Abstract
Strong evidence suggests that the early composition of the oral microbiota of neonates plays an important role for the postnatal development of the oral health or immune system. However, the relationship between the maternal microbiome and the initial neonatal microbiome remains unclear. In this study, 25 pregnant women and their neonates were recruited, and the samples were collected from the maternal oral cavity, amniotic fluid, placenta and neonatal oral cavity. High-throughput sequencing of 16S rRNA was performed using the Illumina MiSeq platform to analyze the correlation with microbial community structure between the maternal and the neonatal oral cavity. The results indicated that the number of shared OTUs was up to 635 in four groups. The PCoA showed that there were certain similarities in the microbial community structure of the four groups. The dominant bacterial genera of the shared OTUs were consistent with human oral microbes, including Streptococcus, Fusobacterium and Prevotella. The results showed that there might be a correlation between the maternal and neonatal oral microbiome, through the amniotic fluid and placenta.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Strong evidence suggested that the early composition of the microbiota of neonates played an important role for the postnatal development of the immune system [1]. It is thought that the initial microbial exposure is important in defining the successional trajectories leading to more complex and stable adult ecosystems [2], the composition of the very first human microbiota could have long-lasting effects, up to months [3] or even years [4]. Therefore, if the development of the oral microflora is different due to the influence of the initial microflora, the development of the postnatal immune system may also be different. The composition of oral microbiota in early days of life seems to be a very important factor for achieving and maintaining good health in the years to come. It follows that it is fundamental to identify more thoroughly the oral ecosystem of the newborn.
Maternal microbiome is considered to be a key factor in the initial colonization and development of the neonatal microbiome, which plays an important role in children’s physical and neurocognitive development [1, 5,6,7]. In addition, a diverse group of oral microbiome can be transmitted to the murine placenta by haematogenous transmission, and most of these species are associated with adverse pregnancy outcomes in humans [8]. This suggests that the initial oral neonatal microbiome could be acquired from the mother before birth [9,10,11] and that aberrant microbiome communities in early life could lead to disease through an altered development of the immune system. Amniotic fluid makes the environment in which the fetus lives, and placenta is the organ that exchanges substances between mother and fetus. New points suggest that amniotic fluid and placenta are not sterile [12]. Via traditional PCR, culture methods etc., non-pathogenic commensal microorganisms, including Firmicutes, Tenericutes, Proteobacteria, Bacteroidetes, and Fusobacteria phyla were found in amniotic fluid and placenta, which was similar to the oral microbiota [12]. These illustrates that the initial oral microorganisms in children may be derived from maternal amniotic fluid and placenta. Though some studies try to compare the similarity of oral microbial community between the newborn and the mother, even the gut microbiota, in the term of the initial stage of development on human oral microbial community is still unclear [1].
In this study, we analyzed the bacterial composition in the maternal oral cavity, amniotic fluid, placenta and neonatal oral cavity, and compared the microbiomes derived from the four sites to explore the correlation of the microbial communities between the mother and the neonate. The overall microbiome will provide more comprehensive information on the microbiome transmission between the mother and foetus, which can not only provide guidance for maternal and child health maintenance during pregnancy, but provide a theoretical basis for preventing from adverse pregnancy outcomes also.
Materials and Methods
Study Samples
The volunteers were recruited from the Gansu Provincial Maternity and Child-care Hospital (China), whose mode of delivery was caesarean section. The inclusion criteria were as follows: (1) Healthy full-term pregnant women (> 37 weeks of gestation) who were aged between 22 and 36, (2) without diabetes or other systemic disease related to the pregnancy, and (3) without antimicrobial therapy in the previous 6 months. 25 volunteers were finally included. A total of 100 samples from maternal oral cavity (MO), amniotic fluid (AF), placenta (PL) and neonatal oral cavity (NO), were taken.
Protocol of Samples Collection
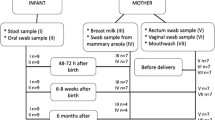
All sample collections were performed by professionals in sterile operating rooms, which strictly follow the principles of surgical asepsis. Sampling method: (1) The volunteers should gargle before the operation and the oral samples were collected by using sterile cotton swabs to wipe the oral cavity of the pregnant women, (2) 2 mL of the amniotic fluid which should be transparent and clear was collected during the operation, (3) The oral samples were collected by using sterile cotton swabs to wipe the oral cavity of the newborn within 2 min of birth, (4) The placental tissue sample was obtained by surgical scissors and all were placed in sterile tubes. The samples were acquired under aseptic conditions and stored at − 80 °C for subsequent experiments.
DNA Extraction
The total sample DNA was extracted using E.Z.N.A.TM Bacterial DNA Kit (D3350, Omega, Inc., USA) according to the manufacturer’s instructions. The total DNA was eluted in 50 µL of elution buffer and stored at − 80 °C before high-throughput sequencing by LC-Bio Technology Co., Ltd, Hang Zhou, Zhejiang Province, China.
PCR Amplification and High-Throughput Sequencing
The V3-V4 region of the bacterial 16S ribosomal DNA (rDNA) was amplified with the forward primer 338F (5′-ACTCCTACGGGAGGCAGCAG-3′) and the reverse primer 806R (5′-GGACTACHVGGGTWTCTAAT-3′). The 5′ ends of the primers were tagged with specific barcodes. PCRs were run with 25 ng of template DNA, 12.5 µL of PCR Premix (NEB, M0536L), 2.5 µL of each primer (1 µM) and deionized water to a final volume of 25 µL. PCR conditions were 30 s at 98 °C, 35 cycles of (10 s at 98 °C, 30 s at 54 °C/52 °C, 45 s at 72 °C), and 10 min at 72 °C. The PCR products were subjected to agarose gel electrophoresis, and the target PCR product was purified. The 16S rDNA of the sample was sequenced on the Illumina MiSeq platform provided by LC-Bio according to the manufacturer’s recommendations.
Data Analysis
The purified PCR products were sequenced on the Illumina MiSeq sequencing platform (Illumnia, Inc., San Diego, CA, USA) at LC-Bio. Paired-end reads were assigned to samples based on their unique barcode and truncated by removing the barcode and primer sequence. Paired-end reads were merged using FLASH [13]. Quality filtering of the raw tags was performed under specific filtering conditions to obtain the high-quality clean tags according to fqtrim (v0.94) [14], and chimeric sequences were filtered using Vsearch software (v2.3.4). Operational Taxonomic Units (OTUs) were clustered with a 97% similarity cut-off using Vsearch (v2.3.4). Representative sequences were chosen for each OTU, and taxonomic data were then assigned to each representative sequence using the RDP (Ribosomal Database Project) classifier. Multiple sequence alignments were conducted using the Mafft software (v7.310) to analyse the phylogenetic relationship of different OTUs to determine the differences in the dominant species of the various groups. OTU abundance was normalized using a standard sequence number corresponding to the sample with the least sequences. Alpha diversity (such as Chao1 and Shannon) was applied to analyze species diversity and was calculated with QIIME (Version 1.8.0). Beta diversity analysis was used to evaluate the species complexity differences of the samples and was calculated by principal coordinate analysis (PCoA). Cluster analyses were conducted using QIIME software (Version 1.8.0).
Result
The Overall Situation in OTU Lever
After filtering, the total number of sequences obtained for NO, MO, AF and PL were 360134, 318,227, 326,554 and 234,966, respectively, the rare fraction curve based on chao1 index was showed in Sup Fig. 1. A total of 1815 OTUs were obtained at the 97% similarity level [15]. The Venn diagram showed that 635 OTUs were shared among the four groups and the shared OTUs comprised approximately 50% of the total OTUs in each group, especially in MO and NO sharing OTUs more (Fig. 1a). In addition, the number of unique OTUs in each group was 117 (NO), 80 (MO), 146 (AF), and 142 (PL) (Fig. 1a). The relative abundance of the shared OTUs in each group indicated that there was no significant difference between the NO and MO groups or between the AF and PL groups (Fig. 1b). The dominant genera of the shared in 4 groups OTUs were Streptococcus, Prevotella, Fusobacterium, Neisseria, Sphingobium, Haemophilus and Saccharibacteria (Fig. 1c), which were mainly attributed to the following phyla: Actinobacteria, Proteobacteria, Firmicutes, Bacteroidetes, Fusobacteria and Candidatus.
Alpha and Beta Diversity of the Microbial Communities of the Four Groups
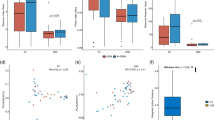
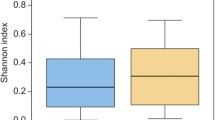
Chao1 index and Shannon index were used to measure the richness and diversity of the groups. The PL group showed the most abundant microbial communities than the MO, NO, and AF groups (Fig. 2a). The diversity of the microbial communities (Shannon) was similar in each group (Fig. 2b). As shown in Fig. 2c, beta diversity was similar in all groups except for PL based on the weighted UniFrac distance matrix in the PCoA plot. A significant difference was detected in PC1 values between the PL group and the other groups (P < 0.05).
Cluster Analysis
A multiple sample similarity tree based on the majority of sequences (93.9–97.4%) was constructed to identify the similarities and differences of microbial community structures among the four groups at the phyla level (Fig. 3a) and genus lever (Fig. 3b). The NO and AF groups were clustered together, indicating that the microbial communities between the two groups were more similar than those between the other groups. The microbial communities of the PL group were more similar to the NO and AF groups compared to the MO group. These indicated that there was no significant difference between the dominant phyla and the dominant genus in the four groups, especially in the AF and NO. Additionally, the four groups were not well separated from each other, indicating that there was no significant difference between the four groups. At the same time, we conducted principal component analysis (PCA) on 30 dominant genera (Sup Fig. 2). Based on the 30 dominant genera of the four groups, the PCA showed that the four groups were overlapped together, further indicating similar microbial composition between the four groups.
The majority of sequences in the phyla level were Proteobacteria, Bacteroidetes, Firmicutes, Actinobacteria, Fusobacteria, SR1, Candidatus_Saccharibacteria, Spirochaetes and Cyanobacteria. The majority of sequences in the genus level were affiliated with 22 genera, including Sediminibacterium, Streptococcus, Haemophilus, Prevotella, Sphingomonas, Neisseria, SR1 and Fusobacterium, Alloprevotella, Lautropia, Porphyromonas, Veillonella, Saccharibacteria, Granulicatella, Actinomyces, Rothia, Reyranella, Corynebacterium, Staphylococcus, Capnocytophaga, Acinetobacter and unclassified bacteria.
Comparison of Microbial Communities at the Phylum and Genus Levels
The differences among the 22 dominant genera were analyzed. Of 22 dominant genera, 13 were same in the four groups. Further analysis showed that 8 phyla presented a similar relative abundance between the AF and NO groups. In addition, the microbial community composition was similar between the AF and NO groups at the genus level. Streptococcus, Haemophilus, Prevotella, Sphingomonas, Neisseria, SR1 and Fusobacterium were relatively abundant in all groups. The relative abundance of Sediminibacterium in MO group was the lowest, compared to other groups. Streptococcus (17.78%), Haemophilus (10.64%) and Veillonella (6.07%) were the most abundant in the MO group, whereas the relative abundance of Sediminibacterium (6.73%) was the lowest. Sphingomonas (7.14%) was the most abundant in the PL group, compared to other groups. Prevotella (6.12%) was the most abundant in the NO group, compared to other groups. In addition, we selected the core microbial in four groups (Tab 1): The core microbial community of NO group is Sediminibacterium, Sphingomonas; the core microbial community of MO group is Prevotella, Sphingomonas, Neisseria, Actinomyces; the core microbial community of AF group is Sphingomonas, Staphylococcus; the core microbial community of PL group is Sediminibacterium, Sphingomonas. The core-bacteria common to the four groups are Sphingomonas. And the core bacteria common to NO group and PL group are Sediminibacterium.
The Major Functional Pathways in All the Four Types of Samples
Though analysis of picrust2, there were only a few significant differences in functions among the four groups, but no significant differences in top 10 dominant functions (P > 0.05), indicating that the dominant functions were similar (Sup Fig. 3).
Discussion
Most of the previous studies on the initial flora of newborn only focused on relatively just one site, such as the similarity of single flora of maternal and infant oral cavity. However, in this study, we analyzed the microbial samples of maternal oral cavity (MO), amniotic fluid (AF), placenta (PL) and neonatal oral cavity (NO). The results showed that there was no significant difference between maternal oral cavity and neonatal oral cavity. So, we concluded that there was a close relationship between mother and infant oral microflora. At the same time, there was no significant difference in the microbial community richness of mother’s oral cavity, amniotic fluid and newborn’s oral cavity, and the microbial community diversity (Shannon) of each group was similar. Therefore, this result suggests a microbiota connection between the mother’s oral cavity and the newborn’s oral cavity. The fetus in womb can obtain a beneficial initial flora, and this plays a vital role in the child’s oral health and even overall health.
A large number of experiments have isolated microorganisms from placenta, fetal membranes, amniotic fluid and cord blood [16, 17]. Therefore, pregnancy is considered to be the beginning of fetal exposure to bacteria [11, 18]. In our study, a total of 365 genera, 22 dominant genera and 8 dominant phyla were identified. It is worth mentioning that all samples in this study were from caesarean mothers and were conducted by sterile operation in a sterile operating room. To a great extent, it avoids the pollution of experimental technology and makes our experimental results more reliable. At the same time, we found that NO, MO, AF and PL groups of core bacteria Sphingomonas existed in the gingival groove of adult patients with periodontitis, suggesting that genus Sphingomonas may be the medium of mother-to-child transmission [19]. More importantly, studies have found that in the case of oral diseases (gingivitis or periodontitis), bacteria in the oral cavity may reach amniotic fluid through transient bacteremia [20], indicating that maternal microbes may be transmitted to the amniotic fluid with blood. Amniotic fluid is essential for the maturity of the embryo and the fetus. It is an important medium for fetus oral microbial transmission and a protective fluid for the fetus. In early development, amniotic fluid was an extension of the fetal extracellular matrix [21]. With the presence of placenta and fetal blood vessels, water and solutes from maternal plasma diffuse into the amniotic fluid [22]. This suggests that the relationship between the composition of oral microorganisms and amniotic fluid is closest. It indicates that the microorganisms in the amniotic fluid may be swallowed by the fetus and planted in the fetal mouth before birth. The maternal oral flora can be colonized into the newborn’s mouth through the amniotic fluid and the placenta by means of blood-borne dissemination. The above results all indicate that there is a certain correlation between maternal-infant oral microbes. The maternal oral microbes may enter the placenta and amniotic fluid through blood, and it indirectly affects or determines the early formation of fetal oral microbes. In the later development, immunity and postnatal growth, for the same reason, it plays an important role.
Streptococcus is a key bacterium for early colonization [23], and their adhesion to the inner surface of the oral cavity promotes colonization of more bacteria [24], prompting the establishment of oral biofilms in the early stages of human life. This study found that Streptococcus was 9.22% of newborn oral samples and indicated that Streptococcus as an early oral colonization had an irreplaceable role in the establishment of the initial flora. In the present study, Streptococcus, Prevotella and Neisseria were prevalent in the maternal oral cavity, amniotic fluid, placenta and neonatal oral cavity. Previous studies based on animal models have found that maternal bacteria in saliva can be transmitted to the placenta, including Streptococcus, Neisseria, Veillonella, Prevotella, Porphyromonas, and Capnocytophaga gingivalis [25]. Bearfield et al. observed that oral-derived Streptococcus and Fusobacterium in amniotic fluid [26]. Moreover, Vesty et al. found that Neisseria, Prevotella, Streptococcus and Veillonella were detected from all salivary samples in a study of different DNA extraction methods for oral flora structure [27]. These indicated that some of the maternal oral flora could be colonized in the neonatal oral cavity through the placenta and amniotic fluid.
In addition, to better understand the spread of oral microbes from mother to infant, the large sample size, metagenomics (which can be accurate to the species level, and at the same time can specify the gene function) and other methods are necessary to be applied to subsequent experimental studies. At the same time, RT-PCR or animal experiments should be involved in order to verify the exact genus of bacteria transmitted from mother to infant, such as Sphingomonas. In order to develop strategies to improve or optimize the oral health of pregnant women, improve the long-term oral health of human beings and prevent the occurrence of oral diseases.
References
Dominguez-Bello MG, Costello EK, Contreras M, Magris M, Hidalgo G, Fierer N, Knight R (2010) Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci USA 107:11971–11975. https://doi.org/10.1073/pnas.1002601107
Biasucci G, Benenati B, Morelli L, Bessi E, Boehm G (2008) Cesarean delivery may affect the early biodiversity of intestinal bacteria. J Nutr 138:1796s–1800s. https://doi.org/10.1093/jn/138.9.1796S
Gronlund MM, Lehtonen OP, Eerola E, Kero P (1999) Fecal microflora in healthy infants born by different methods of delivery: permanent changes in intestinal flora after cesarean delivery. J Pediatr Gastroenterol Nutr 28:19–25. https://doi.org/10.1097/00005176-199901000-00007
Salminen S, Gibson GR, McCartney AL, Isolauri E (2004) Influence of mode of delivery on gut microbiota composition in seven year old children. Gut 53:1388–1389. https://doi.org/10.1136/gut.2004.041640
Neu J (2015) Developmental aspects of maternal-fetal, and infant gut microbiota and implications for long-term health. Matern Health Neonatol Perinatol 1:6. https://doi.org/10.1186/s40748-015-0007-4
Rautava S, Luoto R, Salminen S, Isolauri E (2012) Microbial contact during pregnancy, intestinal colonization and human disease. Nat Rev Gastroenterol Hepatol 9:565–576. https://doi.org/10.1038/nrgastro.2012.144
Yang I, Corwin EJ, Brennan PA, Jordan S, Murphy JR, Dunlop A (2016) The infant microbiome: implications for infant health and neurocognitive development. Nurs Res 65:76–88. https://doi.org/10.1097/nnr.0000000000000133
Fardini Y, Chung P, Dumm R, Joshi N, Han YW (2010) Transmission of diverse oral bacteria to murine placenta: evidence for the oral microbiome as a potential source of intrauterine infection. Infect Immun 78:1789–1796. https://doi.org/10.1128/iai.01395-09
Neu J (2012) Diversity of microbes in amniotic fluid. In: Yearbook of neonatal and perinatal medicine, vol 2012, pp 55–56. https://doi.org/10.1016/j.ynpm.2012.05.017
DiGiulio DB, Romero R, Amogan HP, Kusanovic JP, Bik EM, Gotsch F, Kim CJ, Erez O, Edwin S, Relman DA (2008) Microbial prevalence, diversity and abundance in amniotic fluid during preterm labor: a molecular and culture-based investigation. PLoS ONE 3:e3056. https://doi.org/10.1371/journal.pone.0003056
Jimenez E, Fernandez L, Marin ML, Martin R, Odriozola JM, Nueno-Palop C, Narbad A, Olivares M, Xaus J, Rodriguez JM (2005) Isolation of commensal bacteria from umbilical cord blood of healthy neonates born by cesarean section. Curr Microbiol 51:270–274. https://doi.org/10.1007/s00284-005-0020-3
Aagaard K, Ma J, Antony KM, Ganu R, Petrosino J, Versalovic J (2014) The placenta harbors a unique microbiome. Sci Transl Med 6:237ra265. https://doi.org/10.1126/scitranslmed.3008599
Magoc T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27:2957–2963. https://doi.org/10.1093/bioinformatics/btr507
Rognes T, Flouri T, Nichols B, Quince C, Mahe F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Romano-Keeler J, Weitkamp JH (2015) Maternal influences on fetal microbial colonization and immune development. Pediatr Res 77:189–195. https://doi.org/10.1038/pr.2014.163
Oh KJ, Lee SE, Jung H, Kim G, Romero R, Yoon BH (2010) Detection of ureaplasmas by the polymerase chain reaction in the amniotic fluid of patients with cervical insufficiency. J Perinat Med 38:261–268. https://doi.org/10.1515/jpm.2010.040
Jimenez E, Marin ML, Martin R, Odriozola JM, Olivares M, Xaus J, Fernandez L, Rodriguez JM (2008) Is meconium from healthy newborns actually sterile? Res Microbiol 159:187–193. https://doi.org/10.1016/j.resmic.2007.12.007
van Winkelhoff AJ, Rurenga P, Wekema-Mulder GJ, Singadji ZM, Rams TE (2016) Non-oral gram-negative facultative rods in chronic periodontitis microbiota. Microb Pathog 94:117–122. https://doi.org/10.1016/j.micpath.2016.01.020
Sampaio-Maia B, Monteiro-Silva F (2014) Acquisition and maturation of oral microbiome throughout childhood: an update. Dent Res J (Isfahan) 11:291–301
Tong XL, Wang L, Gao TB, Qin YG, Qi YQ, Xu YP (2009) Potential function of amniotic fluid in fetal development—novel insights by comparing the composition of human amniotic fluid with umbilical cord and maternal serum at mid and late gestation. J Chin Med Assoc 72:368–373. https://doi.org/10.1016/s1726-4901(09)70389-2
Brace RA (1995) Progress toward understanding the regulation of amniotic fluid volume: water and solute fluxes in and through the fetal membranes. Placenta 16:1–18. https://doi.org/10.1016/0143-4004(95)90077-2
Kreth J, Merritt J, Qi F (2009) Bacterial and host interactions of oral streptococci. DNA Cell Biol 28:397–403. https://doi.org/10.1089/dna.2009.0868
Jenkinson HF, Lamont RJ (1997) Streptococcal adhesion and colonization. Crit Rev Oral Biol Med 8:175–200. https://doi.org/10.1177/10454411970080020601
Han YW, Ikegami A, Bissada NF, Herbst M, Redline RW, Ashmead GG (2006) Transmission of an uncultivated Bergeyella strain from the oral cavity to amniotic fluid in a case of preterm birth. J Clin Microbiol 44:1475–1483. https://doi.org/10.1128/jcm.44.4.1475-1483.2006
Bearfield C, Davenport ES, Sivapathasundaram V, Allaker RP (2002) Possible association between amniotic fluid micro-organism infection and microflora in the mouth. BJOG 109:527–533. https://doi.org/10.1111/j.1471-0528.2002.01349.x
Vesty A, Biswas K, Taylor MW, Gear K, Douglas RG (2017) Evaluating the impact of DNA extraction method on the representation of human oral bacterial and fungal communities. PLoS ONE 12:e0169877. https://doi.org/10.1371/journal.pone.0169877
Acknowledgements
This study was supported financially by the Area Fund Projects of the Chinese National Natural Science Fund (31560159 and 31360124/C0309), Nature Science Foundation of Gansu Province (17JR5RA217), the Science and Technology Support Project in Gansu Province (17JR5RA274 and 144WCGA167), and the Key Colleges and Universities Focus on Nurturing Project (31920180017, 31920180128). This study was fully sponsored by Gansu genenbio-yes Biotechnology Cd., Ltd and Gansu MeiTa Biomedicine Co., Ltd. The funders played no role in the study design, data collection and interpretation or in the decision to submit the work for publication.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that no conflict of interest exists in the study.
Informed Consent and Ethical Approval
Informed consent forms were signed by all participants included the mothers have provided the informed consent for their neonates in the study. All procedures performed in studies involving human participants met the ethical standards of the Ethics Committee of the School of Stomatology Lanzhou University (LZUE-201700012) and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wu, S., Yu, F., Ma, L. et al. Do Maternal Microbes Shape Newborn Oral Microbes?. Indian J Microbiol 61, 16–23 (2021). https://doi.org/10.1007/s12088-020-00901-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12088-020-00901-7