Abstract
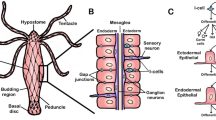
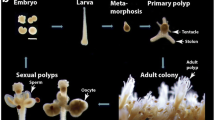
Hydra, a freshwater diploblast, with a simple but defined body plan, an organized nervous system, and the presence of stem cells, is one of the oldest model organisms used in biology. It exhibits many embryonic features even as an adult, a spectacular ability of regeneration, and lack of organismal aging. Hydra can provide insights into how complex animal forms evolved and is waiting to be better utilized in teaching.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Suggested Reading
M J Ratcliff, The Trembley effect or the birth of marine zoology, Int, J, Dev. Biol., 56, pp.425–436, 2012.
U Technau and R E Steele, Evolutionary crossroads in developmental biology, Development, 138, pp.1447–1458, 2011.
D E Martinez and D Bridge, Hydra, the everlasting embryo confronts aging, Int. J. Dev. Biol., 56, pp.479–487, 2011.
E N Browne, The production of new hydranths in hydra by the insertion of small grafts, J. Exp. Zool, 7, pp.1–23, 1909.
V Kadu, S S Ghaskadbi, and S Ghaskadbi, Induction of secondary axis in hydra revisited, Int. J. Mol. Cell. Med., 1, pp.11–20, 2012.
N Vargesson, Positional information — A concept underpinning our understanding of developmental biology, Dev. Dyn., DOI https://doi.org/10.1002/dvdy.116.
S Ghaskadbi, Cell signaling molecules in hydra: insights into evolutionarily ancient functions ofsignaling pathways, Int. J. Dev. Biol., 64, pp.141–149, 2020.
R Londhe, L S Krishnapati, and S Ghaskadbi, Description and phylogenetic characterization of hydra from Naukuchiatal (Uttarakhand, India) and comparison with other hydra strains, Curr. Sci., 113, pp.1739–1745, 2017.
V Patwardhan and S Ghaskadbi, Invertebrate alternatives for toxicity testing: Hydra stakes its claim, ALTEX Proc., 2, pp.69–76, 2013.
T Sugiyama and T Fujisava, 1977, Genetic analysis of developmental mechanisms in hydra I. Sexual reproduction of Hydra magnipapillata and isolation of mutants, Dev. Growth Diff., 19, pp. 187–200, 1977.
S Hoffmeister and H Chica Schaller, A new biochemical marker for foot-specific cell differentiation in hydra, Roux’s Dev Biol., 194, pp.453–461, 1985.
Acknowledgements
I am grateful to Vidya Patwardhan and Satyajit Rath for critical reading of the draft manuscript and suggesting changes. I thank Dr Sujata Deshpande, Ms Rohini Londhe, and Mr Ma-hadeo Daware for various inputs. Over the past two decades, my hydra laboratory has been funded by the Departments of Science & Technology (DST) and Biotechnology (DBT), Science and Engineering Research Board (SERB), Government of India, and MACS-Agharkar Research Institute. Surendra Ghaskadbi is an Emeritus Scientist of Council for Scientific and Industrial Research, New Delhi
Author information
Authors and Affiliations
Corresponding author
Additional information
Surendra Ghaskadbi is a developmental biologist particularly interested in cell signaling during pattern formation and evolution of developmental mechanisms. Over the past two decades he has reintroduced hydra as a model system for teaching and research in India.
Rights and permissions
About this article
Cite this article
Ghaskadbi, S. Hydra: A Powerful Biological Model. Reson 25, 1197–1213 (2020). https://doi.org/10.1007/s12045-020-1039-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12045-020-1039-2