Abstract
Background
Loss of terminal differentiation markers and gain of stem cell-like properties are a major hallmark of cancer malignant progression. Identification of novel biomarkers representing tumor developmental progeny and predictive of patients’ prognosis would greatly benefit clinical cancer management.
Methods
Human embryonic stem cells were induced to differentiate into hepatocytes along hepatic lineages. Transcriptomic data from different liver developmental stages were analyzed combining with the RNA-seq data from The Cancer Genome Atlas (TCGA) project. Kaplan–Meier survival analysis and Cox regression analyses were used to analyze the clinical significance in HCC patients.
Results
A shifted expression pattern of claudin (CLDN) family genes were identified to be closely associated with liver development and tumor progression. Claudins with hepatic features were found to be significantly down-regulated and predicted better prognosis in HCC patients. Conversely, another set of claudins with embryonic stem cell features were found to be significantly up-regulated and predicted worse prognosis in HCC patients. A claudin signature score system was further established by combining the two sets of claudin genes. The newly established claudins signature could robustly predict HCC patients’ prognosis in the training, testing, and independent validation cohorts.
Conclusions
In the present study, we developed a novel embryonic developmental claudins signature to monitor the extent of tumor dedifferentiation in HCC from an in vitro hepatocyte differentiation model. The claudins signature might present a great potential in predicting prognostic significance in HCC as cell surface biomarkers, and provide novel therapeutic targets for precision oncology further in the clinic.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Liver cancer is the fifth most prevalent cancer and the second leading cause of cancer death in the worldwide scale. It has high metastasis and recurrence rate, and the 5-year survival rate is relatively low (30–40%) [1]. Lack of suitable biomarkers for early detection and limited treatment strategies are the major causes of high mortality [2]. Increasing evidences suggested that critical molecules which regulate embryonic stem cell pluripotency and differentiation are usually aberrantly-expressed in the tumor tissues [3]. The developmental networks can also induce retro-differentiation or trans-differentiation between different stem cell progenies and cellular lineages, which constitute the cellular heterogeneity of liver cancer [4]. Like many other tumors, HCC also gains embryonic-like properties, such as elevated expression of alpha-fetoprotein (AFP), which should only appear in fetal liver development [5]. Poorly differentiated HCCs were usually characterized by activation stem or hepatic progenitor cell markers, and loss of hepatocyte terminal differentiation markers [6]. However, the lack of significant prognostic value or difficult for directly targeting of these biomarkers remain major restrictions in clinical practice. Therefore, it is particularly important to identify new valuable biomarkers to monitor the extent of tumor developmental progeny and stratify patient prognosis, with potential for further therapeutic targeting.
With the advance of stem cell technology, it is now possible to direct human embryonic stem cells (hESCs) to differentiate into defined cellular lineages, for example, the human hepatocytes. hESCs were induced to differentiate into human hepatocytes along hepatic lineages in vitro. Through analyzing the transcriptomic data from different developmental stages, we found a significant expression pattern shift of Claudin family genes during hepatocyte differentiation. The claudin multigene family encodes tetraspan membrane proteins which form the crucial structure of tight junctions and play important roles in maintaining cell polarity in epithelial and regulating paracellular permeability sheets. All claudins encode 20–27 kDa proteins with four transmembrane domains, two extracellular loops where the first one is significantly longer than the second one, and a short carboxyl intracellular tail [7]. The claudins family are critical in multiple biology processes probably including tumor plasticity and embryonic stem cell pluripotency [8]. The claudins interact with multiple proteins and are intimately involved in a large amount of signaling transduction [9]. In addition, mutation of some claudin genes has been causatively associated with human diseases and claudin genes have been found to be up- and down-regulated in various cancers [10]. In the present study, we identified two distinct groups of claudins from an intro hepatocyte differentiation model. By screening the RNA-seq data from the TCGA database, a hepatic claudins signature and an embryonic claudins signature with prognostic significance were established. The two claudins gene signatures were then integrated to form a score system, which was further used to monitor HCC tumor dedifferentiation and predict patients’ prognosis. Kaplan–Meier survival analysis from the training, testing and independent validation cohort proved that the newly established embryonic developmental claudins signature could robustly predict clinical outcome of HCC patients.
Material and methods
Generation of hepatocyte-like cells
The differentiation protocol for obtaining hepatocyte-like cells was conducted as described in previous studies [11, 12]. In brief, hESCs were passaged onto feeder-free system until a confluence of 50–70% was attained. Then cells cultured in RPMI-1640 (Life Technologies, Carlsbad, CA, USA) supplemented with 100 ng/ml activin A (R&D Systems, Minneapolis, MN) and 25 ng/mL Wnt3 a (R&D Systems, Minneapolis, MN, USA) for 3 days. To induce hepatic endoderm, cells were grown in KO/DMEM medium (Life Technologies, Carlsbad, CA, USA) supplemented with 25 nm/mL keratinocyte growth factor (KGF, R&D Systems, Minneapolis, MN, USA) and 2% fetal bovine serum (Gibco, Grand Island, NY, USA) for 2 days, and then further cultured in the KO/DMEM medium containing 20% SR, 1 mM glutamine, 1% nonessential amino acids, 0.1 mM 2-mercaptoethanol, and 1% DMSO for 4–7 days. The final maturation step to obtain hepatocyte-like cells involved culturing the cells in mature medium containing 10% FBS, 10 ng/mL hepatocyte growth factor (HGF, R&D Systems, Minneapolis, MN, USA), 20 ng/mL Oncostatin M (OSM, R&D Systems, Minneapolis, MN, USA) and 0.5 μM dexamethasone (DEX, R&D Systems, Minneapolis, MN, USA) for 7 more days. The validation of representative hepatic markers and the examination of hepatic functions were performed according to previous literatures [13, 14].
Clinical cohort and RNA-seq data sets
We obtained mRNA expression data and clinical-pathological data of liver hepatocellular carcinoma from the RNA-seq LIHC project of TCGA (https://tcgadata.nci.nih.gov/tcga/). The data was downloaded from the University of California Santa Cruz cancer genomics data portal UCSC Xena (https://xena.ucsc.edu/). The LIHC project contains 50 para-tumor liver tissues tissue samples and 377 primary liver cancer tissue samples (6 missing mRNA expression data). Samples from TCGA data set were divided randomly into training (TCGA-LIHC Cohort I, n = 189) and testing cohorts (TCGA-LIHC Cohort II, n = 182). A total of 232 samples with RNA-Seq mRNA expression data and clinical-pathological data were obtained from the LIRI-JP project of ICGC in ICGC Data Portal (https://dcc.icgc.org/) as an independent validation cohort. These samples belong to a Japanese population primarily infected with HBV/HCV [15]. All the tumor tissues collected from the TCGA-LIHC project and the LIRI-JP project were pathologically confirmed as hepatocellular carcinoma. We used the normalized FPKM-UQ given in the RNA-seq gene expression file. Detailed clinical background information of the patients could be found in Table S1.
Statistical analysis and signature score generation
The expression of claudins gene family between HCC primary tumor tissues and the para-tumor liver tissues were shown based on the normalized expression value of RNA-seq data. Independent student’s t test was used to compare the mean expression level of two different groups. One-way ANOVA test was used to compare means between three and more subgroups. ROC curve was used to estimate the diagnosis value of claudins signatures. Kaplan–Meier survival curves of the different risk groups were plotted and the log-rank (or log-rank for trend) p value of the survival difference was calculated between them. The tests were performed in GraphPad Prism 8.0 (La Jolla, CA, USA). Cox regression analyses and logistic analysis were used to assess association with overall survival using SPSS v21 (IBM, Inc., Chicago, IL, USA). p value less than 0.05 was considered statistically significant. A hepatic claudins signature (HCS) and a embryonic claudins signature (ECS) were generated by taking into account the expression of individual claudins family genes and their clinical association with patient overall survival time. The HCS score or ECS score were calculated according to the expression of each signature gene. For the HCS score, HCC patient with overexpression (defined as the normalized expression value above median in the HCC primary tumor tissues) of each hepatic claudins signature gene will be given “1” score. Likewise, for the ECS score, HCC patient with overexpression (defined as the normalized expression value above median in the HCC primary tumor tissues) of each embryonic claudins signature gene will be given “1” score. Patients with HCS score greater than 2 was defined as “High HCS group”, and with HCS score less than and including 2 was defined as “Low HCS group”. Similarly, Patients with ECS score greater than 2 was defined as “High ECS group”, and with ECS score less than and including 2 was defined as “Low ECS group”. Patients in “Low HCS group” and “High ECS group” were defined as “High risk group” and patients in “High HCS group” and “Low ECS group” were defined as “Low risk group”. The sum of the claudins signatures genes forms the final scores. To integrate the two distinct claudins signatures, claudins signature scores (CSS) were generated based on their HCS score and ECS score (CSS = HCS-ECS). The cytoscape v3.6.1 was used to establish a network connection of protein–protein interaction with claudins signatures and other closely associated genes. Gene ontology analysis and enrichment analysis were performed using Metascape (https://metascape.org) [16].
Results
Identification of two distinct groups of claudins from an in vitro hepatocyte differentiation model
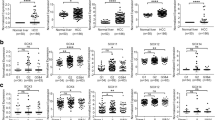
To investigate the association between liver development and carcinogenesis, we established an in vitro hepatocyte differentiation model. Human embryonic stem cells (hESCs) were induced to differentiate into hepatocytes along hepatic lineages. The embryonic stem cells, liver progenitor cells, and the mature hepatocytes from different developmental stages were collected, respectively, and the transcriptomic RNA-sequencing was used to profile the gene expressions (Fig. 1a). Hierarchical clustering analysis reveals two different pattern of gene expression during hepatocyte differentiation. One subgroup of genes was highly expressed in mature hepatocytes but decreased in embryonic stem cells and liver progenitor cells. Conversely, another set of genes was highly expressed in embryonic stem cells and liver progenitor cells but decreased in mature hepatocytes. Considering the easy detection and therapeutic targeting of cell surface proteins, we further focused on the membrane-bound proteins from the two patterns of genes. Interestingly, enrichment of the claudins family genes was found both in the two subgroups of genes. Two distinct groups of claudins with converse expression pattern were identified. The CLDN-6, -7, -10, -19, -4, -9 were found to be highly expressed in the hESCs or liver progenitor cells. Conversely, CLDN-1, -2, -14, -3, -5, -16, -15 were found to be highly expressed in the mature hepatocytes (Fig. 1b). Considering the important roles of the claudins gene family in cancer progression, we tried to screen the aberrantly expressed claudin family genes with prognostic significance in HCC patients. We randomly divided the TCGA liver cancer project into a training cohort (TCGA-LIHC Cohort I, n = 189) and a testing cohort (TCGA-LIHC Cohort II, n = 182). The mRNA expression data and clinical information were downloaded using UCSC XENA. The demographics of these cohorts were well balanced, and the clinical-pathological information was shown in Supplementary Table S1. The relative expression of all 21 claudins gene family members was compared in the 189 HCC patients from TCGA-LIHC Cohort I and 50 para-tumor liver tissues from TCGA-LIHC project. CLDN1, CLDN2, CLDN8, CLDN10, CLDN11, CLDN12, CLDN14, CLDN16 and CLDN23 were found to be significantly down-regulated in HCC. On the contrary, CLDN6, CLDN7, CLDN15, CLDN18, CLDN19, CLDN20, CLDN22 and CLDN25 were found to be significantly up-regulated in HCC (Supplementary Table S2). Kaplan–Meier survival analysis showed that CLDN2, CLDN5, CLDN6, CLDN8, CLDN14, CLDN16, CLDN18, CLDN19, CLDN22 and CLDN25 were significantly relevant to patient overall survival (Supplementary Table S2). The Hepatic-like and Embryonic-like claudins signature were established based on both significant gene expression alteration and clinical prognosis. Taken together, CLDN2, CLDN8, CLDN14, CLDN16, which were highly expressed in mature hepatocytes and down-regulated in HCC, were defined as a “Hepatic claudins signature” (HCS) (Fig. 1c). CLDN6, CLDN18, CLDN19, CLDN25, which were highly expressed in hESCs or liver progenitor cells, were defined as an “Embryonic claudins signature” (ECS) (Fig. 1d, Supplementary Fig. S1).
Identification of two distinct groups of claudins from an in vitro hepatocyte differentiation model. a Human embryonic stem cells (hESCs) were induced to differentiate into hepatocytes along hepatic lineages. b Distribution and expression pattern of claudins in embryonic stem cells or liver progenitor cells compared with mature hepatocytes during hepatocyte differentiation. c The normalized expression of hepatic claudins signature genes (CLDN2, CLDN8, CLDN14, CLDN16) was compared between 50 para-tumor liver tissues and 189 HCC tissues from the TCGA-LIHC Cohort I. d The normalized expression of embryonic claudins signature genes (CLDN6, CLDN18, CLDN19, CLDN25) were compared between 50 para-tumor liver tissues and 189 HCC tissues from the TCGA-LIHC Cohort I. e The normalized expressions of hepatic claudins signature genes were compared between HCC patient subgroups with different tumor grade. f The normalized expressions of embryonic claudins signature genes were compared between HCC patient subgroups with different tumor grade. Independent student’s t test. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. ns not significant
The claudins signature genes associated with tumor grade in HCC
In clinical pathology, tumor grade usually represents the degree of the similarity between tumor tissues and their normal counterparts. Since the claudins were found to be involved in embryonic development, we speculate whether the specified claudin signature genes were related to tumor grade and tumor differentiation. The relative expressions of the hepatic claudins and the embryonic claudins were examined in the HCC patients with a different histological grade from the TCGA-LIHC Cohort I. As shown in Fig. 1a, e progressive decrease of hepatic claudins was found from low-grade tumors to high-grade tumors. Conversely, a progressive increase of embryonic claudins was found from low-grade tumors to high-grade tumors (Fig. 1f). These results indicated that the hepatic claudins are highly expressed in well-differentiated tumors, and the embryonic claudins are highly expressed in poorly-differentiated tumors.
Correlations of claudins signature with stem/progenitor cell markers and hepatocyte differentiation markers
Considering the two claudins signatures showed different expression pattern during liver development, HCC patients with different claudins signatures might represent different developmental characteristics. To test our hypothesis, we established score systems to quantitatively define the claudins signatures in HCC patients. Patient with overexpression (defined as the normalized expression value above the median level in the tumor tissues) of each claudins signature gene will be given “1” score. The scores for each hepatic claudins signature (HCS) genes will be added to form a “HCS score”, and scores for each embryonic claudins signature (ECS) genes will be added to form an “ECS score”. Representative hepatocyte terminal differentiation markers (G6PC, PCK1, TAT, ADH1B) and liver cancer stem cell or progenitor markers (AFP, KRT19, CLIC3, G6PC3) were examined in the TCGA database [17,18,19,20,21]. A progressive increase of hepatocyte terminal differentiation markers and decrease of liver cancer stem cell or progenitor markers could be found from the HCC patients with low HCS score to patients with high HCS score (Fig. 2a,b). Conversely, A progressive decrease of hepatocyte terminal differentiation markers and increase of liver cancer stem cell or progenitor markers could be found from the HCC patients with low ECS score to patients with high ECS score (Fig. 2c, d). In addition, Pearson’s χ2 test revealed that the claudins signatures were also significantly associated with tumor grade (HCS, p = 0.045; ECS, p = 0.026) in clinical HCC patients (Supplementary Table S6). These discoveries indicated that the claudins signatures have a strong correlation with the developmental progeny and differentiation status in HCC.
The claudins signatures were correlated with HCC differentiation. a The normalized expressions of liver cancer hepatocyte terminal differentiation markers in HCC patients with different HCS value. b The normalized expressions of liver cancer stem cell or progenitor markers in HCC patients with different HCS value. c The normalized expressions of liver cancer hepatocyte terminal differentiation markers in HCC patients with different ECS value. d The normalized expressions of liver cancer stem cell or progenitor markers in HCC patients with different ECS value. One-way ANOVA test. p value less than 0.05 was considered statistically significant
PPI network prediction and enrichment analysis of the claudins signatures
Considering the claudins family members are transmembrane proteins that involved in multiple vital cellular processes, the PPI network of claudins signatures genes and the 100 closest interaction genes were formed using STRING database in Cytoscape 3.6.1. The related genes of claudins signatures formed a comprehensive network, which was closely associated with the critical factor of embryonic development including POU5F1, SOX2, FOXA2 and SMAD2 (Supplementary Fig. S2a, Supplementary Table S3 and Supplementary Table S4). The phosphorylation status of all claudins available from the UniProt database (https://www.uniprot.org/) were listed in Supplementary Table S5). Gene ontology and enrichment analysis also revealed the signaling pathways and function categories significantly associated with claudins signatures related genes, which included pathways or biological processes closely related to embryonic development (Supplementary Fig. S2b).
Prognostic value of claudins signatures and their association with clinical-pathological features in HCC
To investigate the clinical significance of the two claudins signatures, the patients were further classified into different subgroups according to their HCS score or ECS score. The “High HCS group” was defined with HCS value greater than 2, and the “Low HCS group” was defined with HCS value less than and including 2. The same classification was also adapted to the ECS counterparts. Kaplan–Meier survival analysis showed that the “Low HCS group” had significantly worse overall survival than the “High HCS group” in the TCGA-LIHC Cohort I. The progressive decrease in mean survival time could also be found with the synchronous decrease of HCS scores in different subgroups of HCC patients (Fig. 3a). Meanwhile, the “Low ECS group” had significantly better overall survival than the “High ECS group” in the TCGA-LIHC Cohort I. Significant increase in mean survival time could also be found with the decrease of ECS scores (Fig. 3d). Cox proportional hazards regression analysis further indicated the two distinct claudins signatures as promising predictors of patient overall survival in both univariate analysis (HCS: HR = 0.406, 95% CI 0.184–0.894, p = 0.025; ECS: HR = 1.818, 95% CI 1.086–3.034, p = 0.023) and multivariate analysis (HCS: HR = 0.374, 95% CI 0.168–0.834, p = 0.016; ECS: HR = 2.365, 95% CI 1.245–4.494, p = 0.009) (Table 1).
The prognostic significance of claudins signatures genes in multiple HCC clinical cohorts. a The patients in the training cohort (TCGA-LIHC cohort I, n = 189) were divided into “High HCS group” and “Low HCS group” according to the HCS value. Kaplan–Meier survival curves of the two risk groups (upper panel) and of subgroups with different HCS value (lower panel) were plotted and the log-rank p value of the survival difference calculated between them. b Similar analysis was performed in the testing cohort (TCGA-LIHC cohort II, n = 182) (c) and validated in an independent validation cohort (LIRI-JP cohort, n = 232). d The patients in the training cohort (TCGA-LIHC cohort I, n = 189) were divided into “High ECS group” and “Low ECS group” according to the HCS value. Kaplan–Meier survival curves of the two risk groups (upper panel) and of subgroups with different HCS value (lower panel) were plotted and the log-rank p value of the survival difference calculated between them. e Similar analysis was performed in the testing cohort (TCGA-LIHC cohort II, n = 182) (f) and validated in an independent validation cohort (LIRI-JP Cohort, n = 232). p value less than 0.05 was considered statistically significant
Independent validation of claudins signatures and their clinical significance
The claudins signatures were further tested in two independent clinical cohorts for validation. Similar to the training cohort TCGA-LIHC cohort I, the claudins signatures could also significantly stratified the testing cohort TCGA-LIHC cohort II for overall survival with the HCS score (HR = 0.469, 95% CI 0.232–0.824, p = 0.008) (Fig. 3b, Table 1) and ECS score (HR = 3.128, 95% CI 1.923–5.088, p = 0.000) (Fig. 3e, Table 1). In a second independent validation LIRI-JP cohort, using the same risk score in the TCGA-LIHC cohort I, the claudins signatures were also able to significantly stratify patients for overall survival with the HCS score (HR = 0.599, 95% CI 0.373–0.962, p = 0.034) (Fig. 3c, Table 1) and ECS score (HR = 1.941, 95% CI 1.213–3.106, p = 0.006) (Fig. 3f, Table 1). Considering most of the HCC patients in Asia were associated with HBV/HCV infection, the prognostic significance of claudins signature were further examined in HCC patients with different etiologies. As shown in Table 2, univariate and multivariate cox regression analysis revealed that the claudins signature significantly predicted prognosis of HCC patients in both HBV/HCV group and non-virus group (NBNC). In addition, Pearson’s χ2 test revealed that the claudins signatures were also significantly associated with tumor grade (HCS, p = 0.030; ECS, p = 0.045) in the validation TCGA-LIHC cohort II (Supplementary Table S6). These results further suggested that two opposite newly established claudins signatures could robustly predict HCC patient’s overall survival and prognosis in multiple clinical cohorts. Logistic analysis and corresponding forest map further proved the contribution of these signatures to the evaluation of prognosis in HCC patients (Supplementary Fig. S3).
Integration of the claudins signatures in predicting the survival risk and differentiation status of HCC patients
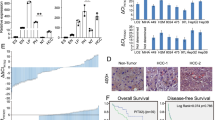
To better characterize the comprehensive claudins signature and predict the clinical outcomes of HCC patients, the two distinct claudins signatures were integrated together. An integrative score system (claudins signature score) was generated using the ECS score minus HCS score (CSS = ECS-HCS). Patients overlapped in “Low HCS group” and “High ECS group” were defined as “High risk group” and patients overlapped in “High HCS group” and “Low ECS group” were defined as “Low risk group”. Kaplan–Meier survival analysis showed that the “High risk group” had significantly worse overall survival than the “Low risk group” all in training, testing and independent validation cohorts. Moreover, the claudins signature score could also significantly stratify the overall survival of HCC patients in the training TCGA-LIHC cohort I, testing TCGA-LIHC cohort II and the independent validation cohort LIRI-JP (Fig. 4a–c). A significant negative correlation of hepatocyte terminal differentiation markers (G6PC and PCK1) with the claudins signature scores, and a significant positive correlation of HCC stem/progenitor cell markers (AFP and CLIC3) with the claudins signature scores could be found in HCC patients (Fig. 4d). ROC curve analyses revealed that the claudin gene signature (claudins signature score, CSS) has a slight advantage over serum AFP in HCC diagnosis (Fig. 4e). However, the serum AFP level could not predict the prognosis of HCC patients (Fig. 4f). To further test whether the claudins signature also applicable in HCC patients treated with targeted therapy, we extracted the sub-group of patients treated with sorafenib form the TCGA-LIHC database and performed Kaplan–Meier survival analysis according to the same criteria. As shown in Supplementary Fig. S4a, high “Embryonic claudins signature” (ECS) score significantly predicted both poor overall survival and disease-free survival of HCC patients treated with sorafenib. Accordingly, “Hepatic claudins signature” (HCS) score stratified HCC patients treated with sorafenib with better overall survival and disease-free survival but did not reach statistical significance (Supplementary Fig. S4b). When combining the ECS score and the HCS score together, the “claudins signature score” (CSS) robustly predicted both overall survival and disease-free survival of HCC patients sub-group treated with sorafenib (Supplementary Fig. S4c). In addition, Kaplan–Meier survival analysis further showed that the claudins signature score (CSS) could also significantly stratify the prognosis of HCC patients both with HCV infection and without HCV infection (Supplementary Fig. S5). These findings indicated that our newly established claudins signature could robustly represent HCC tumor developmental progeny and predict patient’s prognosis.
Integrated claudins signatures contributed to prognosis and differentiation status in multiple HCC clinical cohorts. Two groups patients “High risk group” and “Low risk group” in the training cohort (TCGA-LIHC cohort I, n = 189) were defined according to rules mentioned in the preceding part of the text. a Kaplan–Meier survival curves of the two risk groups were plotted and the log-rank p value of the survival difference calculated between them (upper panel). b Similar analysis was performed in the testing cohort (TCGA-LIHC cohort II, n = 182) (upper panel) (c) and validated in an independent validation cohort (LIRI-JP cohort, n = 232) (upper panel). The patients in the training cohort (TCGA-LIHC cohort I, n = 189) were divided into subgroups with different CSS value. a Kaplan–Meier survival curves were plotted and the log-rank p value of the survival difference calculated between them (lower panel). b Similar analysis was performed in the testing cohort (TCGA-LIHC cohort II, n = 182) (lower panel), c and validated in an independent validation cohort (LIRI-JP cohort, n = 232) (lower panel). p value less than 0.05 was considered statistically significant. d The normalized expressions of liver cancer stem cell or progenitor markers (G6PC, PCK1, AFP, CLIC3) in HCC patients with different CSS value. One-way ANOVA test. p value less than 0.05 was considered statistically significant. e ROC curve analyses were performed to examine the diagnostic value of the claudin gene signature (claudins signature score, CSS) and serum AFP in HCC patients and healthy controls from the TCGA database. f Kaplan–Meier survival curves were plotted and the log-rank P value of the survival difference calculated to test the prognostic value of serum AFP in HCC patients from the TCGA database. p value less than 0.05 was considered statistically significant
Discussion
Hepatocellular carcinoma (HCC) is one of the most common cancers and the second critical cause of cancer death all over the world, with high metastasis rate and recurrence rate [22]. Poorly differentiated tumors preserving lineage characteristics of their developmental precursor cells indicated the strong link between tumor aggressiveness and embryonic development [23]. In the present study, we established an in vitro hepatocyte differentiation model to investigate the association between liver development and carcinogenesis. Our study utilizing TCGA data and ICGC data on hepatocellular carcinoma patients had three overall objectives: (1) to obtain an initial overall view of claudins family in normal live tissues versus primary HCC samples; (2) to mine the tight junction relationship between claudins and tumor development, differentiation status and prognosis significance in HCC patients; and (3) to help provide direction for future studies of precise therapy by showcasing the efficiency of examining molecular differences and assess patients’ risk by different molecular subtypes. Through analyzing the combined data from the hepatocyte differentiation model and TCGA database, we identified two distinct claudins signatures representing different tumor developmental progeny and clinical outcome in HCC patients. Taking into account the differential expression of individual claudins family genes and their clinical associations with patient overall survival time, four claudins family members with hepatic properties were selected to form a hepatic claudins signature. Accordingly, another four claudins family members with embryonic properties were selected to form an embryonic claudins signature. To better quantify our findings, we defined a novel risk points-scoring system generated by claudins signature genes. A progressive increase of HCC stem/progenitor markers was accompanied by HCC patients of lower HCS score and higher ECS score. Conversely, hepatocyte terminal differentiation markers were found to be progressively decreased in the patients with higher HCS score and lower ECS score. The prognostic significance of the claudins signatures were verified in all there training, testing and independent validation cohorts, and the results indicated that the claudins signatures could robustly predict patients’ overall survival time.
Gain of stem cell-like properties is widely accepted as a major cancer hallmark, which substantially contributed to tumor recurrence and metastasis. From an in vitro hepatocyte differentiation model, we identified the combination of claudins family genes with the potential to predict HCC patient prognosis. We found different claudins showed unique expression patterns during liver development. Some of the claudins were highly expressed the embryonic/liver progenitor stem cells (The CLDN-6, -7, -10, -19, -4, -9), and others were highly expressed in the differentiated hepatocytes (CLDN-1, -2, -14, -3, -5, -16, -15). For the embryonic claudins signature genes, CLDN-6 was recently identified as a tumor-associated antigen and proposed to be the therapeutic target for Chimeric antigen receptor (CAR)-T cells [24]. CLDN-18 was involved in lung stem and progenitor cell homeostasis and tumorigenesis. (CAR)-T cells directed to CLDN-18 were under intensive investigation for the treatment of gastric cancer [25]. Mutation of CLDN-19 was found to disrupt retinal neurogenesis and visual function, and CLDN-25 has contributed to proper tight junction strand morphology [26]. For the hepatic claudins signature genes, silencing of CLDN-2 promoted early dissemination of lung cancer cells from primary tumors [27]. CLDN-8 was found to promote colorectal cancer cell proliferation, migration, and invasion by activating MAPK/ERK signaling [28]. EZH2-mediated silencing of CLDN-14 led to the aggressive phenotype of HCC, and provided novel prognostic biomarker [29]. CLDN-16 was also reported to be susceptibility genes involved in chemical-induced carcinogenesis in lung cancer [30]. We noticed that mRNA analysis is relatively restrictive in clinical use. We have already got evidence that the claudin (e.g. CLDN-6) is highly expressed at the protein level in HCCs, and predicted poor prognosis of the patients (unpublished data). As claudins are membrane-bound proteins, it is difficult to directly detect them in the blood. However, with the great technological advances in liquid biopsy diagnosis, we believe further isolation of circulating tumor cells from patient blood samples will become achievable. As claudins are membrane proteins, it will be possible to directly characterize the claudins signature with flow cytometry, and further predict the clinical outcome of the patients. Above all, our present findings indicated that using a combination of claudins family genes can comprehensively indicate the differentiation status of HCC patients and predict patient prognosis. With the advances of liquid biopsy technology, characterization of the circulating tumor cells with our newly identified risk points-scoring system generated by claudins signature genes will help clinical diagnosis and prognosis in HCC patients.
Abbreviations
- HCC:
-
Hepatocellular carcinoma
- RNA-seq:
-
RNA sequencing
- TCGA:
-
The cancer genome atlas
- ICGC:
-
International Cancer Genome Consortium
- hESCs:
-
Human embryonic stem cells
- ROC:
-
Receiver operating characteristic
- AUC:
-
Area under curve
- HCS:
-
Hepatic claudins signature
- ECS:
-
Embryonic claudins signature
- CSS:
-
Claudins signature scores
- PPI:
-
Protein–protein interaction
- HR:
-
Hazard ratio
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin 2015;65:87–108. https://doi.org/10.3322/caac.21262
El-Serag HB. Hepatocellular carcinoma. N Engl J Med 2011;365:1118–1127. https://doi.org/10.1056/NEJMra1001683
Bass AJ, Watanabe H, Mermel CH, Yu S, Perner S, Verhaak RG, et al. SOX2 is an amplified lineage-survival oncogene in lung and esophageal squamous cell carcinomas. Nat Genet 2009;41:1238–1242. https://doi.org/10.1038/ng.465
Yimlamai D, Christodoulou C, Galli GG, Yanger K, Pepe-Mooney B, Gurung B, et al. Hippo pathway activity influences liver cell fate. Cell 2014;157:1324–1338. https://doi.org/10.1016/j.cell.2014.03.060
Sell S. Alpha-fetoprotein, stem cells and cancer: how study of the production of alpha-fetoprotein during chemical hepatocarcinogenesis led to reaffirmation of the stem cell theory of cancer. Tumour Biol 2008;29:161–180. https://doi.org/10.1159/000143402
Ramaswamy S, Tamayo P, Rifkin R, Mukherjee S, Yeang CH, Angelo M, et al. Multiclass cancer diagnosis using tumor gene expression signatures. Proc Natl Acad Sci USA 2001;98:15149–15154. https://doi.org/10.1073/pnas.211566398
Gunzel D, Fromm M. Claudins and other tight junction proteins. Compr Physiol 2012;2:1819–1852. https://doi.org/10.1002/cphy.c110045
Lal-Nag M, Morin PJ. The claudins. Genome Biol 2009;10:235. https://doi.org/10.1186/gb-2009-10-8-235
Gonzalez-Mariscal L, Betanzos A, Nava P, Jaramillo BE. Tight junction proteins. Prog Biophys Mol Biol 2003;81:1–44. https://doi.org/10.1016/S0079-6107(02)00037-8
Kwon M. Emerging roles of claudins in human cancer. Int J Mol Sci 2013;14:18148–18180. https://doi.org/10.3390/ijms140918148
Lin G, Xie Y, Ouyang Q, Qian X, Xie P, Zhou X, et al. HLA-matching potential of an established human embryonic stem cell bank in China. Cell Stem Cell 2009;5:461–465. https://doi.org/10.1016/j.stem.2009.10.009
Xie P, Sun Y, Ouyang Q, Hu L, Tan Y, Zhou X, et al. Physiological oxygen prevents frequent silencing of the DLK1-DIO3 cluster during human embryonic stem cells culture. Stem Cells 2014;32:391–401. https://doi.org/10.1002/stem.1558
Gramignoli R, Tahan V, Dorko K, Venkataramanan R, Fox IJ, Ellis EC, et al. Rapid and sensitive assessment of human hepatocyte functions. Cell Transplant 2014;23:1545–1556. https://doi.org/10.3727/096368914X680064
Zabulica M, Srinivasan RC, Vosough M, Hammarstedt C, Wu T, Gramignoli R, et al. Guide to the assessment of mature liver gene expression in stem cell-derived hepatocytes. Stem Cells Dev 2019;28:907–919. https://doi.org/10.1089/scd.2019.0064
Fujimoto A, Furuta M, Totoki Y, Tsunoda T, Kato M, Shiraishi Y, et al. Whole-genome mutational landscape and characterization of noncoding and structural mutations in liver cancer. Nat Genet 2016;48:500–509. https://doi.org/10.1038/ng.3547
Zhou Y, Zhou B, Pache L, Chang M, Khodabakhshi AH, Tanaseichuk O, et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 2019;10:1523. https://doi.org/10.1038/s41467-019-09234-6
Su JS, Tsai TF, Chang HM, Chao KM, Su TS, Tsai SF. Distant HNF1 site as a master control for the human class I alcohol dehydrogenase gene expression. J Biol Chem 2006;281:19809–19821
Sgodda M, Aurich H, Kleist S, Aurich I, König S, Dollinger MM, et al. Hepatocyte differentiation of mesenchymal stem cells from rat peritoneal adipose tissue in vitro and in vivo. Exp Cell Res 2007;313:2875–2886
Zhang K, Zhang L, Liu W, Ma X, Cen J, Sun Z, et al. In vitro expansion of primary human hepatocytes with efficient liver repopulation capacity. Cell Stem Cell 2018;23:806–819.e4. https://doi.org/10.1016/j.stem.2018.10.018
Leonce J, Brockton N, Robinson S, Venkatesan S, Bannister P, Raman V, et al. Glucose production in the human placenta. Placenta 2006;27(Suppl A):S103–S108
Money TT, King RG, Wong MH, Stevenson JL, Kalionis B, Erwich JJ, et al. Expression and cellular localization of chloride intracellular channel 3 in human placenta and fetal membranes. Placenta 2007;28:429–436
Venook AP, Papandreou C, Furuse J, de Guevara LL. The incidence and epidemiology of hepatocellular carcinoma: a global and regional perspective. Oncologist 2010;15:5–13. https://doi.org/10.1634/theoncologist.2010-S4-05
Lee K, Lee KB, Jung HY, Yi NJ, Lee KW, Suh KS, et al. The correlation between poor prognosis and increased yes-associated protein 1 expression in keratin 19 expressing hepatocellular carcinomas and cholangiocarcinomas. BMC Cancer 2017;17:441. https://doi.org/10.1186/s12885-017-3431-1
Reinhard K, Rengstl B, Oehm P, Michel K, Billmeier A, Hayduk N, et al. An RNA vaccine drives expansion and efficacy of claudin-CAR-T cells against solid tumors. Science 2020. https://doi.org/10.1126/science.aay5967
Zhou B, Flodby P, Luo J, Castillo DR, Liu Y, Yu FX, et al. Claudin-18-mediated YAP activity regulates lung stem and progenitor cell homeostasis and tumorigenesis. J Clin Invest 2018;128:970–984. https://doi.org/10.1172/JCI90429
Wang SB, Xu T, Peng S, Singh D, Ghiassi-Nejad M, Adelman RA, et al. Disease-associated mutations of claudin-19 disrupt retinal neurogenesis and visual function. Commun Biol 2019;2:113. https://doi.org/10.1038/s42003-019-0355-0
Du W, Xu X, Niu Q, Zhang X, Wei Y, Wang Z, et al. Spi-B-mediated silencing of claudin-2 promotes early dissemination of lung cancer cells from primary tumors. Cancer Res 2017;77:4809–4822. https://doi.org/10.1158/0008-5472.CAN-17-0020
Cheng B, Rong A, Zhou Q, Li W. CLDN8 promotes colorectal cancer cell proliferation, migration, and invasion by activating MAPK/ERK signaling. Cancer Manag Res 2019;11:3741–3751. https://doi.org/10.2147/CMAR.S189558
Li CP, Cai MY, Jiang LJ, Mai SJ, Chen JW, Wang FW, et al. CLDN14 is epigenetically silenced by EZH2-mediated H3K27ME3 and is a novel prognostic biomarker in hepatocellular carcinoma. Carcinogenesis 2016;37:557–566. https://doi.org/10.1093/carcin/bgw036
Fan J, Zhu M, Wang Y, Li Z, Zhang J, Wang L, et al. Genome-wide analysis of expression quantitative trait loci identified potential lung cancer susceptibility variants among Asian populations. Carcinogenesis 2019;40:263–268. https://doi.org/10.1093/carcin/bgy165
Acknowledgements
We thank Prof. Lu and Prof. Lin’s group from the Institute of Reproductive and Stem Cell Engineering, School of Basic Medical Science, Central South University, Changsha, China, for kindly establish the in vitro hepatocyte differentiation model for us.
Funding
This work was supported by National Natural Science Foundation of China (81702400); Guangdong Province Universities and Colleges Pear River Scholar Funded Scheme (2018). The funders had no role in the design of the study and collection, analysis, and interpretation of data and in writing the manuscript.
Author information
Authors and Affiliations
Contributions
ML and N-FM initiated and designed the project; F-EK, Y-QT, Y-FG, J-QM, YZ performed statistical analyses; F-EK, Y-QT, M-ML, WC, W-JZ, H-LL performed bioinformatic analyses; S-SL and LH provided the HCC clinical samples and the relevant clinical information; N-FM, and X-YG provided valuable comments; F-EK and ML wrote the manuscript, and all authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest in this work.
Ethical approval
The study was approved by the Research Ethics Committees of Affiliated Cancer Hospital & Institute of Guangzhou Medical University. The experiments regarding the establishment of human embryonic stem cells and induced in vitro differentiation were approved and guided by the ethical committee of CITIC-Xiangya Reproductive & Genetic Hospital. The blastocysts, which were donated for the study after obtaining written informed consents from the patients undergoing assisted reproductive technology treatment, came from various clinical useless embryos, including poor quality embryos, abnormal embryos after pre-implantation genetic diagnosis (PGD), zygotes with the abnormal pronuclear number and partheno-activated embryos [11].
Informed consent
Informed consent was obtained from all patients for being included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
12072_2020_10035_MOESM1_ESM.tif
Supplementary file1 Supplementary Fig. S1 Flowchart of selecting claudins signature genes in predicting HCC prognosis (TIF 6588 kb)
12072_2020_10035_MOESM2_ESM.tif
Supplementary file2 Supplementary Fig. S2 Protein interaction of the claudins signatures genes and relevant enrichment analysis. (a) PPI network of claudins signatures genes and the 100 closest interaction genes were formed by using STRING database in Cytoscape 3.6.1. (b) Gene ontology and enrichment analysis also revealed the signaling pathways and function categories significantly associated with claudins signatures related genes, which included pathways or biological processes closely related to embryonic development (TIF 13711 kb)
12072_2020_10035_MOESM3_ESM.tif
Supplementary file3 Supplementary Fig. S3 Forest map (Logistic analysis) of claudins signatures and common clinical character in multiple HCC clinical cohorts. (a) Forest map (Logistic analysis) of hepatic and embryonic claudins signatures and other clinical characteristics in the training cohort (TCGA-LIHC Cohort I, n=189). (b) Similar analysis was presented in the testing cohort (TCGA-LIHC Cohort II, n=182) (c) and validated in an independent validation cohort (LIRI-JP Cohort, n=232). HR less than 1 was considered as low risk and HR more than 1 was considered as high risk. P value less than 0.05 was considered statistically significant. (TIF 10632 kb)
12072_2020_10035_MOESM4_ESM.tif
Supplementary file4 Supplementary Fig. S4 Kaplan-Meier survival analysis of overall survival and disease-free survival in HCC patients treated with sorafenib from the TCGA-LIHC database. (a) The sub-group of patients treated with sorafenib form the TCGA-LIHC database were extracted, and Kaplan-Meier survival analysis was performed according to the “Embryonic claudins signature” (ECS) score (b) “Hepatic claudins signature” (HCS) score, and (c) “Claudins signature score” (CSS) (TIF 946 kb)
12072_2020_10035_MOESM5_ESM.tif
Supplementary file5 Supplementary Fig. S5 Kaplan-Meier survival analysis of overall survival in HCC patients with or without HCV infection from the TCGA-LIHC database (TIF 4125 kb)
Rights and permissions
About this article
Cite this article
Kong, FE., Tang, YQ., Gong, YF. et al. Identification of prognostic claudins signature in hepatocellular carcinoma from a hepatocyte differentiation model. Hepatol Int 14, 521–533 (2020). https://doi.org/10.1007/s12072-020-10035-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12072-020-10035-z