). Six isolates were found as pan-drug-resistant E. coli. A total of 48 (69.56%) and 7 (10.14%) isolates were positive for the presence of blaTEM and blaCTX-M-G9 genes, respectively, whereas 2 (2.90%) isolates each were found positive for blaSHV, blaOXA, and blaCTX-M-G1 genes. Among APEC associated virulence genes, iss (79.71%) was the most predominant, followed by tsh (50.72%), ast (30.43%), cvaf (26.08%), pap (23.18%), vat (8.69%) and stx-1 (1.44%). Thirty-two isolates harboured class I integrons, either with or without ESBL genes. Conclusively, the isolates under study showed pan and multiple-drug resistance, specifically against fluoroquinolone drugs. ESBL production was mediated principally through blaTEM and blaCTX-M-G9. Multiple virulence factors, toxins, and carriage & spread factor render these as zoonotically potential pathogens for humans.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pathogenic Escherichia coli (E. coli) produces intestinal and extraintestinal infections in the target hosts [1]. Among these, E. coli which causes extra intestinal infections in birds, especially in domestic poultry, is known as avian pathogenic E. coli (APEC). APEC can invade multiple organs, causing peritonitis, perihepatitis, air sacculitis, omphalitis, pericarditis etc. in chickens and the infection is collectively termed colibacillosis’ [2]. These E. coli strains primarily colonise the respiratory and intestinal tracts of birds. The colonisation, tissue invasion, and systematic survival of APEC are determined by several virulence factors. Their genes are encoded by virulence-associated genes (VAGs), viz., iutA, iss, papC, iucD, tsh, irp-2, ompT, hlyF, iron, cva/cvi, and astA etc. [3]. All the genes are not found in a given strain, but, according to the genetic criteria, the pathogenicity of the APEC strain is determined by the presence of combination of virulence genes, which provides them with an advantage of systematic survival [4] over potentially pathogenic E. coli strains residing in the intestinal tract [5].
The principal mode of treatment for APEC infection is the administration of antimicrobials. Despite of government guidelines, antimicrobials are commonly used as feed supplements in the poultry industry. The overuse of antimicrobials is a major driving force for the augmentation of AMR [6].
The emergence of AMR is a natural phenomenon in microorganisms, shared by the use of common antimicrobial agents in both humans and animals. Among the several forms of AMR, extended spectrum β-lactamases (ESBLs) production is a major concern in Gram negative bacteria, specifically in E. coli. ESBLs are enzymes that mediate resistance to extended-spectrum (third generation) cephalosporins (e.g., ceftazidime, cefotaxime, and ceftriaxone) and monobactams (e.g., aztreonam) but do not affect cephamycins (e.g., cefoxitin and cefotetan) or carbapenems (e.g., meropenem or imipenem). There are more than 200 different types of ESBL enzymes, of which TEM, SHV and CTX-M types are more prevalent among E. coli [7, 8]. Ubiquitous presence and wide host range made this bacterial species a suitable candidate organism for the spread of antimicrobial resistance. The drug resistance profile can be transferred from poultry to humans and poses a potential risk to human health. Shared homology in virulence genes in APEC and other extraintestinal E. coli (ExIEC) poses risk of the development of severe human diseases such as haemorrhagic colitis and haemolytic uremic syndrome [9]. Both virulence genes and antibiotic resistance genes of these pathogens may be transmitted to humans through several genetic mechanisms, including mobile genetic elements class I integrons [10]. This may increase the severity of the disease and complicate the therapeutic strategy [11].
Keeping the importance of APEC in mind, the present study was carried out to see the AMR status of E. coli in chickens reared for consumption in Gujarat, India. In addition to that, molecular detection of antimicrobial resistance genes, virulence genes and class I integrons (as carriage and spreading factors) was also investigated as zoonotic spread potential.
Material and Methods
Ethical Approval
As the tissue samples were collected from postmortem of dead chickens, ethical approval for the work was not required.
Study Location and Sample Collection
The present study was conducted in Banaskantha (24.085560° N, 72.144234° E) district, Gujarat, India, during 2019–21. Necropsy was conducted on the chickens, which died due to suspected E. coli infection. Clinical picture and necropsy findings, viz., exhibiting pericarditis, perihepatitis, egg peritonitis, omphalitis, tracheitis, intestinal haemorrhage, etc. were taken into account. A total of 102 tissue samples were collected aseptically in MacConkey broth and transported to laboratory under standard conditions. The details of the samples are shown in Table 1.
Isolation and Identification of Avian Pathogenic E. coli (APEC) Strains
Primary isolation and identification of E. coli were carried out as per Edwards & Ewing [12]. Briefly, samples were inoculated on MacConkey agar medium and incubated aerobically at 37 °C overnight. Lactose-fermenting colonies were purified and sub-cultured on Eosin methylene blue (EMB) agar to detect metallic sheen. Biochemical tests, viz., oxidase, catalase, oxidation-fermentation, indole test, methyl red test, Vogus Proskauer test, citrate utilisation test, and sugar utilisation pattern on TSI slants, were employed for the identification of E. coli [12] (Table 1). Genomic DNA was extracted from freshly grown cultures by the boiling method. In brief, 3 to 5 bacterial colonies were picked up and suspended in 200 µl of deionized water, followed by boiling at 95 °C for 15 min in a thermal cycler and centrifugation at 4000 rpm for 5 min. The supernatant was then used as the DNA template for further molecular characterization. The isolates were confirmed by amplification of the E. coli specific genomic region of 16srRNA using primers described by Lee et al. [13] (Supplementary Table 1).
Phenotypic Antibiotic Susceptibility Testing
Mueller–Hinton agar (Hi Media, India) was used to perform antibiotic susceptibility test (AST) of E. coli isolates. A total of 12 different types of antibiotic discs (Hi Media, India) (Table 2), which represent different antibiotic classes and are commonly used in the poultry sector, were tested following the standard Kirby-Bauer disk diffusion method, 1966 and interpretation of results was made according to the recommendations of the CLSI (CLSI VET -08, 2018)/ manufacturer’s guidelines [14].
Molecular Detection of Antimicrobial Resistance (ESBL Genes) and Virulence Genes
Genotypic antimicrobial resistance of E. coli isolates was carried out by amplifying β-Lactamase producing genes (blaTEM, blaSHV, blaOXA, and blaCTX-M groups 1, 2, and 9) by PCR using various primers listed in supplementary table 1 [15, 16]. Similarly, APEC-associated virulence genes like pap, ast, tsh, iss, vat, cva [17], stx1 and stx2 genes [18] were detected by PCR using respective primers, including the int1 gene, which was detected as per Kar et al. [11] (supplementary table 2).
The PCR reaction mixture was prepared in a total reaction volume of 25 μl for each gene, containing 12.5 μl of the 2 × PCR master mixture, 10 pmol of forward and reverse primers, 2 μl template DNA and nuclease free water to make a volume of 25 μl. The cardinal PCR temperatures and cyclic conditions for each PCR have been depicted in supplementary table 3. The amplified products were electrophoresed in a 1.5%–2% agarose gel stained with ethidium bromide (0.5 μg/ml), and the image was documented by the gel documentation system.
Results and Discussion
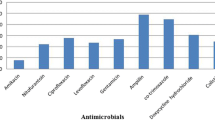
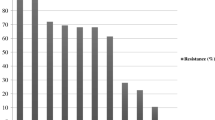
On necropsy, egg peritonitis, perihepatitis, pericarditis, tracheitis, air sacculitis, hepatomegaly, and hemorrhagic intestine were the visible findings. On the basis of microscopic, cultural, and biochemical tests, 69 (67.64%) isolates were identified as E. coli. Further, all 69 isolates were confirmed by PCR employing universal eubacterial primers (SRV3) targeting the 16S rRNA gene. For the in vitro antibiotic susceptibility test, out of 69 isolates, a very high degree of resistance was shown against fluoroquinolone antibiotics, i.e., enrofloxacin (72.46%), levofloxacin (69.56%), & ciprofloxacin (66.66%), followed by amoxyclav (63.77%) and tetracycline (59.42%). whereas lower resistance to ampicillin + sulbactam (34.78%) and ceftriaxone + sulbactam (36.23%) was observed (Fig. 1). Out of them, 6 (8.69%), 5 (7.25%) and 3 (4.34%) were found resistant to 8, 9, and 10 drugs, respectively. Six isolates were found sensitive to none of the tested antibiotics and showed resistance to 11–12 antibiotics, therefore being declared pan-drug resistant isolates (Table 3).
In addition to this, these isolates were subjected to detection of ESBL enzymes by targeting blaTEM, blaSHV, blaOXA, and blaCTX-M group 1, 2, and 9 genes using target gene specific primer pairs (Fig. 2). It revealed that a total of 48 (69.56%) and 7 (10.14%) isolates were positive for the presence of blaTEM and blaCTX-M-G9 genes, respectively, whereas only 2 (2.90%) isolates were found positive for blaSHV, blaOXA, and blaCTX-M-G1 genes out of 69 isolates, and none of the isolates were found positive for blaCTX-M-G2 (Fig. 2; Table 3).
Agarose gel electrophoresis of A E. coli genus specific PCR showing Lane L: Gel pilot mid range ladder (Qiagen); Lane 1,2,3,4: Positive samples with amplicon of the size 203 bp B BlaTEM gene for genus specific PCR showing Lane L: 100 bp plus ladder (Qiagen); Lane 3,4,5,7: Positive samples with amplicon of the size 800 bp; Lane 1,2,6: negative sample C BlaSHV gene for genus specific PCR showing Lane L: 100 bp plus ladder (Qiagen); Lane 1,2,3: Positive samples with amplicon of the size 713 bp D CTX M-9 gene for genus specific PCR showing Lane L: 100 bp ladder (Qiagen); Lane 1,2,3,4: Positive samples with amplicon of the size 561 bp. In all the plates lane P is positive control and lane N is non template control
The frequency of various virulence genes in 69 E. coli isolates revealed that 55 (79.71%), 35 (50.72%), 21 (30.43%), 18 (26.08%), 16 (23.18%), 6 (8.69%), and 1 (1.38%) isolates were found positive for iss, tsh, ast, cvaf, pap, vat, and stx1 genes, respectively (Figs. 3 and 4). But, 8 isolated were negative for any virulence gene. The different combinations of virulence genes have been shown in Table 4. Simultaneous detection of iss, tsh, cva (N = 8); iss, tsh, pap (N = 8); iss, ast, tsh (N = 7) showed that these were the most prevalent virulence factor combinations in these isolates. Out of six virulence factors tested, the gene iss was found in the highest number of isolates (N = 54), followed by tsh (N = 35). Whereas ast, pap, and cva genes were detected in 19, 16, and 12 isolates, respectively. Genes of cva, vat (N = 06 each), and stx1 were the least prevalent genes among the isolates. The total of 9, 27, 10, and 15 isolates showed positive for 4, 3, 2, and 1 virulence gene(s), respectively. A total of 32 isolates were detected with the int1 gene (Fig. 4), out of which five were associated with non ESBL producers, but in the others, it had the co-occurrence of one or more ESBL genes. Mainly, the blaTEM, and blaCTX-M g9 ESBL genes were found to be associated with the int1 gene (Table 3).
Agarose gel electrophoresis of A pap gene for genus specific PCR showing Lane L: Gel pilot 100 bp plus ladder (Qiagen); Lane 3,8,9,10: Positive samples with amplicon of the size 501 bp B iss gene for genus specific PCR showing Lane L: Gel pilot 100 bp plus ladder (Qiagen); Lane 1,3,4,5,6,7,8,9,10,11,12,13,15: Positive samples with amplicon of the size 323 bp C tsh gene for genus specific PCR showing Lane L: Gel pilot 100 bp plus ladder (Qiagen); Lane 6: Positive samples with amplicon of the size 824 bp D ast gene for genus specific PCR showing Lane L: Gel pilot 50 bp plus ladder (Qiagen); Lane 1,2,3,4: Positive samples with amplicon of the size 116 bp E cvaf gene for genus specific PCR showing Lane L: Gel pilot mid-range ladder (Qiagen); Lane 1,2,3,4,5: Positive samples with amplicon of the size 1181 bp F vat gene for genus specific PCR showing Lane L: Gel pilot 100 bp plus ladder (Qiagen); Lane 2,3,4: Positive samples with amplicon of the size 981 bp. In all the plates lane P is positive control and lane N is non template control
Agarose gel electrophoresis of (A) int1 gene PCR showing Lane L: Gel pilot 100 bp plus ladder (Qiagen); Lane 1: Positive samples with amplicon of the size 769 bp; Lane 2 negative sample; lane P is positive control and lane N is non template control. (B) stxI gene PCR showing Lane L: Gel pilot 100 bp plus ladder (Qiagen); Lane 1: Positive samples with amplicon of the size 302 bp; Lane 2–6 negative sample; lane N is non template control; positive control was not used in this PCR. (C)
In the recent past, there has been increased concern about one health approach to tackle anti-microbial resistance [6], and the poultry sector constitutes an important component of the human-animal interface that can transmit antibiotic-resistant bacteria in an interdependent manner. The presence of resistance against newer classes of antibiotics has been increasingly documented in E. coli isolated from diseased [19] and healthy [20] poultry birds from India and neighbouring countries [4, 7]. The present study was carried out in the dry and hot region of Gujarat State, India, and the prevalence of E. coli was recorded at 67.64% (69/102). The prevalence rate approximated the results obtained in Jordan [21] and India [22]. Though almost 100% positive samples have also been reported for E. coli [4, 19], all these reports indicate that this is among the most prevalent pathogens among young poultry birds, and veterinarians are bound to use antibiotics as an obvious choice of treatment. The emergence of AMR is inevitable. AMR is a dynamic phenomenon and therefore, patterns of AMR must be constantly monitored. In recent reports from several parts of the world, like Nepal [4], Bangladesh [7, 23], and Italy [24], either ampicillin or tetracycline were found to be the most resistant drugs against APEC. But our work revealed that enrofloxacin was the most resistant antibiotic, and the fluoroquinolone group was the most resistant group of antibiotics. Similar findings have been revealed by several workers, and 81.94–87.0% fluroquinolone resistance was reported [21, 25,26,27], and one report stated as high as 97% of ciprofloxacin resistance [28]. This change in resistance pattern is conspicuous and clearly indicates widespread use of this antibiotic class for prophylaxis and treatment in the area. Further, the frequency of other resistant classes of antibiotics is also indicative of the proportion of their use in the poultry sector. Earlier, there were reports of colistin resistant [28] and multidrug resistant avian pathogenic E. coli, which were isolated from broiler chickens in Bangladesh [29], Iran [27], Qatar [28], and even India [19, 20]. Most importantly, the present quantum of resistant strains against last resort antibiotics like colistin and pan drug resistance is of great concern to human health. Though there are guidelines for restricted use of last resort drugs from different agencies of the government, but an official ban has been notified against colistin only [30].
When the genes responsible for ESBL production were tested, the occurrence of blaTEM and blaCTXM as the most significant ESBL synthesizing genes was found to be consistent with the findings of other workers [24, 26, 31]. It has been reported that blaCTX-M has become the dominant ESBL gene group [32], but present findings denote that blaTEM is still the most widespread ESBL genes group. The frequency of blaSHV was in line with findings of several researchers among poultry APEC [8, 21, 24] and APEC isolated from migratory birds [7]. Detection of blaOXA seems a rare phenomenon and was reported previously by only a few workers [26], who also detected blaOXA at a very low frequency. Genes viz., blaTEM and blaCTXM-9 were found to be associated with the pan drug resistant and multi drug resistant strains, except for one isolate (Number 26), where none of the tested genes were detected. The major reason behind this is the presence of at least 12 ESBL groups among Gram negative bacteria [32], which may produce at least 200 different types of ESBL enzymes.
The detection of iss as most prevalent virulence factor [19] and tsh as another significant factor is supported by earlier studies [2, 8, 21, 24]. Corroborating to present findings, pap was found among a significant number of APEC [4, 23]. Whereas in other studies, it was found in a lesser number of isolates [24] or in one isolate only [21]. Likewise, ast and vat were found to be less prevalent factors, was corroborated with earlier findings [21, 24]. As indicated in results, the total of 9, 27, 10, and 15 isolates were found positive for 4, 3, 2, and 1 virulence gene(s), respectively in various combinations. As for ESBL genes, there are several varieties of virulence genes but the preponderance of adhesins (tsh and pap) and protectins (iss) makes them essential genes for APEC [2, 3]. Though it has been mentioned that detection of five genes [3, 4] denotes a strain as APEC, the occurrence of these factors in different combinations makes it equivocal to denote any strain conclusively as APEC and differentiate it with faecal E. coli [5]. This fact was further substantiated by Tomaz et al. [26], who also found a non-significant association between virulence genes, APEC phenotyping, and AST pattern. But the presence of the stx1 gene in one isolate (isolate no. -30) and the int1 gene in 46.37% of isolates denotes the increasing zoonotic potential of the APEC. The stx1 gene is the Shiga toxin gene, which is frequently isolated in cattle E. coli [1, 9] within pathotype Shiga toxigenic E. coli, and only seldom reported from poultry birds in India [33] with diarrhoea. Antibiotic resistance, including ESBL production, may be carried and spread by integrons, which are important genetic elements. In our work, class I integrons were detected mostly along with ESBL gene, but their association with strains carrying the lesser number of virulence factors and AMR genes is an important finding which correspond to the previous report that these elements might also be associated with non-pathogenic E. coli and antibiotic non exposed strains [34] and considered among the potent factors of antibiotic spread. Apart from this work, class I integrons was detected in the veterinary sector in India by few workers only [11] and their association with human uropathogenic E. coli [10] underscore the importance of this study.
Conclusions
Avian pathogenic E. coli is predominantly associated as a causative agent of avian colibacillosis; hence, antibiotics are used in the poultry industry as therapeutic and preventive measures, which leads to the development of different forms of AMR. The current status of antibiotic usage shows higher resistance to fluoroquinolone antibiotics. The report also showed an abundance of ESBL producing APEC, which arose principally through the possession of blaTEM and blaCTX-M 9 genes, along with blaSHV and blaOXA. Most isolates carried serum resistance (iss) and temperature sensitive haemagglutinin (tsh) as the main virulence marker genes. The association of class I integrons with ESBL and virulence markers rendered these isolates potentially zoonotic, and they could easily be spread to humans through the food chain or via the environment, causing serious public health concerns.
References
Kaper J, Nataro J, Mobley HLT (2004) Pathogenic Escherichia coli. Nat Rev Microbiol 2:123–140. https://doi.org/10.1038/nrmicro818
Kathayat D, Lokesh D, Ranjit S, Rajashekara G (2021) Avian pathogenic Escherichia coli (APEC): an overview of virulence and pathogenesis factors, zoonotic potential, and control strategies. Pathogens 10:467. https://doi.org/10.3390/pathogens10040467
Dziva F, Stevens MP (2008) Colibacillosis in poultry: unravelling the molecular basis of virulence of avian pathogenic Escherichia coli in their natural hosts. Avian Pathol 37:355–366. https://doi.org/10.1080/03079450802216652
Subedi M, Luitel H, Devkota B, Bhattarai RK, Phuyal S, Panthi P et al (2018) Antibiotic resistance pattern and virulence genes content in avian pathogenic Escherichia coli (APEC) from broiler chickens in Chitwan, Nepal. BMC Vet Res 14:113–118. https://doi.org/10.1186/s12917-018-1442-z
Magray SN, Wani SA, Kashoo ZA, Bhat MA, Adil S, Farooq S et al (2018) Serological diversity, molecular characterisation and antimicrobial sensitivity of avian pathogenic Escherichia coli (APEC) isolates from broiler chickens in Kashmir. Anim Prod Sci, India. https://doi.org/10.1071/AN17065
Sen A, Bandopadhyay S, Misri J, Chauhan HC, Anand Kumar P, Vaid RK et al (2022) Antimicrobial resistance in humans and livestock population in India. Indian J Anim Sci 92:665–681. https://doi.org/10.56093/ijans.v92i6.96034
Islam MS, Sobur MA, Rahman S, Ballah FM, Levy S, Siddique MP et al (2022) Detection of blaTEM, blaCTX-M, blaCMY, and blaSHV genes among Extended-Spectrum Beta-Lactamase-producing Escherichia coli isolated from migratory birds travelling to Bangladesh. Microb Ecol 83:942–950. https://doi.org/10.1007/s00248-021-01803-x
Solà-Ginés M, Cameron-Veas K, Badiola I, Dolz R, Majó N, Dahbi G et al (2015) Diversity of multi-drug resistant Avian Pathogenic Escherichia coli (APEC) causing outbreaks of colibacillosis in broilers during 2012 in Spain. PLoS One 10:e0143191. https://doi.org/10.1371/journal.pone.0143191
Ferens WA, Hovde CJ (2011) Escherichia coli O157:H7: animal reservoir and sources of human infection. Foodborne Pathog Dis 8:465–487. https://doi.org/10.1089/fpd.2010.0673
Sun J, Zheng F, Wang F, Wu K, Wang Q, Chen Q et al (2013) Class 1 Integrons in urinary isolates of extended-spectrum β-Lactamase-producing Escherichia coli and Klebsiella pneumoniae in Southern China during the past five years. Microb Drug Resist. https://doi.org/10.1089/mdr.2012.0130
Kar D, Bandyopadhyay S, Bhattacharyya D, Samanta I, Mahanti A, Nanda PK et al (2015) Molecular and phylogenetic characterization of multidrug resistant extended spectrum beta-lactamase producing Escherichia coli isolated from poultry and cattle in Odisha, India. Infect Genet Evol 29:82–90. https://doi.org/10.1016/j.meegid.2014.11.003
Edwards PR, Ewing WE (1972) Identification of Enterbacteriaceae, 3rd edn. Burgess Publishing CO, Minneapolis, pp 288–291
Lee DH, Zo YG, Kim SJ (1996) Non-radioactive method to study genetic profiles of bacterial communities by PCR single strand conformation polymorphism. Appl Environ Microbiol 62:3112–3120. https://doi.org/10.1128/aem.62.9.3112-3120.1996
Clinical and Laboratory Standards Institute (2018) Performance standards for antimicrobial disk and dilution susceptibility tests for bacteria isolated from animals. VET08, 4th Editon. table 7A: 122.
Dallenne C, Da Costa A, Decre D, Favier C, Arlet G (2010) Development of a set of multiplex PCR assays for the detection of genes encoding important beta-lactamases in Enterobacteriaceae. J Antimicrob Chemother 65:490–495. https://doi.org/10.1093/jac/dkp498
Koovapra S, Bandyopadhyay S, Das G, Bhattacharyya D, Banerjee J, Mahanti A et al (2016) Molecular signature of extended spectrum β-lactamase producing Klebsiella pneumoniae isolated from bovine milk in eastern and north-eastern India. Infect Genet Evol 44:395–402. https://doi.org/10.1016/j.meegid.2016.07.032
Cordoni G, Woodward MJ, Wu H, Alanazi M, Wallis T, La Ragione RM (2016) Comparative genomics of European avian pathogenic Escherichia coli (APEC). BMC Genomics 17:960. https://doi.org/10.1186/s12864-016-3289-7
Blanco M, Blanco JE, Mora A et al (2004) Serotypes, virulence genes, and intimin types of Shiga toxin (verotoxin)-producing Escherichia coli isolates from cattle in Spain and identification of a new intimin variant gene (eae-xi). J Clin Microbiol 42(2):645–651. https://doi.org/10.1128/jcm.42.2.645-651.2004
Krishnegowda DN, Singh BR, Mariappan AK, Munuswamy P, Singh KP, Sahoo M et al (2022) Molecular epidemiological studies on avian pathogenic Escherichia coli associated with septicemia in chickens in India. Microb Pathog 162:105313. https://doi.org/10.1016/j.micpath.2021.105313
Bhardwaj K, Shenoy SM, Baliga S, Unnikrishnan B, Baliga BS, Shetty VS (2021) Characterization of antibiotic resistant phenotypes and linked genes of Escherichia coli and Klebsiella pneumoniae from healthy broiler chickens, Karnataka, India (Research Note). Poult Sci 100:101094. https://doi.org/10.1016/j.psj.2021.101094
Ibrahim RA, Cryer TL, Lafi SQ, Basha EA, Good L, Tarazi YH (2019) Identification of Escherichia coli from broiler chickens in Jordan, their antimicrobial resistance, gene characterization and the associated risk factors. BMC Vet Res 15:159. https://doi.org/10.1186/s12917-019-1901-1
Samanta I, Joardar SN, Das PK, Das P, Sar TS, Dutta TK et al (2014) Virulence repertoire, characterization, and antibiotic resistance pattern analysis of Escherichia coli isolated from backyard layers and their environment in India. Avian Dis 8:39–45. https://doi.org/10.1637/10586-052913-Reg.1
Levy S, Islam MS, Sobur MA, Talukder M, Rahman MB, Khan MFR et al (2020) Molecular detection of Avian Pathogenic Escherichia coli (APEC) for the first time in layer farms in Bangladesh and their antibiotic resistance patterns. Microorganisms 8:1021. https://doi.org/10.3390/microorganisms8071021
Gambi L, Rossini R, Menandro ML, Franzo G, Valentini F, Tosi G et al (2022) Virulence factors and antimicrobial resistance profile of Escherichia coli isolated from laying hens in Italy. Animals 12:1812. https://doi.org/10.3390/ani12141812
Gundran RS, Cardenio PA, Villanueva MA, Sison FB, Benigno CC, Kreausukon K et al (2019) Prevalence and distribution of blaCTX-M, blaSHV, blaTEM genes in extended- spectrum β- lactamase- producing Escherichia coli isolates from broiler farms in the Philippines. BMC Vet Res 15:227. https://doi.org/10.1186/s12917-019-1975-9
Tohmaz M, Askari BM, Rahmani KH, Tabar GH (2022) Antimicrobial resistance, virulence associated genes and phylogenetic background versus plasmid replicon types: the possible associations in avian pathogenic Escherichia coli (APEC). BMC Vet Res 18:421. https://doi.org/10.1186/s12917-022-03496-x
Bakhshi M, Bafghi MF, Astani A, Ranjbar VR, Zandi H, Vakili M (2017) Antimicrobial resistance pattern of Escherichia coli isolated from chickens with colibacillosis in Yazd, Iran. J Food Qual Hazard Control 4:74–78
Johar A, Al-Thani N, Al-Hadidi SH, Dlissi E, Mahmoud MH, Eltai NO (2021) Antibiotic resistance and virulence gene patterns associated with Avian Pathogenic Escherichia coli (APEC) from broiler chickens in Qatar. Antibiotics 10:564. https://doi.org/10.3390/antibiotics10050564
Rahman MA, Rahman AKMA, Islam MA, Alam MM (2017) Antimicrobial resistance of Escherichia coli isolated from milk, beef and chicken meat in Bangladesh. Bangladesh J Vet Med 15:141–146. https://doi.org/10.3329/bjvm.v15i2.35525
Government of India Gazette notification on colistin ban by ministry of health and family welfare (department of health and family welfare), New Delhi, the 19th July, 2019.
Tofani S, Albini E, Blasi F, Cucco L, Lovito C, Maresca C et al (2022) Assessing the load, virulence and antibiotic-resistant traits of ESBL/Ampc Escherichia coli from broilers raised on conventional, antibiotic-free, and organic farms. Antibiotics 11:1484. https://doi.org/10.3390/antibiotics11111484
Castanheira M, Simner PJ, Bradford PA (2021) Extended-spectrum β-lactamases: an update on their characteristics, epidemiology and detection. JAC-Antimicrob 3:092. https://doi.org/10.1093/jacamr/dlab092
Dutta TK, Roychoudhury P, Bandyopadhyay S, Wani SA, Hussain I (2011) Detection & characterization of Shiga toxin producing Escherichia coli (STEC) & enteropathogenic Escherichia coli (EPEC) in poultry birds with diarrhea. Indian J Med Res 133:541–545
Racewicz P, Majewski M, Biesiada H et al (2022) Prevalence and characterisation of antimicrobial resistance genes and class 1 and 2 integrons in multi resistant Escherichia coli isolated from poultry production. Sci Rep 12:6062. https://doi.org/10.1038/s41598-022-09996-y
Acknowledgements
The authors are grateful to the Vice-Chancellor, Kamdhenu University, and the Principal, College of Veterinary Science and A. H., Sardarkrushinagar, for providing the necessary funding and facilities for the work.
Funding
No special funding was given for carrying out this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest related to this article.
Ethical Approval
No ethical approval was required as samples were obtained from dead birds.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Patel, S.S., Patel, A.C., Mohapatra, S.K. et al. Antibiotic Resistance and Virulence Gene Patterns Associated with Multi Drug Resistant Avian Pathogenic Escherichia coli (APEC) Isolated from Broiler Chickens in India. Indian J Microbiol 64, 917–926 (2024). https://doi.org/10.1007/s12088-023-01132-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12088-023-01132-2