Abstract
The BRD4 protein is associated with various diseases, which has been an attractive target for the treatment of cancer and inflammation. This paper is a follow-up to our previous studies, in which we report the structure-based design, synthesis, and evaluation of a new class of small-molecule BRD4 bromodomain inhibitors bearing a benzo[d]isoxazol scaffold. The SARs focused on exploration of the 2′ or 3′ position to afford novel inhibitors that may avoid potential metabolically unstable site. The most potent inhibitor 13f exhibited high binding affinity to BRD4(1) with a ΔTm value of 7.8 °C as evaluated in thermal shift assay (TSA). The potent activity was also demonstrated by a peptide competition assay with an IC50 value of 0.21 μM. The docking studies revealed the binding mode of the compounds with the active site of BRD4(1). In addition, in silico predictions indicated that these compounds possessed good drug-likeness and pharmacokinetic profile.
Graphic abstract
This paper is a follow-up to our previous studies, in which we report the structure-based design, synthesis, and evaluation of a new class of small-molecule BRD4 bromodomain inhibitors bearing a benzo[d]isoxazol scaffold.

This paper is a follow-up to our previous studies, in which we report the structure-based design, synthesis, and evaluation of a new class of small-molecule BRD4 bromodomain inhibitors bearing a benzo[d]isoxazol scaffold.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Acetylation of histone is a classical post-translational modification in the field of epigenetics, which plays an important role in the regulation of chromatin structure. ε-N-acetylation of lysine residues on the amino-terminal tails of histones was discovered 30 years ago and has been associated with the opening of chromatin architecture.1 The opened structure can be accessed by DNA and RNA polymerases as well as transcription factors, resulting in the activation of gene transcription. Acetylation level on histones is highly regulated by histone acetyltransferases (HATs; enzymes that produce or write acetylation marks) and histone deacetylases (HDACs; enzymes that erase acetylation marks), wherein bromodomains act as readers of the acetyl lysine to participate in this regulation process.2,3
The human proteome encodes 61 bromodomains, which are existed in 46 diverse nuclear and cytoplasmic proteins. These bromodomains can be phylogenetically divided into eight distinct families.4 The BET family consists of BRD2, BRD3, BRD4, and BRDT. These proteins bind to the acetylated lysine via their tandem BD1 (expressed as BRD4(1)) and BD2 (expressed as BRD4(2)) domains. The BRD4 protein has been associated with several human diseases and conditions including cancers such as acute myeloid leukemia (AML),5,6 MLL,7 gastrointestinal stromal tumor (GIST),8 triple-negative breast cancer,9,10 prostate cancer,11,12,–13 neuroblastoma,14 pancreatic cancer,15 cholangiocarcinoma,16 as well as inflammations.17,18,–19
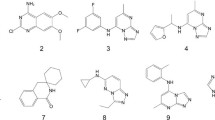
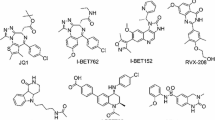
The first BRD4 bromodomain inhibitor was a triazolobenzodiazepine derivative named as (+)-JQ1 (1). This compound has been widely used as a probe for exploring the function of BET inhibition.20 Another triazolobenzodiazepine derivative GSK525762 (I-BET762) was reported by Nicodeme et al. (Figure 1),18 which has entered clinical trials for neoplasms.21,22 The second class were 3,5-dimethylisoxazole containing BRD4 inhibitors such as I-BET151.23 The 3,5-dimethylisoxazole was an effective warhead and has been introduced to different BET inhibitors.24,25,–26 We previously reported the design of a class of inhibitors containing a benzo[cd]indol-2(1H)-one scaffold, as exemplified by compound 2,27 and a class of inhibitors containing a benzo[d]isoxazol scaffold, as exemplified by compound 3 (Figure 1).28 Compounds 3 demonstrated good therapeutic effects in a C4-2B prostate cancer xenograft tumor model in mice.28 In this paper, we aimed to extend the SAR studies on the 2′ and 3′ position of compound 3, and a series of new compounds was designed, synthesized and evaluated as BRD4(1) inhibitors.
2 Experimental
2.1 General chemistry
Melting points were recorded on X-6 melting point apparatus (Beijing Tech instrument, Beijing, China). TLC analysis was performed on GF254 pre-coated silica gel plate (Qingdao Haiyang Chemical, Qingdao, China) and visualized with UV light. 1H and 13C nuclear magnetic resonance (NMR) spectroscopy was performed on a Bruker AV-400 or AV-500 instrument (Bruker, Fällanden, Switzerland) with DMSO-d6 or CDCl3 as the solvent. Tetramethylsilane (TMS) was used as an internal standard. The coupling constants (J) are expressed in hertz (Hz), the chemical shifts (δ) are reported in parts per million (ppm). Signal multiplicities are represented as singlet (s), broad singlet (brs), doublet (d), dd (double doublet), triplet (t), and multiplet (m). The ESI-MS were recorded on an Agilent 1200 HPLC-MSD mass spectrometer (Agilent, California, USA). All solvents and reagents were commercially available and used without further purification.
2.2 General procedures for the preparation of intermediates 9 and 12a–e
2.2.1 Ethyl 2-(4-bromo-2-(chlorosulfonyl)phenoxy) acetate (12c)
Step 1: Synthesis of ethyl 2-(4-bromophenoxy)acetate (11c). To a solution of 4-bromophenol (10a) (3.0 g, 17.34 mmol) in DMF (10 mL) was added K2CO3 (7.19 g, 52.02 mmol) followed by the addition of ethyl chloroacetate (2.55 g, 20.81 mmol). The mixture was stirred at 80 °C for 4 h. After the reaction was completed (monitored by TLC), water was added to the mixture. The precipitated solid was filtered and dried to give the title compound as a white solid. Yield: 99%. 1HNMR (CDCl3, 400 MHz), δ 7.39 (d, 2H, J = 9.0 Hz, 3′, 5′-ArH), 6.80 (d, 2H, J = 9.0 Hz, 2′, 6′-ArH), 4.59 (s, 2H, –OCH2CO–), 4.27 (q, 2H, J = 7.0 Hz, –COOCH2–), 1.30 (t, 3H, J = 7.3Hz, –CH2CH3); ESI-MS (m/z): 283.3 [M + Na]+. Step 2: Synthesis of ethyl 2-(4-bromo-2-(chlorosulfonyl)phenoxy)acetate (12c). To an ice-cooled solution of compound 11c (400 mg, 1.54 mmol) in CH2Cl2 (2 mL) was added a solution of chlorosulfonic acid (1.08 g, 9.26 mmol) in CH2Cl2 (4 mL) over a period of 2 min. The reaction was stirred at room temperature for 45 min. Thionyl chloride (2.39 g, 20.07 mmol) was added slowly to the reaction and stirred for additional 3.5 h. After the reaction was completed (monitored by TLC), the mixture was poured into ice-water and extracted with ethyl acetate. The organic layer was washed with water, dried over Na2SO4 and concentrated under reduced pressure to give the title compound as a white solid. Yield: 82%. M.p. 124–126 °C; 1H NMR (CDCl3, 400 MHz) δ 8.11 (d, 1H, J = 2.4 Hz, 3′-ArH), 7.74 (dd, 1H, J = 8.9, 2.4 Hz, 5′-ArH), 6.88 (d, 1H, J = 8.9 Hz, 6′-ArH), 4.86 (s, 2H, –OCH2CO–), 4.28 (q, 2H, J = 7.1 Hz, –OCH2CH3), 1.29 (t, 3H, J = 7.1 Hz, –CH2CH3); 13C NMR (126 MHz, CDCl3) δ 167.20, 154.78, 139.66, 133.60, 132.44, 115.85, 113.42, 66.33, 62.15, 14.24; ESI-MS (m/z): 380.9 [M + Na]+.
2.2.2 5-Bromo-2,3-dihydrobenzofuran-7-sulfonyl chloride (12e)
To an ice-cooled solution of compound 11d (400 mg, 2.01 mmol) in CH2Cl2 (2 mL) was added a solution of chlorosulfonic acid (1.4 g, 12.06 mmol) in CH2Cl2 (4 mL) over a period of 2 min. The reaction was stirred at room temperature for 2.5 h. On completion of the reaction, the mixture was poured into ice water and the CH2Cl2 was evaporated under reduced pressure. The precipitated solid was filtered and dried to give the target compound as a white solid. Yield: 75%. M.p. 123–124 °C. 1H NMR (400 MHz, CDCl3) δ 7.81–7.74 (m, 1H, ArH), 7.60 (d, J = 1.7 Hz, 1H, ArH), 4.91 (t, J = 8.8 Hz, 2H, –OCH2CH2–), 3.35 (t, J = 8.8 Hz, 2H, –CH2CH2Ar); 13C NMR (126 MHz, CDCl3) δ 156.82, 135.08, 134.02, 128.72, 126.88, 111.89, 74.58, 28.94; ESI-MS (m/z): 321.0 [M + Na]+.
The procedures for the preparation of intermediates 9, 11d, 12a–b and 12d were described in supplementary information.
2.3 General procedures for the preparation of compounds 13a–h
2.3.1 5-Bromo-2-ethoxy-N-(6-methoxy-3-methylbenzo[d]isoxazol-5-yl)benzenesulfonamide (13a)
To a solution of compound 9 (100 mg, 0.56 mmol) in CH2Cl2 (6 mL) was added 5-bromo-2-ethoxybenzenesulfonyl chloride (184.9 mg, 0.62 mmol) followed by the addition of pyridine (0.3 mL). The resulting mixture was stirred at 43 °C for 3 h. Upon completion of the reaction (monitored by TLC, ethyl acetate/light petroleum (1:1, v/v)), the mixture was cooled and diluted with 1 mol/L HCl (8 mL) and water (12 mL), which was then extracted with ethyl acetate (3 × 20 mL). The organic layer was washed successively with water and brine, dried over Na2SO4 and concentrated under reduced pressure. The residue was purified by silica gel chromatography to give the title compound as a white solid. Yield: 65%. M.p. 173–174 °C; 1H NMR (400 MHz, CDCl3) δ 7.95 (d, J = 2.5 Hz, 1H, 6′-ArH), 7.71 (s, 1H, 7-ArH), 7.58–7.50 (m, 2H, 4′-ArH, –NH–), 6.90 (s, 1H, 4-ArH), 6.83 (d, J = 8.8 Hz, 1H, 3′-ArH), 4.18 (q, J = 7.0 Hz, 2H, –CH2–), 3.83 (s, 3H, 6-OCH3), 2.52 (s, 3H, 3-CH3), 1.56 (t, J = 7.0 Hz, 3H, –CH2CH3); 13C NMR (101 MHz, CDCl3) δ 160.89, 155.21, 155.02, 152.28, 137.49, 133.05, 128.42, 123.38, 115.17, 114.44, 112.38, 112.18, 91.92, 65.33, 56.28, 14.58, 10.07; ESI-MS (m/z): 441.0 and 443.0 [M + H]+.
2.3.2 5-Bromo-2-hydroxy-N-(6-methoxy-3-methylbenzo[d]isoxazol-5-yl)benzenesulfonamide (13b)
White solid. Yield: 66%. M.p. 202–203°C; 1H NMR (400 MHz, CDCl3) δ 7.69 (s, 1H, 7-ArH), 7.57 (d, J = 2.4 Hz, 1H, 6′-ArH), 7.45 (dd, J = 8.9, 2.4 Hz, 1H, 4′-ArH), 6.87 (s, 1H, 4-ArH), 6.78 (d, J = 8.9 Hz, 1H, 3′-ArH), 3.69 (s, 3H, 6-OCH3), 2.55 (s, 3H, 3-CH3); 13C NMR (101 MHz, DMSO) δ 161.48, 155.73, 155.06, 154.94, 136.79, 130.88, 127.82, 123.13, 119.35, 117.90, 114.08, 108.60, 92.38, 56.19, 9.46; ESI-MS (m/z): 412.9 and 414.9 [M + H]+.
2.3.3 5-Bromo-N-(6-methoxy-3-methylbenzo[d]isoxazol-5-yl)-2-(trifluoromethoxy)benzene sulfonamide (13c)
White solid. Yield: 66%. M.p. 165–166 °C; 1H NMR (500 MHz, CDCl3) δ 8.04 (d, J = 2.4 Hz, 1H, 6′-ArH), 7.72 (s, 1H, 7-ArH), 7.66 (dd, J = 8.8, 2.4 Hz, 1H, 4′-ArH), 7.30 (brs, 1H, –NH–), 7.21 (dd, J = 8.8, 1.7 Hz, 1H, 3′-ArH), 6.90 (s, 1H, 4-ArH), 3.81 (s, 3H, 6-OCH3), 2.53 (s, 3H, 3-CH3); 13C NMR (101 MHz, CDCl3) δ 161.48, 155.21, 152.77, 145.22, 137.60, 134.11, 132.11, 124.05 (–OCF3), 122.15, 121.45 (–OCF3), 120.61 (–ArCOCF3), 120.59 (–ArCOCF3), 120.57 (–ArCOCF3), 120.55 (–ArCOCF3), 119.08, 118.84 (–OCF3), 116.24 (–OCF3), 115.20, 113.88, 91.99, 56.21, 10.05; ESI-MS (m/z): 480.9 and 482.9 [M + H]+.
2.3.4 5-Bromo-N-(6-methoxy-3-methylbenzo[d]isoxazol-5-yl)-2-propoxybenzenesulfonamide (13d)
White solid. Yield: 67%. M.p. 159–160 °C; 1H NMR (400 MHz, CDCl3) δ 7.93 (d, J = 2.4 Hz, 1H, 6′-ArH), 7.71 (s, 1H, 7-ArH), 7.54 (dd, J = 8.8, 2.4 Hz, 1H, 4′-ArH), 7.49 (brs, 1H, –NH–), 6.89 (s, 1H, 4-ArH), 6.84 (d, J = 8.8 Hz, 1H, 3′-ArH), 4.07 (t, J = 6.5 Hz, 2H, –OCH2CH2–), 3.81 (s, 3H, 6-OCH3), 2.52 (s, 3H, 3-CH3), 2.02–1.87 (m, 2H, –CH2CH2–), 1.15 (t, J = 7.4 Hz, 3H, –CH2CH3); 13C NMR (101 MHz, DMSO) δ 161.49, 155.55, 155.49, 154.96, 137.04, 131.30, 129.67, 123.01, 117.96, 115.73, 114.14, 110.27, 92.36, 70.43, 56.31, 21.49, 10.29, 9.46; ESI-MS (m/z): 455.0 and 457.0 [M + H]+.
2.3.5 Ethyl 2-(4-bromo-2-(N-(6-methoxy-3-methylbenzo[d]isoxazol-5-yl)sulfamoyl)phe noxy)acetate (13e)
White solid. Yield: 69%. M.p. 174–176 °C; 1H NMR (400 MHz, CDCl3) δ 8.11 (s, 1H, –NH–), 7.93 (d, J = 2.5 Hz, 1H, 6′-ArH), 7.73 (s, 1H, 7-ArH), 7.57 (dd, J = 8.7, 2.5 Hz, 1H, 4′-ArH), 6.87 (s, 1H, 4-ArH), 6.78 (d, J = 8.8 Hz, 1H, 3′-ArH), 4.73 (s, 2H, –OCH2CO–), 4.32 (q, J = 7.1 Hz, 2H, –OCH2CH3), 3.71 (s, 3H, 6-OCH3), 2.53 (s, 3H, 3-CH3), 1.33 (t, J = 7.1 Hz, 3H, –OCH2CH3); 13C NMR (101 MHz, CDCl3) δ 167.50, 161.43, 155.18, 153.92, 153.63, 137.18, 132.81, 130.31, 123.00, 115.18, 115.16, 115.06, 113.80, 91.93, 66.49, 62.11, 56.16, 14.15, 10.04; ESI-MS (m/z): 499.0 and 501.0 [M + H]+.
2.3.6 5-Bromo-N-(6-methoxy-3-methylbenzo[d]isoxazol-5-yl)-2,3-dihydrobenzofuran-7-sulfon amide (13f)
White solid. Yield: 74%. M.p. 189–190 °C; 1H NMR (400 MHz, CDCl3) δ 7.69 (s, 1H, 7-ArH), 7.64 (s, 1H, 6′-ArH), 7.46–7.32 (m, 2H, 4′-ArH, –NH–), 6.90 (s, 1H, 4-ArH), 4.71 (t, J = 8.8 Hz, 2H, –OCH2–), 3.84 (s, 3H, 6-OCH3), 3.22 (t, J = 8.7 Hz, 2H, –CH2Ar–), 2.52 (s, 3H, 3-CH3); 13C NMR (101 MHz, CDCl3) δ 161.04, 156.13, 155.21, 152.45, 132.83, 131.95, 130.01, 123.13, 121.96, 115.08, 112.51, 111.72, 91.88, 73.58, 56.37, 28.90, 10.06; ESI-MS (m/z): 439.0 and 441.0 [M + H]+.
2.3.7 N-(6-methoxy-3-methylbenzo[d]isoxazol-5-yl)benzo[c][1, 2, 5]thiadiazole-4-sulfonamide (13g)
White solid. Yield: 69%. M.p. 230–231 °C; 1H NMR (400 MHz, DMSO) δ 9.76 (s, 1H, –NH–), 8.37 (d, J = 8.7 Hz, 1H, 7′-ArH), 7.99 (d, J = 7.0 Hz, 1H, 5′-ArH), 7.75 (t, J = 8.0 Hz, 1H, 6′-ArH), 7.71 (s, 1H, 7-ArH), 7.04 (s, 1H, 4-ArH), 2.96 (s, 3H, 6-OCH3), 2.50 (s, 3H, 3-CH3); 13C NMR (101 MHz, DMSO) δ 162.01, 156.52, 155.05, 154.96, 149.10, 131.92, 130.73, 128.66, 126.15, 122.30, 120.85, 114.28, 92.11, 55.44, 9.50; ESI-MS (m/z): 377.0 [M + H]+.
2.3.8 N-(6-methoxy-3-methylbenzo[d]isoxazol-5-yl)quinoline-8-sulfonamide (13h)
White solid. Yield: 70%. M.p. 212–213 °C; 1H NMR (500 MHz, DMSO) δ 9.17 (m, 1H, 2′-ArH), 9.10 (brs, 1H, –NH–), 8.56 (d, J = 8.3 Hz, 1H, 7′-ArH), 8.28 (d, J = 8.3 Hz, 1H, 5′-ArH), 8.21 (d, J = 7.2 Hz, 1H, 4′-ArH), 7.80–7.75 (m, 1H, 6′-ArH), 7.73 (s, 1H, 7-ArH), 7.65 (t, J = 7.7 Hz, 1H, 3′-ArH), 7.07 (s, 1H, 4-ArH), 3.21 (s, 3H, 6-OCH3), 2.46 (s, 3H, 3-CH3); 13C NMR (101 MHz, DMSO) δ 160.91, 154.91, 154.16, 151.23, 142.66, 137.08, 135.68, 134.11, 130.66, 128.44, 125.47, 123.59, 122.66, 115.62, 114.13, 92.28, 55.77, 9.45; ESI-MS (m/z): 370.0 [M + H]+.
2.4 General procedures for the preparation of compounds 14, 14a–c
2.4.1 2-(4-Bromo-2-(N-(6-methoxy-3-methylbenzo[d]isoxazol-5-yl)sulfamoyl)phenoxy)acetic acid (14)
To a solution of 2 mol/L NaOH (12 mL) was added compound 13e (500 mg, 1.0 mmol). The mixture was stirred at room temperature for 4 h. On completion of the reaction (monitored by TLC), the mixture was treated with 1 mol/L dilute hydrochloric acid to adjust the pH value to 5-6. The precipitated solid was filtered to obtain the target compound as a white solid. Yield: 98%. M.p. 204–205 °C; 1H NMR (400 MHz, DMSO-d6) δ 9.25 (brs, 1H, –COOH), 7.76 (dd, J = 8.9, 2.5 Hz, 1H, 4′-ArH), 7.66 (s, 1H, 7-ArH), 7.64 (d, J = 2.5 Hz, 1H, 6′-ArH), 7.24 (s, 1H, 4-ArH), 7.15 (d, J = 8.9 Hz, 1H, 3′-ArH), 4.90 (s, 2H, –OCH2CO–), 3.63 (s, 3H, 6-OCH3), 2.46 (s, 3H, 3-CH3); 13C NMR (101 MHz, DMSO) δ 169.92, 161.62, 155.90, 154.99, 154.54, 136.95, 130.74, 129.95, 122.61, 118.83, 116.37, 114.22, 111.34, 92.53, 65.85, 56.36, 9.46; ESI-MS (m/z): 468.9 and 470.9 [M–H]–.
2.4.2 2-(4-Bromo-2-(N-(6-methoxy-3-methylbenzo[d]isoxazol-5-yl)sulfamoyl)phenoxy)-N- methylacetamide (14a)
To a solution of compound 14 (80 mg, 0.17 mmol) in DMF (5 mL) were added diisopropylethylamine (DIPEA) (76.8 mg, 0.59 mmol) and HATU (96.8 mg, 0.25 mmol). The resulting mixture was stirred at room temperature for 15 min followed by the addition of methylamine hydrochloride (19.5 mg, 0.29 mmol). The mixture was stirred at room temperature for additional 5 h. After the reaction was completed, it was diluted with water (10 mL) and the precipitated solid was filtered. The crude product was purified by silica gel chromatography to obtain the target compound as a white solid. Yield: 75%. M.p. 183–184 °C; 1H NMR (400 MHz, CDCl3) δ 7.84 (d, J = 2.4 Hz, 1H, 6′-ArH), 7.70 (s, 1H, 7-ArH), 7.59 (dd, J = 8.8, 2.4 Hz, 1H, 4′-ArH), 7.44 (brs, 1H, –CONH–), 7.34 (s, 1H, -SO2NH–), 6.87 (s, 1H, 4-ArH), 6.85 (d, J = 8.9 Hz, 1H, 3′-ArH), 4.64 (s, 2H, –OCH2CO–), 3.73 (s, 3H, 6-OCH3), 2.90 (d, J = 4.8 Hz, 3H, –NHCH3), 2.53 (s, 3H, 3-CH3); 13C NMR (101 MHz, CDCl3) δ 167.34, 161.57, 155.16, 153.43, 153.08, 137.96, 133.29, 128.57, 122.26, 115.30, 115.04, 114.34, 113.79, 92.06, 67.94, 56.43, 26.10, 10.04; ESI-MS (m/z): 484.0 and 486.0 [M + H]+.
2.4.3 2-(4-Bromo-2-(N-(6-methoxy-3-methylbenzo[d]isoxazol-5-yl)sulfamoyl)phenoxy)-N-ethylacetamide (14b)
White solid. Yield: 70%. M.p. 195–196 °C; 1H NMR (400 MHz, CDCl3) δ 7.85 (d, J = 2.4 Hz, 1H, 6′-ArH), 7.70 (s, 1H, 7-ArH), 7.59 (dd, J = 8.8, 2.4 Hz, 1H, 4′-ArH), 7.44 (s, 1H, –CONH–), 7.33 (s, 1H, -SO2NH–), 6.87 (s, 1H, 4-ArH), 6.84 (d, J = 8.8 Hz, 1H, 3′-ArH), 4.63 (s, 2H, –OCH2CO–), 3.74 (s, 3H, 6-OCH3), 3.43–3.34 (m, 2H, –CH2CH3), 2.53 (s, 3H, 3-CH3), 1.21 (t, J = 7.3 Hz, 3H, –CH2CH3); 13C NMR (101 MHz, CDCl3) δ 166.40, 161.52, 155.16, 153.42, 152.95, 137.94, 133.30, 128.55, 122.33, 115.31, 114.76, 114.32, 113.75, 92.06, 67.93, 56.45, 34.34, 14.50, 10.06; ESI-MS (m/z): 498.0 and 500.0 [M + H]+.
2.4.4 2-(4-Bromo-2-(N-(6-methoxy-3-methylbenzo[d]isoxazol-5-yl)sulfamoyl)phenoxy)-N- propylacetamide (14c)
White solid. Yield: 65%. M.p. 180–181 °C; 1H NMR (400 MHz, CDCl3) δ 7.85 (d, J = 2.4 Hz, 1H, 6′-ArH), 7.70 (s, 1H, 7-ArH), 7.59 (dd, J = 8.8, 2.4 Hz, 1H, 4′-ArH), 7.44 (s, 1H, –CONH–), 7.32 (s, 1H, -SO2NH–), 6.87 (s, 1H, 4-ArH), 6.85 (d, J = 8.8 Hz, 1H, 3′-ArH), 4.64 (s, 2H, –OCH2CO–), 3.75 (s, 3H, 6-OCH3), 3.31 (q, J = 6.8 Hz, 2H, –NHCH2CH2–), 2.53 (s, 3H, 3-CH3), 1.65–1.54 (m, 2H, –CH2CH2CH3), 0.94 (t, J = 7.4 Hz, 3H, –CH2CH3); 13C NMR (101 MHz, CDCl3) δ 166.51, 161.50, 155.16, 153.39, 152.90, 137.94, 133.30, 128.53, 122.35, 115.31, 114.64, 114.29, 113.76, 92.07, 67.90, 56.45, 41.16, 22.56, 11.37, 10.06; ESI-MS (m/z): 512.0 and 514.0 [M + H]+.
2.5 Docking studies
Preparation of the Receptor. The protein of compound 3 in complex with BRD4(1) (PDB ID: 5Y8Y) was utilized for generation of grid. The protein was prepared using the Protein Preparation Wizard within Maestro (version 11.5, implanted in Schrödinger). The bond orders were assigned using CCD database, the hydrogens were added, and the missing side chains were filled in with Prime. The original waters beyond 5.0 Å from ligand were deleted. In addition, the solvent water molecules near the 6-OCH3 position were also deleted. The important inner water molecules were kept. The states of ligands were generated using Epik and the pH value was set as 7.0 ± 2.0. The water orientations were optimized via the H-bond assignment using PROPKA program and the pH value was set as 7.0. OPLS3 force field was used to minimize the energy of the added hydrogens.
Preparation of the Ligands. The ligands were prepared using the LigPrep program with default parameters. The energy minimization was performed using an OPLS3 force field.
Individual Docking. The docking studies were carried out using the Ligand Docking module. The grid was defined similar in size to the original ligand. No constraints were applied, and all parameters were kept as default. The compounds were docked using Glide SP mode.
2.6 Biochemical assays
2.6.1 Protein expression and purification
The expression and purification of BRD4 BD1 proteins were carried out as previously described.27,28 The experimental detail was described in supplementary information.
2.6.2 Thermal shift assay (Tm)
Thermal shift assays were performed using the Bio-Rad CFX96 Real-Time PCR system. Each biochemical reaction consists 10 μM protein, 200 μM compound, a fluorescent dye (SYPRO Orange, ABI, Sigma) that was added at a dilution of 1:1000, and a buffer (10 mM HEPES, 150 mM NaCl, 50% (v / v) glycerin as well as deionized water at a pH of 7.5). The reaction mixture (20 μL) was added to the 96-well plate. Next, the mixture was centrifuged at 1000 r/min for 1 min at room temperature and incubated on ice for 30 min in the dark. The excitation and emission filters of the SYPRO Orange dye were set at 465 nm and 590 nm, respectively. Raising the temperature from 30 °C to 80 °C at a rate of 0.3 °C/min. The signal of fluorescence was read at an interval of 0.3 °C. All experiments were performed in triplicate. The melting temperature (Tm) was calculated by fitting the melting curve to Boltzmann equation using GraphPad Prism. ΔTm represents the difference between the calculated Tm values for the tested reaction and the blank reaction.
2.6.3 AlphaScreen peptide displacement assay
The interaction between the protein of BRD4 BD1 bromodomain and ligand was evaluated by AlphaScreen technique (PerkinElmer). The biochemical reaction consisted of 50 nM protein, 50 nM peptide, the indicated concentration of compounds, deionized water, and the buffer (50 mM MOPS, 0.05 mM CHAPS, 50 mM NaF, 0.1 mg/mL BSA and deionized water at a pH value of 7.4). All components were mixed in a 384-well plate (ProxiPlate-384 Plus, PerkinElmer, USA) for 1 h at room temperature. The nickel receptor beads and streptavidin donor beads were dilute with buffer to indicated concentration, and were added to the above 384-well plate at a final concentration of 5 μg/mL in the dark environment. The plates were sealed with foil to avoid light and incubated for 1.5 h at room temperature. The signal was read on an EnSpire microplate reader (PerkinElmer, USA). The protein of BRD4 bromodomain binds to the nickel acceptor beads. The biotinylated acetylated histone H4 peptide binds to streptavidin donor beads. When the donor beads were excited by a 680 nm laser beam, it will emit 1O2. The 1O2 can activate the acceptor beads to release 520 nm photons as signals. All experiments were performed on the same plate in triplicate. The peptide used above is a C-terminal biotinylated tetra-acetylated histone peptide (bH4KAc4) with the sequence of H-YSGRGK(Ac)GGK(Ac)GLGK(Ac)GGAK(Ac) RHRK-Biotin-OH (synthesized by Genscript). The inhibitory curves (Figure S1, Supplementary Information) and half-maximal inhibitory concentration (IC50) values were calculated using GraphPad Prism 7 software. The raw data read from the microplate reader was firstly transformed to logarithms, and then analyzed using the nonlinear regression of curve fit. The dose-response inhibition equation was chosen to generate the presented curves and IC50 values.
3 Results and Discussion
3.1 Chemistry
The benzo[d]isoxazol containing scaffold 9 can be synthesized via multistep reactions from commercially available 2, 4-dimethoxyaniline (4).28 As outlined in Scheme 1, the amino group of 4 was protected with acetic anhydride to give compound 5, which could be converted to compound 6 via a Friedel-Crafts reaction in the presence of a lewis acid of AlCl3. Next, the ketone group of 6 was reacted with hydroxylamine hydrochloride, which afforded the oxime product 7. Compound 7 could undergo a cyclization in the presence of DMF-DMA under high temperature to afford the benzo[d]isoxazol scaffold 8. Finally, the acetyl group of 8 was hydrolyzed by hydrochloric acid to give the aniline 9.
The synthesis of different sulfonyl chlorides was described in Scheme 2. The commercially available 4-bromophenol (10a) was reacted with different haloalkanes to give the intermediates 11a–c. Treatment of 10a or 11a–c with chlorosulfonic acid gave the sulfonyl chlorides 12a–d. The 5-bromo-2, 3-dihydrobenzofuran-7-sulfonyl chloride (12e) was prepared using the 2, 3-dihydrobenzofuran (10b) as the starting material. The compound 10b was treated with Br2 to afford the bromine substituted intermediate 11d,29 followed by the treatment of chlorosulfonic acid to generate target intermediate 12e.
Finally, the scaffold of 9 was reacted with various sulfonyl chlorides in a one-plot procedure to obtain the target compounds 13a–h (Scheme 3). Hydrolysis of 13e with NaOH solution afforded compound 14, which was coupled with available amines to generate the target compounds 14a–c (Scheme 3).
3.2 SARs studies
This work began with an analysis of the co-crystal structure of 3-BRD4(1) complex as reported in our previous work.28 According to the structural analysis, the 3-methylbenzo[d]isoxazole scaffold resided in the binding pocket and formed direct or indirect hydrogen bonds with key residues (Figure 2). It is obvious that 2′-methoxy of compound 3 extends to the solvent region, which guarantees the chemical space for modification. The CYP-mediated O-demethylation is a prevalent metabolic pathway for many drugs.30,31 Thus, the methoxy group is a potential metabolic group in the body. The replacement of 2′-methoxy may improve the metabolic stability of compounds (Figure 2). This article aims to test the feasibility of modification on 2′ or 3′ position, which will extend the structure-activity relationships (SARs).
We preliminarily sought to evaluate whether the length of the alkyl chain affects the binding affinity. When an ethyl group was introduced at R1 position, the resulting compound 13a exhibited potent binding activity with a thermo shift (ΔTm) value of 7.2 °C and an IC50 value of 0.47 μM (Table 1). When shortening the carbon chain, a compound 13b was obtained. This compound had a higher ΔTm value of 8.4 °C compared with 13a. However, the IC50 value (1 μM) was not corresponding to the ΔTm value, which may attribute to the poor solubility. It was noted that the introduction of a trifluoromethyl group (compound 13c) at R1 position led to a 2-fold decrease in potency compared with 13a. The local steric hindrance produced by trifluoromethoxy group may lead to the change of the relative orientation of the aryl ring as well as the decrease on activity. Docking studies also revealed that the scores of 13c (− 6.182, − 6.427, − 60.583) were significantly worse than that of 13a (− 6.726, − 6.782, − 63.101) (Table S1, Supplementary Information). Next, we continued to extend the carbon chain. Introduction of propyl on R1 position led to compound 13d, which restored the activity with a ΔTm value of 5.7 °C and an IC50 value of 0.54 μM.
Next, we tried to test the feasibility of the introduction of heteroatom-containing chains at R1 position, which may be suitable for extending to the solvent region of the protein. When the oxygen-linked ethyl acetate group (compound 13e) was introduced at the R1 position, the IC50 value was still maintained at 0.42 μM. However, the compound did not respond well in TSA experiment. Hydrolyzing the ethyl acetate group of compound 13e to acetic acid group gave the compound 14, which exhibited an IC50 value of 0.66 μM and ΔTm value of 6.9 °C (Table 1). The above results show that R1 position is tolerated for modifications. Further studies on the exploration of the chemical space of the 2′ position were carried out. We designed and synthesized several derivatives including methanamide (compound 14a), acetamide (compound 14b), propanamide (compound 14c) substitutions at the 2′ side chain. All the three compounds displayed high ΔTm values as evaluated in TSA assay. These compounds also showed submicromole binding activities with the IC50 values ranging from 0.35 μM to 0.75 μM. Among the three compounds, 14b showed stronger activity with IC50 value of 0.35 μM and ΔTm value of 7.2 °C.
It was noteworthy that most of the above compounds possessed substituents of linear chains. However, the linear chains are flexible, which may lead to unstable conformations. The variable conformations do not benefit binding affinity. Therefore, we designed several compounds with fused rings to immobilize the conformation. According to the crystal structure of 3-BRD4(1) complex, the 3′ position was also suitable for small substitution (Figure 2). We also noted that the oxygen atom in 2′-methoxy group formed a hydrogen bond with the adjacent water molecule. Therefore, we reserved the oxygen atom at 2′ position and fused the 2′ and 3′ position to afford a new 2, 3-dihydrobenzofuran fragment. The resulting compound 13f displayed potent binding affinity with a ΔTm value of 7.8 °C. The strong activity was also demonstrated by the IC50 value of 0.21 μM, which was only 0.6 fold lower than that of 3 (Table 1). This result met the original expectations to replace the methoxy group with other substituents to avoid potential in vivo metabolism while maintaining the activity with less than 1 fold loss compared with the previous inhibitor 3. We also synthesized two additional compounds 13g–h with fused rings. However, both compounds almost lost the binding potencies, probably due to the rigidity and planarity of the fused rings, making it not accommodate in the binding pocket. The data reported here suggest a feasible direction for further chemical modification on the 2′ position or 3′ position.
3.3 Molecular docking studies of compounds with BRD4(1)
To understand potential binding interactions of the compounds to BRD4(1), several representative compounds were chosen for molecular docking studies. The crystal structure of 3-BRD4(1) complex (PDB ID: 5Y8Y) was used for generating the grid. The overall binding mode predicted by molecular docking was shown in Figure 3. The 3-methylbenzo[d]isoxazole scaffold of all compounds resided in the binding pocket. The oxygen of isoxazole moiety formed a hydrogen bond with Asn140 (Figure 3A). The nitrogen of isoxazole formed a hydrogen bond with a conserved water molecule, which further bound to Tyr97 (Figure 3A). The 5-bromo-2,3-dihydrobenzofuran fragment of 13f attached to the sulfonamide could occupy the sub-pocket referred to as the WPF shelf (Figure 3B).25 The Br substituent directed towards Ile146 while the furan ring exposed to the solvent (Figure 3B). The sulfonamide oxygen also formed a hydrogen bond with the nearby water molecule. The representative compounds 13a and 14b displayed similar binding mode to that of 13f.
(a) Molecular docking of compound 13f (magenta stick) in BRD4(1) (PDB code 5Y8Y; protein, light green ribbon; key residues, pink thin sticks; water molecules, red spheres). The isoxazole moiety forms two H-bonds (yellow dashed lines) to Asn140 and a conserved water molecule. (b) Molecular docking of compound 13f (magenta stick) in BRD4(1) (PDB code 5Y8Y; protein, grey surface; key residues, cyan surface; water molecules, red spheres). The 3-methylbenzo[d]isoxazole moiety of 13f resides in the binding pocket. (c) Overlaying the docked structures of compound 13a, 14b and 13f in BRD4(1). Structures were prepared using the PyMOL program.
It was noteworthy that the fused 2, 3-dihydrobenzofuran ring exhibited fewer conformations than compounds with long linear chains in the docking process, which may contribute to the improved binding activity. The docking scores of 13a, 14b, 13f and 3 are listed in Table S1 (Supplementary Information). Three kinds of binding energy scores, such as docking score, glide gscore, and glide emodel score, were selected for comprehensive evaluation. Compound 13f displayed similar docking scores (− 6.766, − 6.816, − 61.32) compared with 3 (− 6.876, − 6.922, − 62.285). Compounds 13a (− 6.726, − 6.782, − 63.101) and 14b (− 6.358, − 6.407, − 64.341) showed slightly bad scores compared with 13f and 3. In summary, the hydrogen bond and hydrophobic interactions taken together with the good shape complementary contribute to the high binding affinity.
3.4 In silico studies of drug-likeness and pharmacokinetic profile
The drug-likeness and pharmacokinetic profile are closely related to the success of drug development. Generally, a qualified clinical candidate often meets some drug-like criteria. Therefore, the representative compounds 13a, 14b and 13f were selected for calculation using the online SwissADME server.32 Firstly, the drug-like profiles of the compounds were analyzed using several classic filters (Table 2). These rules were established based on the basic physicochemical parameters, such as molecular weight (Mw), number of rotatable bonds (NRB), number of the hydrogen-bond acceptor (NHBA), number of hydrogen bond donor (NHBD), topological polar surface area (TPSA) and LogP, which were used to evaluate the chance for a molecule to be an orally available candidate. A summary of the physicochemical parameters of 13a, 14b and 13f was listed in supporting Table S2 (Supplementary Information). Lipinski rule33 (also known as rule-of-five) is a classic drug-like filter, which can be summarized as the following criteria: NHBD ≤ 5, NHBA ≤ 10, LogP ≤ 5, NRB ≤ 10, and Mw ≤ 500. Afterwards, several new rules such as Ghose,34 Egan35 Muegge (Bayer)36 and Veber37 rules had been established. The results indicated that only compound 14b showed 1 violation against the Ghose ruler, other compounds 13a and 13f met all of the rules (Table 2). The bioavailability score predicts the probability that a compound will have bioavailability (F) > 10% in rat or measurable Caco-2 permeability.38 For anions, the score is 0.11 when TPSA > 150 Å2; 0.56 if TPSA between 75 and 150 Å2; 0.85 if TPSA < 75 Å2.38 For the rest compounds, the score is 0.55 if it passes the five rules, and 0.17 if it fails.38 All three compounds exhibited promising bioavailability score of 0.55 (Table 2).
The pharmacokinetic profile was predicted using parameters including gastrointestinal absorption (GIA),39 blood−brain barrier (BBB) permeation and P-glycoprotein (P-gp) substrate.32 All tested compounds except 14b exhibit high gastrointestinal absorption, which enhance the potential for high bioavailability (Table 3). Furthermore, the three compounds are not permeable of BBB, which is expected to have a lower incidence of adverse effect on the central nervous system (CNS) (Table 3). P-glycoprotein (P-gp) is an ATP-dependent pump located on the cell membrane, which can protect cells via the efflux of invading compounds. If the drug is the substrate of P-gp, the activity of the drug will be limited in vivo or at the cellular level. All three compounds are not substrates of P-gp according to the parameters. Finally, the PAINS (pan-assay interference structures) and Brenk alerts were used to identify the problematic structure in medicinal chemistry. Brenk40 consists of 105 substructures, which may be toxic or metabolically unstable. PAINS41 can help to identify compounds with interfering substructures that have a potent response in many protein assays regardless of the specific target. It was noteworthy that none of the tested compounds triggered any alerts as predicted by the two filters.
4 Conclusions
In conclusion, a series of novel BRD4 bromodomain inhibitors bearing a benzo[d]isoxazol scaffold was designed through a structure-based drug design strategy. The facile and efficient synthetic routes were developed for the synthesis and optimization of all compounds. The SAR studies suggest a feasible direction for chemical exploration on the 2′ or 3′ position. Molecular docking studies revealed the binding patterns and the structural characteristics required for high affinity for the BRD4(1). The optimal inhibitor 13f exhibited high binding affinity to BRD4(1) with a ΔTm value of 7.8 °C and an IC50 of 0.21 μM. In addition, the compound 13f displayed acceptable drug-likeness and pharmacokinetic profiles as predicted by the silico models. All the data suggest a promising starting point for further optimization of the compounds as potent BRD4 inhibitors and exploration of the biological functions.
References
Marushige K 1976 Activation of chromatin by acetylation of histone side chains Proc. Natl. Acad. Sci. U. S. A. 73 3937
Dawson M A and Kouzarides T 2012 Cancer epigenetics: from mechanism to therapy Cell 150 12
Brennan P, Filippakopoulos P and Knapp S 2012 The therapeutic potential of acetyl-lysine and methyl-lysine effector domains Drug Discov. Today: Ther. Strategies 9 e101
Filippakopoulos P, Picaud S, Mangos M, Keates T, Lambert J-P, Barsyte-Lovejoy D, Felletar I, Volkmer R, Müller S, Pawson T, Gingras A-C, Arrowsmith C H and Knapp S 2012 Histone recognition and large-scale structural analysis of the human bromodomain family Cell 149 214
Zuber J, Shi J W, Wang E, Rappaport A R, Herrmann H, Sison E A, Magoon D, Qi J, Blatt K, Wunderlich M, Taylor M J, Johns C, Chicas A, Mulloy J C, Kogan S C, Brown P, Valent P, Bradner J E, Lowe S W and Vakoc C R 2011 RNAi screen identifies Brd4 as a therapeutic target in acute myeloid leukaemia Nature 478 524
Reyes-Garau D, Ribeiro L M and Roué G 2019 Pharmacological targeting of BET bromodomain proteins in acute myeloid leukemia and malignant lymphomas: From molecular characterization to clinical applications Cancers 11 1483
Dawson M A, Prinjha R K, Dittmann A, Giotopoulos G, Bantscheff M, Chan W-I, Robson S C, Chung C-W, Hopf C, Savitski M M, Huthmacher C, Gudgin E, Lugo D, Beinke S, Chapman T D, Roberts E J, Soden P E, Auger K R, Mirguet O, Doehner K, Delwel R, Burnett A K, Jeffrey P, Drewes G, Lee K, Huntly B J and Kouzarides T 2011 Inhibition of BET recruitment to chromatin as an effective treatment for MLL-fusion leukaemia Nature 478 529
Mu J, Sun P, Ma Z and Sun P 2019 BRD4 promotes tumor progression and NF-κB/CCL2-dependent tumor-associated macrophage recruitment in GIST Cell Death Dis. 10 935
Tian Y, Wang X, Zhao S, Liao X, Younis M R, Wang S, Zhang C and Lu G 2019 JQ1-loaded polydopamine nanoplatform inhibits c-MYC/programmed cell death ligand 1 to enhance photothermal therapy for triple-negative breast cancer ACS Appl. Mater. Inter. 11 46626
Shu S K, Lin C Y, He H H, Witwicki R M, Tabassum D P, Roberts J M, Janiszewska M, Huh S J, Liang Y, Ryan J, Doherty E, Mohammed H, Guo H, Stover D G, Ekram M B, Peluffo G, Brown J, D’Santos C, Krop I E, Dillon D, McKeown M, Ott C, Qi J, Ni M, Rao P K, Duarte M, Wu S Y, Chiang C M, Anders L, Young R A, Winer E P, Letai A, Barry W T, Carroll J S, Long H W, Brown M, Liu X S, Meyer C A, Bradner J E and Polyak K 2016 Response and resistance to BET bromodomain inhibitors in triple-negative breast cancer Nature 529 413
Faivre E J, McDaniel K F, Albert D H, Mantena S R, Plotnik J P, Wilcox D, Zhang L, Bui M H, Sheppard G S, Wang L, Sehgal V, Lin X, Huang X, Lu X, Uziel T, Hessler P, Lam L T, Bellin R J, Mehta G, Fidanze S, Pratt J K, Liu D, Hasvold L A, Sun C, Panchal S C, Nicolette J J, Fossey S L, Park C H, Longenecker K, Bigelow L, Torrent M, Rosenberg S H, Kati W M and Shen Y 2020 Selective inhibition of the BD2 bromodomain of BET proteins in prostate cancer Nature 578 306
Asangani I A, Dommeti V L, Wang X J, Malik R, Cieslik M, Yang R D, Escara-Wilke J, Wilder-Romans K, Dhanireddy S, Engelke C, Iyer M K, Jing X J, Wu Y M, Cao X H, Qin Z S, Wang S M, Feng F Y and Chinnaiyan A M 2014 Therapeutic targeting of BET bromodomain proteins in castration-resistant prostate cancer Nature 510 278
Asangani I A, Wilder-Romans K, Dommeti V L, Krishnamurthy P M, Apel I J, Escara-Wilke J, Plymate S R, Navone N M, Wang S M, Feng F Y and Chinnaiyan A M 2016 BET bromodomain inhibitors enhance efficacy and disrupt resistance to AR antagonists in the treatment of prostate cancer Mol. Cancer Res. 14 324
Huang M, Zeki J, Sumarsono N, Coles G L, Taylor J S, Danzer E, Bruzoni M, Hazard F K, Lacayo N J, Sakamoto K M, Dunn J C Y, Spunt S L and Chiu B 2020 Epigenetic targeting of TERT-associated gene expression signature in human neuroblastoma with TERT overexpression Cancer Res. 80 1024
He S, Dong G, Li Y, Wu S Y, Wang W and Sheng C 2020 Potent dual BET/HDAC inhibitors for efficient treatment of pancreatic cancer Angew. Chem. Int. Ed. 59 3028
Fehling S C, Miller A L, Garcia P L, Vance R B and Yoon K J 2020 The combination of BET and PARP inhibitors is synergistic in models of cholangiocarcinoma Cancer Lett. 468 48
Huang B, Yang X D, Zhou M M, Ozato K and Chen L F 2009 Brd4 coactivates transcriptional activation of NF-κB via specific binding to acetylated RelA Mol. Cell. Biol. 29 1375
Nicodeme E, Jeffrey K L, Schaefer U, Beinke S, Dewell S, Chung C-W, Chandwani R, Marazzi I, Wilson P, Coste H, White J, Kirilovsky J, Rice C M, Lora J M, Prinjha R K, Lee K and Tarakhovsky A 2010 Suppression of inflammation by a synthetic histone mimic Nature 468 1119
Jiang F, Hu Q, Zhang Z, Li H, Li H, Zhang D, Li H, Ma Y, Xu J, Chen H, Cui Y, Zhi Y, Zhang Y, Xu J, Zhu J, Lu T and Chen Y 2019 Discovery of benzo[cd]indol-2(1H)-ones and pyrrolo[4,3,2-de]quinolin-2(1H)-ones as bromodomain and extra-terminal domain (BET) inhibitors with selectivity for the first bromodomain with potential high efficiency against acute gouty arthritis J. Med. Chem. 62 11080
Filippakopoulos P, Qi J, Picaud S, Shen Y, Smith W B, Fedorov O, Morse E M, Keates T, Hickman T T, Felletar I, Philpott M, Munro S, McKeown M R, Wang Y, Christie A L, West N, Cameron M J, Schwartz B, Heightman T D, La Thangue N, French C A, Wiest O, Kung A L, Knapp S and Bradner J E 2010 Selective inhibition of BET bromodomains Nature 468 1067
A dose escalation study to investigate the safety, pharmacokinetics (PK), pharmacodynamics (PD) and clinical activity of GSK525762 in subjects with relapsed, refractory hematologic malignancies ClinicalTrials.gov NCT01943851
Dose escalation and dose expansion study of GSK525762 in combination with androgen deprivation therapy and other agents in subjects with castrate-resistant prostate cancer ClinicalTrials.gov NCT03150056
Seal J, Lamotte Y, Donche F, Bouillot A, Mirguet O, Gellibert F, Nicodeme E, Krysa G, Kirilovsky J, Beinke S, McCleary S, Rioja I, Bamborough P, Chung C W, Gordon L, Lewis T, Walker A L, Cutler L, Lugo D, Wilson D M, Witherington J, Lee K and Prinjha R K 2012 Identification of a novel series of BET family bromodomain inhibitors: binding mode and profile of I-BET151 (GSK1210151A) Bioorg. Med. Chem. Lett. 22 2968
Bamborough P, Diallo H, Goodacre J D, Gordon L, Lewis A, Seal J T, Wilson D M, Woodrow M D and Chung C W 2012 Fragment-based discovery of bromodomain inhibitors part 2: optimization of phenylisoxazole sulfonamides J. Med. Chem. 55 587
Hewings D S, Fedorov O, Filippakopoulos P, Martin S, Picaud S, Tumber A, Wells C, Olcina M M, Freeman K, Gill A, Ritchie A J, Sheppard D W, Russell A J, Hammond E M, Knapp S, Brennan P E and Conway S J 2013 Optimization of 3,5-dimethylisoxazole derivatives as potent bromodomain ligands J. Med. Chem. 56 3217
Ran X, Zhao Y J, Liu L, Bai L C, Yang C Y, Zhou B, Meagher J L, Chinnaswamy K, Stuckey J A and Wang S M 2015 Structure-based design of γ-carboline analogues as potent and specific BET bromodomain inhibitors J. Med. Chem. 58 4927
Xue X Q, Zhang Y, Liu Z X, Song M, Xing Y L, Xiang Q P, Wang Z, Tu Z C, Zhou Y L, Ding K and Xu Y 2016 Discovery of benzo[cd]indol-2(1H)-ones as potent and specific BET bromodomain inhibitors: structure-based virtual screening, optimization, and biological evaluation J. Med. Chem. 59 1565
Zhang M, Zhang Y, Song M, Xue X, Wang J, Wang C, Zhang C, Li C, Xiang Q, Zou L, Wu X, Wu C, Dong B, Xue W, Zhou Y, Chen H, Wu D, Ding K and Xu Y 2018 Structure-based discovery and optimization of benzo[d]isoxazole derivatives as potent and selective BET inhibitors for potential treatment of castration-resistant prostate cancer (CRPC) J. Med. Chem. 61 3037
Albaster R J, Cottrell I F, Marley H and Wright S H B 1988 The synthesis of 5-substituted 2,3-dihydrobenzofurans Synthesis-Stuttgart 950
Zhang Z, Zhu M and Tang W 2009 Metabolite identification and profiling in drug design: current practice and future directions Curr. Pharm. Des. 15 2220
Zhang Z and Tang W 2018 Drug metabolism in drug discovery and development Acta Pharm. Sin. B 8 721
Daina A, Michielin O and Zoete V 2017 SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules Sci. Rep. 7 42717
Lipinski C A, Lombardo F, Dominy B W and Feeney P J 1997 Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings Adv. Drug Deliver. Rev. 23 3
Ghose A K, Viswanadhan V N and Wendoloski J J 1999 A knowledge-based approach in designing combinatorial or medicinal chemistry libraries for drug discovery. 1. A qualitative and quantitative characterization of known drug databases J. Comb. Chem. 1 55
Egan W J, Merz K M and Baldwin J J 2000 Prediction of drug absorption using multivariate statistics J. Med. Chem. 43 3867
Muegge I, Heald S L and Brittelli D 2001 Simple selection criteria for drug-like chemical matter J. Med. Chem. 44 1841
Veber D F, Johnson S R, Cheng H Y, Smith B R, Ward K W and Kopple K D 2002 Molecular properties that influence the oral bioavailability of drug candidates J. Med. Chem. 45 2615.
Martin Y C 2005 A bioavailability score J. Med. Chem. 48 3164
Daina A and Zoete V 2016 A BOILED-Egg to predict gastrointestinal absorption and brain penetration of small molecules ChemMedChem 11 1117
Brenk R, Schipani A, James D, Krasowski A, Gilbert I H, Frearson J and Wyatt P G 2008 Lessons learnt from assembling screening libraries for drug discovery for neglected diseases ChemMedChem 3 435
Baell J B and Holloway G A 2010 New substructure filters for removal of pan assay interference compounds (PAINS) from screening libraries and for their exclusion in bioassays J. Med. Chem. 53 2719
Acknowledgements
This work was partially supported by the Natural Science Foundation of Jiangsu Province (No. BK20190246) and the Natural Science Foundation of the Jiangsu Higher Education Institutions of China (No.19KJB350011). The authors would like to thank Dr. L. Zhu, National Fine Chemicals Quality Supervision and Inspection Center, for recording NMR spectra and LC–MS measurements. The authors gratefully acknowledge the technical support of docking studies and bioassays from Dr. Y. Xu at Guangzhou Institutes of Biomedicine and Health, Chinese Academy of Science.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, M., Liu, Z., Wang, L. et al. Synthesis, evaluation and in silico studies of novel BRD4 bromodomain inhibitors bearing a benzo[d]isoxazol scaffold. J Chem Sci 133, 11 (2021). https://doi.org/10.1007/s12039-020-01874-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12039-020-01874-2