Abstract
Bone marrow (BM) microenvironment plays an important role in normal and malignant hematopoiesis. As a consequence of interaction with the leukemic cells, the stromal cells of the bone marrow become deregulated in their normal function and gene expression. In our study, we found that mesenchymal stem cells (MSC) from BM of chronic myeloid leukemia (CML) patients have defective osteogenic differentiation and on interaction with K562 CML cells, the normal MSC showed reduced osteogenic differentiation. On interaction with K562 cells or its secreted factors, MSC acquired phenotypic abnormalities and secreted high levels of IL6 through NFκB activation. The MSC derived secreted factors provided a survival advantage to CML cells from imatinib induced apoptosis. Thus, a therapy targeting stromal cells in addition to leukemia cells might be more effective in eliminating CML cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
BM microenvironment stromal cells have been extensively studied for their role in normal hematopoiesis and modifications in the stromal cells were reported to disrupt HSC maintenance and differentiation (Morrison and Scadden 2014). BM stromal cells support and regulate normal and leukemic stem cells by providing signaling cues for the maintenance, proliferation, survival, migration, homing and differentiation (Levesque et al. 2010; Lo Celso and Scadden 2011). Leukemic cells were proposed to hijack and manipulate the normal BM microenvironment and render it more supportive for leukemic cells and less suitable for normal hematopoiesis (Krause and Scadden 2015) and play an important role in chemoresistance development (Kumar et al. 2017). BM stroma was implicated in leukemia initiation where dysfunctional osteoprogenitor cells or osteoblasts in the mouse BM resulted in deregulation of normal hematopoiesis, leading to myelodysplastic syndrome (MDS) which further gave rise to acute myeloid leukemia (AML) (Kode et al. 2016; Raaijmakers et al. 2010). Mutations have also been reported in MSC isolated from patients with hematologic malignancies. MSC from MDS and AML were reported to carry cytogenetic abnormalities which were distinct from those seen in leukemic cells (Blau et al. 2011). However, MSC derived from CML patients did not have BCR-ABL fusion gene present in the hematopoietic cells (Wohrer et al. 2007). MSC isolated form hematological malignancies also carry functional differences compared to MSC derived from healthy individuals (Campioni et al. 2009; Huang et al. 2015). In addition, stromal cells are a major source of cytokines and growth factors. High levels of pro-survival cytokines were identified in the bone marrow aspirates of cancer patients (Civini et al. 2013; Semesiuk et al. 2013) which might support the cancer cell proliferation and cancer chemoresistance.
In our study, we found that MSC isolated form bone marrow of CML patients have defective osteogenic differentiation, altered cell surface marker expression and modified gene expression. Interaction with CML cells also resulted in high IL6 expression in MSC and paracrine factors from MSC in turn chemoprotected CML cells from imatinib mesylate (IM) induced apoptosis.
Materials and methods
Isolation of bone marrow derived MSC
Bone marrow aspirates were obtained from patients referred for bone marrow biopsy following protocols approved by local ethical committee and after informed consent from the patients included in the study. Bone marrow samples of BCR-ABL positive CML patients were obtained at chronic phase during diagnosis (7 samples) or patients at remission stage but on imatinib therapy for 2 months (4 samples) from both male and female patients with a median age of 40 years. Bone marrow from patients who were confirmed to have no hematologic malignancies after bone marrow analysis were considered as control samples (Control-MSC). MSC were isolated from the bone marrow samples after red cell lysis and plated in growth media as described previously (Sonowal et al. 2013). The non-adherent cells were removed and the adherent spindle shaped cells were expanded further and characterized.
Differentiation of MSC
MSC were differentiated into adipocytes, osteoblasts or chondrocytes as described previously (Mawrie et al. 2016; Somaiah et al. 2015). Briefly, the cells were cultured in media supplemented with dexamethasone, isobutyl methyl xanthine, indomethacin and insulin to induce adipogenic differentiation and dexamethasone, β-glycerolphosphate, and ascorbic acid (Sigma) to induce osteogenic differentiation for 14 days. Adipogenic differentiation was determined by Oil Red O staining or gene expression analysis of adipo specific gene ADIPONECTIN (ADIPOQ) and osteogenic differentiation was detected by alizarin Red staining (Sigma) or expression levels of osteogenic genes OSTEOCALCIN (OCN) or Bone Sialoprotein (BSP). Osteogenic differentiation was also determined by histochemical staining for alkaline phosphatase (Sigma). Osteogenic differentiation was quantified by eluting Alizarin red with cetylpyridinium chloride and absorbance measurement at 562 nm. Chondrogenic differentiation was performed by generating micromass cultures in a 12-well plate according to the manufacturer’s instructions (Thermofisher scientific). Chondrocyte differentiation was analyzed by Safranin O staining after 21 days of differentiation.
Colony forming unit (CFU-F) analysis
One hundred cells per well were seeded in a 6-well plate and cultured in growth media containing 20% FBS for 14 days. Colonies larger than 50 cells were counted.
Senescence assay
Cells were seeded in a 12 well plate and allowed to grow until 80% confluent. The cells were stained for β-galactosidase activity using histochemical staining kit according to the manufacturer’s instructions (Sigma). The stained cells were visualized under microscope and documented.
Flow cytometry
MSC were trypsinized and stained with anti-human fluorescent conjugated antibodies (BD Biosciences) against the indicated cell surface receptors. The cells were incubated at 4°C for 30 min. The cells were washed and analyzed with FACS calibur (BD Biosciences). Propidium iodide was used for live/dead discrimination.
MSC co-culture with CML K562 cells
For co-culture experiments, MSC were seeded and allowed to attach for 24 h. K562 cells (National Centre for Cell Sciences, Pune, India) were added to the MSC and allowed to grow for indicated time periods. K562 cells were removed from the co-cultured MSC by quick trypsinization for 30 s. For flow cytometric and gene expression analysis, MSC were further trypsinized and used (Suppl. File 1)
Collection of conditioned media
To collect conditioned media from MSC, MSC were seeded and grown to 90% confluence. Fresh complete growth media (RPMI) was added to the cells and conditioned media containing the secreted factors was collected after 3 h and used immediately or frozen at −80°C until required.
To collect conditioned media from K562 cells, the cells were resuspended in fresh growth media (RPMI) at a density of 8 × 105 cells/ml. The cells were incubated for 3 h at 37°C, the cells were separated from the conditioned media by centrifugation. The resulting supernatant containing the secreted factors was used immediately or frozen at −80°C until use.
Mitochondrial ROS analysis
Mitochondrial ROS production was analyzed in MSC by staining the cells with Mitosox Red Mitochondrial Superoxide Indicator (ThermoFisher Scientific) according to the manufacturer’s instructions. The stained cells were analyzed by flow cytometry and mean fluorescent intensity was calculated for each condition.
Phospho protein analysis
The phosphorylated form of NFκB was analyzed by flow cytometry. The cells were fixed with 2% formaldehyde, permeabilized with 100% ice-cold methanol and incubated with anti-phospho NFκB antibody (pS529, BD Biosciences) for 1 h at room temperature. The cells were washed and analyzed with FACS calibur.
RNA extraction and real-time PCR
The cell pellet was resuspended in TriZol reagent (Thermo Fisher Scientific) and total RNA was extracted using TRIzol Plus RNA Purification Kit (Thermo Fisher Scientific) according to the manufacturer’s instructions. The resulting RNA was reverse transcribed using high capacity cDNA synthesis kit and oligodT primers at 37°C for 120 min. Real-time PCR was performed using Power SyBr Green reagents in an ABI 7500 system (Thermo Fisher Scientific). The gene expression levels in each sample was normalized to their respective GAPDH expression levels. The fold change in the expression levels compared to the control was calculated using 2−ΔΔCt method.
Quantification of secreted IL6 level
IL6 secreted levels in the conditioned media of MSC subjected to various conditions were determined using Human IL6 ELISA Kit (Thermo Fisher Scientific) as per the manufacturer’s instructions. MSC were treated with inhibitors or co-cultured with K562 cells for the indicated time period. Fresh serum free media was added to the cells and incubated for 24 h at 37°C, the conditioned media was collected, centrifuged and secreted IL6 levels were determined.
Exosome depletion
Exosome release in the MSC were blocked by treatment of MSC with 10μm of GW4869 (Sigma) overnight in serum free media as described earlier (Kang et al. 2015). The media was replaced with fresh media and conditioned media was collected after 6 h and frozen at −80°C until required.
Data analysis
Flow cytometric data was analyzed using FlowJo software. Statistical analysis was performed with SPSS software and values of p < 0.05 were considered statistically significant. The comparison between different treatment groups was done using paired samples T test and patient samples from different groups was analyzed by Mann-Whitney non-parametric variables test.
Results
CML bone marrow derived MSC show functional defects
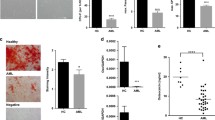
In hematologic malignancies such as CML, the cancer cells utilize the signals from bone marrow microenvironment to proliferate and acquire chemoresistance. In order to understand the changes incurred by the stromal cells during CML, mesenchymal stem cells were isolated from bone marrow of CML patients at the chronic stage during diagnosis. CML derived MSC (CML-MSC) were spindle shaped, could be passaged in culture up to 8 passages and showed normal CFU-F ability (26.5 ± 4.5 colonies per 100 cells seeded). When induced with specific differentiation media, CML-MSC differentiated into adipocytes, chondrocytes and osteoblasts (Fig. 1a, b). However, they showed a significantly reduced osteogenic differentiation potential (Fig. 1b-d) compared to control MSC as determined by alizarin red staining and OSTEOCALCIN expression. To determine whether the reduced osteogenic differentiation of CML-MSC was due to its interaction with the CML cells, control-MSC were co-cultured with CML cell line K562, their gene expression and differentiation potential was analyzed. When control-MSC were co-cultured with K562 cells, there was a significant reduction in the expression levels of adipogenic marker gene ADIPOQ and osteogenic gene BSP prior to differentiation induction (Fig. 1e). When subjected to directed differentiation, the K562 co-cultured control-MSC showed significantly reduced differentiation into osteoblasts similar to that observed in the CML patient derived MSC (Fig. 1f). However, IM treatment which is the first line of therapy for CML did not affect the adipogenic and osteogenic differentiation potential of MSC (Fig. 1g). CML-MSC in culture did not show any senescent phenotype when stained for β-galactosidase activity (Fig. 1h).
CML derived MSC have reduced osteogenic differentiation potential. a MSC isolated from bone marrow of CML patients (CML-MSC) were differentiated into adipocytes and chondrocytes. Adipogenic differentiation was determined by Oil Red O staining and chondrogenic differentiation by Safranin-O staining. b-d CML-MSC and control-MSC (CON-MSC) were differentiated into osteoblasts and stained with alizarin-Red. b Microscopic image showing osteoblasts derived from CML-MSC and CON-MSC stained with alizarin red. c Alizarin red levels or (d) OSTEOCALCIN (OCN) transcript levels in osteo differentiated CON-MSC and CML-MSC. e, f Control-MSC were co-cultured without (CON) or with K562 cells for 72 hours (MSC + K) and (e) expression levels of ADIPOQ and BSP were determined by real-time PCR. f Control-MSC (CON) and MSC co-cultured with K562 cells (MSC + K) were subjected to adipogenic and osteogenic differentiation by addition of induction media. Adipogenic differentiation was determined by oil red O (ORO) staining and osteogenic differentiation was detected by alizarin red staining (AZR). ORO and AZR staining in individual samples were quantified colorimetrically. g Control-MSC were treated without (CON) or with imatinib (10 μM) for 48 h (IM tr) and subjected to adipogenic and osteogenic differentiation. Adipogenic, osteogenic differentiation was detected by ORO, AZD staining respectively and quantified. h Representative microscopic image showing control late passage MSC (CON) and CML-MSC stained histochemically for β-galactosidase activity. Blue stain represents the senescent cells. Values are mean ± SE, *p < 0.05, **p < 0.005, ***p < 0.0005, n ≥ 3
CML cells modify the cell surface phenotype of MSC
The cell surface antigen expression profile of CML-MSC were similar to the control-MSC, however, they showed significantly reduced expression levels of CD13, CD73 and CD90 (Fig. 2a). To test further, when conditioned media from K562 cells were added to the control-MSC, there was a significant reduction in the cell surface expression levels of CD73 and CD90 in conditioned media treated control-MSC compared to the untreated cells (Fig. 2b, c). On the other hand, when cultured in direct contact with K562 cells, the co-cultured MSC showed downregulated cell surface expression of CD13, CD44, CD90 and CD95 (Fig. 2d, e). When tested further, the reduction in CD90 expression in MSC during co-culture with CML cells was also observed at the transcript level (Fig. 2f). To understand whether CML cells induce oxidative stress on MSC, the transcript levels of ROS scavenging enzymes MnSOD and CAT was analyzed in MSC co-cultured with K562 CML cells. There was a significant increase in transcript levels of ROS scavenging enzymes MnSOD and CAT (Fig. 2g) which correlated with the reduced ROS levels in these cells (Fig. 2h).
Interaction with K562 CML cells and its paracrine factors modified cell surface antigen expression in MSC. a Cell surface expression of CD13, CD73, CD90, CD95 and CD105 in control-MSC (CON-MSC) and CML-MSC was analyzed by flow cytometry. Mean (geometric) fluorescent intensity (MFI) was calculated for each marker against its isotype control. b, c Control-MSC were cultured in conditioned media derived from K562 cells for one week and their cell surface gene expression in control-MSC (CON) and conditioned media treated MSC (MSC + CM) was analyzed by flow cytometry. MFI of analyzed markers was normalized to control-MSC. c Representative flow cytometry histogram showing cell surface antigen expression levels in CON and MSC + CM conditions. Grey line represents the isotype control, blue and red line represents the stained cells. d, e Control-MSC were co-cultured without (CON) or with K562 cells (MSC + K) for one week and their cell surface gene expression profile was determined by flow cytometry. MFI was calculated for each antigen and normalized to control-MSC. e Representative flow cytometry histograms showing the cell surface antigen expression in CON (blue line), MSC + K (red line) and isotype control (grey line). f, g RNA was extracted from control-MSC (CON) and K562 co-cultured MSC (MSC + K) and reverse transcribed into cDNA. mRNA expression levels of (f) CD90, (g) CAT and MnSOD was analyzed by real-time PCR. (h) Mitochondrial ROS levels in CON and MSC + K were analyzed by staining with mitosox red and MFI of mitosox was normalized to unstained cells. Values are mean ± SE, *p < 0.05, **p < 0.005, ***p < 0.0005, n ≥ 3
CML cells modify stromal cells for chemoprotection
We further found that CML derived MSC had high IL6 mRNA levels compared to the control-MSC (Fig. 3a). Similarly, interaction of control-MSC with K562 cells resulted in increased expression of IL6 and TNFα mRNA in the stromal cells (Fig. 3b). When MSC were co-cultured with the K562 cells, they secreted significantly high levels of IL6 post exposure to the leukemic cells. This increase in IL6 was mediated by an increased phosphorylation of NFκB in MSC during its interaction with the CML cells (Fig. 3c, d). Treatment of MSC with TNFα significantly induced IL6 secretion whereas addition of NFκB inhibitor BAY 11–7082, downregulated the TNFα induced IL6 secretion (Fig. 3e). In the same way, when K562 co-cultured MSC were treated with BAY 11–7082, secreted IL6 level was downregulated (Fig. 3f, Suppl. File 1). IL6 levels increased when MSC were differentiated into osteoblasts, but IL6 levels did not predict osteogenic differentiation ability of MSC (Fig. 3g, h).
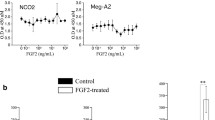
IL6 expression in MSC was induced through contact with K562-CML cells. a IL6 transcript levels were determined in control-MSC (CON-MSC) and CML-MSC by real-time PCR. b Control-MSC were co-cultured without (CON) or with K562 cells (MSC + K) and mRNA levels of TNFα (TNFA) and IL6 was determined by real-time PCR. c Control-MSC were co-cultured with K562 and phosphorylated NFκB levels was determined by flow cytometry in control (CON) and co-cultured (MSC + K) MSC. d Representative flow cytometric histogram showing the phospho NFκB analysis in CON (blue line) and MSC + K (red line). Grey line in the histogram is isotype control. e Secreted IL6 levels was determined by ELISA in conditioned media collected from K562 cells (K), control-MSC (CON), MSC treated with NFκB inhibitor BAY 11–7082 (10μM; MSC + BAY 11tr), MSC treated with TNFα (10 ng/ml; MSC-TNFa tr) or MSC treated with both TNFα and BAY 11–7082 (MSC-TNFa + BAY 11 tr). f Conditioned media was collected from control-MSC (CON), MSC co-cultured with K562 cells for 48 hours (MSC + K) or MSC treated with NFκB inhibitor BAY 11–7082 during co-culture with K562 cells (MSC + K + BAY 11). Secreted IL6 levels was determined by ELISA. Represented IL6 levels are per 1 × 105 cells/ml. g MSC were differentiated into adipocytes (AD) or osteoblasts (OS) for one week and IL6 mRNA expression level was determined by real-time PCR. h IL6 mRNA expression level was determined in MSC isolated from 10 different bone marrow donors. The respective MSC were differentiated into osteoblasts and their percentage of osteogenic differentiation was determined by alkaline phosphatase staining. The x-axis represents the different donors and bar shows the IL6 expression levels in different MSC on the primary y-axis and the line represents the osteogenic differentiation percentage in the respective MSC in the secondary y-axis. Values are mean ± SE, *p < 0.05, **p < 0.005, ***p < 0.0005, n ≥ 3
Since interaction with CML cells significantly increased NFκB phosphorylation and IL6 secretion by MSC, we tested to see whether factors from MSC could chemoprotect K562 cells from IM treatment. When K562 cells were treated with IM in the presence of conditioned media from control-MSC, there was a significant reduction in the percentage of apoptotic cells compared to the cells treated with IM without the conditioned media (Fig. 4a). Since we found that IL6 secreted levels in MSC increased post co-culture with the K562 cells, we tested the effect of IL6 receptor (IL6R) blocking in chemoprotection against IM. K562 cells incubated with MSC derived conditioned media were treated with IL6R blocking monoclonal antibody tocilizumab during treatment with IM. IL6R blocking did not chemosensitize K562 cells to IM treatment, when conditioned media from MSC were present. Similarly, blocking the release of exosomes into the MSC derived conditioned media did not affect the chemosensitivity of K562 cells. However, when conditioned media collected from MSC co-cultured with K562 cells was added, IM induced apoptosis was further inhibited in K562 cells. (Fig. 4a, b).
Paracrine factors from MSC which were in contact with K562 cells chemoprotected K562 cells from imatinib induced cell death. a K562 cells were left untreated (UNTR) or treated with IM alone (CON), or in the presence of control-MSC derived conditioned media (CM), conditioned media from control-MSC co-cultured with K562 cells (CC-CM), tocilizumab along with CM (CM + TZB) or exosome depleted conditioned media from MSC (ED-CM). Apoptosis in treated K562 cells was analyzed by annexin-V/PI staining after 48 hours. b Representative flow cytometric dot plots showing apoptosis in different conditions as shown in (a). Values are mean ± SD, ***p < 0.0005, n ≥ 3; n.s-not significant
Thus, K562 CML cells obtained survival advantage by modifying the differentiation ability and gene expression in the microenvironment stromal cells.
Discussion
In the current study, we investigated the effect of CML cells on the stromal cells of the BM microenvironment. An important role for BM stromal cells in leukemia progression, relapse and chemoresistance has been documented by several studies. MSC from hematologic malignancies have been shown to have functional abnormalities such as reduced osteogenic differentiation in multiple myeloma (Xu et al. 2012), acute myeloid leukemia (Geyh et al. 2016), increased osteogenesis in primary myelofibrosis (Martinaud et al. 2015) and increased adipogenic differentiation in acute lymphoblastic leukemia (Lopez et al. 2014). In our current study, we showed that MSC isolated from CML patients had reduced osteogenic differentiation and this was confirmed during co-culture with K562 cells. We found that the interaction of MSC with the leukemic cells down regulated the expression of osteogenic marker BSP and reduced their osteogenic differentiation potential. However, imatinib, which is a protein tyrosine kinase inhibitor used to treat CML, did not have any significant effect on the osteogenic and adipogenic differentiation of MSC. Osteoblasts are important components of the bone marrow niche to maintain HSC (Calvi et al. 2003; Zhang et al. 2003), and reduction in osteoblast number or dysfunctional osteoprogenitor cells were less supportive of normal hematopoiesis and more supportive of malignant hematopoietic cells (Kode et al. 2014; Krevvata et al. 2014; Raaijmakers et al. 2010). Thus, a leukemia induced reduction in osteogenic differential potential of MSC may result in BM microenvironment which is hostile for normal HSC but conducive to the leukemia cells proliferation.
Moreover, interaction with the leukemia cells also modified the cell surface phenotype of MSC and cell surface expression represents functional characteristics. MSC isolated from patients with hematologic malignancies expressed low CD90, which correlated with their reduced immunomodulatory properties (Campioni et al. 2009; Jaganathan et al. 2010). In our study, MSC isolated from CML patients showed reduced CD13, CD73 and CD90 expression. However, reduction in CD90 expression was also observed when leukemia cells derived conditioned media or leukemia cells were added to the MSC culture. A significant decrease also in transcript level of CD90 was observed in MSC co-cultured with the leukemia cells. With several studies suggesting the role of CD90 in cancer progression (Kumar et al. 2016), CD90 expression in stromal cells might be an important molecule to study in detail to understand its importance in leukemia progression and chemoresistance. MSC co-cultured with K562 cells also showed reduced expression of CD44, an adhesion molecule that is important for normal hematopoiesis.
An aberrant expression of cytokines was reported in acute myeloid leukemia, multiple myeloma and in some cases of lymphoma (Arnulf et al. 2007; Huang et al. 2015; Wallace et al. 2001). Schmidt et al. identified a high expression of angiogenic cytokines such as VEGF, PlGF, IL6 in the BM of CML mouse (Schmidt et al. 2011). High levels of IL6 along with transforming growth factor-α (TGFα) were detected in the plasma of CML patients (Nievergall et al. 2014) and high levels of TNFα and IL6 were reported in BM plasma of BCR-ABL mouse providing a preferential growth advantage to LSC (Zhang et al. 2012). Similarly, our study found significantly high levels of IL6 mRNA expression in MSC isolated from CML patients and in MSC co-cultured with K562 cells which might have immune modulatory functions as also reported by others (Giallongo et al. 2016). Even though MSC isolated from CML patients were separated from the leukemia cells and cultured alone, the IL6 expression levels remained significantly higher than the control cells, suggesting sustained aberrations in gene expression. Schmidt et al. reported NFκB activation to be responsible for secretion of inflammatory cytokines by MSC (Schmidt et al. 2011) and similarly in our study, an increased phosphorylation of NFκB was identified in MSC co-cultured with CML cells which led to increased IL6 secretion. Increased IL6 secretion through NFκB activation was confirmed by treatment with NFκB inhibitor BAY 11–7082 which brought the IL6 amount to basal levels. This increased NFκB activation might be responsible for reduced osteogenic differentiation (Chang et al. 2013) of CML-MSC and MSC interacting with K562 cells. Although increase in IL-6 mRNA levels during osteogenesis was found in our study, no significant correlation was observed between the IL-6 mRNA levels and osteogenic differentiation potential of MSC as reported by others (Pricola et al. 2009).
The conditioned media from CML cells was sufficient to offer survival advantage against IM induced cell death. Although high IL6 levels were seen in conditioned media of CML-MSC, inhibition of IL6 receptors (secreted and cell surface bound) through the monoclonal antibody tocilizumab even at high concentrations did not chemosensitize CML cells to IM treatment. Saini et al. reported that the pro-survival effects offered by IL6 did not depend on IL6 receptor expression but rather on iNOS pathway (Saini et al. 2014) which might explain the lack of apoptotic effect that was observed when IL6 receptor was inhibited in CML cells in our study. Also, the chemoprotective effect of MSC derived conditioned media were not mediated through exosomes but might be through FGF2 as reported by Traer et al. (Traer et al. 2014). Taken together, interaction of leukemia cells with the stromal cells modifies the stromal cells by activating NFκB pathway and its osteogenic differentiation potential, which might support CML cell proliferation and chemoprotection (Fig. 5). Cross-talk between cancer cells and stromal cells which facilitated cancer cell proliferation and chemoresistance development have been reported by several studies (Geyh et al. 2016; Kumar et al. 2017; Schroeder et al. 2016). As a result of this reciprocal interaction, the stromal cells showed aberrant gene expression profiles and leukemia cells acquired proliferative advantage and chemoprotection. Thus, utilizing a combinatorial therapeutic approach to target stromal cells to abolish the aberrant cross-talk between the leukemia cells and stromal cells might lead to better patient outcome.
Abbreviations
- ADIPOQ:
-
Adiponectin
- ALP:
-
Alkaline phosphatase
- AML:
-
Acute myeloid leukemia
- AZR:
-
Alizarin red
- BM:
-
Bone marrow
- BSP:
-
Bone sialoprotein
- CAT:
-
Catalase
- CML:
-
Chronic myeloid leukemia
- HSC:
-
Hematopoietic stem cells
- IL6:
-
Interleukin 6
- IM:
-
Imatinib mesylate
- MnSOD:
-
Manganese superoxide dismutase
- MSC:
-
Mesenchymal stem cells
- NFκB:
-
Nuclear factor kappa B
- OCN:
-
Osteocalcin
- ORO:
-
Oil red O
- PlGF:
-
Placental growth factor
- TNFα:
-
Tumor necrosis factor alpha
- VEGF:
-
Vascular endothelial growth factor
References
Arnulf B, Lecourt S, Soulier J, Ternaux B, Lacassagne MN, Crinquette A, Dessoly J, Sciaini AK, Benbunan M, Chomienne C, Fermand JP, Marolleau JP, Larghero J (2007) Phenotypic and functional characterization of bone marrow mesenchymal stem cells derived from patients with multiple myeloma. Leukemia 21(1):158–163. https://doi.org/10.1038/sj.leu.2404466
Blau O, Baldus CD, Hofmann WK, Thiel G, Nolte F, Burmeister T, Turkmen S, Benlasfer O, Schumann E, Sindram A, Molkentin M, Mundlos S, Keilholz U, Thiel E, Blau IW (2011) Mesenchymal stromal cells of myelodysplastic syndrome and acute myeloid leukemia patients have distinct genetic abnormalities compared with leukemic blasts. Blood 118(20):5583–5592. https://doi.org/10.1182/blood-2011-03-343467
Calvi LM, Adams GB, Weibrecht KW, Weber JM, Olson DP, Knight MC, Martin RP, Schipani E, Divieti P, Bringhurst FR, Milner LA, Kronenberg HM, Scadden DT (2003) Osteoblastic cells regulate the haematopoietic stem cell niche. Nature 425(6960):841–846. https://doi.org/10.1038/nature02040
Campioni D, Rizzo R, Stignani M, Melchiorri L, Ferrari L, Moretti S, Russo A, Bagnara GP, Bonsi L, Alviano F, Lanzoni G, Cuneo A, Baricordi OR, Lanza F (2009) A decreased positivity for CD90 on human mesenchymal stromal cells (MSCs) is associated with a loss of immunosuppressive activity by MSCs. Cytometry B Clin Cytom 76B(3):225–230. https://doi.org/10.1002/cyto.b.20461
Chang J, Liu F, Lee M, Wu BJM, Ting K, Zara JN, Soo C, Al Hezaimi K, Zou WP, Chen XH, Mooney DJ, Wang CY (2013) NF-kappa B inhibits osteogenic differentiation of mesenchymal stem cells by promoting beta-catenin degradation. Proc Natl Acad Sci U S A 110(23):9469–9474. https://doi.org/10.1073/pnas.1300532110
Civini S, Jin P, Ren JQ, Sabatino M, Castiello L, Jin JJ, Wang H, Zhao YL, Marincola F, Stroncek D (2013) Leukemia cells induce changes in human bone marrow stromal cells. J Transl Med 11. https://doi.org/10.1186/1479-5876-11-298
Geyh S, Rodriguez-Paredes M, Jager P, Khandanpour C, Cadeddu RP, Gutekunst J, Wilk CM, Fenk R, Zilkens C, Hermsen D, Germing U, Kobbe G, Lyko F, Haas R, Schroeder T (2016) Functional inhibition of mesenchymal stromal cells in acute myeloid leukemia. Leukemia 30(3):683–691. https://doi.org/10.1038/leu.2015.325
Giallongo C, Romano A, Parrinello NL, La Cava P, Brundo MV, Bramanti V, Stagno F, Vigneri P, Chiarenza A, Palumbo GA, Tibullo D, Di Raimondo F (2016) Mesenchymal stem cells (MSC) regulate activation of granulocyte-like myeloid derived suppressor cells (G-MDSC) in chronic myeloid leukemia patients. PLoS One 11(7). https://doi.org/10.1371/journal.pone.0158392
Huang JC, Basu SK, Zhao X, Chien S, Fang M, Oehler VG, Appelbaum FR, Becker PS (2015) Mesenchymal stromal cells derived from acute myeloid leukemia bone marrow exhibit aberrant cytogenetics and cytokine elaboration. Blood Cancer J 5. https://doi.org/10.1038/bcj.2015.17
Jaganathan BG, Tisato V, Vulliamy T, Dokal I, Marsh J, Dazzi F, Bonnet D (2010) Effects of MSC co-injection on the reconstitution of aplastic anemia patient following hematopoietic stem cell transplantation. Leukemia 24(10):1791–1795. https://doi.org/10.1038/leu.2010.164
Kang K, Ma RL, Cai WF, Huang W, Paul C, Liang JL, Wang YH, Zhao TJ, Kim HW, Xu MF, Millard RW, Wen ZL, Wang YG (2015) Exosomes secreted from CXCR4 overexpressing mesenchymal stem cells promote cardioprotection via Akt signaling pathway following myocardial infarction. Stem Cells Int. https://doi.org/10.1155/2015/659890
Kode A, Manavalan JS, Mosialou I, Bhagat G, Rathinam CV, Luo N, Khiabanian H, Lee A, Murty VV, Friedman R, Brum A, Park D, Galili N, Mukherjee S, Teruya-Feldstein J, Raza A, Rabadan R, Berman E, Kousteni S (2014) Leukaemogenesis induced by an activating beta-catenin mutation in osteoblasts. Nature 506(7487):240–244. https://doi.org/10.1038/nature12883
Kode A, Mosialou I, Manavalan SJ, Rathinam CV, Friedman RA, Teruya-Feldstein J, Bhagat G, Berman E, Kousteni S (2016) FoxO1-dependent induction of acute myeloid leukemia by osteoblasts in mice. Leukemia 30(1):1–13. https://doi.org/10.1038/leu.2015.161
Krause DS, Scadden DT (2015) A hostel for the hostile: the bone marrow niche in hematologic neoplasms. Haematologica 100(11):1376–1387. https://doi.org/10.3324/heamatol.2014.113852
Krevvata M, Silva BC, Manavalan JS, Galan-Diez M, Kode A, Matthews BG, Park D, Zhang CA, Galili N, Nickolas TL, Dempster DW, Dougall W, Teruya-Feldstein J, Economides AN, Kalajzic I, Raza A, Berman E, Mukherjee S, Bhagat G, Kousteni S (2014) Inhibition of leukemia cell engraftment and disease progression in mice by osteoblasts. Blood 124(18):2834–2846. https://doi.org/10.1182/blood-2013-07-517219
Kumar A, Bhanja A, Bhattacharyya J, Jaganathan BG (2016) Multiple roles of CD90 in cancer. Tumour Biol. https://doi.org/10.1007/s13277-016-5112-0
Kumar A, Bhattacharyya J, Jaganathan BG (2017) Adhesion to stromal cells mediates imatinib resistance in chronic myeloid leukemia through ERK and BMP signaling pathways. Sci Rep 7(1):9535. https://doi.org/10.1038/s41598-017-10373-3
Levesque JP, Helwani FM, Winkler IG (2010) The endosteal 'osteoblastic' niche and its role in hematopoietic stem cell homing and mobilization. Leukemia 24(12):1979–1992. https://doi.org/10.1038/leu.2010.214
Lo Celso C, Scadden DT (2011) The haematopoietic stem cell niche at a glance. J Cell Sci 124(21):3529–3535. https://doi.org/10.1242/jcs.074112
Lopez AV, Garcia MNV, Melen GJ, Martinez AE, Moreno IC, Garcia-Castro J, Orellana MR, Gonzalez AGZ (2014) Mesenchymal stromal cells derived from the bone marrow of acute lymphoblastic leukemia patients show altered bmp4 production: correlations with the course of disease. PLoS One 9(1). https://doi.org/10.1371/journal.pone.0084496
Martinaud C, Desterke C, Konopacki J, Pieri L, Torossian F, Golub R, Schmutz S, Anginot A, Guerton B, Rochet N, Albanese P, Henault E, Pierre-Louis O, Souraud JB, de Revel T, Dupriez B, Ianotto JC, Bourgeade MF, Vannucchi AM, Lataillade JJ, Le Bousse-Kerdiles MC (2015) Osteogenic potential of mesenchymal stromal cells contributes to primary myelofibrosis. Cancer Res 75(22):4753–4765. https://doi.org/10.1158/0008-5472.CAN-14-3696
Mawrie D, Kumar A, Magdalene D, Bhattacharyya J, Jaganathan BG (2016) Mesenchymal stem cells from human extra ocular muscle harbor neuroectodermal differentiation potential. PLoS One 11(6). https://doi.org/10.1371/journal.pone.0156697
Morrison SJ, Scadden DT (2014) The bone marrow niche for haematopoietic stem cells. Nature 505(7483):327–334. https://doi.org/10.1038/nature12984
Nievergall E, Reynolds J, Kok CH, Watkins D, Biondo M, Busfield SJ, Vairo G, Yeung DT, Lopez A, Hiwase DK, White DL, Hughes TP (2014) High plasma levels of tgf-alpha and IL-6 at diagnosis predict early molecular response failure and transformation in CML. Blood 124(21):1788
Pricola KL, Kuhn NZ, Haleem-Smith H, Song YJ, Tuan RS (2009) Interleukin-6 maintains bone marrow-derived mesenchymal stem cell stemness by an ERK1/2-dependent mechanism. J Cell Biochem 108(3):577–588. https://doi.org/10.1002/jcb.22289
Raaijmakers MHGP, Mukherjee S, Guo SQ, Zhang SY, Kobayashi T, Schoonmaker JA, Ebert BL, Al-Shahrour F, Hasserjian RP, Scadden EO, Aung Z, Matza M, Merkenschlager M, Lin C, Rommens JM, Scadden DT (2010) Bone progenitor dysfunction induces myelodysplasia and secondary leukaemia. Nature 464(7290):852–U858. https://doi.org/10.1038/nature08851
Saini AS, Shenoy GN, Rath S, Bal V, George A (2014) Inducible nitric oxide synthase is a major intermediate in signaling pathways for the survival of plasma cells. Nat Immunol 15(3):275–282. https://doi.org/10.1038/ni.2806
Schmidt T, Kharabi Masouleh B, Loges S, Cauwenberghs S, Fraisl P, Maes C, Jonckx B, De Keersmaecker K, Kleppe M, Tjwa M, Schenk T, Vinckier S, Fragoso R, De Mol M, Beel K, Dias S, Verfaillie C, Clark RE, Brummendorf TH, Vandenberghe P, Rafii S, Holyoake T, Hochhaus A, Cools J, Karin M, Carmeliet G, Dewerchin M, Carmeliet P (2011) Loss or inhibition of stromal-derived PlGF prolongs survival of mice with imatinib-resistant Bcr-Abl1(+) leukemia. Cancer Cell 19(6):740–753. https://doi.org/10.1016/j.ccr.2011.05.007
Schroeder T, Geyh S, Germing U, Haas R (2016) Mesenchymal stromal cells in myeloid malignancies. Blood Res 51(4):225–232
Semesiuk NI, Zhylchuk A, Bezdenezhnykh N, Lykhova A, Vorontsova AL, Zhylchuk VE, Kudryavets YI (2013) Disseminated tumor cells and enhanced level of some cytokines in bone marrow and peripheral blood of breast cancer patients as predictive factors of tumor progression. Exp Oncol 35(4):295–302
Somaiah C, Kumar A, Mawrie D, Sharma A, Patil SD, Bhattacharyya J, Swaminathan R, Jaganathan BG (2015) Collagen promotes higher adhesion, survival and proliferation of mesenchymal stem cells. PLoS One 10(12). https://doi.org/10.1371/journal.pone.0145068
Sonowal H, Kumar A, Bhattacharyya J, Gogoi PK, Jaganathan BG (2013) Inhibition of actin polymerization decreases osteogeneic differentiation of mesenchymal stem cells through p38 MAPK pathway. J Biomed Sci 20. https://doi.org/10.1186/1423-0127-20-71
Traer E, Javidi-Sharifi N, Agarwal A, Dunlap J, English I, Martinez J, Tyner JW, Wong M, Druker BJ (2014) Ponatinib overcomes FGF2-mediated resistance in CML patients without kinase domain mutations. Blood 123(10):1516–1524. https://doi.org/10.1182/blood-2013-07-518381
Wallace SR, Oken MM, Lunetta KL, Panoskaitsis-Mortari A, Masellis AM (2001) Abnormalities of bone marrow mesenchymal cells in multiple myeloma patients. Cancer 91(7):1219–1230. https://doi.org/10.1002/1097-0142(20010401)91:7<1219::Aid-Cncr1122>3.0.Co;2-1
Wohrer S, Rabitsch W, Shehata M, Kondo R, Esterbauer H, Streubel B, Sillaber C, Raderer M, Jaeger U, Zielinski C, Valent P (2007) Mesenchymal stem cells in patients with chronic myelogenous leukaemia or bi-phenotypic ph plus acute leukaemia are not related to the leukaemic clone. Anticancer Res 27(6B):3837–3841
Xu S, Evans H, Buckle C, De Veirman K, Hu J, Xu D, Menu E, De Becker A, Broek IV, Leleu X, Camp BV, Croucher P, Vanderkerken K, Van Riet I (2012) Impaired osteogenic differentiation of mesenchymal stem cells derived from multiple myeloma patients is associated with a blockade in the deactivation of the Notch signaling pathway. Leukemia 26(12):2546–2549. https://doi.org/10.1038/leu.2012.126
Zhang JW, Niu C, Ye L, Huang HY, He X, Tong WG, Ross J, Haug J, Johnson T, Feng JQ, Harris S, Wiedemann LM, Mishina Y, Li LH (2003) Identification of the haematopoietic stem cell niche and control of the niche size. Nature 425(6960):836–841. https://doi.org/10.1038/nature02041
Zhang B, Ho YW, Huang Q, Maeda T, Lin A, Lee SU, Hair A, Holyoake TL, Huettner C, Bhatia R (2012) Altered microenvironmental regulation of leukemic and normal stem cells in chronic myelogenous leukemia. Cancer Cell 21(4):577–592. https://doi.org/10.1016/j.ccr.2012.02.018
Acknowledgements
This work was supported by grants from Indian Council of Medical Research (ICMR) to BGJ, JB and Department of Biotechnology (DBT) to BGJ. ICMR, DBT did not have any role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: AK JB BGJ. Performed majority of experiments: AK. Performed experiments: TA AS. Analyzed the data: AK BGJ. Wrote the manuscript: AK JB BGJ. All authors have read and approved the final version of the manuscript.
Corresponding author
Electronic supplementary material
ESM 1
(PDF 96 kb)
Rights and permissions
About this article
Cite this article
Kumar, A., Anand, T., Bhattacharyya, J. et al. K562 chronic myeloid leukemia cells modify osteogenic differentiation and gene expression of bone marrow stromal cells. J. Cell Commun. Signal. 12, 441–450 (2018). https://doi.org/10.1007/s12079-017-0412-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12079-017-0412-8