Abstract
Peanut (Arachis hypogaea L.) is one of the important oil crops worldwide. Revealing the molecular basis of seed oil synthesis and analyzing key candidate genes may have important implications for enhancing peanut production. In this study, we investigated the dynamic changes in the oil accumulation rate and gene expression levels in developing peanut seeds. A Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis revealed that the up-regulated differentially expressed genes (DEGs) among three varieties at 25–35 days after flowering (DAF) were involved in fatty acid synthesis, carbon metabolism, and pyruvate metabolism. Between 25 and 35 DAF, the up-regulated DEGs in the high-oil varieties, but not in the low-oil variety, were related to acetyl-CoA synthesis. We examined the differences in the varieties at the same stages. We analyzed the lipid metabolism-related genes by protein–protein interaction (PPI), and finally identified 57 DEGs that may influence the peanut oil content. These candidate genes included three genes encoding acetyl-CoA carboxylase (ACCase), one gene encoding ketoacyl-ACP synthase III (KASIII), one gene encoding hydroxyacyl-ACP dehydrase (HAD), and one gene encoding enoyl-ACP reductase (EAR), which form the Type II fatty acid synthase complex. Additionally, we also found that 11 transcription factors may affect oil content. These identified candidate genes may help to further clarify the molecular basis of the oil accumulation in peanut seeds.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Peanut (Arachis hypogaea L.) is an important economic crop and oil crop worldwide and has an oil content as high as 40%–60% (Yol et al. 2017). Peanut characteristics include a balanced fatty acid and a substantial abundance of specific antioxidants as well as monounsaturated and polyunsaturated fatty acids (Akhtar et al. 2014). Although the peanut oil content is generally higher than that of other oil crops, there are considerable differences among peanut varieties. Several breeding programs worldwide, including in China, are currently committed to developing peanut varieties with high oil contents (Chen et al. 2016). The consumer demand for peanuts has increased as the global population has increased. Therefore, a top priority among peanut breeders and producers in China is increasing the peanut oil content and cultivating excellent peanut varieties.
With the development of molecular biology, the molecular mechanism of oil biosynthesis in plants has been gradually unveiled (Harwood et al. 2013; Weselake et al. 2009). The genes and pathways involved in lipid biosynthesis have been extensively studied in oil crops, including soybean (Bates et al. 2009; Yang et al. 2019; Zhao et al. 2019), sesames (Wang et al. 2014; Wei et al. 2015) and Brassica napus (Liao et al. 2019; Tan et al. 2018; Tang et al. 2020). The biosynthesis of plant oil, which requires a series of complex physiological and biochemical processes involving a multi-enzyme system, is one of the most important metabolic activities in plants (Hills 2004). Fatty acid biosynthesis is the first step in lipid biosynthesis, which is catalyzed by a fatty acid synthase (FAS) complex in plastids. The FAS complex, which determines the fatty acid carbon chain length, contains four enzymes, namely β-ketoacyl-ACP synthetase, β-ketoacyl-ACP reductase, β-hydroxyacyl-ACP dehydratase, and enoyl-ACP reductase (Haslam and Kunst 2013). Additionally, Acetyl-CoA carboxylase is an important rate-limiting enzyme in FA de novo synthesis, which was reported that ACCase gene expression can promote fatty acid biosynthesis (Baud et al. 2003; Li et al. 2011; Mu et al. 2008; Slabas and Fawcett 1992). Transcription factors (TFs) are important components involved in transcriptional regulatory network functioning in lipid metabolism, and they activate some key enzymes related to fatty acid biosynthesis to induce oil accumulation. The ABI3, FUS3, LEC1, and LEC2 genes encode the master regulators of gene networks governing most seed developmental mechanisms (To et al. 2006).
With the development of next-generation, high-throughput sequencing technologies, RNA sequencing (RNA-seq) is extensively used to unravel the molecular basis of lipid biosynthesis in many plant species (Chen et al. 2019; Gupta et al. 2016; Shahid et al. 2019; Wang et al. 2019; Yang et al. 2019; Yin et al. 2013). Transcriptome analyses via RNA-seq are commonly performed to compare gene expression among diverse peanut species. For example, an earlier transcriptome analysis determined the lipid-related gene expression patterns in developing peanut seeds as well as the molecular mechanism underlying lipid metabolism, while also revealing the often rate-limiting enzymes for lipid storage in peanuts with varying oil contents (Yin et al. 2013). Furthermore, the transcriptome analysis indicated that the genetically controlled podding period is critical for peanut seed development. The identification of differentially expressed genes (DEGs) related to oil synthesis may clarify the genotypic differences in peanut oil biosynthesis and pod filling, thereby elucidating the genetic basis of peanut seed development and oil accumulation (Gupta et al. 2016). Notably, more than 2,500 genes related to oil metabolism were identified during genome and transcriptome analyses in 2019. Most of the genes underwent expression-level changes in the early seed developmental stage and were unexpressed during maturation (Chen et al. 2019).
In this study, the seed oil content was determined for different developmental stages of three peanut varieties (including two high-oil varieties and one low-oil variety) to examine oil accumulation in peanut seeds. The transcriptomes of the samples collected at 15, 25, 35, 45, 55, 65, and 85 days after flowering (DAF) were sequenced using RNA-seq to determine the transcriptome dynamic changes in developing peanut seeds and to identify key genes that determine seed oil content in peanut seeds. This study provides insights into the molecular mechanisms underlying oil accumulation and the factors determining seed oil contents in peanut.
Results
Dynamic Changes to the Oil Content in Developing Peanut Seeds
This study was performed using the following three peanut genotypes with varying oil contents: W191 (Ha, high oil variety a; 58% oil content), YH15 (Hb, high oil variety b; 54% oil content) and JT1 (L, low oil variety; 44% oil content). The oil content of the peanut seeds was determined between 35 and 95 DAF. Seven seed developmental stages corresponding to the major events occurring during the seed development were analyzed, including the pod expansion and formation stage, the full pod stage, and the late pod ripening stage (Fig. 1a). More specifically, 35–45 DAF corresponded to the pod expansion and formation stage (pod filling), 45–65 DAF corresponded to the early and mid-maturation stages (seed filling), and 65–95 DAF corresponded to the late maturation stage (seed desiccation). The seed oil contents in the three genotypes began to increase as early as 35 DAF and continued to increase until 55 DAF (Fig. 1b). Thus, the period between 35 and 55 DAF was critical for the rapid accumulation of oil. After 55 DAF, oil accumulated at a slower rate. There were significant differences between the high- and low-oil varieties in the same period (Fig. 1b, Supplementary Table 1).
Differences in the seed phenotypes at various developmental stages among the peanut cultivars. a Phenotypes of W191 (Ha), YH15 (Hb), and JT1 (L) seeds at different developmental stages (35–95 DAF). b Dynamic changes of the total lipid content of three peanut varieties. Data are means of three repeats in all cases and error bars represent SD (n = 3)
Global Transcriptome Analysis of Peanut Seeds
On the basis of the changes to the oil accumulation of peanut seeds with high and low oil contents, we selected six seed developmental stages for further analyses, including the main stages: pod expansion or embryo stage (25–45 DAF), seed filling stage (45–65 DAF), late dehydration and ripening stage (85 DAF), and pods with minute embryos at 25 DAF. We also examined the aerial pegs at 15 DAF. The materials we selected for analyses cover the important stages for seed development and oil accumulation, enabling the characterization of the molecular mechanism underlying these processes. The transcriptome sequence was assembled using Trinity v2.0.6, and the data for individual RNA-seq libraries were aligned to the Tifrunner.gnm2.J5K5 reference genome (www.Peanutbase.org). In this study, all samples were analyzed using the BGISEQ-500 platform, with an average of 6.42 Gb data generated per sample. The average rate of alignment to the genome for the samples was 93.43%, and the average alignment rate for the gene set was 74.51%. A total of 7,789 genes were predicted. Moreover, 60,289 expressed genes were detected, of which 52,620 and 7,669 genes were known and predicted, respectively. A total of 68,083 new transcripts were detected, of which 52,034 were new alternative splicing subtypes of known protein-coding genes, 7,789 were new protein-coding transcripts, and the remaining 8,260 were long-chain non-coding RNAs. The number of genes with high (FPKM > 10), moderate (1 < FPKM < 10), and low (FPKM < 1) expression levels was similar among the different varieties during the three stages between 15 and 35 DAF (Fig. 2a). Interestingly, between 45 and 85 DAF, more highly expressed genes (FPKM > 10) were detected in the low-oil variety (JT1, L) than in the two high-oil varieties (W191, Ha; YH15, Hb), and the number of genes increased as the JT1 (L) seeds developed. The Pearson correlation coefficient (PCC) for the FPKM of biological replicates of the same variety and same stage samples was 0.91–0.99, which indicates that the data were highly reproducible (Supplementary Table 2). The tissues/stages that were highly correlated were expected to have relatively similar transcriptomes and functions/activities (Fig. 2b). These analyses reflected a sufficient transcriptome coverage over the peanut seed developmental period and confirmed the assembled transcriptome was appropriate for the subsequent analysis.
Global transcriptome analysis of three peanut genotypes (Ha: W191; Hb: YH15; L: JT1) at seven developmental stages (15, 25, 35, 45, 55, 65, and 85 DAF). a The stacked histogram of gene expression in seven developmental stages of three peanut varieties. The x axis represents the sample name, the y axis represents the number of genes, and different colors indicate different levels of expression. b Pearson correlation coefficient (PCC) analysis of the RNA-seq data for seven developmental stages of three peanut varieties
Analysis of Differential Gene Expression during Seed Development
A functional analysis of genes was completed to identify the enriched biological processes and activities during different developmental stages for the three peanut genotypes. We used Venn diagrams to identify the unique and common differentially expressed genes at adjacent developmental stages for three cultivars. A total of 2,058, 2,049, 317, 12, 73 and 916 common DEGs were detected with up-regulated expression levels, whereas 2,678, 8,872, 441, 17, 122 and 423 common DEGs were identified with down-regulated expression levels during the 15 vs 25 DAF, 25 vs 35 DAF, 35 vs 45 DAF, 45 vs 55 DAF, 55 vs 65 DAF, and 65 vs 85 DAF comparisons for the three genotypes, respectively (Fig. 3a, b). An analysis of the enriched gene ontology (GO) terms and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways, which was completed to functionally characterize the DEGs, revealed that the common and up-regulated DEGs at 25–35 DAF were significantly related to lipid metabolism (fatty acid biosynthetic process, fatty acid metabolic process, and glycerolipid metabolic process), carbon metabolism, and pyruvate metabolism (Supplementary Fig. 1a, b). These findings combined with the phenotypic data (Fig. 1b) of the differences in the oil contents at 35 DAF indicated that this time-point may be critical for oil accumulation. The common and up-regulated DEGs at 35–45 DAF were significantly associated with cotyledon development, seed development, carpel development, and embryo development (Supplementary Fig. 1b). The phenotypic data indicated that the seeds rapidly increased in size at 35–45 DAF (Supplementary Fig. 2). The transcription group analysis explained this developmental feature at the molecular level.
Analysis of differentially expressed genes in various seed developmental stages for three peanut varieties. a The number of up- and down-regulated differentially expressed genes were identified in W191 (Ha), YH15 (Hb), and JT1 (L) (|log2 fold-change|> 1, corrected p-value < 0.05). b Venn diagrams presenting the number of differentially expressed genes during consecutive developmental stages for each variety. c Gene ontology (GO) enrichment analysis of consecutive developmental stages for three peanut varieties. The color scale at the top represents significance (corrected p-value)
A functional enrichment analysis of the unique DEGs between the high- and low-oil varieties at each stage revealed that the up-regulated DEGs at 25–35 DAF were significantly involved in the fatty acid biosynthetic process, fatty acid metabolic process, glycerolipid metabolic process, acetyl-CoA biosynthetic process, and lipid biosynthetic process in the two high-oil varieties (Ha, W191 and Hb, YH15) (Fig. 3c). In the low-oil variety (L, JT1), the unique and up-regulated DEGs at 25–35 DAF were significantly related to fatty acid degradation, fatty acid alpha-oxidation, and fatty acid metabolic process (Fig. 3c). The unique and up-regulated DEGs at 35–45 DAF were significantly associated with fatty acid metabolic process and lipid metabolic process in the two high-oil varieties (Fig. 3c). Conversely, the unique and up-regulated DEGs at 35–45 DAF were significantly involved in the regulation of fatty acid beta-oxidation, the regulation of fatty acid oxidation, the fructose metabolic process, and peroxisome organization in the low-oil variety (L, JT1) (Fig. 3c). Accordingly, the reason for the lower oil content of the low-oil variety (L, JT1) may be that fatty acid synthesis processes were no longer dominating but fatty acid oxidation during the 35–45 DAF of seed developmental. In W191 (Ha), the unique and down-regulated DEGs at 45–55 DAF were significantly related to fatty acid alpha-oxidation and lipid oxidation. In YH15 (Hb), the unique and down-regulated DEGs at 45–55 DAF were significantly involved in the fatty acid biosynthetic process (Fig. 3c). These findings may help to explain why the oil content was higher in W191 (Ha) than in YH15 (Hb) after 55 DAF (Fig. 1b). In JT1 (L), the unique and down-regulated DEGs at 45–55 DAF were significantly associated with the lipid biosynthetic process. In the last stage (65–85 DAF), the common down-regulated DEGs were significantly involved in photosynthesis, fatty acid biosynthesis, fatty acid metabolism, and glycerolipid metabolism (Supplementary Fig. 1a). From the transcriptome level, it was confirmed that the oil accumulation rate was basically unchanged in the later stage of seed ripening. On the basis of above results, we speculated that 35 DAF and 45 DAF was two crucial time-point for the oil accumulation leading to the oil content differences among the examined varieties.
Differentially Expressed Gene Sets between Peanut Varieties
To further identify the key genes affecting the differences between the high- and low-oil varieties, DEGs at the same developmental stages were analyzed (Fig. 4a, b). After eliminating the duplicated DEGs, 21,513 and 11,418 DEGs were respectively identified in the Ha vs L and Hb vs L comparisons (Fig. 4a, b), with each gene significantly differentially expressed (i.e., |log2 fold-change|> 1, corrected p-value < 0.05) in at least one period. A subsequent KEGG pathway classification analysis revealed that the 21,513 and 11,418 DEGs included 840 and 503 genes related to lipid metabolism, respectively (Fig. 4c, d, Supplementary Table 3 and Table 4). A Venn diagram analysis indicated that 376 DEGs (Fig. 4e, Supplementary Table 5) were shared. An additional PPI analysis and screening of 376 DEGs detected 57 highly correlated genes (Supplementary Table 6). A further cluster analysis of the 57 DEGs confirmed that the differences between the high- and low-oil varieties were highly significant (Fig. 5c and Supplementary Table 7). The expression levels of 36 and 21 genes were respectively up- and down-regulated in the high-oil varieties (Ha and Hb), relative to the expression levels in the low-oil variety (L).
Analysis of differentially expressed genes (DEGs) in the comparisons of Ha (W191) vs L (JT1) and Hb (YH15) vs L (JT1). a The number of up- and down-regulated genes in Ha (W191) relative to the corresponding expression levels in L (JT1), at each seed developmental stage. b The number of up- and down-regulated genes in Hb (YH15) relative to the corresponding expression levels in L (JT1), at each seed developmental stage. c KEGG pathway classification of differentially expressed genes in Ha vs L. d KEGG pathway classification of differentially expressed genes in Hb vs L. e Venn diagram analysis indicating 376 lipid-related common DEGs
Expression analysis of 57 lipid-related DEGs in the comparisons at each developmental stage of Ha (W191) vs L (JT1) and Hb (YH15) vs L (JT1). KEGG enrichment analysis of 21 up-regulated a and 36 down-regulated b lipid-related DEGs. Heatmap c of the relative expression of 57 lipid-related DEGs in Ha, Hb, and L at each developmental stage
The enriched Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways among the up-regulated DEGs (Fig. 5a) were as follows: fatty acid biosynthesis (36% of the genes), fatty acid metabolism (36% of the genes), pyruvate metabolism (22% of the genes) and carbon metabolism (16% of the genes). The enriched KEGG pathways among the down-regulated DEGs (Fig. 5b) were as follows: glycerophospholipid metabolism (38% of the genes), fatty acid degradation (24% of the genes), and glycerolipid metabolism (24% of the genes). On the basis of the gene ontology (GO) enrichment analysis of the up-regulated genes (Supplementary Fig. 3a), 53% were enriched in lipid metabolic process (GO:0,006,629), 47% were enriched in lipid biosynthetic process (GO:0,008,610), 31% were enriched in fatty acid biosynthetic process (GO:0,006,633), 36% were enriched in cellular lipid metabolic process (GO:0,044,255), 11% were enriched in acetyl-CoA carboxylase activity (GO:0,003,989). Regarding the down-regulated DEGs (Supplementary Fig. 3b), the top20 enriched GO terms include “lipid catabolic process (GO:0,016,042)”. Overall, the up-regulated DEGs comparing high-oil varieties (Ha and Hb) with low-oil (L) were mainly related to fatty acid biosynthesis and lipid metabolic process, and down-regulated DEGs were not only related to fatty acid biosynthesis and lipid metabolic process, but also related to fatty acid degradation and lipid catabolic process. The expression characteristics of these genes may be responsible for the low oil content of L (JT1). On the basis of these findings, we identified ACCase genes (Arahy.70V125, Arahy.KBA53Y, and Arahy.L14V1E), KASIII gene (Arahy.U0ISBZ), HAD gene (Arahy.KP3Y1K), and ER gene (Arahy.KJ0WPF) as candidate genes involved in determination of the seed oil content.
Identification of Transcription Factor Families in Peanut
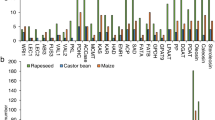
Many transcription factors (TFs) are reported to be important for lipid biosynthesis. In our study, 3,655 putative TF-encoding genes belonging to 58 major TF families were analyzed in peanut (Supplementary Table 10). The TFs potentially involved in oil synthesis were from the ABI3VP1 (291 TFs), AP2-EREBP (253 TFs), and C2C2-Dof (70 TFs) families. Of these TFs 11 DEGs were significantly down-regulated in JT1 (L) in at least one development stage and regarded as candidate genes for high oil content in peanut. The significance levels of these 11 TF genes of the TF families ABI3VP1 (Arahy.XD3PB3, Arahy.6FQ15B, Arahy.54SH3C), AP2-EREBP (Arahy.WY3QF4, Arahy.DRB5KP, Arahy.804PS6, Arahy.GP7L65, Arahy.45NR7C, BGI_novel_G002657), and C2C2-Dof (Arahy.5HWL40, BGI_novel_G001230) were shown in a heatmap (Supplementary Fig. 4 and Supplementary Table 13).
Analysis of the Expression of DEGs at the mRNA Level
To validate whether the differences in the RNA-seq data for the DEGs accurately reflected the transcription levels in vivo, a qRT-PCR assay was conducted to determine the DEG expression levels in the three peanut cultivars at four critical periods. The 2−ΔΔCt method was used to analyze the expression data. The relative expression levels for the DEGs (Fig. 6), including the candidate genes Arahy.KP3Y1K, Arahy.70V125, Arahy.KBA53Y, Arahy.L14V1E, Arahy.U0ISBZ, Arahy.KJ0WPF, and Arahy.GJWA3D, were correlated with the dynamic changes in the RNA-seq data (FPKM values). Details regarding the qRT-PCR primers were provided in Supplementary Table 11.
qRT-PCR verification of DEGs in three varieties. Transcript levels of 18 different genes including the candidate genes a Arahy.H4HC75 (accC), b Arahy.KBA53Y (accB), c Arahy.70V125 (accA), e Arahy.L14V1E (ACACA), g Arahy.U0ISBZ (KASIII/FabH), h Arahy.KP3Y1K (HAD/FabZ), and i Arahy.KJ0WPF (ER/FabI). The y-axis shows the relative gene expression levels of RNA-Seq (left) and qRT-PCR (right). FPKM of RNA-seq are shown as columns and qRT-PCR as lines: Ha (red), Hb (green) and L (blue). The qRT-PCR expression data correspond a 2 − ΔΔCT fold change. Data are means of three repeats in all cases and error bars represent SD (n = 3)
Discussion
The seed oil content is an important characteristic that considerably affects the efficiency of peanut production. Breeding new varieties with high oil contents is currently one of the main goals of peanut breeders. To explore the dynamic changes to the oil content in different peanut varieties and to clarify the underlying mechanisms, we measured the seed oil contents of high- and low-oil peanut varieties during seven seed developmental stages. According to the characteristics of oil accumulation and seed development in peanut, seven key periods were selected for transcriptome sequencing and analysis.
In this study, the seed oil accumulation dynamics in three peanut varieties with different genetic backgrounds were basically the same. Oil accumulated in peanut seed kernels at 35 DAF, and the oil content at this time-point was 54.0%–66.4% of that in mature seeds. Since then, the oil content of 45 DAF and 55 DAF seeds increased rapidly. The oil content of 55 DAF seeds reached 92.5%-99.2% of that of mature seeds, reaching the peak period of oil synthesis. The oil accumulation rate was basically unchanged after 65 days and reached the maximum value in the mature stage (Fig. 1b). This is consistent with the results of previous studies (Pickett 1950). The RNA-seq technique was used to determine the transcriptome changes in three peanut genotypes (with varying oil contents) at different developmental stages in this study. We analyzed the differences in consecutive seed developmental stages in peanut varieties with different genetic backgrounds. The identified DEGs were functionally characterized. The common and up-regulated DEGs at 25–35 DAF were significantly related to lipid metabolism (fatty acid biosynthetic process, fatty acid metabolic process, and glycerolipid metabolic process), carbon metabolism, and pyruvate metabolism (Supplementary Fig. 1a, b). These findings combined with the phenotypic data (Fig. 1b) of the differences in the oil contents at 35 DAF indicated that this time-point may be critical for oil accumulation. Moreover, the unique and up-regulated DEGs of the high-oil varieties (Ha and Hb) at 25–35 DAF were significantly involved in the fatty acid biosynthetic process, fatty acid metabolic process, glycerolipid metabolic process, acetyl-CoA biosynthetic process, and lipid biosynthetic process, whereas the unique and up-regulated DEGs of the low-oil variety were not significantly involved in acetyl-CoA biosynthetic process, which may explain the low oil content of L at 35 DAF (Fig. 3c). Because acetyl-CoA carboxylase is the key rate-limiting enzyme for fatty acids synthesis, which are the key raw materials for TAG synthesis. The unique and up-regulated DEGs at 35–45 DAF were significantly associated with fatty acid metabolic process and lipid metabolic process in the two high-oil varieties (Fig. 3c). However, the unique and up-regulated DEGs of the low-oil variety (L) at 35–45 DAF were associated with fatty acid oxidation and lipid catabolism (Fig. 3c), which have a certain competitive relationship with lipid accumulation. Previous studies confirmed that preventing lipid catabolism positively affects lipid accumulation and leads to increased oil contents (Fan et al. 2013; Kong et al. 2017; Slocombe et al. 2009; Trentacoste et al. 2013). In oilseed rape (Brassica napus L.), Kelly et al. (2013) found that SUGAR-DEPENDENT1 triacylglycerol (SDP1) lipase family may be involved in oil catabolism during seed development, and shown that using an SDP1 RNAi strategy can result in up to a approximately 8% increase in oil yield on a per seed, per plant or a per unit planted area basis, in the greenhouse, with relatively little adverse impact on seed vigour. Based on the analysis of the above phenotypic and transcriptome data, we speculated that 35 DAF and 45 DAF was two crucial time-point for the oil accumulation leading to the oil content differences among the examined varieties.
During a comparative transcriptome analysis of two high-oil varieties (Ha and Hb) and a low-oil variety (L), we detected 840 and 503 DEGs related to the lipid metabolism pathway, respectively. Interestingly, we found that there were more DEGs in carbohydrate metabolism than in lipid metabolism (Fig. 4c, d). It may be that carbohydrate metabolism provide carbon sources for fatty acid synthesis (Bates et al. 2013). Additionally, the 376 common DEGs (including 48 novel genes) reflected the similarities in the examined two high-oil varieties, which were integrated into pathway of lipid biosynthesis (Fig. 7). We found that the lipid-related differential genes in high-oil varieties (Ha and Hb) compared with the low-oil variety (L) were up-regulated in FA de novo synthesis. We identified 57 DEGs that were highly correlated among the 376 genes according to a PPI analysis (https://string-db.org/), of which the up-regulated DEGs in the high-oil varieties (Ha and Hb) compared with the low-oil variety (L) were associated with the fatty acid synthesis and lipid metabolism (Fig. 5a and Supplementary Fig. 3a), whereas the down-regulated DEGs were involved in fatty acid degradation pathway and lipid catabolic process (Fig. 5b and Supplementary Fig. 3b), which was consistent with the results of a previous analysis (Fig. 3c). According to the above analysis, we identified genes of encoding ACCase (Arahy.70V125, Arahy.KBA53Y, and Arahy.L14V1E). The observed ACCase genes expression patterns were essentially the same in the two high-oil varieties, with expression levels that were significantly higher than that in the low-oil variety (L) (Fig. 7). This implies that genes of encoding ACCase enhance the fatty acid content, and is closely related to the seed oil content. The relative expression levels of the ACCase genes determined by qRT-PCR (Fig. 6) were basically consistent with the corresponding FPKM values (RNA-seq data). Page et al. (1994) who first demonstrated the importance of acetyl-CoA carboxylase in controlling lipid biosynthesis. The expression of ACCase promotes fatty acid biosynthesis, which positively regulate the oil content (Baud et al. 2003; Klaus et al. 2004; Li et al. 2011; Mu et al. 2008; Slabas and Fawcett 1992). The genes of encoding KASIII/FabH (Arahy.U0ISBZ), HAD/FabZ (Arahy.KP3Y1K), and ER/FabI (Arahy.KJ0WPF) were preliminarily screened based on the transcriptome and protein–protein interaction analyses in this study. The expression levels of the three genes were verified by qRT-PCR, with the resulting data basically consistent with the FPKM values. In plants, fatty acid chain lengths and saturation are determined by the Type II FAS complex in plastids (Yu et al. 2015). The Type II FAS responsible for de novo fatty acid synthesis functions synergistically with ACCase (Yasuno et al. 2004). The FAS complex comprises the following four enzymes: β-ketoacyl-[ACP]-synthetase (KASI, II, or III), β-ketoacyl-[ACP]-reductase (KAR), β-hydroxyacyl-[ACP]-dehydrase (HAD), and enoyl-[ACP]-reductase (ER). These enzymes are involved in the condensation, reduction, dehydration, and re-reduction of fatty acids, respectively. The genes encoding the Type II FAS complex enzymes in peanut were previously cloned and verified (Li et al. 2009). The KASIII gene, which encodes the initial enzyme mediating fatty acid chain elongations, catalyzes the condensation reaction involving malonyl-ACP and acetyl-CoA to yield ketobutyryl-ACP. In spinach homogenates, KASIII activity is correlated with the FAS rate, and KASIII may have a rate-limiting role in FAS (Jaworski et al. 1989). In 1990, some researchers reported that KASIII gradually plays an important role in fatty acid synthesis (Walsh et al. 1990).
Lipid biosynthesis pathway including de novo fatty acid synthesis and elongation, and TAG synthesis. A total of 87 DEGs were identified from 376 lipid-related DEGs, which were placed in the pathway. The first seven columns represent the seven time points of Ha vs L (from left to right: 15–65DAF and 85DAF) and the last seven columns represent the seven time points of Hb vs L (from left to right: 15–65DAF and 85DAF), and the rows indicates the different lipid-related DEGs, which were placed near the enzyme name to show the differences in the expression of enzyme-encoding genes at different stages. Each block represents the relative expressions of DEGs using the log2 fold change of relative intensity in high-oil varieties (Ha and Hb) compared with low-oil variety (L). Red represents up-regulation of genes, while blue represents down-regulation of genes
There is increasing evidence that many TFs participate in lipid synthesis, during which they activate some key enzymes related to oil biosynthesis to induce oil accumulation (Baud and Lepiniec 2010; Courchesne et al. 2009). The ABI3, FUS3, -LEC1, and LEC2 genes encode the master regulators of gene networks governing most seed developmental mechanisms (To et al. 2006; Wang and Perry 2013). A recent study confirmed that ABI3 is an important regulator of oil accumulation and revealed for the first time the direct correlation between ABI3 overexpression and increased lipid contents in plants (Pouvreau et al. 2020). The main effects of LEC1, LEC2, and FUS3 on lipid accumulation are due to their direct or indirect regulation of the expression of the WRINKLED1 (WRI1) TF gene (Marchive et al. 2014). The WRI1 TF gene belongs to the AP2/EREBP family (Cernac and Benning 2004; Riechmann et al. 2016). This TF directly regulates the accumulation and storage of oil in seeds and mediates the production of enough oil for developing seeds (Cernac et al. 2006). The overexpression of WRI1 in A. thaliana can increase the fatty acid content, but also regulates pyruvate kinase, ACCase, the biotin carboxyl carrier protein, and enoyl-ACP reductase, which affect glycolysis and the fatty acid biosynthesis pathway (Baud et al. 2009; Courchesne et al. 2009; Liu et al. 2010; Maeo et al. 2009). The protein encoded by the WRI1a gene can actively regulate oil accumulation in soybean (Chen et al. 2018a). Other studies demonstrated that Dof (DNA binding with one finger), which is a plant-specific TF, regulates fatty acid synthesis (Diaz et al. 2002; Ibanez-Salazar et al. 2014). The 28 Dof genes in soybean are involved in the regulation of oil synthesis. The overexpression of GmDof4 and GmDof11 can increase the expression of genes encoding ACCase and long-chain acetyl-CoA synthetase, while also increasing the total fatty acid and lipid contents in transgenic A. thaliana seeds (Wang et al. 2007). The fatty acid synthesis-related TF genes identified in this study belonged to the ABI3VP1, AP2-EREBP, and C2C2-Dof families, with 291, 253, and 70 predicted genes, respectively. Among these genes, the expression levels of 11 genes differed significantly among the varieties, including three ABI3VP1 genes, six AP2-EREBP genes, and two C2C2-Dof genes (Supplementary Fig. 4).
The mechanism underlying the oil formation and accumulation in peanut and other oil crops is very complex because of the joint regulation of a series of genes and the fact, that it is affected by both genetic factors and external factors. The rapid development and subsequent application of molecular biology research techniques have enabled in-depth investigations of the enzyme systems associated with the oil biosynthesis pathway, which have revealed the general mechanism regulating the oil biosynthesis in oil crops. However, peanut is an amphidiploid plant species and its large genome (> 2.5 GB) complicate molecular studies. In this study, we identified 57 candidate genes encoding the enzymes and 11 TFs affecting oil accumulation during the seed developmental in peanut varieties with high and low oil contents. These genes were analyzed in terms of their effects on the oil content and their expression levels and may play a major role in the determination of the final oil content in peanut varieties. Further studies should confirm the results of this study, while also examining protein expression levels, protein modifications, and enzyme activities.
Conclusions
We analyzed the oil accumulation and transcriptome dynamic process during the seed development of three peanut varieties. We speculated that two crucial time-points, 35 and 45 DAF, were responsible for the oil content differences among the examined varieties. We finally identified 57 candidate genes that affected peanut oil content. The up-regulated lipid DEGs of comparing two high-oil varieties with a low-oil variety were significantly related to fatty acid synthesis, of which three DEGs encoded acetyl CoA, and three DEGs belonged to KASIII, HAD and ER in type II FAS. We combined phenotypic data and differential expression analysis to reveal the molecular basis of peanut seed oil synthesis, which lay a foundation to further understand the seed oil accumulation process and provide candidate genes for molecular breeding to improve peanut seed oil content.
Materials and Methods
Plant Materials and Sample Collection
Three peanut varieties, W191 (Ha, high oil; 58% oil content), YH15 (Hb, high oil; 54% oil content), and JT1 (L, low oil; 44% oil content), with varying mature seed oil contents were used in this study. The three varieties were provided by the Henan Academy of Agricultural Sciences, Zhengzhou, China. The plants were grown at the Henan Modern Agricultural Research and Development Base. Each variety was planted with 10 rows, 40 plants per row, and the row spacing was 40 cm, which was managed uniformly in the field. During the early stage of flowering, undesirable pollination was prevented via artificial bud removal. When the plants reached the full flowering stage, the flowers were maintained on plants for 3 days and marked with a string. After that, the new buds were removed for 15 days to prevent undesirable pollination and ensure the accuracy and uniformity of the samples. After the gynophores entered the soil and developed into pods, we collected the seeds at different time-points (from 25 to 95 DAF). Additionally, we also selected and examined the aerial pegs at 15 DAF to analyze the transcriptome activity before pods formed. We selected freshly harvested seeds with similar phenotypes at the same developmental stages for analyzing the total oil contents (three repeats of each ten uniform pods per sample) and extracting RNA. All samples collected for the transcriptome analysis were immediately frozen in liquid nitrogen and then stored at -80 °C.
Analysis of the Seed Lipid Content
For each developmental stage, 4 g seeds per biological repetition were collected for an analysis of oil contents. The dried seeds were ground to a fine powder with a grinder, after which the Soxhlet extraction- residue method was used to quantify the oil content: (1) The weight of the filter paper dried to constant weight is M1, and the crushed sample is 1–2 g with the filter paper dried to constant weight. The sample package was dried in an oven at 105 ± 2 °C for 3 h, and the quality of the sample package M2 was weighed after cooling. (2) The sample package was placed in an extraction tube with anhydrous diethyl ether, and the sample package was completely dipped with anhydrous diethyl ether. The next day, the soaked sample bag was put into the extraction bottle and several glass balls. In addition, anhydrous diethyl ether was re-added to the extraction tube to make the sample package completely soaked. Connect all parts of the instrument, turn on the condensed water, then heat it in a water bath at 70–80 °C, reflux and extract for 8 h. (3) Take out the sample package and place it in a ventilated place to make the ether volatilize completely. The sample package was dried in an oven at 105 ± 2 °C for 2 h, then removed, and the residual package M3 was weighed after cooling. \(\mathrm{Oil content }(\mathrm{\%})=\frac{M2-M3}{M2-M1}\times 100\%\). GraphPad Prism v.7.0 was used to compute analyses of variances (ANOVA) followed by Tukey’s multiple range comparison tests; differences at p < 0.05 were considered statistically significant.
Total RNA Isolation, Library Construction, and Transcriptome Sequencing
Total RNA was isolated using the TaKaRa MiniBEST Plant RNA Extraction Kit (Takara, Japan). The minimum total amount of RNA used for the RNA-seq analysis was 3 µg, and the RNA concentration calibrated to 50 ng/µL for all samples. The RNA quality was checked using the 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). The RNA used for the subsequent analyses satisfied the following criteria: OD260/OD280 ≥ 1.8, 28S/18S ≥ 1, and RNA integrity number ≥ 7. The RNA-seq raw data have been submitted to the NCBI database (BioProject with the SRA accession number: PRJNA685603).
Bioinformatic Analysis of Transcriptome Data
Raw reads were filtered with SOAPnuke v1.4.0 (Chen et al. 2018b ). Specifically, reads containing adapters, reads with poly-N sequences, and low-quality reads were eliminated to obtain the clean reads, which were then assembled with Trimmomatic v0.36. The clean reads were mapped to the reference genome using HISAT2 v2.1.0 (Kim et al. 2015). Additionally, Bowtie2 v2.2.5 (Langmead and Salzberg 2012) was used to align the clean reads to the reference coding gene set, after which gene expression levels were calculated with RSEM v1.2.8 (Li and Dewey 2011). A Perl script was used to calculate the length of the reference transcriptional group sequence. The transcripts of each sample were reconstructed using StringTie v1.0.4 (http://ccb.jhu.edu/software/stringtie). The reconstruction information for all samples was integrated with Cuffmerge (http://cole-trapnell-lab.github.io/cufflinks/), after which the integrated transcripts were compared with the reference notes using Cuffcompare. The transcripts with class code types ‘u’, ‘i’, ‘o’, and ‘j’ were defined as new transcripts. The protein coding potential of the new transcripts was predicted using CPC v0.9-r2 (http://cpc.cbi.pku.edu.cn). Finally, the new transcripts with a predicted protein coding potential were added to the reference gene sequence to complete the reference sequence information for subsequent analyses. Genes were functionally characterized by screening available databases (e.g., NR and KEGG) using the BLAST algorithm v2.2.23. Blast2GO v2.5.0 and NR annotations were used to assign GO terms to genes. The PCCs for the biological replicates were calculated using the cor function of the R software. Moreover, DEseq2 (Love et al. 2014) was used to detect significant DEGs, which satisfied the following criteria: fold-change > 2 or < − 2 and an adjusted p-value < 0.05. The GO and KEGG pathway enrichment analyses were performed using Phyper, which is a function of the R program. The significance of the GO terms and KEGG pathways was corrected using p-value, with a rigorous threshold (corrected p-value < 0.05).
Quantitative Real-Time PCR Analysis
Total RNA was extracted using the TaKaRa MiniBEST Plant RNA Extraction Kit (Takara), after which 1 µg total RNA was used as the template for the synthesis of first-strand cDNA using the Reverse Transcription System (Promega, USA). A qRT-PCR assay was completed using the PowerUp™ SYBR™ Green Master Mix (ABI, USA). Alcohol dehydrogenase class III (ADH3) (Forward Primer: 5′-GACGCTTGGCGAGATCAACA-3′, Reverse Primer: 5′-AACCGGACAACCACCACATG-3′) was selected as the internal reference control. The PCR program was as follows: 50 ℃ for 2 min; 95 ℃ for 2 min; 40 cycles of 95 ℃ for 15 s and 60 ℃ for 1 min. A melting curve analysis was performed to verify primer specificity (95 ℃ for 15 s, 60 ℃ for 1 min, and 95 ℃ for 15 s).
Abbreviations
- ACCase:
-
Acetyl-CoA carboxylase
- DAF:
-
Days after flowering
- DEG:
-
Differently expressed gene
- ER:
-
Enoyl-ACP reductase
- FAD2:
-
Fatty acid desaturase 2
- FAS:
-
Fatty acid synthase
- FPKM:
-
Fragments per kilobase per million
- HAD:
-
Hydroxyacyl-ACP dehydrase
- KAR:
-
Ketoacyl-ACP reductase
- KASI:
-
Ketoacyl-ACP synthase I
- KASII:
-
Ketoacyl-ACP synthase II
- KASIII:
-
Ketoacyl-ACP synthase III
- MCMT:
-
Malony-CoA: ACP malonyltransferase
- PPI:
-
Protein–Protein Interaction
- RNA-seq:
-
RNA sequencing
- TAG:
-
Triacylglycerol
- TFs:
-
Transcription factors
References
Akhtar S, Khalid N, Ahmed I et al (2014) Physicochemical characteristics, functional properties, and nutritional benefits of peanut oil: a review. Crit Rev Food Sci Nutr 54(12):1562–1575. https://doi.org/10.1080/10408398.2011.644353
Bates PD, Durrett TP, Ohlrogge JB et al (2009) Analysis of acyl fluxes through multiple pathways of triacylglycerol synthesis in developing soybean embryos. Plant Physiol 150:55–72. https://doi.org/10.1104/pp.109.137737
Bates PD, Stymne S, Ohlrogge J (2013) Biochemical pathways in seed oil synthesis. Curr Opin Plant Biol 16(3):358–364
Baud S, Guyon V, Kronenberger J et al (2003) Multifunctional acetyl-CoA carboxylase 1 is essential for very long chain fatty acid elongation and embryo development in Arabidopsis. Plant J 33(1):75–86. https://doi.org/10.1046/j.1365-313X.2003.016010.x
Baud S, Wuille`me S, To A, et al (2009) Role of WRINKLED1 in the transcriptional regulation of glycolytic and fatty acid biosynthetic genes in Arabidopsis. Plant J 60(6):933–947. https://doi.org/10.1111/j.1365-313X.2009.04011.x
Baud S, Lepiniec L (2010) Physiological and developmental regulation of seed oil production. Prog Lipid Res 49(3):235–249. https://doi.org/10.1016/j.plipres.2010.01.001
Cernac A, Andre C, Hoffmann-Benning S et al (2006) WRI1 is required for seed germination and seedling establishment. Plant Physiol 141(2):745–757. https://doi.org/10.1104/pp.106.079574
Cernac A, Benning C (2004) WRINKLED1 encodes an AP2/EREB domain protein involved in the control of storage compound biosynthesis in Arabidopsis. Plant J 40(4):575–585. https://doi.org/10.1111/j.1365-313X.2004.02235.x
Chen L, Zheng YH, Dong ZM et al (2018a) Soybean (Glycine max) WRINKLED1 transcription factor, GmWRI1a, positively regulates seed oil accumulation. Mol Genet Genomics 293(2):401–415. https://doi.org/10.1007/s00438-017-1393-2
Chen XP, Li HJ, Pandey MK et al (2016) Draft genome of the peanut A-genome progenitor (Arachis duranensis) provides insights into geocarpy, oil biosynthesis, and allergens. P Natl Acad Sci USA 113(24):6785–6790. https://doi.org/10.1073/pnas.1600899113
Chen XP, Lu Q, Liu H et al (2019) Sequencing of Cultivated Peanut, Arachis hypogaea, Yields Insights into Genome Evolution and Oil Improvement. Mol Plant 12(7):920–934. https://doi.org/10.1016/j.molp.2019.03.005
Chen YX, Chen YS, Shi CM et al (2018b) SOAPnuke: a MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. GigaScience 7(1):1–6. https://doi.org/10.1093/gigascience/gix120
Courchesne NM, Parisien A, Wang B et al (2009) Enhancement of lipid production using biochemical, genetic and transcription factor engineering approaches. J Biotechnol 141:31–41. https://doi.org/10.1016/j.jbiotec.2009.02.018
Diaz I, Vicente-Carbajosa J, Abraham Z et al (2002) The GAMYB protein from barley interacts with the DOF transcription factor BPBF and activates endosperm-specific genes during seed development. Plant J 29(4):453–464. https://doi.org/10.1046/j.0960-7412.2001.01230.x
Fan JL, Yan CS, Zhang XB et al (2013) Dual role for phospholipid:diacylglycerol acyltransferase: enhancing fatty acid synthesis and diverting fatty acids from membrane lipids to triacylglycerol in Arabidopsis leaves. Plant Cell 25(9):3506–3518. https://doi.org/10.1105/tpc.113.117358
Gupta K, Kayam G, Faigenboim-Doron A et al (2016) Gene expression profiling during seed-filling process in peanut with emphasis on oil biosynthesis networks. Plant Sci 248:116–127. https://doi.org/10.1016/j.plantsci.2016.04.014
Harwood JL, Ramli US, Tang MG et al (2013) Regulation and enhancement of lipid accumulation in oil crops: The use of metabolic control analysis for informed genetic manipulation. Eur J Lipid Sci Tech 115(11):1239–1246. https://doi.org/10.1002/ejlt.201300257
Haslam TM, Kunst L (2013) Extending the story of very-long-chain fatty acid elongation. Plant Sci 210:93–107. https://doi.org/10.1016/j.plantsci.2013.05.008
Hills MJ (2004) Control of storage-product synthesis in seeds. Curr Opin Plant Biol 7(3):302–308. https://doi.org/10.1016/j.pbi.2004.03.003
Ibanez-Salazar A, Rosales-Mendoza S, Rocha-Uribe A et al (2014) Over-expression of Dof-type transcription factor increases lipid production in Chlamydomonas reinhardtii. J Biotechnol 184:27–38. https://doi.org/10.1016/j.jbiotec.2014.05.003
Jaworski JG, Clough RC, Barnum SR (1989) A Cerulenin Insensitive Short Chain 3-Ketoacyl-Acyl Carrier Protein Synthase in Spinacia oleracea Leaves. Plant Physiol 90:41–44. https://doi.org/10.2307/4272031
Kelly AA , Shaw E, Powers SJ et al (2013) Suppression of the SUGAR-DEPENDENT1 triacylglycerol lipase family during seed development enhances oil yield in oilseed rape (Brassica napus L.). Plant Biotechnol J 11(3):355–361. https://doi.org/10.1111/pbi.12021
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12(4):357–360. https://doi.org/10.1038/nmeth.3317
Klaus D, Ohlrogge JB, Neuhaus HE et al (2004) Increased fatty acid production in potato by engineering of acetyl-CoA carboxylase. Planta 219(3):389–396. https://doi.org/10.1007/s00425-004-1236-3
Kong FT, Liang YX, Legeret B et al (2017) Chlamydomonas carries out fatty acid beta-oxidation in ancestral peroxisomes using a bona fide acyl-CoA oxidase. Plant J 90(2):358–371. https://doi.org/10.1111/tpj.13498
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9(4):357–359. https://doi.org/10.1038/nmeth.1923
Liao P, Woodfield HK, Harwood JL et al (2019) Comparative transcriptomics analysis of Brassica napus L. during seed maturation reveals dynamic changes in gene expression between embryos and seed coats and distinct expression profiles of Acyl-CoA-Binding proteins for lipid accumulation. Plant Cell Physiol 60(12): 2812–2825. https://doi.org/10.1093/pcp/pcz169
Li B, Dewey CN (2011) RSEM, accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12(1):323. https://doi.org/10.1186/1471-2105-12-323
Li MJ, Li AQ, Xia H et al (2009) Cloning and sequence analysis of putative type II fatty acid synthase genes from Arachis hypogaea L. J Biosciences 34(2):227–238. https://doi.org/10.1007/s12038-009-0027-1
Li ZG, Yin WB, Song LY et al (2011) Genes encoding the biotin carboxylase subunit of acetyl-CoA carboxylase from Brassica napus and parental species: cloning, expression patterns, and evolution. Genome 54(3):202–211. https://doi.org/10.1139/G10-110
Liu J, Hua W, Zhan GM et al (2010) Increasing seed mass and oil content in transgenic Arabidopsis by the overexpression of wri1-like gene from Brassica napus. Plant Physiol Bioch 48(1):9–15. https://doi.org/10.1016/j.plaphy.2009.09.007
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Maeo K, Tokuda T, Ayame A et al (2009) An AP2-type transcription factor, WRINKLED1, of Arabidopsis thaliana binds to the AW-box sequence conserved among proximal upstream regions of genes involved in fatty acid synthesis. Plant J 60(3):476–487. https://doi.org/10.1111/j.1365-313X.2009.03967.x
Marchive C, Nikovics K, To A et al (2014) Transcriptional regulation of fatty acid production in higher plants: Molecular bases and biotechnological outcomes. Eur J Lipid Sci Tech 116(10):1332–1343. https://doi.org/10.1002/ejlt.201400027
Mu J, Tan H, Zheng Q et al (2008) LEAFY COTYLEDON1 is a key regulator of fatty acid biosynthesis in Arabidopsis. Plant Physiol 148(2):1042–1054. https://doi.org/10.1104/pp.108.126342
Page RA et al (1994) Biochim Biophys Acta 1210:369–372
Pickett TA (1950) Composition of developing peanut seed. Plant Physiol 25(2):210–224. https://doi.org/10.1104/pp.25.2.210
Pouvreau B, Blundell C, Vohra H et al (2020) A versatile high throughput screening platform for plant metabolic engineering highlights the major role of ABI3 in lipid metabolism regulation. Front Plant Sci 11:288. https://doi.org/10.3389/fpls.2020.00288
Riechmann JL, Heard J, Martin G et al (2016) Arabidopsis Transcription Factors: Genome-Wide comparative analysis among eukaryotes. Science 290:2105–2110. https://doi.org/10.1126/science.290.5499.2105
Shahid M, Cai GQ, Zu F et al (2019) Comparative transcriptome analysis of developing seeds and silique wall reveals dynamic transcription networks for effective oil production in Brassica napus L. Int J Mol Sci 20(8):1982. https://doi.org/10.3390/ijms20081982
Slabas AR, Fawcett T (1992) The biochemistry and molecular biology of plant lipid biosynthesis. Plant Mol Biol 19(1):169–191. https://doi.org/10.1007/BF00015613
Slocombe SP, Cornah J, Pinfield-Wells H et al (2009) Oil accumulation in leaves directed by modification of fatty acid breakdown and lipid synthesis pathways. Plant Biotechnol J 7(7):694–703. https://doi.org/10.1111/j.1467-7652.2009.00435.x
Tan HL, Zhang JH, Qi X et al (2018) Correlation analysis of the transcriptome and metabolome reveals the regulatory network for lipid synthesis in developing Brassica napus embryos. Plant Mol Biol 99:31–44. https://doi.org/10.1007/s11103-018-0800-3
Tang S, Zhao H, Lu SP et al (2020) Genome- and transcriptome-wide association studies provide insights into the genetic basis of natural variation of seed oil content in Brassica napus. Mol Plant 14:1–18. https://doi.org/10.1016/j.molp.2020.12.003
To A, Valon C, Savino G et al (2006) A network of local and redundant gene regulation governs Arabidopsis Seed Maturation. Plant Cell 18(7):1642–1651. https://doi.org/10.1105/tpc.105.039925
Trentacoste EM, Shrestha RP, Smith SR et al (2013) Metabolic engineering of lipid catabolism increases microalgal lipid accumulation without compromising growth. P Natl Acad Sci USA 110(49):19748–19753. https://doi.org/10.1073/pnas.1309299110
Walsh MC, Klopfenstein WE, Harwood JL (1990) The short chain condensing enzyme has a widespread occurrence in the fatty acid synthetases from higher plants. Phytochemistry 29(12):3797–3799. https://doi.org/10.1016/0031-9422(90)85334-C
Wang FF, Perry SE (2013) Identification of direct targets of FUSCA3, a key regulator of Arabidopsis seed development. Plant Physiol 161(3):1251–1264. https://doi.org/10.1104/pp.112.212282
Wang HW, Zhang B, Hao YJ et al (2007) The soybean Dof-type transcription factor genes, GmDof4 and GmDof11, enhance lipid content in the seeds of transgenic Arabidopsis plants. Plant J 52(4):716–729. https://doi.org/10.1111/j.1365-313X.2007.03268.x
Wang LH, Yu S, Tong CB et al (2014) Genome sequencing of the high oil crop sesame provides insight into oil biosynthesis. Genome Biol 15(2):R39. https://doi.org/10.1186/gb-2014-15-2-r39
Wang LH, Zhang YX, Li DH et al (2019) Gene expression profiles that shape high and low oil content sesames. BMC Genet 20(1):1982. https://doi.org/10.1186/s12863-019-0747-7
Wei X, Liu KY, Zhang YX, Feng Q et al (2015) Genetic discovery for oil production and quality in sesame. Nat Commun 6:8609. https://doi.org/10.1038/ncomms9609
Weselake RJ, Taylor DC, Rahman MH et al (2009) Increasing the flow of carbon into seed oil. Biotechnol Adv 27(6):866–878. https://doi.org/10.1016/j.biotechadv.2009.07.001
Yang SN, Miao L, He JB et al (2019) Dynamic transcriptome changes related to oil accumulation in developing soybean seeds. Int J Mol Sci 20(9):2202. https://doi.org/10.3390/ijms20092202
Yasuno R, von Wettstein-Knowles P, Wada H (2004) Identification and molecular characterization of the beta-ketoacyl-[acyl carrier protein] synthase component of the Arabidopsis mitochondrial fatty acid synthase. J Biol Chem 279(9):8242–8251. https://doi.org/10.1074/jbc.M308894200
Yin DM, Wang Y, Zhang XG et al (2013) De novo assembly of the peanut (Arachis hypogaea L.) seed transcriptome revealed candidate unigenes for oil accumulation pathways. PLoS One 8(9):e73767. https://doi.org/10.1371/journal.pone.0073767
Yol E, Ustun R, Golukcu M et al (2017) Oil content, oil yield and fatty acid profile of groundnut germplasm in mediterranean climates. J Am Oil Chem Soc 94(6):787–804. https://doi.org/10.1007/s11746-017-2981-3
Yu N, Xiao WF, Zhu J et al (2015) The Jatropha curcas KASIII gene alters fatty acid composition of seeds in Arabidopsis thaliana. BIOL PLANTARUM 59:773–782. https://doi.org/10.1007/s10535-015-0555-5
Zhao JZ, Bi RR, Li SX et al (2019) Genome-wide analysis and functional characterization of AcylCoA:diacylglycerol acyltransferase from soybean identify GmDGAT1A and 1B roles in oil synthesis in Arabidopsis seeds. J Plant Physiol 242:1–9. https://doi.org/10.1016/j.jplph.2019.153019
Acknowledgements
This work was supported by China Agriculture Research System (CARS-13), Henan Provincial Agriculture Research System, China (S2012-5), Science-Technology Foundation for Outstanding Young Scientists of Henan Academy of Agricultural Sciences (Grant no. 2020YQ08), Fund for Distinguished Young Scholars from Henan Academy of Agricultural Sciences (2019JQ02), Henan Provincial Young Talents Supporting Project (2020HYTP044). The funding agencies played no role in the design of the study and collection, analysis, and interpretation of data and in writing the manuscript. The authors also thank for editorial assistance.
Author information
Authors and Affiliations
Contributions
JW analyzed transcriptome data, made pictures and wrote manuscripts. FYQ offered plant materials, conceived the idea for the work and revised the manuscript. XYZ provided funding and reviewed the manuscript. ZZ, ZQS, MDT, XW, BYH, WZD did critical revision of the manuscript.
Corresponding author
Ethics declarations
Research Involving Human and Animal Participants
The research does not involve human and/or animal experimentation.
Conflict of Interest
There are no conflicts of interest.
Additional information
Communicated by: Chiedozie Egesi
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, J., Qi, F., Zheng, Z. et al. Global Transcriptome Analyses Provide Into Several Fatty Acid Biosynthesis-related Genes in Peanut (Arachis hypogaea L.). Tropical Plant Biol. 14, 267–282 (2021). https://doi.org/10.1007/s12042-021-09285-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12042-021-09285-4