Abstract
Wilson disease (WD) is an autosomal recessive disorder of copper metabolism. The gene responsible for WD was discovered in 1993 and is located on chromosome 13 at 13q14.3. It encodes a copper-specific transporting P-type ATPase. Early diagnosis can improve treatment outcome and decrease the rate of disability or even mortality. We used Sanger sequencing to identify mutation hot spots in 55 northern Vietnamese with a clinical diagnosis of WD. Mutations were screened and detected by direct DNA sequencing. A total of 26 different ATP7B gene mutations were identified, including seven novel mutations (five nonsense and two missense mutations). The most frequent mutations were p.Ser105Ter (24.55%), p.Arg778Leu (5.45%) and p.Thr850Ile (4.55%). Mutation detection rate in exon 2 was 34.55% and ranked first, followed by exon 8 with 16.36%, and exon 18 with 10.91% each, thus, exons 2, 8 and 18 are the mutation hot spots for northern Vietnamese WD patients. These findings were different from previous studies in Asia. Our research established a suitable strategy for ATP7B gene testing in northern Vietnamese WD patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Progressive hepatolenticular degeneration also known as Wilson disease (WD; OMIM: 277900) was first defined as a syndrome in 1912. It is a rare autosomal recessive genetic disorder of copper metabolism in which excessive amounts accumulate in the body, particularly in the liver, brain and eyes (Kayser–Fleischer ring in the cornea). Biochemical indicators for the disease include low serum concentrations of ceruloplasmin (\({<}20\,\hbox {g/L}\)) and elevated excretion of urinary copper (\({>}100\,\mu \hbox {g/24}\,\hbox {h}\)) (Sternlieb 1990). Early-onset presentations in infancy and late disease-onset manifestations in adults older than 70 years of age are now well recognized (Figus et al. 1995; Vajro et al. 2013).
WD is caused by mutations in the ATP7B gene, discovered in 1993, that encodes a copper-specific transporting P-type ATPase and the gene is located on chromosome 13 at 13q14 (Bull et al. 1993; Tanzi et al. 1993).
The widely cited prevalence figure of one in 30,000 for WD with a heterozygous carrier frequency of one in 90 was estimated in 1984 and thus, predates the identification of ATP7B as the causative gene. This prevalence estimate was at least partly based on assumptions, and has been questioned (Bull et al. 1993). More recent data from population screening of WD in the UK suggested a potentially higher rate of ATP7B heterozygote mutation carriers, predicting one in 7021 prevalence in the UK population (Coffey et al. 2013). Results from biochemical and genetic prevalence studies suggest that WD might be much more common than previously estimated and may vary by population (Duc et al. 1998; Terada et al. 1998). Early diagnosis can improve treatment outcome and decrease the rate of disability and mortality (Bull et al. 1993; Tanzi et al. 1993).
The ATP7B gene consists of 21 exons and 20 introns. It is \({\sim }7.5\,\hbox {kb}\) in size and encodes a 1465 amino acid–protein that consists of six copper-binding sites (exons 2–5), eight transmembrane domains of copper channel (exons 6–8, 12–13, 19–20) and the ATP-binding domain (exons 10–11 and 14–18). The copper transportation is provided by converting ATP hydrolysis energy in the ATP-binding domain (Terada et al. 1998).
Untill now more than 500 mutations were identified in the ATP7B gene, as detailed in the database of the University of Alberta, Canada (www.wilsondisease.med.ualberta.ca/database.asp). Mutations in ATP7B are scattered in the whole gene, but some hot spots were reported to be varying in different populations. The mutation hot spots in Europeans and North Americans were identified in exon 14, p.His1069Gln (Thomas et al. 1995; Duc et al. 1998; Riordan and Williams 2001; Kucinskas et al. 2008). In some Asian countries, such as China, Korea and Taiwan, the hot spot lies in exon 8 with the p.Arg778Leu mutation (Yoo 2002; Liu et al. 2004; Wan et al. 2006; Li et al. 2013; Diao et al. 2014; Wei et al. 2014). Identification of mutation hot spots may significantly reduce the time of genetic test processing. However, in Vietnam, no such study was conducted previously. In this paper, we used Sanger sequencing to identify the mutation spectrum in northern Vietnamese WD patients to investigate whether any mutation hot spot exist that would facilitate genetically diagnosis confirmation.
Materials and methods
Patients
Fifty-five WD patients from unrelated families in northern Vietnam were enrolled in this study. All WD patients were diagnosed and treated at the National Pediatrics Hospital in Hanoi from 2010 to 2015. Diagnosis of WD was based on many clinical symptoms and signs, including acute or chronic liver failure and/or typical neurological symptoms, or the presence of Kayser–Fleischer ring and biochemical parameters, such as low serum ceruloplasmin (<0.2 \(\hbox {g/L}\)) and high level of urinary copper (\({>}100\,\mu \hbox {g/24}\, \hbox {h}\)) (Roberts et al. 2008). Forty healthy northern Vietnamese individuals were enrolled as controls. Informed consent was obtained from all the patient families for molecular analysis and this study was approved by the ethical committees of Hanoi Medical University (IRB00003121 Hanoi Med U IRB, Hanoi, Vietnam).
DNA extraction
Genomic DNA samples were extracted from peripheral blood collected in EDTA-coated tubes using Wizard genomic DNA purification kit (Promega, Madison, USA) following the manufacturer’s recommendations.
Polymerase chain reaction (PCR)
The full-length gene was amplified by using primer pairs for exons 1–21 (IDT, Coralville, USA). PCR was performed using GoTaq Green Master Mix (Promega) with 100 ng of genomic DNA in a mix containing 10 pmol of each primer, \(12.5\,\mu \hbox {L}\) of 2\(\times \) GoTaq Green Master mix in a total volume of \(25\,\mu \hbox {L}\). The thermocycle programme consisted of an initial denaturation at \(94{^{\circ }}\hbox {C}\) for 5 min, followed by 35 cycles at \(94{^{\circ }}\hbox {C}\) for 30 s, \(56{^{\circ }}\hbox {C}\) for 30 s and \(72{^{\circ }}\hbox {C}\) for 30 s, with a final extension at \(72{^{\circ }}\hbox {C}\) for 5 min. The size and quantity of PCR products were verified by electrophoresis in 2% (w/v) agarose gel.
DNA sequencing
PCR products were directly sequenced using an Advant 3100 automated sequencer (Applied Biosystems, Foster City, USA). Sequences were aligned and inspected using a reference sequence from GenBank (NM_0000.53). DNA sequencing was used to detect variations of the entire coding region of ATP7B gene from the 55 patients and 40 healthy controls.
Results
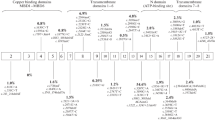
We investigated 55 patients with WD and detected 26 different ATP7B gene mutations, including 17 missense, six nonsense mutations, two frame-shift deletions and one frame-shift insertion. Of these, seven mutations were not previously reported (novel mutations) that included two missense (p.Asp1027His and p.Asn1270Asp) and five nonsense mutations (p.Glu45Ter, p.Met119Ter, p.Lys867Ter, p.Glu905Ter and p.Leu1159Ter) (table 1). All novel missense mutations were tested for the possibilities being pathogenic in nature using Alamut Visual ver. 2.7 (Interactive Biosoftware, Rouen, France) and were not found in 80 alleles of 40 healthy controls. Additionally, five novel variants were detected (p.Glu583Gln, p.Pro1098Gln, p.Gly1099Asp, p.Gly1213Asp and p.Cys980Ser) where in silico analysis was either predicted to be tolerated, partly based on the conservation of amino acid residues or in one case (p.Cys980Ser) predicted to be benign by PolyPhen-2 (table 2). The result of sequencing and distribution of the novel mutations and variants are shown in figure 1. Four different mutations were identified on exons 2, 8 or 18, three on exon 14 or 16, two on exon 5 or 11 and one on exons 10, 12, 13 or 20; whereas, no mutation was found on exons 1, 3, 4, 6, 7, 9, 15, 17, 19, 21 and in the promoter region. In exon 2, four mutations were found in 19 patients (one patient with p.Glu45Ter and p.Ser105Ter), with a detection rate of 34.55% (19/55) and ranked first; two other most frequent exon mutations, exon 8 detected in nine patients, exon 18 detected in six patients, accounted for 16.36% (9/55) and 10.91% (6/55), respectively. Overall, p.Ser105Ter is the most frequent mutation in this study. It was detected in 18 cases with nine homozygous, eight compound heterozygous and one single heterozygous, at a detection rate of 32.73% (18/55) and an allele frequency of 24.55% (27/110). p.Arg778Leu, the common mutation in Chinese population stayed in exon 8 (Li et al. 2013; Wei et al. 2014) was found in four compound and two single heterozygous cases, accounted for 10.9% (6/55) of cases and 5.45% (6/110) of studied alleles became the second most frequent mutation. Following p.Thr850Ile reveled in four compound and one single heterozygous cases, contributed for 9.1% of cases and 4.55% of studied alleles. Interestingly, we found a novel mutation p.Glu905Ter on exon 11 which was prevalent in Vietnamese population. Homozygous mutation p.Glu905Ter was identified in two patients accounted for cases and allele frequency of 3.64%. We found that, exons 2, 8 and 18 were thus, recognized as hot spots for WD mutation detection in this study. The total mutation detection rate on these three exons was 52.73% (29/55). The most frequent mutations are p.Ser105Ter (24.55%), p.Arg778Leu (5.45%) and p.Thr850Ile (4.55%).
In addition to the mutations, six single-nucleotide polymorphisms (SNPs) were identified and their details are provided in table 3. These base substitutions were defined as polymorphisms because they were predicted as polymorphisms by Alamut Visual ver. 2.7 (Interactive Biosoftware, Rouen, France), or existed in healthy controls and were demonstrated previously (Haas et al. 1999; Gu et al. 2003; Wan et al. 2006; Gupta et al. 2007).
Discussion
Copper is an essential component of many enzymes such as lysyl oxidase, superoxide dismutase, dopamine-\(\upbeta \)-hydroxylase and cytochrome C oxidase. These copper-dependent enzymes are needed for diverse processes of oxidase metabolism including respiration, free-radical detoxification, neurotransmitter synthesis, maturation of connective tissue and iron uptake (Yuan et al. 1995; Linder and Hazegh-Azam 1996). However, copper is only required in trace amount; accumulation of copper can damage plasma membranes, peroxisomes, mitochondria, microtubules, enzymes and even DNA (Duc et al. 1998). Typical presentations of WD include neuropsychiatric and hepatic dysfunctions, whereas a typical presentation is extremely variable. Diagnosis relies typically on a high clinical suspicion, typical neurological symptoms, presence of Kayser–Fleischer rings, and reduced serum ceruloplasmin concentration. The conventional value of \({<}0.20\,\hbox {g}/\hbox {L}\) is not a universal diagnostic value. Age of the subjects and analytical variations should be considered when interpreting these levels. Patients with inconclusive findings require further investigations including 24-h urinary free-copper excretion, penicillamine challenge test, liver copper measurement, and more recently detection of gene mutations. Direct molecular diagnosis remains the most decisive test.
Early diagnosis and treatment of WD are associated with better outcome. ATP7B gene testing is proved as a suitable method for prenatal diagnosis and neonatal screening (Roberts et al. 2008). At present, more than 500 mutations in the ATP7B gene are listed in the WD mutation database. In this study, we identified 26 different mutations including seven novel mutations in 55 WD patients from northern Vietnam. Mutations exist as compound heterozygous, homozygous and single heterozygous forms. The mutation detection rate of exon 2 was 34.55% and ranked first, followed by exon 8 with 16.36% and exon 18 with 10.91%, we recognized exons 2, 8 and 18, which can cover 52.73% of mutations as the hot spots for northern Vietnamese WD patients. Our result was different from previous studies in Asian populations. Most mutations located on exons 8, 12, 13 and 16 in northern Chinese covering 60.5–74% (Wu et al. 2001; Liu et al. 2004); on exons 8, 11 and 18 in Korean covering 59.8–71.4% (Yoo 2002; Park et al. 2007); and on exons 5, 8, 12, 13 and 18 in Japanese with coverage of 59.8–71.4% (Shimizu et al. 1999; Okada et al. 2000) mutations. Thus, these three exons 2, 8 and 18 should be screened first in our upcoming ATP7B genetic testing. We could not find any trace of mutations in exons 1, 3, 4, 6, 7, 9, 15, 17, 19, 21 and promoter region in this series. Perhaps in these regions where there are large deletion and duplication mutations that are not detectable by sequencing, stayed as heterozygous, which should be examined and detected by other method such as MLPA, particularly, when a single heterozygous mutation has been detected at sequencing. This is a limitation of our study, which had limited funding. Our study for the first time provides the mutation spectrum in WD in a Vietnamese population.
The sequencing results and distribution of novel mutations and variants in ATB7B gene. DNA sequences were shown with highlighted letters indicating for the substitution nucleotides of (a) nonsense, (b) missense mutations and (c) variants, respectively. (d) Illustration for distribution of novel mutations and variants in ATB7B gene.
The p.Ser105Ter mutation on exon 2 was most common in our cohort, accounting for 24.55% of diagnosed alleles. The second most frequent mutations were p.Arg778Leu on exon 8 with 5.45%, following the mutation p.Thr850Ile on exon 10 accounted for 4.55% of studied alleles. These findings were different from the results of previous studies, in other continents, as well as in related regions in Asia. The p.His1069Gln is the most common mutation in European and North American populations (Thomas et al. 1995; Duc et al. 1998; Riordan and Williams 2001; Kucinskas et al. 2008), and the common mutation in Indian population, the p.Cys271Ter (Aggarwal et al. 2013; Mukherjee et al. 2014), but we could not detect any case in our patients. p.Arg778Leu was recognized as the most frequent mutation in Chinese (Liu et al. 2004; Li et al. 2013; Diao et al. 2014; Wei et al. 2014), Korean (Yoo 2002; Park et al. 2007) and Taiwanese (Wan et al. 2006) populations. This mutation was ranked as second in our cohort but the allele frequency (5.45%) is much lower than those of other previous studies in Asia (Liu et al. 2004: 74%; Li et al. 2013: 21.5%; Yoo 2002: 37.9%). Also, other relatively common mutations in these studies, such as p.Pro992Leu, p.Thr935Met or p.Ala874Val, were not found in our study. On the other hand, the most common mutation in our cohort, p.Ser105Ter was found only in a few cases in the Chinese population (Liu et al. 2004; Mak et al. 2008). But we detected novel mutation which was prevalent in this study as p.Glu905Ter (3.64%). In addition, five novel variants were detected in our study where in silico analysis was either predicted to be tolerated, partly based on the conservation of amino acid residues or in one case (p.Cys980Ser) predicted to be benign by PolyPhen-2. Although, none of these novel variants were present in the 1000 Genomes or the ExAC databases. Currently, these variants, at the best can be considered as possibly pathogenic and might be relatively common in Vietnamese population but this remains to be proven in the future.
In summary, we have revealed the mutation spectrum of the ATP7B gene in northern Vietnamese WD patients with seven novel mutations identified. Our study provided an additional data for understanding mutation patterns in the ATP7B gene worldwide. Direct sequencing has proved a sensitive, specific and relatively low invasive method and it is increasingly used for ATP7B gene testing for early diagnosis confirmation and prenatal diagnosis of WD. Similar analysis in Vietnamese patients with WD from other regions of Vietnam is warranted to provide a better assessment of the mutation spectrum of this disorder in Vietnam. We recommend screening of exons 2, 8 and 18 which can cover 52.73% of mutations. This finding is expected to reduce time and costs of mutation screening significantly.
References
Aggarwal A., Chandhok G., Todorov T., Parekh S., Tilve S., Zibert A. et al. 2013 Wilson disease mutation pattern with genotype-phenotype correlations from Western India: confirmation of p.C271* as a common Indian mutation and identification of 14 novel mutations. Ann. Hum. Genet. 77, 299–307.
Bull P. C., Thomas G. R., Rommens J. M., Forbes J. R. and Cox D. W. 1993 The wilson disease gene is a putative copper transporting P-type ATPase similar to the Menkes gene. Nat. Genet. 5, 327–337.
Coffey A. J., Durkie M., Hague S., McLay K., Emmerson J., Lo C. et al. 2013 A genetic study of Wilson’s disease in the United Kingdom. Brain 136, 1476–1487.
Diao S. P., Hong M. F., Huang Y. Q., Wei Z. S., Su Q. X., Peng Z. X. et al. 2014 Identification and characterization of a novel splice-site mutation in the Wilson disease gene. J. Neurol. Sci. 345, 154–158.
Duc H. H., Hefter H., Stremmel W., Castaneda-Guillot C., Hernandez A., Cox D. W. et al. 1998 His1069Gln and six novel Wilson disease mutations: analysis of relevance for early diagnosis and phenotype. Eur. J. Hum. Genet. 6, 616–623.
Figus A., Angius A., Loudianos G., Bertini C., Dessi V., Loi A. et al. 1995 Molecular pathology and haplotype analysis of Wilson disease in Mediterranean populations. Am. J. Hum. Genet. 57, 1318–1324.
Gu Y. H., Kodama H., Du S. L., Gu Q. J., Sun H. J. and Ushijima H. 2003 Mutation spectrum and polymorphisms in ATP7B identified on direct sequencing of all exons in Chinese Han and Hui ethnic patients with wilson’s disease. Clin. Genet. 64, 479–484.
Gupta A., Maulik M., Nasipuri P., Chattopadhyay I., Das S. K., Gangopadhyay P. K. et al. 2007 Molecular diagnosis of wilson disease using prevalent mutations and informative single nucleotide polymorphism markers. Clin. Chem. 53, 1601–1608.
Haas R., Gutierrez-Rivero B., Knoche J., Boker K., Manns M. P. and Schmidt H. H. 1999 Mutation analysis in patients with wilson disease: identification of 4 novel mutations. Hum. Mutat. 14, 88.
Kucinskas L., Jeroch J., Vitkauskiene A., Sakalauskas R., Petrenkiene V., Kucinskas V. et al. 2008 High frequency of the \(\text{ c }.3207\text{ C }>\text{ A }\) (p.H1069Q) mutation in ATP7B gene of Lithuanian patients with hepatic presentation of Wilson’s disease. World J. Gastroenterol. 14, 5876–5879.
Li K., Zhang W. M., Lin S., Wen L., Wang Z. F., Xie D. et al. 2013 Mutational analysis of ATP7B in north Chinese patients with wilson disease. J. Hum. Genet. 58, 67–72.
Linder M. C. and Hazegh-Azam M. 1996 Copper biochemistry and molecular biology. Am. J. Clin. Nutr. 63, 797–811.
Liu X. Q., Zhang Y. F., Liu T. T., Hsiao K. J., Zhang J. M., Gu X. F. et al. 2004 Correlation of ATP7B genotype with phenotype in Chinese patients with Wilson disease. World J. Gastroenterol. 10, 590–593.
Mak C. M., Lam C. W., Tam S., Lai C. L., Chan L. Y., Fan S. T. et al. 2008 Mutational analysis of 65 Wilson disease patients in Hong Kong Chinese: identification of 17 novel mutations and its genetic heterogeneity. J. Hum. Genet. 53, 55–63.
Mukherjee S., Dutta S., Majumdar S., Biswas T., Jaiswal P., Sengupta M. et al. 2014 Genetic defects in Indian Wilson disease patients and genotype-phenotype correlation. Parkinsonism Relat. Disord. 20, 75–81.
Okada T., Shiono Y., Hayashi H., Satoh H., Sawada T., Suzuki A. et al. 2000 Mutational analysis of ATP7B and genotype–phenotype correlation in Japanese with wilson’s disease. Hum. Mutat. 15, 454–462.
Park S., Park J. Y., Kim G. H., Choi J. H., Kim K. M., Kim J. B. et al. 2007 Identification of novel ATP7B gene mutations and their functional roles in Korean patients with wilson disease. Hum. Mutat. 28, 1108–1113.
Riordan S. M. and Williams R. 2001 The wilson’s disease gene and phenotypic diversity. J. Hepatol. 34, 165–171.
Roberts E. A., Schilsky M. L. and American Association for Study of Liver Disease (AASLD) 2008 Diagnosis and treatment of wilson disease: an update. Hepatology 47, 2089–2111.
Shimizu N., Nakazono H., Takeshita Y., Ikeda C., Fujii H., Watanabe A. et al. 1999 Molecular analysis and diagnosis in Japanese patients with wilson’s disease. Pediatr. Int. 41, 409–413.
Sternlieb I. 1990 Perspectives on wilson’s disease. Hepatology 12, 1234–1239.
Tanzi R. E., Petrukhin K., Chernov I., Pellequer J. L., Wasco W., Ross B. et al. 1993 The wilson disease gene is a copper transporting ATPase with homology to the Menkes disease gene. Nat. Genet. 5, 344–350.
Terada K., Schilsky M. L., Miura N. and Sugiyama T. 1998 ATP7B (WND) protein. Int. J. Biochem. Cell. Biol. 30, 1063–1067.
Thomas G. R., Forbes J. R., Roberts E. A., Walshe J. M. and Cox D. W. 1995 The Wilson disease gene: spectrum of mutations and their consequences. Nat. Genet. 9, 210–217.
Vajro P., Maddaluno S. and Veropalumbo C. 2013 Persistent hypertransaminasemia in asymptomatic children: a stepwise approach. World J. Gastroenterol. 19, 2740–2751.
Wan L., Tsai C. H., Tsau Y., Hsu C. M., Lee C. C. and Tsai F. J. 2006 Mutation analysis of Taiwanese wilson disease patients. Biochem. Biophys. Res. Commun. 345, 734–738.
Wei Z., Huang Y., Liu A., Diao S., Yu Q., Peng Z. et al. 2014 Mutational characterization of ATP7B gene in 103 wilson’s disease patients from Southern China: identification of three novel mutations. Neuroreport 25, 1075–1080.
Wu Z. Y., Wang N., Lin M. T., Fang L., Murong S. X. and Yu L. 2001 Mutation analysis and the correlation between genotype and phenotype of Arg778Leu mutation in Chinese patients with Wilson disease. Arch. Neurol. 58, 971–976.
Yoo H. W. 2002 Identification of novel mutations and the three most common mutations in the human ATP7B gene of Korean patients with wilson disease. Genet. Med. 4, 43–48.
Yuan D. S., Stearman R., Dancis A., Dunn T., Beeler T. and Klausner R. D. 1995 The menkes/wilson disease gene homologue in yeast provides copper to a ceruloplasmin-like oxidase required for iron uptake. Proc. Natl. Acad. Sci. USA 92, 2632–2636.
Acknowledgements
We thank the patients and their families for their voluntary involvement in this study. This work was supported by National Foundation for Science and Technology Development (NAFOSTED) research fund, Vietnam.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Arun Kumar
Rights and permissions
About this article
Cite this article
Pham, L.A.T., Nguyen, T.T., Nga Le, H.B. et al. Genetic analysis of 55 northern Vietnamese patients with Wilson disease: seven novel mutations in ATP7B . J Genet 96, 933–939 (2017). https://doi.org/10.1007/s12041-017-0857-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12041-017-0857-9