Abstract
Helicobacter is an economically important genus within the phylum Proteobacteria and include many species which cause many diseases in humans. With the conventional methods, it is difficult to identify them easily due to the high genetic similarity among its species. In the present study, 361 16S rRNA (rrs) gene sequences belonging to 45 species of genus Helicobacter were analyzed. Out of these, 264 sequences of 10 clinically relevant species (including Helicobacter pylori) were used. rrs gene sequences were analyzed to obtain a phylogenetic framework tree, in silico restriction enzyme analysis and species-specific conserved motifs. Protein sequences of another housekeeping gene, hsp60 were also subjected to phylogenetic analysis to supplement the data obtained using rrs sequences. Using these approaches, six out of ten species (including H. pylori) were easily segregated, whereas four species namely H. bilis, H. cinaedi, H. felis and Candidatus H. heilmannii were found to be heterogeneous. The above approaches have also helped in segregating unclassified sequences, thus proving them as an easy diagnostic method for identifying members of genus Helicobacter up to species level.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
16S rRNA gene (rrs) is used as a standard molecular marker and has been used extensively for characterizing microbes [1] including those that are pathogenic [2–4]. The genus Helicobacter comprises of Gram-negative, spiral shaped bacteria which are known for their high virulence and for colonising the gastric mucosa of humans. It belongs to class ε-Proteobacteria and family Helicobacteraceae [5]. This genus, separated from Campylobacter [6], comprises of more than 45 species. The pathogenic species belonging to this genus could be categorized into either gastric (that colonize the stomach) or enterohepatic (that infect the intestine and/or the liver) species [7].
Helicobacter pylori (type species) is the most common pathogen among all the species of this genus with its infection having a global prevalence of around 50 % [8, 9]. More than 450 genomes of H. pylori have already been sequenced making it the most extensively studied species of this genus.
To diagnose the pathogenic species of Helicobacter, various laboratory methods are used for preliminary identification. Urease testing is much more rapid and less costly, but it has been found to be inaccurate in case of gastrointestinal bleeding [10, 11]. Serology for H. pylori and rapid urease tests (RUT) for non-pylori urease-positive species such as H. heilmannii, can be relatively insensitive due to the patchy nature of H. heilmannii colonization and low numbers of bacteria present when compared with H. pylori in case of RUT [12]. Polymerase Chain Reaction (PCR) and Multi Locus Sequence Analysis (MLSA) have also been employed but can produce erroneous results [13–15]. These shortcomings emphasize the need for a preliminary identification method which can supplement the existing laboratory methods. Recently novel biomarkers have been identified for rapid identification of pathogenic bacteria especially those which possess multiple copies of rrs gene [16–21].
Based on the comparative analysis of the rrs sequences, three molecular tools have been used to define the genetic variability among the closely related species within bacterial domain [4, 15, 22–25]. In the present study as well, these tools i.e., (1) marker enzymes using in silico restriction digest of DNA, (2) phylogenetic framework tree and (3) species-specific conserved motifs have been used to define the genetic variability among the Helicobacter species. Another housekeeping gene, hsp60 has been used as the basis for phylogenetic analysis to validate the results obtained using rrs sequences. Discrepant sequences were also investigated to reduce redundancy in the database. This can improve the accuracy as well as provide a systematic approach for characterizing the strains unambiguously.
Materials and Methods
Sequence Data
Out of the 45 species reported, 10 clinically relevant species with a significant number of rrs gene sequences were considered as master species for the detailed analysis. These include isolates from H. pylori (96 sequences), Candidatus H. heilmannii (36 sequences), H. canadensis (18 sequences), H. cinaedi (28 sequences), H. felis (17 sequences), H. bilis (20 sequences), H. hepaticus (9 sequences), H. pullorum (19 sequences), H. macacae (11 sequences) and H. cetorum (10 sequences). In the present study, out of the available 361 16S rDNA sequences, 264 sequences belonging to this genus were analysed and downloaded from the RDP (Ribosomal Database Project) (Table 1) [26]. 45 protein sequences of hsp60 (Heat Shock Protein) gene for 6 closely associated species were also downloaded from NCBI.
Phylogenetic Analysis
Phylogenetic analysis was carried out using the 16S rDNA sequences and sequences of Hsp60. CLUSTAL_X (version 2.0.11) [27] was used for aligning the sequences of each master species with Wolinella succinogenes ATCC 29543 (NR_025942) as the outgroup. Evolutionary distances were estimated by Kimura [28] using DNADIST (for rrs sequences) and PROTDIST (for Hsp60 sequences) of the PHYLIP 3.6 package [29]. The program NEIGHBOR was used to construct phylogenetic tree using neighbor-joining method and statistical analysis was carried out using SEQBOOT and CONSENSE, with 100 replicates of the data set. From each species-specific phylogenetic tree, sequences that clustered together were aligned and a consensus sequence for each clade was obtained using JALVIEW sequence editor [30]. The sequence close to the consensus sequence in the clade was chosen as its representative or master sequence and in total, 41 representative sequences were selected to determine the genetic variability among the Helicobacter species.
Species Specific Conserved Motifs
The online MEME (Multiple EM for Motif Elicitation) program [31] was used to find out the species specific signature sequences or motifs. In order to obtain maximum number of motifs, the default setting was modified from 3 to 20 motifs, with the width ranging between 30 and 50 nucleotides. The uniqueness of each motif was checked using BLASTN search against NCBI database.
In-Silico Restriction Enzyme Analysis
Restriction pattern was obtained for the 10 master data sets using www.biophp.org. Uniqueness of a restriction enzyme for a particular species was investigated.
Results
In the present study, 264 16S rDNA sequences of genus Helicobacter were analyzed to construct a phylogenetic framework, to identify species-specific conserved motifs and for in silico RE analysis.
Phylogenetic Framework Generation
96 rrs sequences of H. pylori were considered in the present analysis. All but 2 of the total 96 strains were found to be distributed into 10 distinct clades in the phylogenetic tree for H. pylori (Fig. S1). Each clade consisted of 6-13 strains with bootstrap ranging from as low as 2 to as high as 100. Similar analysis was done on other species as well (Figs. S2-S10) and many low bootstrap values were observed in species specific trees (except for H. macacae, H. hepaticus and H. cetorum) indicating high level of heterogeneity within the species. Representative sequences were selected from each species tree that could define the range of genetic variability present in rrs sequences. A total of 41 such sequences were selected for constructing a phylogenetic framework tree (Fig. S11). The phylogenetic framework showed clear segregation of all the species except H. bilis & H. cinaedi and H. felis & Candidatus H. heilmannii.
Validation of Framework Tree
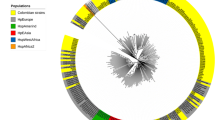
The phylogenetic framework was validated with the data sets of 10 species to check the credibility of the constructed framework. New phylogenetic trees for each species were constructed by using the framework as well as species specific sequences as the input sequences (Fig. 1). Except for a few, strains of all the species were observed to form distinct clusters with their own master sequences in their validation tree (Figs. S12-S20). This proved the validity of the framework for identification of uncharacterized Helicobacter strains.
Phylogenetic tree of 96 16S rDNA sequences of Helicobacter pylori (black) and 41 framework sequences (red). A neighbor joining analysis with Kimura correction and bootstrap support was performed on the 16S rDNA sequences belonging to H. pylori (89 shown in black excluding for those used as framework sequences) along with 41 of the phylogenetic framework. W. succinogenes was chosen as the outgroup. Bootstrap values are given at the nodes (based on 100 resampling). Values in parentheses are the accession numbers (NCBI) (http://www.ncbi.nlm.nih.gov/)
Sequences of species like H. felis & Candidatus H. heilmannii and H.bilis & H. cinaedi were found to show heterogeneity in their respective validation trees by clustering with each other (Figs. S12, S13, S15 & S17). Whereas species like H. pullorum & H. canadensis and H. pylori & H. cetorum were clearly separated by distinct yet adjacent clades (Figs. 1, S14, S19, S20).
Classification of Uncharacterized Helicobacter Strains
119 of uncharacterized species that were previously identified up to genus level were downloaded from RDP. For this, all these sequences along with those of the framework were used to generate new trees (Fig. 2a, b). Out of 119 sequences, 22 were found to clearly segregate with 7 Helicobacter framework species. Among these 22 sequences, 6 were clustered with H. cinaedi, 8 with H. pylori, 4 with H. bilis, 1 with Candidatus H. heilmannii, 1 with H. cetorum, 1 with H. macacae and 1 with H. pullorum (Table S1).
Phylogenetic tree of 41 framework sequences (red) and 119 of the uncharacterized 16S rDNA Helicobacter sp. sequences (black). The tree was constructed by neighbor joining method with Kimura correction. W. succinogenes was chosen as the outgroup. Bootstrap values are given at the nodes (based on 100 resampling). Values in parentheses are the accession numbers (NCBI) (http://www.ncbi.nlm.nih.gov/). Out of the 119 uncharacterized Helicobacter sequences, a 62 have been presented; b remaining 57 have been presented to achieve clarity in presentation
Validation Using Another Housekeeping Gene, Hsp60
To supplement the results of rrs gene analysis, 45 Hsp60 (heat shock protein) sequences (as per their availability in the database) were analyzed for 8 species (depending upon the availability of sequences). The phylogenetic tree constructed with Campylobacter coli (AAX19049) as the outgroup was found to be homogenous (Fig. S21).
In Silico Restriction Analysis
In the present study, 624 restriction enzymes (REs) were analyzed for the 16S rDNA sequences. Out of the 624 REs analyzed, 72 REs (including isoschizomers) were found to be unique in distinguishing 6 out of the 10 master data sets (H. pylori, H. cinaedi, H. felis, H. canadensis, H. macacae and Candidatus H. heilmannii) from each other (Table 2). However, no unique REs could be found for the remaining 4 species, namely, H. bilis, H. cetorum, H. hepaticus and H. pullorum.
Nucleotide Signature Analysis
Unique motifs or nucleotide signatures were found for 6 species, namely, H. cinaedi, H. hepaticus, H. cetorum, H. macacae, Candidatus H. heilmannii, H. pylori (Table 3) considered for the present study. The validation of the uncharacterized sequences was done using motifs that were found to segregate with particular framework.
Discussion
In the present study three molecular tools, i.e., phylogenetic framework, patterns of in silico RE of DNA and species-specific conserved motifs were employed using rrs gene sequences representing 10 species of genus Helicobacter, including H. pylori. The phylogenetic framework proved to be a powerful tool for investigating the classification of Helicobacter. 6 species (except for H. cinaedi, H. bilis, Candidatus H. heilmannii and H. felis) were found to form distinct clades. H. cinaedi was found to cluster with H. bilis indicating high genetic similarity between the two. Similarly, low genetic variability could be suggested between Candidatus H. heilmannii and H. felis as their strains were found to cluster with each other. Closely related species such as H. pullorum & H. canadensis were found to form distinct yet adjacent clades. Phylogenetic trees were constructed for each species to validate the framework. Segregation was distinctly observed between most species except for H. cinaedi and H. bilis; and Candidatus H. heilmannii and H. felis. Species, H. felis and Candidatus H. heilmannii were previously reported to show heterogeneity [32]. Similar observations were reported for H. bilis and H. cinaedi [33]. In the present study as well, this heterogeneous behavior was observed in their respective validation trees with bootstraps ranging from 32 to 97. Whereas closely related species like H. pullorum & H. canadensis [34] and H. cetorum & H. pylori were clearly separated by distinct yet adjacent clades. This indicated that although phylogenetic analysis of rrs couldn’t distinguish between 4 of the 10 species but was able to clearly segregate other closely related species. Specifically, H. pylori could be distinguished from other Helicobacter species.
Along with the framework sequences, 119 uncharacterized sequences were used as input to generate 2 other phylogenetic trees. 22 out of a total of 119 strains could be distributed into 7 Helicobacter species. Many of the uncharacterized strains did not cluster with the species under study. These unclustered sequences could belong to novel species or the remaining 35 that could not be considered in the present analysis.
For in silico RE analysis, 624 restriction enzymes were used out of which, 72 REs (including isoschizomers) were found to be unique for 7 species. Morphologically similar H. felis & Candidatus H. heilmannii and H. cinaedi & H. bilis that were found to cluster with each other could be segregated on the basis of their unique REs. H. canadensis could be distinguished from its related species, H. pullorum by using the marker enzymes found for the former.
Using the online MEME program, 30–50 nucleotide signatures were analyzed for each species. Unique motifs could be deduced for 6 species using BLASTN. These include both H. cinaedi and Candidatus H. heilmannii that were found to be heterogeneous with H. bilis and H. felis respectively in the framework and their respective validation trees. Both these techniques i.e. in silico restriction enzyme analysis and species specific motifs were found to be instrumental tools that could validate the results of the phylogenetic analysis as well as supplement it. While all the sequences considered in this study were scrutinized using the above three tools, some sequences were found to produce results that were unexpected according to their classification.
The hsp60 gene (GroEL, chaperonin) is a potential phylogenetic marker as it is ubiquitous and conserved in nature [35]. Sequences of the Heat Shock Protein—Hsp60 underwent phylogenetic analysis to supplement the results obtained using rrs gene (Fig. S21). Species that were found to be heterogeneous—H. cinaedi, H. bilis, Candidatus H. heilmannii and H. felis were clearly segregated using this housekeeping gene though they were found to form adjacent clades. H. pullorum was found to act as an outgroup to H. canadensis supporting possible evolution of the latter from the former [34]. The entire study proved to be useful in characterizing sequences belonging to respective species of Helicobacter, including the most dreadful one, H. pylori.
Conclusion
In the present study, the reliability of the widely acknowledged and highly conserved gene, 16S rDNA was scrutinized using molecular tools and Helicobacter as the model organism. The three tools namely, the phylogenetic framework, species specific restriction enzymes and nucleotide signatures based on the conserved rrs gene were found to be reliable, effective and aided in (1) preliminarily identifying characterized as well as uncharacterized strains of Helicobacter and (2) suggesting incorrect classification of some strains in the database so as to reduce its redundancy. Phylogenetic analysis of rrs was found to be quite reliable for identification of six pathogenic species of Helicobacter. Difficulty in the investigation for unique restriction enzymes and nucleotide signatures indicated the high genetic similarity among these species. But these tools were found to be highly successful in discriminating H. pylori from its other relative species.
Hsp60 was found to be a reliable marker and was found to segregate four species (that were found to be heterogeneous using rrs gene) quite clearly. With the availability of more Hsp60 sequences in the databases and other housekeeping genes, it can be used for supplementation of the 16S rDNA data to facilitate the identification of emerging and widespread pathogens like Helicobacter thus reducing the time and efforts to identify and characterize the new strains.
References
Woese CR (1987) Bacterial evolution. Microbiol Rev 51:221–271
Clarridge JE 3rd (2004) Impact of 16S rRNA gene sequence analysis for identification of bacteria on clinical microbiology and infectious diseases. Clin Microbiol Rev 17:840–862. doi:10.1128/CMR.17.4.840-862.2004
Janda JM, Abbott SL (2007) 16S rRNA gene sequencing for bacterial identification in the diagnostic laboratory: pulses, perils and pitfalls. J Clin Microbiol 45:2761–2764. doi:10.1128/JCM.01228-07
Porwal S, Lal S, Cheema S, Kalia VC (2009) Phylogeny in aid of the present and novel microbial lineages: diversity in Bacillus. PLoS One 4:e4438. doi:10.1371/journal.pone.0004438
Owen RJ (1998) Helicobacter—species classification and identification. Br Med Bull 54:17–30. doi:10.1093/oxfordjournals.bmb.a011667
Goodwin CS, Armstrong JA, Chilvers T, Peters M, Collins MD et al (1989) Transfer of Campylobacter pylori and Campylobacter mustelae to Helicobacter gen. nov. as Helicobacter pylori comb. nov. and Helicobacter mustelae comb. nov. respectively. Int J Syst Bacteriol 39:397–405. doi:10.1099/00207713-39-4-397
Solnick JV, O’Rourke J, Lee A, Paster BJ, Dewhirst FE et al (1993) An uncultured gastric spiral organism is a newly identified Helicobacter in humans. J Infect Dis 168:379–385. doi:10.1093/infdis/168.2.379
Parsonnet J (1998) Helicobacter pylori: the size of the problem. Gut 43:S6–S9. doi:10.1136/gut.43.2008.S6
Suerbaum S, Michetti P (2012) Helicobacter pylori infection. N Engl J Med 347:175–186
Schilling D, Demel A, Adamek HE, Nüsse T, Weidmann E et al (2003) A negative rapid urease test is unreliable for exclusion of Helicobacter pylori infection during acute phase of ulcer bleeding—a prospective case control study. Dig Liver Dis 35:217–221. doi:10.1016/S1590-8658(03)00058-6
Laine LA, Nathwani RA, Naritoku W (2005) The effect of GI bleeding on Helicobacter pylori diagnostic testing: a prospective study at the time of bleeding and 1 month later. Gastro Intest Endosc 62:853–859. doi:10.1016/j.gie.2005.09.019
Morris A, Ali MR, Thomsen L, Hollis B (1990) Tightly spiral shaped bacteria in the human stomach: another cause of active chronic gastritis? Gut 31:139–143. doi:10.1136/gut.31.2.139
Chong SKF, Lou Q, Fitzgerald JF, Lee CH (1996) Evaluation of 16S rRNA gene PCR with primers Hp1 and Hp2 for detection of Helicobacter pylori. J Clin Microbiol 34:2728–2730
Sugimoto M, Wu JY, Abudayyeh S, Hoffman J, Brahem H (2009) Unreliability of results of PCR detection of Helicobacter pylori in clinical or environmental samples. J Clin Microbiol 47:738–742. doi:10.1128/JCM.01563-08
Lal D, Verma M, Lal R (2011) Exploring internal features of 16S rRNA gene for identification of clinically relevant species of the genus Streptococcus. Ann Clin Microbiol Antimicrob 10:28. doi:10.1186/1476-0711-10-28
Kalia VC, Kumar P (2015) Genome wide search for biomarkers to diagnose Yersinia infections. Indian J Microbiol 55:366–374. doi:10.1007/s12088-015-0552-6
Kalia VC, Kumar P, Kumar R, Mishra A, Koul S (2015) Genome wide analysis for rapid identification of Vibrio species. Indian J Microbiol 55:375–383. doi:10.1007/s12088-015-0553-5
Kalia VC, Kumar R, Kumar P, Koul S (2015) A genome-wide profiling strategy as an aid for searching unique identification biomarkers for Streptococcus. Indian J Microbiol 56:46–58. doi:10.1007/s12088-015-0561-5
Kekre A, Bhushan A, Kumar P, Kalia VC (2015) Genome wide analysis for searching novel markers to rapidly identify Clostridium strains. Indian J Microbiol 55:250–257. doi:10.1007/s12088-015-0535-7
Kumar R, Koul S, Kumar P, Kalia VC (2016) Searching biomarkers in the sequenced genomes of Staphylococcus for their rapid identification. Indian J Microbiol 56:64–71. doi:10.1007/s12088-016-0565-9
Koul S, Kumar P, Kalia VC (2015) A unique genome wide approach to search novel markers for rapid identification of bacterial pathogens. J Mol Genet Med 9:194. doi:10.4172/1747-0862.1000194
Bhushan A, Joshi J, Shankar P, Kushwah J, Raju SC, Purohit HJ, Kalia VC (2013) Development of genomic tools for the identification of certain Pseudomonas up to species level. Indian J Microbiol 53:253–263. doi:10.1007/s12088-013-0412-1
Bhushan A, Mukherjee T, Joshi J, Shankar P, Kalia VC (2015) Insights into the origin of Clostridium botulinum strains: evolution of distinct restriction endonuclease sites in rrs (16S rRNA gene). Indian J Microbiol 55:140–150. doi:10.1007/s12088-015-0514-z
Kalia VC, Mukherjee T, Bhushan A, Joshi J, Shankar P, Huma N (2011) Analysis of the unexplored features of rrs (16S rDNA) of the genus Clostridium. BMC Genom 12:18. doi:10.1186/1471-2164-12-18
Kalia VC (2015) Let’s explore the latent features of genes to identify bacteria. J Mol Genet Med 9:e105. doi:10.4172/1747-0862.1000E105
Maidak BL, Cole JR, Lilburn TG, Parker CT, Saxman PR, Stredwick JM et al (2000) The RDP (ribosomal database project) continues. Nucl Acids Res 28:173–174
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA et al (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120. doi:10.1007/BF01731581
Felsenstein J (1993) Phylip (Phylogeny Inference Package) version 3.57c. Department of Genetics, University of Washington, Seattle. http://evolution.genetics.washington.edu/phylip.html
Waterhouse AM, Procter JB, Martin DM, Clamp M, Barton GJ (2009) Jalview version 2: a multiple sequence alignment and analysis workbench. Bioinformatics 25:1189–1191. doi:10.1093/bioinformatics/btp033
Bailey TL, Elkan C (1994) Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc Int Conf Intell Syst Mol Biol 2:28–36
O’Rourke JL, Solnick JV, Neilan BA, Seidel K, Hayter R et al (2004) Description of Candidatus Helicobacter heilmannii infections based on DNA sequence analysis of 16S rRNA and urease genes. Int J Syst Evol Microbiol 54:2203–2211. doi:10.1099/ijs.0.63117-0
Vandamme P, Harrington CS, Jalava K, On SLW (2000) Misidentifying Helicobacters: the Helicobacter cinaedi example. J Clin Microbiol 38:2261–2266
Fox JG, Chien CC, Dewhirst FE, Paster BJ, Shen Z et al (2000) Helicobacter canadensis sp. nov. isolated from humans with diarrhea as an example of an emerging pathogen. J Clin Microbiol 38:2546–2549
Segal G, Ron EZ (1996) Regulation and organization of the GroE and dnaK operons in eubacteria. FEMS Microbiol Lett 138:1–10. doi:10.1111/j.1574-6968.1996.tb08126.x
Acknowledgments
A.P., A.R. acknowledge Delhi University Innovative Scheme for providing the fellowship and opportunity for undertaking this project.
Authors’ Information
This work is performed by the undergraduate students at Sri Venkateswara College, University of Delhi.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Puri, A., Rai, A., Dhanaraj, P.S. et al. An In Silico Approach for Identification of the Pathogenic Species, Helicobacter pylori and Its Relatives. Indian J Microbiol 56, 277–286 (2016). https://doi.org/10.1007/s12088-016-0575-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12088-016-0575-7