Abstract
Neurodegenerative diseases constitute a large proportion of disorders in elderly, majority being sporadic in occurrence with \(\sim \)5–10% familial. A strong genetic component underlies the Mendelian forms but nongenetic factors together with genetic vulnerability contributes to the complex sporadic forms. Several gene discoveries in the familial forms have provided novel insights into the pathogenesis of neurodegeneration with implications for treatment. Conversely, findings from genetic dissection of the sporadic forms, despite large genomewide association studies and more recently whole exome and whole genome sequencing, have been limited. This review provides a concise account of the genetics that we know, the pathways that they implicate, the challenges that are faced and the prospects that are envisaged for the sporadic, complex forms of neurodegenerative diseases, taking four most common conditions, namely Alzheimer’s disease, Parkinson’s disease, amyotrophic lateral sclerosis and Huntington disease as examples. Poor replication across studies, inability to establish genotype–phenotype correlations and the overall failure to predict risk and/or prevent disease in this group poses a continuing challenge. Among others, clinical heterogeneity emerges as the most important impediment warranting newer approaches. Advanced computational and system biology tools to analyse the big data are being generated and the alternate strategy such as subgrouping of case–control cohorts based on deep phenotyping using the principles of Ayurveda to overcome current limitation of phenotype heterogeneity seem to hold promise. However, at this point, with advances in discovery genomics and functional analysis of putative determinants with translation potential for the complex forms being minimal, stem cell therapies are being attempted as potential interventions. In this context, the possibility to generate patient derived induced pluripotent stem cells, mutant/gene/genome correction through CRISPR/Cas9 technology and repopulating the specific brain regions with corrected neurons, which may fulfil the dream of personalized medicine have been mentioned briefly. Understanding disease pathways / biology using this technology, with implications for development of novel therapeutics are optimistic expectations in the near future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Neurodegenerative disorders are a collective term used to include a number of diseases such as Alzheimer, Parkinson, multiple system atrophy, progressive supranuclear palsy, dementia with Lewy bodies, amyotrophic lateral sclerosis, Huntington disease, prion disease, spinocerebellar ataxia, spinal muscular atrophy, neuronal ceroid lipofuscinoses, Pick’s disease, frontotemporal dementia etc. All these conditions are characterized by neuronal degeneration and consequent altered functions in different regions across the central and peripheral nervous system with symptoms manifesting in the respective target tissues. Different parts of the brain are comprised of specialized neurons such as dopaminergic, cholinergic, serotoninergic, glutamatergic among others, alterations in which lead to different disease phenotypes (figure 1). Loss of specific neurons either due to intrinsic (genetic) or extrinsic (injury, infection etc.) factor is generally an irreversible process and contribute to a range of pathological features. Underlying these disorders are a common set of clinical features such as involuntary movement and relentless progress in disease (Bertram and Tanzi 2005). Yet another shared feature of all these conditions are that they are generally late onset and sporadic in occurrence with only a small but varying proportion being familial. A strong genetic component is undisputed in the aetiology of the latter group which is also generally characterized by comparatively earlier age of disease onset. However, based on the classical twin/adoption studies and epidemiological surveys, nongenetic factors together with genetic vulnerability are believed to contribute to the more abundant sporadic forms, thus justifying their inclusion under complex disease category.
Conventional genetic analysis tools for linkage and contemporary approaches of next-generation sequencing performed using familial forms of most of these disorders have contributed notably to the discovery of several putative disease causal genes. These have provided novel insights into the pathogenesis of neurodegeneration and the field is still evolving at a dramatic pace and is expected to greatly influence the diagnosis and treatment in future (Pihlstrøm et al. 2017). Conversely, findings from genetic dissection of the common sporadic, complex disease forms have been limited, warranting use of additional strategies. Thus, uncovering additional risk conferring genes/loci may provide newer insights into disease biology with implications for improved/novel therapeutics. This review aims to provide a concise account of the genetics that we know, the pathways that they implicate, the challenges that are faced and the prospects that are envisaged for the sporadic, complex forms of neurodegenerative diseases, taking four most common conditions namely Alzheimer’s disease (AD), Parkinson’s disease (PD), amyotrophic lateral sclerosis (ALS) and Huntington disease (HD) as examples. It may be relevant to mention here that HD is a monogenic disorder, but due to its exceptional inheritance pattern, caused by as yet unclear modifier gene effects, it has been included under complex disease forms. A brief account of the disease per se and the available genetic data are provided separately for each of these four pathologies in the earlier part of this review and the challenges and prospects being more or less similar across these conditions are discussed together in the subsequent section. Major similarities and unique features of these four neurodegenerative disorders are also tabulated in table 1 in electronic supplementary material at http://www.ias.ac.in/jgenet/.
Alzheimer’s disease
AD is a chronic progressive neurodegenerative disease that is responsible for around 60 to 70% cases of dementia. AD is generally a disease in elderly population with a prevalence of 10–30% in the population >65 years of age, and an incidence of 1–3% affecting 20–30 million people worldwide with the incidence shooting up from 0.5% per year in 65–75 age group to 6–8% per year in age group 85 and above (Masters et al. 2015). Late onset sporadic cases comprise >95% of disease burden and <1% are familial, showing an autosomal dominant mode of inheritance and notably a much earlier age of onset cognitive deficits in AD is insidious in onset and progressive in course (Scheltens et al. 2016). Memory impairment, specifically the loss of memory of recent events, is the most frequent feature of AD and is usually its first manifestation followed by deficits in other cognitive domains. Executive dysfunction and impaired visuospatial skills are affected relatively earlier than language function and behavioural symptoms. Noncognitive neurologic deficits (pyramidal and extrapyramidal motor signs and seizures) can occur in late stages of AD. The speed of progression is variable but the average life expectancy after diagnosis is three to nine years.
In AD, there is neuronal cell death in wide areas of the cerebral cortex and hippocampus, starting with the neurons of the frontal and temporal lobes, gradually progressing to other areas of the neocortex at rates that vary considerably between individuals. Pathologically, it is marked by accumulation of insoluble forms of amyloid-\(\upbeta \) (A\(\upbeta )\) as plaques in extracellular spaces, as well as in the walls of blood vessels and aggregation of the microtubule protein tau in neurofibrillary tangles in neurons (Masters et al. 2015). The clinical diagnosis in AD is challenging as comorbidities such as cerebrovascular disease and hippocampal sclerosis are seen in patients. The development of Pittsburgh compound B (PiB)—a C11 radioactive analogue of fluorescent amyloid dye thioflavin T62 has helped in A\(\upbeta \) imaging in patients by positron emission tomography (PET) brain scans. Greater accuracy in diagnosis has been possible with the development of an advanced tracer florbetaben F18. Another method for diagnosis is CSF collection by lumbar puncture from AD patients and profiling for protein biomarkers A\(\upbeta \)42, total tau (T-tau) and phosphorylated tau (P-tau); and the most commonly used assay is specific for P-tau at Thr181 (Olsson et al. 2016).
Familial occurrence together with a greater risk of individuals of developing AD among those with first degree relatives with disease were suggestive of a genetic component in disease aetiology which was confirmed by heritability estimates for AD ranging between 57 and 78% (Meyer and Breitner 1998). Twin studies showing 32.2% concordance between monozygotic and 8.7% between dizygotic twins have revealed gene–environment interactions in the aetiology of AD with genetic determinants being probably lesser contributors (Pedersen et al. 2004). Average life span after disease onset is 8–10 years and there is no treatment available for AD. Cholinesterase inhibitors like donepezil, rivastigmine and galantamine are used to treat the cognitive symptoms (memory loss, confusion and hampered thought processes) associated with AD. Memantine (glutamate antagonist) provides some benefit for patients with moderate to severe dementia. New treatment modalities are urgently needed to prevent, delay or treat the symptoms of AD. Researchers are currently focussing on anti-amyloid approaches (including active and passive immunization strategies), \(\upgamma \)-secretase and \(\upbeta \)-secretase inhibitors and anti-aggregation drugs.
Genetic risk factors
The discovery of A\(\upbeta \) peptides in brains of both AD patients and persons with Down’s syndrome was one of the earliest clues that mutation in a gene on chromosome 21 might also cause AD in persons without Down’s syndrome (Glenner and Wong 1984a, b). As in PD, causal gene discovery was more plausible using the familial forms. Linkage analysis in families with AD identified Amyloid-\(\upbeta \) A4 protein precursor (APP) gene at 21q21.3 coding for the A\(\upbeta \) peptide precursor as a promising candidate (Hardy 2006). Specific mutations in APP seen in early onset AD (EOAD) patients confirmed its role in disease aetiology (Chartier-Harlin et al. 1991) but mutations in APP could not explain all the forms of familial EOAD and thus the search for additional genes/loci continued. Linkage analysis performed in more families discovered two more genes, namely presenilin-1 (PSEN1) (Sherrington et al. 1995) and presenilin-2 (PSEN2) (Levy-Lahad et al. 1995). As of date, a total of 52,241 and 45 mutations are reported in APP, PSEN1 and PSEN2, respectively (listed at http://www.alzforum.org/mutations) accounting for \(\sim \)5–10% of EOAD families. Identification of mutations has not only provided important insights into molecular mechanisms and pathways involved in AD pathogenesis but has also led to the identification of druggable targets (Van Cauwenberghe et al. 2016). In contrast, no mutations in these genes have been identified in the sporadic late onset AD (LOAD) cases. Environmental attributes have been notably elusive and genetic component, even if small, is probably complex and heterogeneous.
APOE (19q13.32) is well-documented and strongly associated risk conferring gene with AD. The presence of APOE immunoreactivity in A\(\upbeta \) amyloid deposits and neurofibrillary tangles linked APOE to AD pathology for the first time (Namba et al. 1991; Wisniewski and Frangione 1992). APOE encodes a glycoprotein involved in catabolism of triglyceride-rich lipoproteins, transport of cholesterol and other lipids, and also in neuronal growth, repair response to tissue injury, nerve regeneration, immune regulation and activation of lipolytic enzymes. APOE contains three major alleles, \(\upvarepsilon 2, \upvarepsilon 3\) and \(\upvarepsilon 4\) with varying frequencies across different ethnic groups. For example, it is 0.07 for APOE \(\upvarepsilon \)2, 0.78 for APOE \(\upvarepsilon \)3 and 0.15 for APOE \(\upvarepsilon \)4 in Americans of European descent (Saunders et al. 1993) versus 0.04% for APOE \(\upvarepsilon \)2, 0.89% for APOE \(\upvarepsilon \)3 and 0.07% for APOE \(\upvarepsilon \)4 in a north Indian cohort (Thelma et al. 2001). APOE2 (Cys112 and Cys158), APOE3 (Cys112 and Arg158) and APOE4 (Arg112 and Arg158) isoforms differ in the total charge present on the respective proteins and their structure which leads to altered binding to both cellular receptors and lipoprotein particles, and possibly changing the stability and rate of production and clearance. APOE binds to A\(\upbeta \) and effectuates the clearance of soluble A\(\upbeta \) and A\(\upbeta \) aggregations, and APOE \(\upvarepsilon \)4 is thought to be less efficient in mediating A\(\upbeta \) clearance and deposition, ultimately contributing to plaque formation (Strittmatter et al. 1993). APOE \(\upvarepsilon \)4 which increases the risk of developing AD in familial and sporadic EOAD and LOAD (with one allele imparting a threefold increase in risk and two alleles a 12 fold increase), is also associated with an earlier age of onset of AD and reported to contribute to \(\sim \)50% of sporadic AD (Strittmatter et al. 1993). On the other hand, APOE \(\upvarepsilon \)2 decreases the risk of developing AD and also delays the age of onset. Human APOE isoforms have been shown to cause isoform dependent decreases (APOE2>APOE3>APOE4) in neuritic plaque load and delayed time of onset of A\(\upbeta \) deposition in several mouse models of AD (Holtzman et al. 1999, 2000). Collectively, these studies indicate that APOE is a major causative or contributing factor for AD pathogenesis. Also studies have shown that T-tau and P-tau levels are increased in both brains and CSF of patients with AD which indicate the role of tau protein in AD pathology. A total of 107 mutations in MAPT (Tau) has been reported in LOAD as of today (http://www.alzforum.org/mutations).
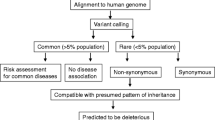
In addition to APOE and MAPT as major susceptibility genes, >20 risk genes/loci for AD have been reported, mostly based on GWASs performed on cohorts of Caucasian ancestry in the last decade. Of note, APOE locus has been the most consistently associated across population groups of which all but three were case–control analysis (table 1). With limited insights from these studies, similar to most of the other studies in complex traits, emphasis shifted from common disease common variant (CDCV) to common disease rare variant (CDRV) hypothesis, the latter facilitated by the emergence of next-generation sequencing (NGS) technologies. Rare variants are expected to have larger size effect than GWAS loci, are easier to characterize functionally and to develop cellular and animal models of disease. Rare variants thus identified in sporadic AD patients by whole exome sequencing (WES) are seen in favourite candidate genes, namely APOE and APP as well as in five other genes of biological relevance (table 2). However, only APOE, APP and SORL1 are identified in GWAS. Meta-analysis of GWASs, identified genes that are mostly involved in endosomal vesicle cycling, immune response and cytoskeletal functions. Importantly, several of them are also involved in A\(\upbeta \) clearance or tau mediated toxicity (Van Cauwenberghe et al. 2016). Taken together, all these findings are still insufficient to explain the complete pathological spectrum of disease, facilitate effective disease risk prediction/prevention thus encouraging continuing efforts.
Parkinson’s disease
The second most common neurodegenerative disorder is PD, which results from specific loss of dopaminergic neurons in the substantial nigra pars compacta in midbrain (Fearnley and Lees 1991) leading to dopamine deficiency. Intraneuronal protein aggregates largely comprised of \(\upalpha \)-synuclein and commonly referred to as Lewy bodies are the pathological signature of PD (Vekrellis et al. 2004). The symptoms generally appears slowly over time. Early in the disease, the most obvious are tremors, rigidity, slowness of movement, and difficulty with walking. Dementia becomes common in the advanced stages of the disease. Depression and anxiety are also common in patients with PD. Other nonmotor symptoms include sensory, sleep and emotional problems. The diagnosis is made using UK brain bank criteria (Hughes et al. 1992) and clinical assessment is done using Movement Disorder Society Unified Parkinson’s Disease Rating Scale (MDS-UPDRS). Exclusion criteria for PD is elaborate and includes history of repeated strokes with stepwise progression of Parkinsonian features, history of repeated head injury, history of definite encephalitis, 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP) exposure, negative response to large doses of levodopa, supranuclear gaze palsy and presence of a cerebral tumour. Assessment of the extent of neuronal damage is done by neuroimaging techniques including PET, single-photon emission CT (SPECT) and novel MRI techniques. Unfortunately, there is no cure for PD and only symptomatic relief is generally achieved by levodopa-based therapies. Motor features which are initially manageable with these therapies worsen with disease progression together with emergence of complications including motor and nonmotor fluctuations, dyskinesia, and psychosis (Hely et al. 1999; Pandey and Srivanitchapoom 2017). Surgery involving placement of microelectrodes for deep brain stimulation has been used to reduce motor symptoms in severe cases where drugs are ineffective (Martinez-Ramirez et al. 2015), but its precise mechanism is unknown (Pandey and Sarma 2015). PD is largely an age dependent disability with a global prevalence of \(\sim \)1% in population >65 years of age and 4–5% at or after 85 years of age (Twelves et al. 2003; Savica et al. 2016) with approximately double the number of males affected (Van Den Eeden et al. 2003). Most PD cases are sporadic in occurrence but \(\sim \)10% of PD cases also exist as Mendelian forms. A small proportion of cases develop PD before the age of 40 years and thus are grouped under EOPD (Van Den Eeden et al. 2003) and another smaller category is comprised of juvenile PD with disease onset at <20 years of age. The prevalence of these two categories varies across ethnic groups (Pringsheim et al. 2014).
Genetic risk factors
As mentioned above, majority of PD cases are sporadic and early evidence of MPTP damaging the nigrostriatal dopaminergic neurons through glial activation were suggestive of a significant environmental attribute (Watanabe et al. 2005). Twin studies in an American cohort showing a concordance of 15.5% in monozygotic twins compared to 11.1% in dizygotic twins substantiated a genetic component as well (Tanner et al. 1999) and this was more or less replicated in a Swedish study (Wirdefeldt et al. 2011). As a consequence of the prolonged lifespan of neurons, it is believed that complex gene–environment interactions generate stress and ultimately results in cell death (Gu et al. 2005). But as in all the other common complex traits, elucidating the genetic contribution to complex forms of PD aetiology has been exigent. To start with, their genetic analysis was limited to a few candidate gene based association studies. Genes from the dopaminergic pathway and those coding for phases 1 and 2 drug metabolising enzymes were the obvious choice based on pharmacological evidence from successful levodopa treatment and the classical MPTP induced animal model of PD suggesting the role of environmental toxicants (Kopin et al. 1986). However, most of these observed associations were not replicated across studies and ethnic groups, and were far from being of any predictive value. On the other hand, genetic analysis of large PD kindred and smaller independent PD families using classical and contemporary approaches of linkage analysis and NGS respectively, have identified >18 PARK genes/loci namely SNCA, PARKIN, SPR, UCHL1, PINK1, DJ-1, LRRK2, ATP13A2, GIGYF2, HTRA2, PLA2G6, FBXO7, ADORA1, VPS35, EIF4G1, DNAJC13, CHCHD2, TMEM230, VPS13C, PTRHD1, PODXL and RIC3 with the last two genes discovered recently in our laboratory (Sudhaman et al. 2016a, b). These PARK genes identified to date have provided significant insights into cellular and molecular pathways involved in PD aetiology thus providing likely leads for identification of novel drug targets/improved therapeutics. Collectively these findings implicate aberrant protein folding, protein degradation, neuro-inflammation, neuro-developmental defects and mitochondrial homeostasis with the involvement of oxidative stress for disease pathology. At this time, it is relevant to note that although recessive forms of PD have a few clinical features that are distinct from typical sporadic PD, such as early onset, severe dystonia, stable cognition and early motor fluctuations (Khan et al. 2005), they are indistinguishable at the pathological level which may suggest the involvement of similar pathways malfunctioning and leading to disease progression. In fact, it was this knowledge that encouraged the extension of association studies of Mendelian genes such as SNCA, LRRK2, PARKIN and PINK1 etc. with sporadic PD cases (Oliveira et al. 2003; Groen et al. 2004; Mellick et al. 2005; Paisán-Ruíz et al. 2005). This marked the beginning of the marginal progress that has been witnessed in genetics of complex forms of PD, with significant associations of common SNPs in these four genes being consistently replicated across studies (Lesage and Brice 2012). This however, could not explain total disease heritability. It was only then that the GWAS approach was employed to revisit sporadic complex forms of PD to identify genomewide susceptibility genes/loci. More than 33 GWAS, mostly in Caucasian populations have been conducted to date. These studies have unravelled significantly associated common variants in >24 independent genes/loci and recent meta-analyses have revealed a few additional genes, namely SIPA1L2, INPP5F, MIR4697, GCH1, VPS13C, DDRGK1, ITPKB, SCN3A, SATB1, IL1R2 etc. (Nalls et al. 2014; Chang et al. 2017) as shown in table 3. However, it is notable that associations have been consistently replicated at only four of these genes, namely GBA, SNCA, MAPT and LRKK2 in independent case–control (Trotta et al. 2012; Mao et al. 2013; Campêlo et al. 2017) and genomewide association studies (GWAS) (Nalls et al. 2014). It may be relevant here to briefly describe the most consistently replicated linkage and/or GWAS based genes which may lend support to the true contribution of these genes to complex PD forms.
Mutations in a-synuclein (SNCA) and leucine-rich repeat kinase 2 (LRRK2) cause familial autosomal dominant forms of the disease. SNCA is the first PD gene identified by linkage studies (Polymeropoulos et al. 1997) but mutations/single-nucleotide variants and copy number variations in this gene contribute a very minor proportion to familial PD cases (Petrucci et al. 2016) compared to their robust association with sporadic cases. A dinucleotide repeat polymorphism in the promoter of SNCA, which increases its expression in vitro has been shown to be associated with increased PD risk (Maraganore et al. 2006). A strong association between specific haplotypes in the SNCA locus and sporadic PD has been demonstrated (Chai and Lim 2013). Even a common variant (rs356168) in the SNCA 3\(^\prime \) UTR is reported to be associated with increased PD risk by increasing the translation and accumulation SNCA protein (Toffoli et al. 2017). Both these types of variants essentially result in the aggregation of the \(\upalpha \)-synuclein protein leading to formation of Lewy bodies and presumably lead to dopaminergic neuronal cell death. On the other hand, mutations in LRRK2 coding for a kinase are the most common cause of sporadic PD, with a mutation frequency ranging from 2 to 40% in different populations (Klein and Westenberger 2012). More than 100 LRRK2 mutations have been so far described in PD families and sporadic cases (Li et al. 2014). PARKIN (Shimizu et al. 1998) and PINK1 (Valente et al. 2004) were identified in autosomal recessive PD families. Unlike familial cases, mutations reported in sporadic forms of PD are heterozygous and several lines of evidence suggest that these mutations are indeed susceptibility factors for PD (Khan et al. 2005; Eggers et al. 2010). PARKIN is an ubiquitin E3 ligase and is involved in mitochondrial maintenance, mitochondrial cytochrome c release, and autophagy of dysfunctional mitochondria (Pickrell and Youle 2015). PINK1 is a kinase that provides protection against mitochondrial dysfunction and regulates mitochondrial morphology via fission/fusion machinery and helps in autophagy (Pickrell and Youle 2015). Glucocerebrosidase (GBA) is another most common known genetic risk factor for sporadic PD (Ran et al. 2016). Interestingly, a few Gaucher disease (GD) patients also develop Parkinsonism (Neudorfer et al. 1996). Mutations in GBA are linked to GD where deficiency of the enzyme glucocerebrosidase causes lysosomal storage disorder. In Ashkenazi Jewish population with PD, 31.3% carried GBA mutations compared with 6.2% in the control individuals (Aharon-Peretz et al. 2004) with Asn370Ser mutation accounting for \(\sim \)70% of the disease burden (Sidransky and Lopez 2012). Another locus consistently associated with sporadic PD is MAPT, where H1 haplotype was shown to confer risk of PD in three Caucasian populations from Ireland, Norway and US (Skipper et al. 2004). Exon 1 of MAPT harbours a gene SAITOHIN (STH), where in a coding variant Gln7Arg (rs62063857) was shown to associate with risk of progressive supranuclear palsy and PD, and is in complete LD with the MAPT 238-bp intron 9 deletion that discriminates H1/H2 (Tobin et al. 2008). Besides, pathways such as ubiquitin–proteasome system, mitochondrial dysfunction, ROS mediated degeneration etc. leading to neuronal toxicity and cell death identified based on familial studies (described above), genes from association studies have enabled identification of additional pathways in the pathogenesis of complex forms of PD. These include immune system, autophagy/lysosomal degradation, microtubule stabilization, axonal transport, synaptic function and endocytosis. Yet genetic basis of majority of cases in this category is unexplained indicating that more genes/pathways remain to be identified.
ALS
ALS, commonly known as motor neuron disease is a progressive degenerative condition of the brain and spinal cord. Motor neurons are grouped into upper populations in the motor cortex and lower populations in the brain stem and spinal cord, with lower motor neurons innervating the muscles (Rowland and Shneider 2001). The core pathological feature in ALS is motor neuron death in the motor cortex and spinal cord, while in ALS with frontotemporal dementia (\(\sim \)10–15% ALS cases), neuronal degeneration is more widespread, occurring throughout the frontal and temporal lobes. ALS and frontotemporal dementia (FTD) are considered to be a part of spectrum. FTD patients are characteristically present with behavioural and speech problems, and ALS patients are present with alteration of voluntary movements caused by degeneration of motor neurons (Guerreiro et al. 2015). Loss of corticospinal motor neurons causes thinning and scarring (sclerosis) leading to muscle stiffness and spasticity, while loss of neurons in brain stem and spinal cord show electrical irritability and spontaneous muscle twitching. With disease progression, there is thinning of the ventral roots and they lose synaptic connectivity to their target muscles leading to denervational atrophy (amyotrophy) of the tongue, oropharynx and limb muscles (Brown and Al-Chalabi 2017). Diaphragm muscles are also affected, which generally leads to death due to respiratory paralysis within 3 to 5 years of diagnosis (Shoesmith et al. 2007). About one-third of ALS cases are bulbar, having difficulty in chewing, speaking, or swallowing. Cognition and behavioural changes are also observed in >30% ALS patients (Hobson and McDermott 2016). The diagnosis of ALS is primarily based on clinical examination using electromyography, MRI, blood/urine testing and muscle biopsy to rule out other neurological disorders which mimic ALS. Specific criteria for the diagnosis of ALS is known as the El Escorial criteria which provides a structured tool to define the level of confidence of a diagnosis in individual patients (Brooks et al. 2000). This classifies patients to different degrees of diagnostic certainty (definite, probable, probable with laboratory confirmation and possible) according to the presence of lower and upper motor neuron signs and their distribution in the four regions, i.e. bulbar, upper limb, thoracic and lower limb. First line of treatment is based on riluzole, a neuroprotective drug that blocks glutamatergic neurotransmission, which has been shown to improve mortality rate by 23% and 15% at 6 and 12 months, respectively (Traynor et al. 2003). In 2015, edaravone, an antioxidant compound was approved in Japan for treatment of ALS. On 5 May 2017, the FDA approved edaravone to extend the survival period of people with ALS (Rothstein 2017). Arimoclomol is another promising compound, which is a coinducer of heat shock protein expression under conditions of cellular stress. The therapeutic potential of this drug is currently under phase II clinical trial for ALS patients with SOD1 mutations (Kalmar et al. 2014).
ALS with a prevalence of 3–5 per 100,000 is an age dependent disorder with an increased incidence and higher prevalence in older populations (Pasinelli and Brown 2006). Monozygotic twin concordance is estimated to be between 60–80% suggesting a major genetic contribution to disease aetiology (Al-Chalabi et al. 2010). About 90% of ALS cases are sporadic with the rest being familial and generally inherited as an autosomal dominant condition (Byrne et al. 2011). Except in a few families with younger age at onset, it is \(\sim \)55 years in both these groups. As in AD and PD, there is genetic heterogeneity in both familial as well as sporadic ALS forms, but pathological and clinical features are similar indicating involvement of common cellular and molecular events leading to motor neuron degeneration. Based on the genes known to date, three processes namely (i) proteostasis and protein quality control; (ii) cytoskeletal dynamics; and (iii) RNA stability, function and metabolism seem to be generally affected (Weishaupt et al. 2016) which may provide leads for new drug discoveries.
Genetic risk factors
A number of genes relevant to ALS pathogenesis identified recently explain \(\sim \)70% of the familial cases and \(\sim \)30% of sporadic cases. This is notably higher compared to the heritability explained in PD and AD. Mutations found in familial ALS are also reported in sporadic forms. More than 50 genes including TARDBP, FUS and VPC have been identified in ALS using approaches ranging from conventional linkage analysis, candidate gene association/GWAS to recent NGS (Li and Wu 2016). Of these, disease causal variants/risk alleles in a few genes such as DCTN1, FIG4 and DAO have been limited to the study families or populations respectively. SOD1 was the first gene identified to be causal for ALS using linkage analysis in affected families (Siddique et al. 1991; Rosen et al. 1993), with mutations accounting for \(\sim \)20% of familial cases and also in \(\sim \)2–4% of the sporadic cases (Chen et al. 2013). Both sporadic and familial ALS cases with different SOD1 mutations show variation in the phenotype, in the age of onset, severity, rate of disease progression and duration of illness. For example: Asp90Ala mutation in exon 4 of SOD1 causes less severe form of the disease where average survival years are >10 after diagnosis (Andersen et al. 1996) whereas Ala4Val mutation in exon 1 has 91% penetrance and patients with this mutation survive for <18 months after the diagnosis (Cudkowicz et al. 1997). SOD1 is a ubiquitous cytosolic homodimeric protein, each subunit containing copper and zinc ions in the active site, its primary function being reduction of superoxide radical to \(\hbox {H}_{2}\hbox {O}_{2}\) (Fukai and Ushio-Fukai 2011). Studies have led to widespread acceptance of the hypothesis that SOD1 mutants acquire a novel toxic property independent of their enzymatic function (Redler and Dokholyan 2012). Both sporadic and familial ALS cases have aggregation of cytoplasmic proteins, prominently in motor neurons. The hallmark proteins of the pathogenic inclusions are SOD-1, TDP-43 or FUS. Almost all cases of ALS and tau-negative FTD have abnormal TDP-43 protein, also known as TDP-43 proteinopathy (Liscic 2017). The genetic findings of ALS and FTD also support a common link, as mutations in the same genes have been found in ALS, FTD, or FTD/ALS patients. Mutations in the TANK-binding kinase 1 gene (TBK1) has presented with features of ALS and FTD worldwide, however additional neurological features have also been described in TBK1 subjects. In a recent publication progressive supranuclear palsy (PSP) like phenotypes and progressive cerebellar syndromes in TBK1 subjects have been reported (Wilke et al. 2017). C9orf72 is another recently discovered gene in ALS linked to 9p chromosomal region. This gene is recognized as the most common form of ALS and FTD, additionally some patients may present with Parkinsonism, essential tremor and restless leg syndrome (Nuytemans et al. 2013b). Notably, an expansion of a hexa-nucleotide repeat GGGGCC (2–23 repeats in healthy individuals) in the noncoding region of this gene has been reported to be most common cause of familial ALS explaining >30% cases and >10% of the sporadic forms (DeJesus-Hernandez et al. 2011; Majounie et al. 2012). The C9orf72 protein has a role in nuclear and endosomal membrane trafficking and autophagy. There is evidence that RNA foci containing this repeat accumulate in the brains and spinal cords of affected people, and this suggested a second possible disease mechanism, involving toxic gain of function by repeat-containing RNA (Zu et al. 2013). Another \(\sim \)5% of sporadic and \(\sim \)3% of familial cases are caused by mutations in the gene coding for TDP-43 (Beleza-Meireles and Al-Chalabi 2009). In addition, modifiers such as 27–33 CAG repeats in ATXN2, increasing the risk of developing ALS (Daoud et al. 2011) and variants in EPHA4 that reduce expression of the axonal guidance improving the overall survival of ALS patients have also been reported (Elden et al. 2010; Van Hoecke et al. 2012). Besides linkage based causal gene discoveries, there have been more than 14 GWASs carried out on ALS, which have identified common variants associated (table 4). Genes such as UNC13A and C9orf72 are robustly replicated but many others including FGGY and ITPR2 have failed to show association in large cohorts (Daoud et al. 2010). These findings from sporadic forms have implicated regulation of chemotaxis and cellular communication/differentiation, intracellular signal transduction, oxidative stress and cytoskeleton organization pathway in addition to validating the involvement of SOD1 and c9orf72 mediated protein/RNA toxicity reported earlier in familial forms. Further, all common variants known to date explain <30% of sporadic cases. However, since most of these studies have been performed in European populations there is scope for identification of additional risk loci using populations with different genome architectures.
Huntington disease
Huntington disease (HD) is also a progressive neurodegenerative disease but unlike the sporadic complex forms of AD, PD and ALS described above, it is the most common monogenic condition in this group of disorders and largely following an autosomal dominant mode of inheritance. However, the variable age of disease onset witnessed among HD patients implying a significant role of modifier genes, warrants it to be considered under complex disease forms. HD is clinically diagnosed by assessment of motor dysfunction, cognitive impairment and neuropsychiatric features, with family history being a major feature. Loss of self and spatial awareness, depression, dementia and anxiety which is devastating to both patients and his or her family (Labbadia and Morimoto 2013) are associated features observed in HD patients. The earliest symptoms are often subtle changes in mood or mental abilities. Noncoordination and an unsteady gait often follow. As the disease advances, uncoordinated, jerky body movements also appears. Mental abilities generally decline into dementia (Ross and Tabrizi 2011). Degeneration of medium spiny neurons of striatum and also other regions of the brain such as cerebellar cortex, thalamus and cerebellum results in the manifestation of the characteristic disease symptoms (Bano et al. 2011), which are assessed by MRI or CT scans. Treatment to alter the course of HD is unavailable. Tetrabenazine treatment reduced chorea in a randomized controlled trial (Huntington Study Group 2006). Other drugs that help to reduce chorea include neuroleptics and benzodiazepines. Compounds such as amantadine or remacemide are still under investigation but have shown preliminary positive results (Mestre et al. 2009). Hypokinesia and rigidity, especially in juvenile cases, can be treated with antiparkinsonian drugs, and myoclonic hyperkinesia can be treated with valproic acid. Genetic counselling is advised if there is a family history of disease. HD has a global prevalence of 5–10 individuals per 100,000 (Roos 2010). The onset of disease is in the prime of adult life, i.e. fourth and fifth decade with disease duration of 15–20 years (Hardiman et al. 2016).
Genetic risk factors
Conventional restriction fragment length polymorphism based analysis in a large Venezuelan family with HD established its linkage at 4p16 (Gusella et al. 1983). However, it was only after the success stories of causal gene identification in Duchenne muscular dystrophy, cystic fibrosis and trinucleotide repeat expansion disorder fragile-X syndrome that the pathological CAG repeats in HTT were identified (MacDonald et al. 2017). The repeat expansion is a gain-of-function mutation leading to protein aggregation and toxicity by neuronal cell death in the striatum. Impairment of proteostasis network leading to synaptic dysfunction, mitochondrial toxicity and axonal transport seem to be the underlying pathological mechanisms. One of the strongest genetic risk factors associated with HD is the length of the CAG trinucleotide repeats in exon 1 of HTT. Normal populations have 9–35 of these repeats with median between 17 and 20, and >40 repeats lead to definite disease manifestations (Andrew et al. 1993; Snell et al. 1993). Repeats >60 results in juvenile Huntington’s disease (JHD) with onset at <20 years of age accounting for \(\sim \)7% of all HD cases (Nance and Myers 2001). It is the intermediate repeat lengths of 36–40 that have a lower disease predisposition and which may lead to incomplete penetrance (McNeil et al. 1997). These repeat sequences are prone to slippage during DNA replication resulting in their expansion and this is witnessed more through males and together with great variability in repeat lengths (Wheeler et al. 2007). The age at onset and severity of the disease are inversely correlated with the number of repeats. Further, HD is also characterized by genetic anticipation which means increased severity in disease and earlier age at onset in subsequent generations. Unlike other classical Mendelian disorders where a clear genotype–phenotype correlation is observed in HD such a prediction of disease phenotype and age at onset is not possible despite the presence of expanded CAG repeats. Two individuals with the same number of repeats in the pathological range can have as many as 20 years of difference in age of disease onset (Gusella and MacDonald 2009). Such a striking variation is currently attributed to interaction of additional genetic factors, i.e. modifier genes present in the genome, which has been substantiated by evidence discussed below. CAG repeat expansion in HTT is detected in 99% of cases with typical HD. In the remaining 1%, the disease has been linked to mutations in PRNP and JPH3 causing HD-like disease 1 and 2 (Pihlstrøm et al. 2017).
Since the discovery of pathogenic trinucleotide repeat expansion mutation in HTT and the observed clinical heterogeneity in HD, there have been a number of studies which investigated the role of genes that modify the disease pathogenicity. The length of CAG repeat in an affected individual was speculated to have a disease modifying effect and therefore, earlier studies focussed on variants at HTT locus itself that could alter the structure, function or expression of Huntingtin protein (Gusella et al. 2014). A recent study conducted on 4068 HD individuals however showed no significant impact. Interestingly, the same group showed that individuals with (CAG)\(_{n}\) expansion in both chromosomes (although a very rare event) have the age of disease onset consistent with the length of longer allele of the two expanded CAG repeat alleles (Lee et al. 2012). The other variants present in HTT haplotypes, like variable CCG repeats next to the CAG repeat, a deletion polymorphism at codon 2642, a deletion of one of the four consecutive GAG codons in exon 58, untranslated sequence of the transcript, intron sequences, and sequences flanking the centromeric and telomeric ends of the gene were also investigated for their modifying effect on age of disease onset but no correlation was observed (Gusella and MacDonald 2009). This was followed by the hypothesis testing approach where genes for establishing association were selected from transcriptome and proteome data obtained from peripheral blood of HD patients (Gusella and MacDonald 2009; Arning 2016). The genes were first sequenced to identify variants, if any, and then tested for association with age of onset. A number of replication studies using these variants have also been conducted lately in larger sample sizes and using appropriate statistical tools but with negligible replication (Arning and Epplen 2012). In addition, a few other candidate genes prioritized based on their role in energy metabolism, neurotransmission, HTT protein interactions and regulation of gene expression in the target tissue were also assessed for their association but results were not promising. These candidate gene-based association findings are summarised in table 5.
These limited insights from early association studies gave an impetus to perform the hypothesis free GWAS. A combined analysis of two sequential GWASs (GWAS1+GWAS2) using 1951 HD mutation positive samples of European ancestry collected at MaHDC (Massachusetts HD Center Without Walls) identified SNP rs146353869 on chromosome 15 to be significantly associated (\(P = 4.36 \times 10^{-9)}\) with the age of disease onset. Association of this SNP was notably replicated in another GWAS on an independent cohort of 2131 individuals (GWAS3) of the same ancestry (\(P=1.35\times 10^{-12})\). Meta-analysis of GWAS1+GWAS2+GWAS3 detected an enhanced association of the index SNP \((P=4.3\times 10^{-20})\) and interestingly, the analysis also identified two more associated SNPs, one on chromosome 15 (rs2140734; \(P=7.1\times 10^{-14})\) and another on chromosome 8 (rs1037699; \(P= 2.7\times 10^{-8})\). Of note, of the two SNPs on chromosome 15 (not in LD), rs146353869 seems to be the risk allele which accelerated the age of disease onset by 6.1 years while rs2140734 delayed the onset by 1.4 years; and risk allele rs1037699 on chromosome 8 hastened the onset by 1.6 years (Lee et al. 2015). In addition to the primary (CAG)\(_{n}\) expansion mediated neurodegeneration, the above detailed association findings are informative, but the actual mechanism(s) by which these modifiers influence the age of onset remains unclear. The major limitations in unravelling the modifiers to explain reduced penetrance or variable age of onset in HD could be clinical and genetic heterogeneity which current diagnostic/analytical tools have failed to address, an idea which has also been proposed by a few others (Genin et al. 2008). In depth analysis of the GWAS findings to identify potential protein-coding variants in LD with the GWAS SNPs and rare variants if any, in yet to be identified relevant genes may provide some leads.
Limitations
A detailed account of the limited insights into disease genetics, poor replication of association findings across studies and populations, an overall inability to establish genotype–phenotype correlations and/or predict risk in the predominant group of complex, sporadic forms of AD, PD, ALS and HD has been presented in the preceding paragraphs. The contrast between these observations and the powerful causal genes discoveries made in familial forms of the same pathologies is striking. Common and rare variants identified by several GWASs and their meta-analyses (Ramanan and Saykin 2013; Chang et al. 2017), emerging WES data and the limited WGS studies (Nicolas et al. 2016) have failed to explain total disease heritability in the sporadic forms of these disorders. Further, rare variants detected in these conditions such as PD (Nuytemans et al. 2013a), AD (Bertram 2016) and ALS (Morgan et al. 2015) are mostly private and confirming their inherited nature (and not a de novo origin) is also rather difficult due to the generally late age of onset of these disorders. Another aspect to be noted is that all these neurodegenerative diseases are tissue specific and sporadic cases may have postzygotic de novo mutations in the target tissue during early development leading to disease manifestation, which will never be detected in blood DNA. A few reports showing such postzygotic brain specific mutations in brain disorders support such a possibility (Beck et al. 2004; Rivière et al. 2012). Thus, a small percentage of disease may always remain unexplained. What these big data obtained from GWASs and NGS imply is then obvious. Even if only protein-coding variants and not regulatory changes were to be considered as predominant contributors to suboptimal functioning of the putative causal gene(s), we would neither be able to catalogue all the possible genomewide and tissue specific disease causing variants (DCVs) nor would we be successful in establishing replicable/reliable genotype–phenotype correlations. To put it in perspective, can even whole genome sequencing of blood DNA from sporadic forms of AD, PD and ALS identify oligogenic/polygenic variants and facilitate risk prediction and disease prevention? Would we be able to identify all the modifiers and their interactions to predict the clinical outcome of trinucleotide expansion present in the pathological range in individuals from HD families? Cohorts with differences in the genetic background may also segregate independent modifiers and may show poor replication. Further, clinical and genetic heterogeneity which characterize these diseases are unlikely to be addressed by attempts to document common susceptibility variants by increasing sample sizes from a few thousands to hundreds of thousands for GWASs and rare variants by advocating WGS. At best what the former could provide would be a few more risk conferring genes and a few more putative contributory pathways, if not already identified by WES based gene discoveries in familial forms. To reiterate, insights from GWAS findings for understanding the disease pathology per se have been negligible compared to those from findings in familial forms. Several loci of minor effects have been identified but very few of them have been functionally validated. Most of these are in the intronic regions and do not result in structural changes, but they may alter the expression levels of the adjacent genes as shown by rare study on functional validation of GWAS identified variant in SNCA (Campêlo et al. 2017). Therefore, a discovery approach combined with their functional validation using appropriate in vitro, cellular/animal models may be rewarding.
With all these constraints, advanced computational and systems biology approaches to analyse the big data are emerging. How the study of molecular, cellular and brain networks provides additional information on the effects of late onset AD-associated genetic variants has been discussed in a recent review highlighting the immune/microglia module to be strongly associated with AD pathophysiology (Gaiteri et al. 2016). At this juncture, it may be just important to follow these limited leads in hand and possibly explore alternate paradigms for unravelling the genetics of these complex neurodegenerative disorders in this era of predictive, preventive, personalized and participatory (P4) medicine. It is now evident that clinical heterogeneity in complex disease forms is the primary obstacle for effective discovery genomics (but not only limited to the four disorders under discussion in this review but also relevant to almost all of the \(\sim \)60% common complex disorders in humans). Developing a strategy to address this concern and obtain homogeneous sample sets may be the most rewarding. One such emerging strategy is ‘Ayurgenomics’—that is a combination of deep phenotyping principles practised in Ayurveda (the Indian traditional system of medicine) and contemporary genome analysis tools (Thelma 2008; Mukerji and Bhavana 2011; Juyal et al. 2012; Govindaraj et al. 2015; Prasher et al. 2016), which may fulfil this need. In this approach, every individual is categorized into one of the specific constitution (prakriti) groups of vata, pitta and kapha predominant or mixed prakriti using the Ayurveda criteria. Further, each of these prakriti groups are also documented for their susceptibility to different common complex traits, e.g. vata prakriti individuals are more prone to neurological disorders (Sharma 2003; Dey and Pahwa 2014). Accordingly, it is hypothesized that applying Ayurveda principles to predict prior risk/treatment outcome in genetic association studies may explain much more disease variance and thus potentially open up more predictive health, the major goal of P4 medicine. A few recent studies have validated these principles using rheumatoid arthritis (RA) as an example. An unpublished GWAS from our lab identified (i) different sets of functionally relevant risk genes with impressive effect sizes in each of the RA subgroups, and (ii) a very good correlation between genetic findings and clinical knowledge of Amavata (RA). Similar findings were also observed in a previous study (Juyal et al. 2012) suggesting that it may indeed be a very promising approach to overcome the current limitation of phenotypic heterogeneity in complex traits.
Prospects
It is clear from the above discussion that advances in discovery genomics and functional analysis of putative determinants with translational potential for complex forms of neurodegenerative diseases are negligible. Levodopa administration in PD, cholinesterase inhibitors in AD, tetrabenazine in HD and riluzole in ALS have been the oldest pharmacotherapies. However, the efficacy of these first lines of treatment have been limited. This may be due to pharmacogenetic differences, drug noncompliance or unknown causes contributing singly or in combination. In PD, mutations in PINK1 and PARKIN cause neuronal death due to mitochondrial dysfunction. By discovering the gene networks that orchestrate this process, a transcription regulator ATF4 has been singled out which plays a central role in mitochondrial metabolism and which may be used as a neuroprotector and may emerge as a new therapeutic strategy (Sun et al. 2013) but such examples of genome based medicine/novel therapeutics are rather few. Given that the central dogma in medicine is diagnosis-prevention-cure-treatment, should wisdom and solution be sought from alternate technologies/experimental advances?
One of the attempted alternate therapy has been replacement of dead dopaminergic neurons with the stem cell-derived neurons. Stem cell therapies have been attempted over the last two decades holding promise for treatment of neurodegenerative disorders, particularly PD and ALS. First clinical trial for tissue replacement using autologous adrenal medullary tissue (Backlund et al. 1985) showed significant clinical improvement. Subsequently, xenograft of porcine mesencephalic tissue was also done in PD patients but without much success (Fink et al. 2000). In another attempt, neural stem cells taken from cortical and subcortical tissue were isolated and expanded and injected into striatum resulting in improvement of clinical phenotypes (Neuman et al. 2009). Bone marrow derived stem cells have also been effectively used in clinical trials of patients with neurodegeneration (Lee et al. 2008) In the next phase, foetal mesencephalic transplantation was used successfully in three open label trials but did not replicate in National Institute of Health double blind trials (http://www.medicalnewstoday.com/articles/113908.php). However, these approaches suffered from limitations such as only a small number of embryonic stem (ES) cells which can be obtained from each abortuse, and thereby the need for more which in turn has associated problems of quality deterioration of already collected neurons. Thus, of the four stem cell populations, namely ES cells, neural stem cells, haematopoietic stem cells and induced pluripotent stem cells (iPSCs) being explored for clinical applications, iPSCs seem to be the most powerful permitting multiple experimentations. With the remarkable combination of iPSCs and genetic information, firstly the possibility, to generate iPSCs using patient-derived fibroblasts differentiated into neurons of choice depending on the neurodegenerative disease under consideration and secondly, to correct the disease causing mutation in a gene using clustered regularly interspaced short palindromic repeats (CRISPR) and its associated protein Cas9 strategy (Hockemeyer and Jaenisch 2016), this technology seems to emerge as the most potential therapeutic intervention for a range of genetic disorders. From functional characterization of genes of therapeutic relevance using gene specific assays to hypothesis free approaches (such as RNA sequencing or methylome sequencing etc.), from gene/genetic variant editing to genome editing using CRISPR/Cas9 in the same genetic background, and the possibility for screening of known or novel therapeutic molecules facilitating personalized medicine are the added advantages that iPSCs offer. The success story of iPSC-derived cells being transplanted in a Japanese woman in her 70s to treat her macular degeneration provided a proof of concept of its use for humans (Cyranoski 2014). Recent reports of iPSC-based dopaminergic neuron transplants in monkeys and sustained improvement after two year of follow up (Callaway 2017); and the first group of patients who received the transplant of cells in phase 1 clinical trial (NCT02452723) launched in March 2016 and results thereof further support the potential application of iPSCs in clinical medicine. A recent study showed the correction of the heterozygous MYBPC3 mutation which causes hypertrophic cardiomyopathy in human preimplantation embryos using this technology (Ma et al. 2017). Although a few challenges similar to earlier transplantation attempts of stem cells including infections, tumourigenesis, graft induced dyskinesia etc. have to be addressed completely, this therapeutic option holds promise. Given that iPSC-CRISPR/Cas9 based therapy becomes a reality in the near future, whole genome sequencing, identification of one or more rare causal variants therein, patient derived iPSC generation and gene/genome editing and repopulating the specific brain regions with the corrected neurons may be expected to provide the much needed effective therapeutic intervention for the debilitating neurodegenerative disorders and fulfil the dreams of personalized medicine. Even as efforts in this direction are ongoing, the concept of moderate sized stem cell banks where a quality checked set of lines carrying different combinations of commonly present HLA alleles is emerging (International stemcell corporation, Australia). In this clinical strategy, only the patient needs to be matched for immune compatibility and the whole process of generating individual patient-derived stem cells is not required. Simultaneously, pathway information for the gene(s) which is identified as putative causal in the patient would enable use of an already known drug molecule, bringing it under the current engagement with repurposed drug(s) which beats the need for new and elaborate FDA approvals etc. Pathway knowledge would also facilitate identification of novel druggable targets therein and subsequent development of new lead molecules, which would always be a preferred treatment option.
References
Abraham R., Moskvina V., Sims R., Hollingworth P., Morgan A., Georgieva L. et al. 2008 A genome-wide association study for late-onset alzheimer’s disease using DNA pooling. BMC Med. Genomics 1, 44.
Aharon-Peretz J., Rosenbaum H. and Gershoni-Baruch R. 2004 Mutations in the glucocerebrosidase gene and Parkinson’s disease in Ashkenazi Jews. N. Engl. J. Med. 351, 1972–1977.
Ahmeti K. B., Ajroud-Driss S., Al-Chalabi A., Andersen P. M., Armstrong J., Birve A. et al. 2013 Age of onset of amyotrophic lateral sclerosis is modulated by a locus on 1p34.1. Neurobiol. Aging 34, 357. e7–19.
Al-Chalabi A., Fang F., Hanby M. F., Leigh P. N., Shaw C. E., Ye W. et al. 2010 An estimate of amyotrophic lateral sclerosis heritability using twin data. J. Neurol. Neurosurg. Psychiatry 81, 1324–1326.
Andersen P. M., Forsgren L., Binzer M., Nilsson P., Ala-Hurula V., Keränen M. L. et al. 1996 Autosomal recessive adult-onset amyotrophic lateral sclerosis associated with homozygosity for Asp90Ala CuZn-superoxide dismutase mutation. A clinical and genealogical study of 36 patients. Brain 119, 1153–1172.
Andrew S. E., Paul Goldberg Y., Kremer B., Telenius H., Theilmann J., Adam S. et al. 1993 The relationship between trinucleotide (CAG) repeat length and clinical features of Huntington’s disease. Nat. Genet. 4, 398–403.
Arning L. 2016 The search for modifier genes in Huntington disease – Multifactorial aspects of a monogenic disorder. Mol. Cell Probes. 30, 404–409.
Arning L. and Epplen J. T. 2012 Genetic modifiers of Huntington’s disease: beyond CAG. Future Neurol. 7, 93–109.
Arning L., Saft C., Wieczorek S., Andrich J., Kraus P. H., Epplen J. T. 2007 NR2A and NR2B receptor gene variations modify age at onset in Huntington disease in a sex-specific manner. Hum. Genet. 122, 175–182.
Arning L., Monté D., Hansen W., Wieczorek S., Jagiello P., Akkad D. A. et al. 2008 ASK1 and MAP2K6 as modifiers of age at onset in Huntington’s disease. J. Mol. Med. 86, 485–490.
Backlund E.-O., Granberg P.-O., Hamberger B., Knutsson E., Mårtensson A., Sedvall G. et al. 1985 Transplantation of adrenal medullary tissue to striatum in parkinsonism. J. Neurosurg. 62, 169–173.
Bano D., Zanetti F., Mende Y. and Nicotera P. 2011 Neurodegenerative processes in Huntington’s disease. Cell Death Dis. 2, e228.
Beck J. A., Poulter M., Campbell T. A., Uphill J. B., Adamson G., Geddes J. F. et al. 2004 Somatic and germline mosaicism in sporadic early-onset alzheimer’s disease. Hum. Mol. Genet. 13, 1219–1224.
Beecham G. W., Martin E. R., Li Y. J., Slifer M. A., Gilbert J. R., Haines J. L. et al. 2009 Genome-wide association study implicates a chromosome 12 risk locus for late-onset Alzheimer disease. Am. J. Hum. Genet. 84, 35–43.
Beecham G. W., Dickson D. W., Scott W. K., Martin E. R., Schellenberg G., Nuytemans K. et al. 2015 PARK10 is a major locus for sporadic neuropathologically confirmed Parkinson disease. Neurology 84, 972–980.
Beleza-Meireles A. and Al-Chalabi A. 2009 Genetic studies of amyotrophic lateral sclerosis: Controversies and perspectives. Amyotroph Lateral Scler. 10, 1–14.
Bertram L. 2016 Next generation sequencing in Alzheimer’s disease. Methods in molecular biology, pp 281–297. Clifton, N.J.
Bertram L. and Tanzi R. E. 2005 The genetic epidemiology of neurodegenerative disease. J. Clin. Invest. 115, 1449–1457.
Bertram L., Lange C., Mullin K., Parkinson M., Hsiao M., Hogan M. F. et al. 2008 Genome-wide association analysis reveals putative Alzheimer’s disease susceptibility loci in addition to APOE. Am. J. Hum. Genet. 83, 623–632.
Biernacka J. M., Chung S. J., Armasu S. M., Anderson K. S., Lill C. M., Bertram L. et al. 2016 Genome-wide gene-environment interaction analysis of pesticide exposure and risk of Parkinson’s disease. Parkinsonism Relat. Disord. 32, 25–30.
Brooks B. R., Miller R. G., Swash M., Munsat T. L. and World Federation of Neurology Research Group on Motor Neuron Diseases. 2000 El Escorial revisited: revised criteria for the diagnosis of amyotrophic lateral sclerosis. Amyotroph Lateral Scler. Other Motor Neuron. Disord. 1, 293–299.
Brown R. H. and Al-Chalabi A. 2017 Amyotrophic lateral sclerosis. N. Engl. J. Med. 377, 162–172.
Byrne S., Walsh C., Lynch C., Bede P., Elamin M., Kenna K. et al. 2011 Rate of familial amyotrophic lateral sclerosis: a systematic review and meta-analysis. J. Neurol. Neurosurg. Psychiatry 82, 623–627.
Callaway E. 2017 Reprogrammed cells relieve Parkinson’s symptoms in trials. Nature (https://doi.org/10.1038/nature.2017.22531).
Campêlo C. L. das C. and Silva R. H. 2017 Genetic variants in SNCA and the risk of sporadic Parkinson’s disease and clinical outcomes: a review. Parkinsons Dis. 2017, 1–11.
Carrasquillo M., Zou F., Pankratz V., Wilcox S., Walker L., Younkin S. et al. 2009 Genetic variation in PCDH11X is associated with susceptibility to late-onset Alzheimer’s disease. Nat. Genet. 41, 192–198.
Chai C. and Lim K.-L. 2013 Genetic insights into sporadic Parkinson’s disease pathogenesis. Curr. Genomics 14, 486–501.
Chang D., Nalls M. A., Hallgrímsdóttir I. B., Hunkapiller J., van der Brug M., Cai F. et al. 2017 A meta-analysis of genome-wide association studies identifies 17 new Parkinson’s disease risk loci. Nat. Genet. 49, 1511–1516.
Chartier-Harlin M.-C., Crawford F., Houlden H., Warren A., Hughes D., Fidani L. et al. 1991 Early-onset Alzheimer’s disease caused by mutations at codon 717 of the \(\upbeta \)-amyloid precursor protein gen. Nature 353, 844–846.
Chattopadhyay B., Baksi K., Mukhopadhyay S. and Bhattacharyya N. P. 2005 Modulation of age at onset of Huntington disease patients by variations in TP53 and human caspase activated DNase (hCAD) genes. Neurosci. Lett. 374, 81–86.
Chen C.-J., Chen C.-M., Pai T.-W., Chang H.-T. and Hwang C.-S. 2016 A genome-wide association study on amyotrophic lateral sclerosis in the Taiwanese Han population. Biomark Med. 10, 597–611.
Chen S., Sayana P., Zhang X. and Le W. 2013 Genetics of amyotrophic lateral sclerosis: an update. Mol. Neurodegener. 8, 28.
Chung S. J., Armasu S. M., Biernacka J. M., Anderson K. J., Lesnick T. G., Rider D. N. et al. 2012 Genomic determinants of motor and cognitive outcomes in Parkinson’s disease.Parkinsonism Relat. Disord. 18, 881–886.
Coon K. D., Myers A. J., Craig D. W., Webster J. A., Pearson J. V., Lince D. H. et al. 2007 A high-density whole-genome association study reveals that APOE is the major susceptibility gene for sporadic late-onset Alzheimer’s disease. J. Clin. Psychiatry 68, 613–618.
Coppedè F., Migheli F., Ceravolo R., Bregant E., Rocchi A., Petrozzi L. et al. 2010 The hOGG1 Ser326Cys polymorphism and Huntington’s disease. Toxicology 278, 199–203.
Cronin S., Berger S., Ding J., Schymick J. C., Washecka N., Hernandez D. G. et al. 2007 A genome-wide association study of sporadic ALS in a homogenous Irish population. Hum. Mol. Genet. 17, 768–774.
Cruchaga C., Karch C. M., Jin S. C., Benitez B. A., Cai Y., Guerreiro R. et al. 2013 Rare coding variants in the phospholipase D3 gene confer risk for Alzheimer’s disease. Nature 505, 550.
Cudkowicz M. E., McKenna-Yasek D., Sapp P. E., Chin W., Geller B., Hayden D. L. et al. 1997 Epidemiology of mutations in superoxide dismutase in amyotrophic lateal sclerosis. Ann. Neurol. 41, 210–221.
Cyranoski D. 2014 Japanese woman is first recipient of next-generation stem cells. Nature (https://doi.org/10.1038/nature.2014.15915).
Daoud H., Belzil V., Desjarlais A., Camu W., Dion P. A. and Rouleau G. A. 2010 Analysis of the UNC13A gene as a risk factor for sporadic amyotrophic lateral sclerosis. Arch. Neurol. 67, 516–517.
Daoud H., Belzil V., Martins S., Sabbagh M., Provencher P., Lacomblez L. et al. 2011 Association of long ATXN2 CAG repeat sizes with increased risk of amyotrophic lateral Sclerosis. Arch. Neurol. 68, 739–742.
Davis M. F., Cummings A. C., D’Aoust L. N., Jiang L., Velez Edwards D. R., Laux R. et al. 2013 Parkinson disease loci in the mid-western Amish. Hum. Genet. 132, 1213–1221.
DeJesus-Hernandez M., Mackenzie I. R., Boeve B. F., Boxer A. L., Baker M., Rutherford N. J. et al. 2011 Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-Linked FTD and ALS. Neuron 72, 245–256.
Deng M., Wei L., Zuo X., Tian Y., Xie F., Hu P. et al. 2013 Genome-wide association analyses in Han Chinese identify two new susceptibility loci for amyotrophic lateral sclerosis. Nat. Genet. 45, 697–700.
Dey S. and Pahwa P. 2014 Prakriti and its associations with metabolism, chronic diseases, and genotypes: possibilities of new born screening and a lifetime of personalized prevention. J. Ayurveda Integr. Med. 5, 15–24.
Dhaenens C.-M., Burnouf S., Simonin C., Van Brussel E., Duhamel A., Defebvre L. et al. 2009 A genetic variation in the ADORA2A gene modifies age at onset in Huntington’s disease. Neurobiol. Dis. 35, 474–476.
Diekstra F. P., Van Deerlin V. M., van Swieten J. C., Al-Chalabi A., Ludolph A. C., Weishaupt J. H. et al. 2014 C9orf72 and UNC13A are shared risk loci for amyotrophic lateral sclerosis and frontotemporal dementia: A genome-wide meta-analysis. Ann. Neurol. 76, 120–133.
Do C. B., Tung J. Y., Dorfman E., Kiefer A. K., Drabant E. M., Francke U. et al. 2011 Web-based genome-wide association study identifies two novel loci and a substantial genetic component for Parkinson’s disease. PLoS Genet. 7, e1002141.
Edwards T. L., Scott W. K., Almonte C., Burt A., Powell E. H., Beecham G. W. et al. 2010 Genome-wide association study confirms SNPs in SNCA and the MAPT region as common risk factors for Parkinson disease. Ann. Hum. Genet. 74, 97–109.
Eggers C., Schmidt A., Hagenah J., Bruggemann N., Klein J. C., Tadic V. et al. 2010 Progression of subtle motor signs in PINK1 mutation carriers with mild dopaminergic deficit. Neurology 74, 1798–1805.
Elden A. C., Kim H.-J., Hart M. P., Chen-Plotkin A. S., Johnson B. S., Fang X. et al. 2010 Ataxin-2 intermediate-length polyglutamine expansions are associated with increased risk for ALS. Nature 466, 1069–1075.
Fearnley J. M. and Lees A. J. 1991 Ageing and Parkinson’s disease: substantia nigra regional selectivity. Brain 114, 2283–2301.
Fink J. S., Schumacher J. M., Ellias S. L., Palmer E. P., Saint-Hilaire M., Shannon K. et al. 2000 Porcine xenografts in Parkinson’s disease and Huntington’s disease patients: preliminary results. Cell Transplant. 9, 273–278.
Foo J. N., Tan L. C., Irwan I. D., Au W.-L., Low H. Q., Prakash K.-M. et al. 2016 Genome-wide association study of Parkinson’s disease in East Asians. Hum. Mol. Genet. 26, ddw379.
Fukai T. and Ushio-Fukai M. 2011 Superoxide dismutases: role in redox signaling, vascular function, and diseases. Antioxid. Redox Signal 15, 1583–606
Fung H.-C., Scholz S., Matarin M., Simón-Sánchez J., Hernandez D., Britton A. et al. 2006 Genome-wide genotyping in Parkinson’s disease and neurologically normal controls: first stage analysis and public release of data. Lancet Neurol. 5, 911–916.
Gaiteri C., Mostafavi S., Honey C. J., De Jager P. L. and Bennett D. A. 2016 Genetic variants in Alzheimer disease — molecular and brain network approaches. Nat. Rev. Neurol. 12, 413–427.
Gaj P., Paziewska A., Bik W., Dbrowska M., Baranowska-Bik A., Styczynska M., Chodakowska-Ebrowska M. et al. 2012 Identification of a late onset alzheimer’s disease candidate risk variant at 9q21.33 in polish patients. J. Alzheimer’s Dis. 32, 157–168.
Genin E., Feingold J. and Clerget-Darpoux F. 2008 Identifying modifier genes of monogenic disease: strategies and difficulties. Hum. Genet. 124, 357–368.
Glenner G. G. and Wong C. W. 1984a Alzheimer’s disease: initial report of the purification and characterization of a novel cerebrovascular amyloid protein. Biochem. Biophys. Res. Commun. 120, 885–890.
Glenner G. G. and Wong C. W. 1984b Alzheimer’s disease and Down’s syndrome: Sharing of a unique cerebrovascular amyloid fibril protein. Biochem. Biophys. Res. Commun. 122, 1131–1135.
Govindaraj P., Nizamuddin S., Sharath A., Jyothi V., Rotti H., Raval R. et al. 2015 Genome-wide analysis correlates Ayurveda Prakriti. Sci. Rep. 5, 15786.
Groen J. L., Kawarai T., Toulina A., Rivoiro C., Salehi-Rad S., Sato C. et al. 2004 Genetic association study of PINK1 coding polymorphisms in Parkinson’s disease. Neurosci. Lett. 372, 226–229.
Grupe A., Abraham R., Li Y., Rowland C., Hollingworth P., Morgan A. et al. 2007 Evidence for novel susceptibility genes for late-onset Alzheimer’s disease from a genome-wide association study of putative functional variants. Hum. Mol. Genet. 16, 865–873.
Gu Z., Nakamura T., Yao D., Shi Z.-Q. and Lipton S. A. 2005 Nitrosative and oxidative stress links dysfunctional ubiquitination to Parkinson’s disease. Cell Death Differ. 12, 1202–1204.
Guerreiro R., Brás J. and Hardy J. 2015 SnapShot: genetics of ALS and FTD. Cell 160, 798–798.e1.
Gusella J. F. and MacDonald M. E. 2009 Huntington’s disease: the case for genetic modifiers. Genome Med. 1, 80.
Gusella J. F., Wexler N. S., Conneally P. M., Naylor S. L., Anderson M. A., Tanzi R. E. et al. 1983 A polymorphic DNA marker genetically linked to Huntington’s disease. Nature 306, 234–238.
Gusella J. F. and MacDonald M. E., Lee J.-M. 2014 Genetic modifiers of Huntington’s disease. Mov. Disord. 29, 1359–1365.
Hamza T. H., Zabetian C. P., Tenesa A., Laederach A., Montimurro J., Yearout D. et al. 2010 Common genetic variation in the HLA region is associated with late-onset sporadic Parkinson’s disease. Nat. Genet. 42, 781–785.
Hardiman O., Doherty C. P., Elamin M. and Bede P. 2016 Neurodegenerative disorders. Springer International Publishing Switzerland.
Hardy J. 2006 A hundred years of Alzheimer’s disease research. Neuron 52, 3–13.
Harold D., Abraham R., Hollingworth P., Sims R., Hamshere M., Pahwa J. S. et al. 2009 Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer’s disease, and shows evidence for additional susceptibility genes. Nat. Genet. 41, 1088–1093.
Hely M. A., Morris J. G., Traficante R., Reid W. G., O’Sullivan D. J. and Williamson P. M. 1999 The sydney multicentre study of Parkinson’s disease: progression and mortality at 10 years. J. Neurol. Neurosurg. Psychiatry 67, 300–307.
Hill-Burns E. M., Wissemann W. T., Hamza T. H., Factor S. A., Zabetian C. P. and Payami H. 2014 Identification of a novel Parkinson’s disease locus via stratified genome-wide association study. BMC Genomics 15, 118.
Hill-Burns E. M., Ross O. A., Wissemann W. T., Soto-Ortolaza A. I., Zareparsi S., Siuda J., Lynch T. et al. 2016 Identification of genetic modifiers of age-at-onset for familial Parkinson’s disease. Hum. Mol. Genet. 25, 3849–3862.
Hirano A., Ohara T., Takahashi A., Aoki M., Fuyuno Y., Ashikawa K. et al. 2015 A genome-wide association study of late-onset Alzheimer’s disease in a Japanese population. Psychiatr. Genet. 1, 139–146.
Hobson E. V. and McDermott C. J. 2016 Supportive and symptomatic management of amyotrophic lateral sclerosis. Nat. Rev. Neurol. 12, 526–538.
Hockemeyer D. and Jaenisch R. 2016 Induced pluripotent stem cells meet genome editing. Cell Stem Cell 18, 573–586.
Hollingworth P., Harold D., Sims R., Gerrish A., Lambert J.-C., Carrasquillo M. M. et al. 2011 Common variants at ABCA7, MS4A6A/MS4A4E, EPHA1, CD33 and CD2AP are associated with Alzheimer’s disease. Nat. Genet. 43, 429–435.
Holtzman D., Bales K., Tenkova T., M. Fagan A., Parsadanian M., J. Sartorius L. et al. 2000 Apolipoprotein E isoform-dependent amyloid deposition and neuritic degeneration in a mouse model of Alzheimer’s disease. Proc. Natl. Acad. Sci. USA 97, 2892–2897.
Holtzman D. M., Bales K. R., Wu S., Bhat P., Parsadanian M., Fagan A. M. et al. 1999 Expression of human apolipoprotein E reduces amyloid-\(\upbeta \) deposition in a mouse model of Alzheime’s diseas. J. Clin. Invest. 103, R15–R21.
Hu X., Pickering E., Liu Y. C., Hall S., Fournier H., Katz E. et al. 2011 Meta-analysis for genome-wide association study identifies multiple variants at the BIN1 locus associated with late-onset Alzheimer’s disease. PLoS One 6, 1–9.
Hu Y., Deng L., Zhang J., Fang X., Mei P., Cao X. et al. 2016 A Pooling Genome-wide association study combining a pathway analysis for typical sporadic Parkinson’s disease in the Han population of Chinese mainland. Mol. Neurobiol. 53, 4302–4318.
Hughes A. J., Daniel S. E., Kilford L. and Lees A. J. 1992 Accuracy of clinical diagnosis of idiopathic Parkinson’s disease: a clinico-pathological study of 100 cases. J. Neurol. Neurosurg. Psychiatry 55, 181–184.
Huntington Study Group 2006 Tetrabenazine as antichorea therapy in Huntington disease: a randomized controlled trial. Neurology 66, 366–372.
International Parkinson Disease Genomics Consortium, Nalls M. A., Plagnol V., Hernandez D. G., Sharma M., Sheerin U.-M. et al. 2011 Imputation of sequence variants for identification of genetic risks for Parkinson’s disease: a meta-analysis of genome-wide association studies. Lancet 377, 641–649.
Jiao B., Liu X., Tang B., Hou L., Zhou L., Zhang F. et al. 2014 Investigation of TREM2, PLD3, and UNC5C variants in patients with Alzheimer’s disease from mainland China. Neurobiol. Aging 35, 2422.e9–2422.e11.
Jonsson T., Atwal J. K., Steinberg S., Snaedal J., Jonsson P. V., Bjornsson S. et al. 2012 A mutation in APP protects against Alzheimer’s disease and age-related cognitive decline. Nature 488, 96–99.
Jonsson T., Stefansson H., Steinberg S., Jonsdottir I., Jonsson P. V., Snaedal J. et al. 2013 Variant of TREM2 associated with the risk of Alzheimer’s disease. N. Engl. J. Med. 368, 107–116.
Jun G., Asai H., Zeldich E., Drapeau E., Haroutunian V., Foroud T. et al. 2016 PLXNA4 is Associated with Alzheimer disease and modulates tau phosphorylation. Ann. Neurol. 76, 379–392.
Juyal R. C., Negi S., Wakhode P., Bhat S., Bhat B. and Thelma B. K. 2012 Potential of ayurgenomics approach in complex trait research: leads from a pilot study on rheumatoid arthritis. PLoS One 7, e45752.
Kalmar B., Lu C.-H. and Greensmith L. 2014 The role of heat shock proteins in amyotrophic lateral sclerosis: the therapeutic potential of arimoclomol. Pharmacol. Ther. 141, 40–54.
Khan N. L., Horta W., Eunson L., Graham E., Johnson J. O., Chang S. et al. 2005 Parkin disease in a Brazilian kindred: manifesting heterozygotes and clinical follow-up over 10 years. Mov. Disord. 20, 479–484.
Klein C. and Westenberger A. 2012 Genetics of Parkinson’s disease. Cold Spring Harb. Perspect Med. 2, a008888.
Kloster E., Saft C., Epplen J. and Arning L. 2013 CNR1 variation is associated with the age at onset in Huntington disease. Eur. J. Med. Genet. 56, 416–419.
Kloster E., Saft C., Akkad D., Epplen J. and Arning L. 2014 Association of age at onset in Huntington disease with functional promoter variations in NPY and NPY2R. J. Mol. Med. 92, 177–184.
Kopin I. J., Burns S. R., Chiueh C. C. and Markey S. P. 1986 MPTP-Induced Parkinsonian syndromes in humans and animals, pp. 519–530. Springer, Boston.
Kwee L. C., Liu Y., Haynes C., Gibson J. R., Stone A., Schichman S. A. et al. 2012 A high-density genome-wide association screen of sporadic ALS in US veterans. PLoS One 7, e32768.
Laaksovirta H., Peuralinna T., Schymick J. C., Scholz S. W., Lai S.-L., Myllykangas L. et al. 2010 Chromosome 9p21 in amyotrophic lateral sclerosis in Finland: a genome-wide association study. Lancet Neurol. 9, 978–985.
Labbadia J. and Morimoto R. I. 2013 Huntington’s disease: underlying molecular mechanisms and emerging concepts. Trends Biochem. Sci. 38, 378–385.
Lambert J.-C., Heath S., Even G., Campion D., Sleegers K., Hiltunen M. et al. 2009 Genome-wide association study identifies variants at CLU and CR1 associated with Alzheimer’s disease. Nat. Genet. 41, 1094–1099.
Lambert J.-C., Ibrahim-Verbaas C. A., Harold D., Naj A. C., Sims R., Bellenguez C. et al. 2013 Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458.
Landers J. E., Melki J., Meininger V., Glass J. D., van den Berg L. H., van Es M. A. et al. 2009 Reduced expression of the Kinesin-associated protein 3 (KIFAP3) gene increases survival in sporadic amyotrophic lateral sclerosis. Proc. Natl. Acad. Sci. USA 106, 9004–9009.
Latourelle J. C., Pankratz N., Dumitriu A., Wilk J. B., Goldwurm S., Pezzoli G. et al. 2009 Genomewide association study for onset age in Parkinson disease. BMC Med. Genet. 10, 98.
Lee J. H., Cheng R., Barral S., Reitz C., Medrano M., Lantigua R. et al. 2011 Identification of novel loci for Alzheimer disease and replication of CLU, PICALM, and BIN1 in Caribbean Hispanic individuals. Arch. Neurol. 68, 320–328.
Lee J. M., Ramos E. M., Lee J. H., Gillis T., Mysore J. S., Hayden M. R. et al. 2012 CAG repeat expansion in Huntington disease determines age at onset in a fully dominant fashion. Neurology 78, 690–695.
Lee J. M., Wheeler V. C., Chao M. J., Vonsattel J. P. G., Pinto R. M., Lucente D. et al. 2015 Identification of genetic factors that modify clinical onset of Huntington’s disease. Cell 162, 516–526.
Lee P., Kim J., Bang O., Ahn Y., Joo I. and Huh K. 2008 Autologous mesenchymal stem cell therapy delays the progression of neurological deficits in patients with multiple system atrophy. Clin. Pharmacol. Ther. 83, 723–730.
Lesage S. and Brice A. 2012 Role of Mendelian genes in “sporadic” Parkinson’s disease. Parkinsonism Relat. Disord. 18, S66–S70.
Levy-Lahad E., Wasco W., Poorkaj P., Romano D. M., Oshima J., Pettingell W. H. et al. 1995 Candidate gene for the chromosome 1 familial Alzheimer’s disease locus. Science 269, 973–977.
Li H., Wetten S., Li L., St Jean P. L., Upmanyu R., Surh L. et al. 2008 Candidate single-nucleotide polymorphisms from a genomewide association study of Alzheimer disease. Arch. Neurol. 65, 45–53.
Li H.-F. and Wu Z.-Y. 2016 Genotype-phenotype correlations of amyotrophic lateral sclerosis. Transl. Neurodegener. 5, 3.
Li J.-Q., Tan L. and Yu J.-T. 2014 The role of the LRRK2 gene in Parkinsonism. Mol. Neurodegener. 9, 47.
Lill C. M., Roehr J. T., McQueen M. B., Kavvoura F. K., Bagade S., Schjeide B.-M. M. et al. 2012 Comprehensive research synopsis and systematic meta-analyses in Parkinson’s disease genetics: The PDGene database. PLoS Genet. 8, e1002548.
Liscic R. M. 2017 Als and Ftd: Insights into the disease mechanisms and therapeutic targets. Eur. J. Pharmacol. 817, 2–6.
Liu X., Cheng R., Verbitsky M., Kisselev S., Browne A., Mejia-Sanatana H. et al. 2011 Genome-wide association study identifies candidate genes for Parkinson’s disease in an Ashkenazi Jewish population. BMC Med. Genet. 12, 104.
Logue M. W., Schu M., Vardarajan B. N., Farrell J., Bennett D. A., Buxbaum J. D. et al. 2014 Two rare AKAP9 variants are associated with Alzheimer’s disease in African Americans. Alzheimer’s Dement. 10, 609–618.
Ma H., Marti-Gutierrez N., Park S.-W., Wu J., Lee Y., Suzuki K. et al. 2017 Correction of a pathogenic gene mutation in human embryos. Nature 548, 413–419.
MacDonald M. E., Ambrose C. M., Duyao M. P., Myers R. H., Lin C., Srinidhi L. et al. 2017 A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell 72, 971–983.
Majounie E., Renton A. E., Mok K., Dopper E. G., Waite A., Rollinson S. et al. 2012 Frequency of the C9orf72 hexanucleotide repeat expansion in patients with amyotrophic lateral sclerosis and frontotemporal dementia: a cross-sectional study. Lancet Neurol. 11, 323–330.
Meyer J. M. and Breitner J. 1998 Multiple threshold model for the onset of alzheimer’s disease in the NAS-NRC twin panel. Am. J. Med. Genet. 81, 92–97.
Mao X., Wang T., Peng R., Chang X., Li N., Gu Y. et al. 2013 Mutations in GBA and risk of Parkinson’s disease: a meta-analysis based on 25 case-control studies. Neurol. Res. 35, 873–878.
Maraganore D. M., de Andrade M., Lesnick T. G., Strain K. J., Farrer M. J., Rocca W. A. et al. 2005 High-resolution whole-genome association study of Parkinson disease. Am. J. Hum. Genet 77, 685–693.
Maraganore D. M., Andrade M. de, Elbaz A., Farrer M. J., Ioannidis J. P., Krüger R. et al. 2006 Collaborative Analysis of \(\upalpha \)-Synuclein gene promoter variability and Parkinson disease. JAMA 296, 661.
Martinez-Ramirez D., Hu W., Bona A. R., Okun M. S. and Shukla A. W. 2015 Update on deep brain stimulation in Parkinson’s disease. Transl. Neurodegener. 4, 12.
Masters C. L., Bateman R., Blennow K., Rowe C. C., Sperling R. A. and Cummings J. L. 2015 Alzheimer’s disease. Nat. Rev. Dis. Prim. 1, 15056.
McNeil S. M., Novelletto A., Srinidhi J., Barnes G., Kornbluth I., Altherr M. R. et al. 1997 Reduced penetrance of the Huntington’s disease mutation. Hum. Mol. Genet. 6, 775–779.
Medway C. W., Abdul-Hay S., Mims T., Ma L., Bisceglio G., Zou F. et al. 2014 ApoE variant p.V236E is associated with markedly reduced risk of Alzheimer’s disease. Mol. Neurodegener. 9, 11.
Mellick G. D., Maraganore D. M. and Silburn P. A. 2005 Australian data and meta-analysis lend support for alpha-synuclein (NACP-Rep1) as a risk factor for Parkinson’s disease. Neurosci. Lett. 375, 112–116.
Mestre T., Ferreira J., Coelho M. M., Rosa M. and Sampaio C. 2009 Therapeutic interventions for disease progression in Huntington’s disease. In Cochrane database of systematic reviews (ed. T. Mestre), pp. CD006455. John Wiley, Chichester.
Metzger S., Bauer P., Tomiuk J., Laccone F., Didonato S., Gellera C. et al. 2006 The S18Y polymorphism in the UCHL1 gene is a genetic modifier in Huntington’s disease. Neurogenetics 7, 27–30.
Metzger S., Rong J., Nguyen H. P., Cape A., Tomiuk J., Soehn A. S. et al. 2008 Huntingtin-associated protein-1 is a modifier of the age-at-onset of Huntington’s disease. Hum. Mol. Genet. 17.
Metzger S., Walter C., Riess O., Roos R. A. C., Nielsen J. E., Craufurd D. et al. 2013 The V471A Polymorphism in autophagy-related gene ATG7 modifies age at onset specifically in Italian Huntington disease patients. PLoS One 8, e68951.
Mez J., Chung J., Jun G., Kriegel J., Bourlas A. P., Sherva R. et al. 2016 Two novel loci, COBL and SLC10A2, for Alzheimer’s disease in African Americans. Alzheimer’s Dement. 1–11.
Miyashita A., Koike A., Jun G., Wang L. S., Takahashi S., Matsubara E. et al. 2013 SORL1 Is Genetically associated with late-onset alzheimer’s disease in Japanese, Koreans and Caucasians. PLoS One 8.
Morgan S., Shoai M., Fratta P., Sidle K., Orrell R., Sweeney M. G. et al. 2015 Investigation of next-generation sequencing technologies as a diagnostic tool for amyotrophic lateral sclerosis. Neurobiol. Aging 36, 1600.e5-1600.e8.
Mukerji M. and Bhavana P. 2011 Ayurgenomics: A new approach in personalized and preventive medicine. Sci. Cult. 77, 10–17.
Naj A. C., Beecham G. W., Martin E. R., Gallins P. J., Powell E. H., Konidari I. et al. 2010 Dementia revealed: novel chromosome 6 locus for Late-onset alzheimer disease provides genetic evidence for Folate-pathway abnormalities. PLoS Genet. 6.
Naj A. C., Jun G., Beecham G. W., Wang L., Narayan B., Buros J. et al. 2011 Common variants in MS4A4/MS4A6E, CD2uAP, CD33, and EPHA1 are associated with late-onset alzheimer’s disease Adam. Nat. Genet. 43, 436–441.
Nalls M. A., Pankratz N., Lill C. M., Do C. B., Hernandez D. G., Saad M. et al. 2014 Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson’s disease. Nat. Genet. 46, 989–993.
Namba Y., Tomonaga M., Kawasaki H., Otomo E. and Ikeda K. 1991 Apolipoprotein E immunoreactivity in cerebral amyloid deposits and neurofibrillary tangles in Alzheimer’s disease and kuru plaque amyloid in Creutzfeldt-Jakob disease. Brain Res. 541, 163–166.
Nance M. and Myers R. H. 2001 Junvenile onset Huntington’s disease- clinical and research perspectives. Ment. Retard Dev. Disabil. Res. Rev. 7, 153–157.
Neudorfer O., Giladi N., Elstein D., Abrahamov A., Turezkite T., Aghai E. et al. 1996 Occurrence of Parkinson’s syndrome in type I Gaucher disease. QJM 89, 691–694.
Neuman T., Rezak M. and Levesque M. F. 2009 Therapeutic microinjection of autologous adult human neural stem cells and differentiated neurons for Parkinson’s disease: five-year post-operative outcome. Open Stem Cell J. 1, 20–29.
Nicolas G., Wallon D., Charbonnier C., Quenez O., Rousseau S., Richard A. C. et al. 2016 Screening of dementia genes by whole-exome sequencing in early-onset alzheimer disease: input and lessons. Eur. J. Hum. Genet. 24, 710–716.
Nuytemans K., Bademci G., Inchausti V., Dressen A., Kinnamon D. D., Mehta A. et al. 2013a Whole exome sequencing of rare variants in EIF4G1 and VPS35 in Parkinson disease. Neurology 80, 982–989.
Nuytemans K., Bademci G., Kohli M. M., Beecham G. W., Wang L., Young J. I. et al. 2013b C9ORF72 intermediate repeat copies are a significant risk factor for Parkinson disease. Ann. Hum. Genet. 77, 351–363.
Oliveira S. A., Scott W. K., Nance M. A., Watts R. L., Hubble J. P., Koller W. C. et al. 2003 Association study of Parkin gene polymorphisms with idiopathic Parkinson disease. Arch. Neurol. 60, 975.
Olsson B., Lautner R., Andreasson U., Öhrfelt A., Portelius E., Bjerke M. et al. 2016 CSF and blood biomarkers for the diagnosis of Alzheimer’s disease: a systematic review and meta-analysis. Lancet Neurol. 15, 673–684.
Paisán-Ruíz C., Lang A. E., Kawarai T., Sato C., Salehi-Rad S., Fisman G. K. et al. 2005 LRRK2 gene in Parkinson disease: mutation analysis and case control association study. Neurology 65, 696–700.
Pandey S. and Sarma N. 2015 Deep brain stimulation: current status. Neurol. India 63, 9–18.
Pandey S. and Srivanitchapoom P. 2017 Levodopa-induced Dyskinesia: Clinical features, pathophysiology, and medical management. Ann. Indian Acad. Neurol. 20, 190–198.
Pankratz N., Wilk J. B., Latourelle J. C., DeStefano A. L., Halter C., Pugh E. W. et al. 2009 Genomewide association study for susceptibility genes contributing to familial Parkinson disease. Hum. Genet. 124, 593–605.
Pankratz N., Beecham G. W., DeStefano A. L., Dawson T. M., Doheny K. F., Factor S. A. et al. 2012 Meta-analysis of Parkinson’s disease: identification of a novel locus, RIT2. Ann. Neurol. 71, 370–384.
Pasinelli P. and Brown R. H. 2006 Molecular biology of amyotrophic lateral sclerosis: insights from genetics. Nat. Rev. Neurosci. 7, 710–723.
Pedersen N. L., Gatz M., Berg S. and Johansson B. 2004 How heritable is Alzheimer’s disease late in life? Findings from Swedish twins. Ann. Neurol. 55, 180–185.
Petrucci S., Ginevrino M. and Valente E. M. 2016 Phenotypic spectrum of alpha-synuclein mutations: new insights from patients and cellular models. Parkinsonism Relat. Disord. 22, S16–S20.
Pickrell A. M. and Youle R. J. 2015 The roles of PINK1, parkin, and mitochondrial fidelity in Parkinson’s disease. Neuron 85, 257–273.
Pickrell J. K., Berisa T., Liu J. Z., Ségurel L., Tung J. Y. and Hinds D. A. 2016 Detection and interpretation of shared genetic influences on 42 human traits. Nat. Genet. 48, 709–717.
Pihlstrøm L., Wiethoff S. and Houlden H. 2017 Genetics of neurodegenerative diseases: an overview. Handb. Clin. Neurol. 145, 309–323.
Poduslo S. E., Huang R., Huang J. and Smith S. 2009 Genome screen of late-onset alzheimer’s extended pedigrees identifies TRPC4AP by haplotype analysis. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 150, 50–55.
Polymeropoulos M. H., Lavedan C., Leroy E., Ide S. E., Dehejia A., Dutra A. et al. 1997 Mutation in the alpha-synuclein gene identified in families with Parkinson’s disease. Science 276, 2045–2047.
Pottier C., Hannequin D., Coutant S., Rovelet-Lecrux A., Wallon D., Rousseau S. et al. 2012 High frequency of potentially pathogenic SORL1 mutations in autosomal dominant early-onset alzheimer disease. Mol. Psychiatry 17, 875–879.
Prasher B., Gibson G. and Mukerji M. 2016 Genomic insights into ayurvedic and western approaches to personalized medicine. J. Genet. 95, 209–228.
Pringsheim T., Jette N., Frolkis A. and Steeves T. D. L. 2014 The prevalence of Parkinson’s disease: a systematic review and meta-analysis. Mov. Disord. 29, 1583–1590.
Ramanan V. K. and Saykin A. J. 2013 Pathways to neurodegeneration: mechanistic insights from GWAS in Alzheimer’s disease, Parkinson’s disease, and related disorders. Am. J. Neurodegener. Dis. 2, 145–175.
Ran C., Brodin L., Forsgren L., Westerlund M., Ramezani M., Gellhaar S. et al. 2016 Strong association between glucocerebrosidase mutations and Parkinson’s disease in Sweden. Neurobiol. Aging 45, 212.e5–212.e11.
Redler R. L. and Dokholyan N. V 2012 The complex molecular biology of amyotrophic lateral sclerosis (ALS). Prog. Mol. Biol. Transl. Sci. 107, 215–262.
Reiman E. M., Webster J. A., Myers A. J., Hardy J., Dunckley T., Zismann V. L. et al. 2007 GAB2 alleles modify Alzheimer’s risk in APOE \(\upvarepsilon \)4 Carriers. Neuron 54, 713–720.
Reitz C., Wang L., Lin C., Larson E. B., Graff-radford N. R., Evans D. et al. 2013 Variants in the ATP-binding cassette transporter (ABCA7), apolipoprotein E4, and the risk of late-onset alzheimer disease in African Americans. J. Am. Med. Assoc. 309, 1483–1492.
Rivière J.-B., Mirzaa G. M., O’Roak B. J., Beddaoui M., Alcantara D., Conway R. L. et al. 2012 De novo germline and postzygotic mutations in AKT3, PIK3R2 and PIK3CA cause a spectrum of related megalencephaly syndromes. Nat. Genet. 44, 934–940.
Roos R. A. C. 2010 Huntington’s disease: a clinical review. Orphanet. J. Rare Dis. 5, 40.
Rosen D. R., Siddique T., Patterson D., Figlewicz D. A., Sapp P., Hentati A. et al. 1993 Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 362, 59–62.
Ross C. A. and Tabrizi S. J. 2011 Huntington’s disease: from molecular pathogenesis to clinical treatment. Lancet Neurol. 10, 83–98.
Rothstein J. D. 2017 Edaravone: a new drug approved for ALS. Cell 171, 725.
Rowland L. P. and Shneider N. A. 2001 Amyotrophic lateral sclerosis. N. Engl. J. Med. 344, 1688–1700.
Rubinsztein D., Leggo J., Chiano M., Dodge A., Norbury G., Rosser E. et al. 1997 Genotypes at the GluR6 kainate receptor locus are associated with variation in the age of onset of Huntington disease. Proc. Natl. Acad. Sci. USA 94, 3872–3876.
Saad M., Lesage S., Saint-Pierre A., Corvol J.-C., Zelenika D., Lambert J.-C. et al. 2011 Genome-wide association study confirms BST1 and suggests a locus on 12q24 as the risk loci for Parkinson’s disease in the European population. Hum. Mol. Genet. 20, 615–627.
Saunders A. M., Strittmatter W. J., De S., St George-Hyslop P. H. S., Pericak-Vance M., Joo S. H. et al. 1993 Association of apolipoprotein E allele epsilon 4 with late-onset familial and sporadic Alzheimer’s disease. Neurology 43, 1467–1472.
Savica R., Grossardt B. R., Bower J. H., Ahlskog J. E. and Rocca W. A. 2016 Time trends in theincidence of Parkinson disease. JAMA Neurol. 73, 981.
Scheltens P., Blennow K., Breteler M. M. B., de Strooper B., Frisoni G. B., Salloway S. et al. 2016 Alzheimer’s disease. Lancet 388, 505–517.
Schymick J. C., Scholz S. W., Fung H.-C., Britton A., Arepalli S., Gibbs J. R. et al. 2007 Genome-wide genotyping in amyotrophic lateral sclerosis and neurologically normal controls: first stage analysis and public release of data. Lancet Neurol. 6, 322–328.
Seshadri S., Fitzpatrick A. L., Ikram M. A., DeStefano A. L. and Breteler M. M. B. 2011 Genome-wide analysis of genetic loci associated with Alzheimer’s Disease. JAMA 303, 1832–1840.
Sharma P. 2003 Charaka Samhita (with English translation), 7th edition. Chaukamba Orientalia, Varanasi.
Shatunov A., Mok K., Newhouse S., Weale M. E., Smith B., Vance C., et al. 2010 Chromosome 9p21 in sporadic amyotrophic lateral sclerosis in the UK and seven other countries: a genome-wide association study. Lancet Neurol. 9, 986–994.
Sherrington R., Rogaev E. I., Liang Y., Rogaeva E. A., Levesque G., Ikeda M. et al. 1995 Cloning of a gene bearing missense mutations in early-onset familial Alzheimer’s disease. Nature 375754–375760.
Sherva R., Baldwin C. T., Inzelberg R., Vardarajan B., Cupples L. A., Lunetta K. et al. 2011 Identification of novel candidate genes for Alzheimer’s disease by autozygosity mapping using genome wide SNP data. J. Alzheimer’s Dis. 23, 349–359.
Shimizu N., Kitada T., Asakawa S., Hattori N., Matsumine H., Yamamura Y. et al. 1998 Mutations in the parkin gene cause autosomal recessive juvenile parkinsonism. Nature 392, 605–608.
Shoesmith C. L., Findlater K., Rowe A. and Strong M. J. 2007 Prognosis of amyotrophic lateral sclerosis with respiratory onset. J. Neurol. Neurosurg. Psychiatry 78, 629–631.
Siddique T., Figlewigz D. A., Pericak-Vance M. A., Haines J. L., Rouleau G. et al. 1991 Linkage of a gene causing familial amyotrophic lateral sclerosis to chromosome 21 and evidence of genetic-locus heterogeneity. N. Engl. J. Med. 324, 1381–1384.
Sidransky E. and Lopez G. 2012 The link between the GBA gene and parkinsonism. Lancet Neurol. 11, 986–998.
Siitonen A., Nalls M. A., Hernández D., Gibbs J. R., Ding J., Ylikotila P. et al. 2017 Genetics of early-onset Parkinson’s disease in Finland: exome sequencing and genome-wide association study. Neurobiol. Aging 53, 195.e7–195.e10.
Skipper L., Wilkes K., Toft M., Baker M., Lincoln S., Hulihan M. et al. 2004 Linkage disequilibrium and association of MAPT H1 in Parkinson disease. Am. J. Hum. Genet. 75, 669–677.
Snell R. G., MacMillan J. C., Cheadle J. P., Fenton I., Lazarou L. P., Davies P. et al. 1993 Relationship between trinucleotide repeat expansion and phenotypic variation in Huntington’s disease. Nat. Genet. 4, 393–397.
Spencer C. C. A., Plagnol V., Strange A., Gardner M., Paisan-Ruiz C., Band G. et al. 2011 Dissection of the genetics of Parkinson’s disease identifies an additional association 5’ of SNCA and multiple associated haplotypes at 17q21. Hum. Mol. Genet. 20, 345–353.
Strittmatter W. J., Saunders A. M., Schmechel D., Pericak-Vance M., Enghild J., Salvesen G. S. et al. 1993 Apolipoprotein E: high-avidity binding to beta-amyloid and increased frequency of type 4 allele in late-onset familial Alzheimer disease. Proc. Natl. Acad. Sci. USA 90, 1977–1981.
Sudhaman S., Muthane U. B., Behari M., Govindappa S. T., Juyal R. C. and Thelma B. K. 2016a Evidence of mutations in RIC3 acetylcholine receptor chaperone as a novel cause of autosomal-dominant Parkinson’s disease with non-motor phenotypes. J. Med. Genet. 53, 559–566.
Sudhaman S., Prasad K., Behari M., Muthane U. B., Juyal R. C. and Thelma B. 2016b Discovery of a frameshift mutation in podocalyxin-like (PODXL) gene, coding for a neural adhesion molecule, as causal for autosomal-recessive juvenile Parkinsonism. J. Med. Genet. 53, 450–456.
Sun X., Liu J., Crary J., Malagelada C., Sulzer D., A Greene L., Levy O. 2013 ATF4 Protects against neuronal death in cellular Parkinson’s disease models by maintaining levels of Parkin. J. Neurosci. 33, 2398–2407.
Tanner C. M., Ottman R., Goldman S. M., Ellenberg J., Chan P., Mayeux R. et al. 1999 Parkinson disease in twins: an etiologic study. JAMA 281, 341–346.
Thelma B. K. 2008 From genomics to ‘Ayurgenomics’. PHISPC series on history of science, philosophy and culture in Indian civilization volume: life and organicism, (ed. N. S. Rangaswamy and D. P. Chattopadhyaya).
Thelma B. K., Juyal R. C., Dodge H. H., Pandav R., Chandra V. and Ganguli M. 2001 APOE polymorphism in a rural older population-based sample in India. Hum. Biol. 73, 135–144.
Tobin J. E., Latourelle J. C., Lew M. F., Klein C., Suchowersky O., Shill H. A. et al. 2008 Haplotypes and gene expression implicate the MAPT region for Parkinson disease: The GenePD Study. Neurology 71, 28–34.
Toffoli M., Dreussi E., Cecchin E., Valente M., Sanvilli N., Montico M. et al. 2017 SNCA 3\(^\prime \) UTR genetic variants in patients with Parkinson’s disease and REM sleep behavior disorder. Neurol. Sci. 38, 1233–1240.
Tosto G., Fu H., Vardarajan B. N., Lee J. H., Cheng R., Reyes-dumeyer D. et al. 2015 F-box\(/\)LRR-repeat protein 7 is genetically associated with Alzheimer’ s disease. Ann. Clin. Transl. Neurol. 810–820.
Traynor B. J., Alexander M., Corr B., Frost E. and Hardiman O. 2003 An outcome study of riluzole in amyotrophic lateral sclerosis. J. Neurol. 250, 473–479.
Trotta L., Guella I., Soldà G., Sironi F., Tesei S., Canesi M. et al. 2012 SNCA and MAPT genes: independent and joint effects in Parkinson disease in the Italian population. Park Relat. Disord. 18, 257–262.
Twelves D., Perkins K. S. M. and Counsell C. 2003 Systematic review of incidence studies of Parkinson’s disease. Mov. Disord. 18, 19–31.
Vacic V., Ozelius L. J., Clark L. N., Bar-Shira A., Gana-Weisz M., Gurevich T. et al. 2014 Genome-wide mapping of IBD segments in an Ashkenazi PD cohort identifies associated haplotypes. Hum. Mol. Genet. 23, 4693–4702.
Valente E. M., Abou-Sleiman P. M., Caputo V., Muqit M. M. K., Harvey K., Gispert S. et al. 2004 Hereditary early-onset Parkinson’s disease caused by mutations in PINK1. Science 304, 1158–1160.
Van Cauwenberghe C., Van Broeckhoven C. and Sleegers K. 2016 The genetic landscape of Alzheimer disease: clinical implications and perspectives. Genet. Med. 18, 421–430.
Van Den Eeden S. K., Tanner C. M., Bernstein A. L., Fross R. D., Leimpeter A., Bloch D. A. et al. 2003 Incidence of Parkinson’s disease: variation by age, gender, and race/ethnicity. Am. J. Epidemiol. 157, 1015–1022.
van Es M. A., Van Vught P. W., Blauw H. M., Franke L., Saris C. G., Andersen P. M. et al. 2007 ITPR2 as a susceptibility gene in sporadic amyotrophic lateral sclerosis: a genome-wide association study. Lancet Neurol. 6, 869–877.
van Es M. A., van Vught P. W., Blauw H. M., Franke L., Saris C. G., Van Den Bosch L. et al. 2008 Genetic variation in DPP6 is associated with susceptibility to amyotrophic lateral sclerosis. Nat. Genet. 40, 29–31.
van Es M. A., Veldink J. H., Saris C. G. J., Blauw H. M., van Vught P. W. J., Birve A. et al. 2009 Genome-wide association study identifies 19p13.3 (UNC13A) and 9p21.2 as susceptibility loci for sporadic amyotrophic lateral sclerosis. Nat. Genet. 41, 1083–1087.
Van Hoecke A., Schoonaert L., Lemmens R., Timmers M., Staats K. A., Laird A. S. et al. 2012 EPHA4 is a disease modifier of amyotrophic lateral sclerosis in animal models and in humans. Nat. Med. 18, 1418–1422.
van Rheenen W., Shatunov A., Dekker A. M., McLaughlin R. L., Diekstra F. P., Pulit S. L. et al. 2016 Genome-wide association analyses identify new risk variants and the genetic architecture of amyotrophic lateral sclerosis. Nat. Genet. 48, 1043–1048.
Vekrellis K., Rideout H. J. and Stefanis L. 2004 Neurobiology of \(\upalpha \)-Synuclein. Mol. Neurobiol. 30, 001–022.
Watanabe Y., Himeda T. and Araki T. 2005 Mechanisms of MPTP toxicity and their implications for therapy of Parkinson’s disease. Med. Sci. Monit. 11, RA17-23.
Weishaupt J. H., Hyman T. and Dikic I. 2016 Common molecular pathways in amyotrophic lateral sclerosis and frontotemporal dementia. Trends Mol. Med. 22, 769–783.
Weydt P., Soyal S. M., Gellera C., DiDonato S., Weidinger C., Oberkofler H. et al. 2009 The gene coding for PGC-1\(\upalpha \) modifies age at onset in Huntington’s Diseas. Mol. Neurodegener. 4, 3.
Wheeler V. C., Persichetti F., McNeil S. M., Mysore J. S., Mysore S. S., MacDonald M. E. et al. 2007 Factors associated with HD CAG repeat instability in Huntington disease. J. Med. Genet. 44, 695–701.
Wijsman E. M., Pankratz N. D., Choi Y., Rothstein J. H., Faber K. M., Cheng R. et al. 2011 Genome-wide association of familial late-onset Alzheimer’s disease replicates BIN1 and CLU and nominates CUGBP2 in interaction with APOE. PLoS Genet. 7.
Wilke C., Baets J., De Bleecker J. L., Deconinck T., Biskup S., Hayer S. N. et al. 2017 Beyond ALS and FTD: the phenotypic spectrum of TBK1 mutations includes PSP-like and cerebellar phenotypes. Neurobiol. Aging 62, e9–e13.
Wirdefeldt K., Gatz M., Reynolds C. A., Prescott C. A. and Pedersen N. L. 2011 Heritability of Parkinson disease in Swedish twins: a longitudinal study. Neurobiol. Aging 32, e1–e8.
Wisniewski T. and Frangione B. 1992 Apolipoprotein E: a pathological chaperone protein in patients with cerebral and systemic amyloid. Neurosci. Lett. 135, 235–238.
Zu T., Liu Y., Banez-Coronel M., Reid T., Pletnikova O., Lewis J., Miller T. M. et al. 2013 RAN proteins and RNA foci from antisense transcripts in C9ORF72 ALS and frontotemporal dementia. Proc. Natl. Acad. Sci. USA 110, E4968–E4977.
Acknowledgements
We gratefully acknowledge Dr Ramesh Juyal for critical reading of the manuscript. We thank funding support vide grant no. BT/PR14500/MED/12/478/2010 from Department of Biotechnology, Govt. of India for work on genetics of Parkinson’s disease and grant no. SR/S2/JCB-44/2011 JC Bose national fellowship to BKT; Senior Research Fellowship to SK from Council of Scientific and Industrial Research, New Delhi; and Junior Research fellowship to NY from University Grants Commission, New Delhi.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kumar, S., Yadav, N., Pandey, S. et al. Advances in the discovery of genetic risk factors for complex forms of neurodegenerative disorders: contemporary approaches, success, challenges and prospects. J Genet 97, 625–648 (2018). https://doi.org/10.1007/s12041-018-0953-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12041-018-0953-5