Abstract
In general, biological macromolecules require significant dynamical freedom to carry out their different functions, including signal transduction, metabolism, catalysis and gene regulation. Effectors (ligands, DNA and external milieu, etc) are considered to function in a purely dynamical manner by selectively stabilizing a specific dynamical state, thereby regulating biological function. In particular, proteins in presence of these effectors can exist in several dynamical states with distinct binding or enzymatic activity. Here, we have reviewed the efficacy of ultrafast fluorescence spectroscopy to monitor the dynamical flexibility of various proteins in presence of different effectors leading to their biological activity. Recent studies demonstrate the potency of a combined approach involving picosecond-resolved Förster resonance energy transfer, polarisation-gated fluorescence and time-dependent stokes shift for the exploration of ultrafast dynamics in biomolecular recognition of various protein molecules. The allosteric protein–protein recognition following differential protein–DNA interaction is shown to be a consequence of some ultrafast segmental motions at the C-terminal of Gal repressor protein dimer with DNA operator sequences OE and OI. Differential ultrafast dynamics at the C-terminal of λ-repressor protein with two different operator DNA sequences for the protein–protein interaction with different strengths is also reviewed. We have also systemically briefed the study on the role of ultrafast dynamics of water molecules on the functionality of enzyme proteins α-chymotrypsin and deoxyribonuclease I. The studies on the essential ultrafast dynamics at the active site of the enzyme α-chymotrypsin by using an anthraniloyl fluorescent extrinsic probe covalently attached to the serine-195 residue for the enzymatic activity at homeothermic condition has also been reviewed. Finally, we have highlighted the evidence that a photoinduced dynamical event dictates the molecular recognition of a photochromic ligand, dihydroindolizine with the serine protease α-chymotrypsin and with a liposome (L-α-phosphatidylcholine).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Molecular recognition of biological macromolecules is fundamental to almost every physiological process, including catalysis, mediation of cell motility, cell replication, transcription and, particularly, the protein associations underlying cellular signal transduction (del Sol et al. 2009; Goodey and Benkovic 2008; Kuriyan and Eisenberg 2007; Smock and Gierasch 2009). Recognition required flexibility of biological macromolecules because it facilitates conformational rearrangements and dynamical interaction leading to a different activity (Changeux and Edelstein 2005; Dunker et al. 2000; Lee et al. 2008). In particular, protein dynamics are now widely accepted as universally occurring in all kingdoms of life (Dunker et al. 2000; Ward et al. 2004) and involved in a wide range of cellular functions including escorting, transport, and regulation of transcription and cell signalling (Dyson and Wright 2005; Tompa et al. 2005). The so-called hub proteins, which bind to many partners and are thus central to protein interaction networks, use conformational dynamics to provide the required plasticity to interact with a large number of different proteins (Dosztanyi et al. 2006; Ekman et al. 2006). Conformational dynamics also increases the plasticity and malleability of proteins and facilitates the interaction of the same protein sequence with several binding partners (Fuxreiter et al. 2008), possibly with opposing activities. Importantly, dynamics play the crucial role to allow kinases and other modifying enzymes to access their targets. Induced dynamics in one of the binding partners rule out a lock-and-key mechanism of binding since the altered conformational dynamics could not bound by a complementary surface (Mittag et al. 2010). Further the dynamics of proteins, which are essential for both folding and function, are known to be strongly dependent on solvent viscosity and friction. However, an increasing number of experiments have demonstrated the importance of a contribution to protein dynamics due to internal friction instead of solvent friction. Recently, the two complementary experimental methods (FRET and PET), simulations and theory revealed the role of internal friction in unfolded proteins and its effects on the functional dynamics of intrinsically disordered proteins (Soranno et al. 2017).
Biomolecular recognition of small molecules, such as ligand and nucleic acids, influences the dynamical fluctuation and are essential for macromolecular associations (Frederick et al. 2007). It is also well known that water associated with biological macromolecules is essential for their structure and function (Luby-Phelps et al. 1988; Rupley and Careri 1991). The mobility/dynamics of water allows a protein to interact with other molecules, enables proton transfer and facilitates a large number of biochemical processes (Fenimore et al. 2002; Swenson et al. 2007). Experiments and molecular dynamics (MD) simulations have indicated that the protein motions are mainly determined by the water dynamics and alteration of hydration dynamics can also contribute significantly to dynamics-driven biomolecular recognition (Tarek and Tobias 2002). Changes in these dynamics upon binding can have significant thermodynamic consequences. The free energy of binding is the sum of the changes in enthalpy and entropy of the interacting molecules and their solvent (Frederick et al. 2007; Tzeng and Kalodimos 2009). The estimation of changes in conformational dynamics due to a recognition of protein from molecular dynamics simulations remains a considerable challenge (Grünberg et al. 2006). Experimental measurement of the conformational dynamics of proteins in their free and complexed states is therefore required. Recent developments in nuclear magnetic resonance (NMR) relaxation methods and analysis now make measurement of conformational dynamics feasible (Tzeng and Kalodimos 2009). However, for the intrinsic limitation of the NMR, the technique explores the importance of slow collective motions (millisecond/microsecond regime) rather than that of fast motion (subpicosecond–nanosecond) of biomolecules for their activity (Williams 1989). Hence, nanosecond to subpicosecond dynamics in biomolecules is less explored and understood (Andreatta et al. 2005). Fluorescence spectroscopic techniques involving picosecond-resolved anisotropy, intermolecular Förster resonance energy transfer (FRET) and time-dependent stokes shift (TDSS), which are briefly discussed in the present review are found to be useful to understand biomolecular dynamics over a broad time scale (Brauns et al. 1999, 2002).
The present review addresses the key issues regarding the binding of the protein to different small molecules such as ligand and DNA, leading to differential dynamics of protein in the complexes and its outcome on the biomolecular recognition for different functionalities. The details of the studies including methodology are out of the scope of the review and can be found in the original publications duly cited here. In this regard, we have provided a brief discussion on the role of ultrafast dynamics in the binding of the GalR protein dimer to the DNA operators (OE and OI), which facilitate conformational and dynamical changes thereby leading to allosteric recognition ensuing protein–protein interactions (Choudhury et al. 2016). Subsequently, we have briefed the key differential segmental dynamics in the interaction of λ-repressor proteins with two DNA operators OR1 and OR2 (Mondol et al. 2012). In addition, the role of ultrafast hydration water in the conformational dynamics of α-chymotrypsin (CHT) protein enabling different enzymatic activities has been discussed (Verma et al. 2011). The effect of modulation of the ultrafast dynamics around a model endonuclease glycoprotein, bovine pancreatic deoxyribonuclease I (DNase I) with the addition of polyethylene glycols (PEG with different molecular weight) has also been highlighted (Singh et al. 2017a). The correlated biomolecular recognition and ultrafast dynamics in and around a model enzyme protein α-chymotrypsin are reviewed in various physiologically relevant conditions (Banerjee and Pal 2008). First, a correlation between dynamics of an essential residue (serine) at the active site of the enzyme with the temperature-dependent catalytic efficiency of the enzyme is mentioned. Second, the photoinduced dynamical-driven molecular recognition of a photochromic ligand, dihydroindolizine (DHI), by the enzyme protein is also deliberated (Bagchi et al. 2016). Finally, we have discussed the dynamical modulation of phosphatidylcholine liposome by photochromic dye dihydroindolizine (DHI) to investigate its efficacy for controlled drug delivery (Singh et al. 2017b).
2 Results and discussions
2.1 Dynamics-driven biomolecular recognition of various proteins upon their interaction with different sequences of DNA
Here we have highlighted the important issues regarding the sequences-dependent binding of the protein to different DNA sites, leading to differential ultrafast dynamics of the protein in the complexes and its outcome on the biomolecular recognition involving additional protein–protein or protein–DNA interaction. In this regard, the structural and dynamical changes of Escherichia coli galactose repressor (GalR) and λ-repressor protein upon recognition of two operator DNA target sequences (i.e. OE, OI are used for GalR and OR1, OR2 for gene regulatory λ-protein) have been discussed (Mondol et al. 2012; Choudhury et al. 2016). First, we have used the single tryptophan (Trp165) of the GalR dimer as an intrinsic fluorescent probe. Polarisation-gated fluorescence spectroscopy of Trp165 reveals changes in local fluctuations of the protein upon interaction with operator DNAs. In order to explore intra-protein dynamics of the protein and its complexes cysteine-reactive extrinsic probe, IAEDANS (5-((((2-Iodoacetyl)amino)ethyl)amino)naphthalene-1-sulfonic acid) is used at the C terminus. The decrease in steady-state and time-resolved emission of Trp165 in GalR dimer upon attachment of IAEDANS at their C terminal is shown in figure 1a. The efficiency of energy transfer is calculated to be 50% as a consequence, both the calculated Förster distance (R0) and the Trp165–IAEDANS distance are found to be same (28.3 Ǻ). To study protein dynamics alterations of GalR upon recognition of different operator DNA sequences, FRET of unlabelled and IAEDANS-labelled GalR bound to OE or OI were deliberated (figure 1b and inset of figure 1b, respectively). A 70 ps component for both OE and OI reflects efficient energy transfer from tryptophan to IAEDANS and as consequences Trp165–IAEDANS distance for GalR–OE, and GalR–OI were calculated to be 20.5 and 20.7 Ǻ. The distribution of donor–acceptor distances in the labelled protein (figure 1c) reveals the internal fluctuation of the protein (Nag et al. 2013), which is found to be less broadening in the case of operator-bound GalR compared to native GaIR. Picoseconds-resolved fluorescence (figure 1d) and polarisation-gated (figure 2a and b) studies of IAEDANS were performed to observe the significant structural rearrangement in the C-terminal domain of the repressor protein upon protein–DNA complexation. The faster fluorescent transient and rotational motion of IAEDANS in the presence of OE/OI clearly reveals structural fluctuations in the C-terminal domain of GalR. In order to confirm the location of the IAEDANS and structural differences between GalR–OE /OI complexes, we used FRET from IAEDANS (donor) to the acceptor FITC (fluorescein-5-isothiocyanate) attached to the operator DNA, which is bound to an N-terminal domain of the GalR. The increase in energy transfer efficiency and a decrease in the distribution of the IAEDANS–FITC distances in the protein–OE complexes compared to OI complexes clearly revealed changes in overall protein dynamics upon interaction with the different operators (figure 2c and d). Hence, an attempt is made to correlate the dynamic changes in the protein dimers with OE and OI with the consequent protein–protein interaction (tetramerisation) to form a DNA loop encompassing the promoter segment.
(a) Picosecond-resolved fluorescence transients of Trp165 in GalR or IAEDANS–GalR, inset depicts steady-state emission spectra of Trp165 in GalR and IAEDANS-labelled GalR (λex = 283 nm). Picosecond-resolved fluorescence transients of Trp165 in unlabelled and labelled GalR in presence of (b) OI and inset of (b) OE; λex = 283 nm, λem = 350 nm. (c) Distribution of donor–acceptor distances in labelled GalR and in complexes with OE and OI. (d) Fluorescence transients of IAEDANS-GalR alone or bound to OE or OI, Inset depicts equivalent steady-state emission spectra (λex = 375 nm, λem = 460 nm for all plots).
Fluorescence anisotropy of IAEDANS–GalR (a) alone and complexed with OI, (b) complexed with OE. (c) Fluorescence transient of IAEDANS–GalR alone or in complex with FITC-labelled OI and FITC-labelled OE is shown in insets; λex = 375 nm, λem = 460 nm. (d) Distribution of donor–acceptor distances in complexes with FITC-labelled OE and OI.
Second, we have briefed the functional outcome of λ-repressor protein due to altered dynamics of bound protein in a DNA-sequences–dependent manner (Mondol et al. 2012). In order to monitor the changes in dynamical flexibility of λ-repressor upon complexation with two operator DNAs, OR1 and OR2, a Förster resonance energy transfer technique has been employed by essentially labelling the C-terminal region of λ-repressor through a fluorescent probe 5-(dimethyl-amino) naphthalene-1-sulfonyl chloride (dansyl chloride). The average lifetime of tryptophan decreases from 2.8 ns in λ-repressor to 1.37 ns in dansyl-modified repressor (figure 3a and b). The efficiency of energy transfer is found out to be 50% and, as a consequence, both the Förster distance (R0) and tryptophan– dansyl distance are estimated to be 14.5 Å. An additional faster component of 20 ps appears in the fluorescence transient of tryptophan upon addition of unlabelled OR1 and OR2 DNA signifies intra-protein structural rearrangement in the C-terminal domain of the protein (figure 3a and b). In order to monitor the dynamical time scales and structural differences between λ-repressor–OR1/OR2 DNA complexes, polarization gated fluorescence anisotropy and FRET studies from dansyl to the intercalated ethidium probe bound to the N-terminal domain of the λ-repressor has been used. The higher energy transfer efficiency from dansyl to EtBr in λ-repressor–OR2 DNA–EtBr complex than λ-repressor–OR1 DNA–EtBr complex (inset of figure 3a and b) and enhanced flexibility due to fast time scales of the C-terminal domain of the λ-repressor after complex formation with OR1 (figure 3c) indicates that the structure of the protein is more compact in the OR2 complex than in the OR1 complex. Thus, an attempt has been made to elucidate that differential dynamics is important for protein–protein interaction between two λ-repressor dimers bound to OR1 and OR2. The interaction was studied by sedimentation equilibrium (figure 3d). Here, OR1 was end labelled with fluorescein and a labelled OR1/repressor complex was formed. This was mixed with an excess of unlabelled OR2/repressor complex and sedimentation equilibrium run was performed. Fitting of the concentration profile to appropriate equations yields dissociation constant of 4.4 μM and 41.6 μM for OR1/repressor–OR2/repressor and OR1/repressor–OR1/repressor, respectively. Finally, it clearly corroborates that the structural and dynamical differences are important for the correct protein–protein interaction.
FRET studies in operator–repressor complexes. (a) Picosecond-resolved fluorescence transients of tryptophan residues in λ-repressor (λ, green), in dansyl-modified λ-repressor (λ-DC; red) and in dansylated λ-repressor-OR1 DNA complex (λ-DC-OR1 DNA, violet) λex-299 nm and λem(Trp)-350 nm, respectively. Picosecond-resolved fluorescence transients of dansyl in λ-repressor-OR1 DNA complex in absence (λ-DC-OR1 DNA, green) and in presence (λ-DC-OR1 DNA-EtBr, red) of EtBr are shown in inset. (b) Picosecond-resolved fluorescence transients of tryptophan residues in λ-repressor (λ; green), in dansyl-modified λ-repressor (λ-DC; red) and in dansylated λ-repressor–OR2 DNA complex (λ-DC-OR2 DNA; violet). Picosecond-resolved fluorescence transients of dansyl in λ-repressor-OR2 DNA complex in absence (λ-DC-OR2 DNA; green) and in presence of EtBr (λ-DC-OR2 DNA-EtBr; red) are shown in inset, λex-375 nm and λem(DC)-515 nm, respectively. (c) Fluorescence anisotropy, r(t) of dansyl in λ repressor–OR1 DNA complex and dansyl in λ-repressor–OR2 DNA complex. (d) Determination of dimer–tetramer association by sedimentation equilibrium. The inset cartoon figure shows the type of association that is being studied here. The red rod represents the operator DNA, while the blue balls represent the protein. Self-association of OR1–repressor complex are shown in inset and association between OR1–repressor complex and the OR2–repressor complex are shown in (d).
2.2 Modulation of conformational dynamics by external milieu leading to differential recognition of protein
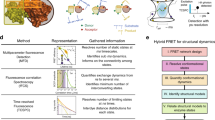
Here, we focus on the effect of external milieu on alteration of protein dynamics, which leads to different activity. The water activity of protein was varied by adding osmotically inert solvents as well as by varying the temperature. Here, an attempt has been made to correlate the dynamical flexibility of protein in presence of external milieu with its activity by using ultrafast spectroscopy (Banerjee and Pal 2008; Singh et al. 2017a; Verma et al. 2011). First, the role of hydration on the functionality, that is, molecular recognition of a proteolytic enzyme α-chymotrypsin (CHT) was deliberated by modulating the water activity with the addition of polyethylene glycol (PEG, MW 400) (Verma et al. 2011). The kinetics of hydrolysis of the substrate peptide Ala-Ala-Phe-7-amido-4-methyl coumarin (AMC) by CHT is found to decrease gradually as the concentration of PEG increases in the solution. It is evident from figure 4a and its inset that increase in PEG concentration decreases both KM and kcat for the substrate AMC in CHT. Further, the overall secondary and tertiary structures of CHT determined from far-UV and near-UV circular dichroism (CD) measurements show no considerable change in the presence of PEG (Verma et al. 2011), which corroborates that the change in catalytic activity induced by PEG are not accompanied by alterations in the secondary and tertiary structures of α-chymotrypsin. Energetics calculations show that ΔGM value becomes more negative, whereas both ΔG# and ΔG #T becomes more positive as PEG concentration is increased in the solution, which indicates that the addition of PEG facilitates the formation of the ES complex (entrance channel) while restricting the formation of the product (exit channel). This decrease is attributed to the thinning of the hydration shell of the enzyme due to the loss of critical water residues as evidenced from volumetric and compressibility measurements (Verma et al. 2011). In order to specifically examine the local hydration dynamics of the enzyme CHT at various concentration of PEG, the steady-state and time-resolved fluorescence spectroscopy has been employed. A chromophoric inhibitor of the enzyme (proflavin) is used to monitor the change in local polarity of the enzyme’s active site, where as local hydration dynamics at the surface of the enzyme CHT is explored by using probe ANS, commonly used to monitor structural changes of proteins and membranes. Figure 4(b) depicts the emission spectrum of proflavin in CHT, and it is found that the fluorescence maximum shifts towards lower wavelength with the addition of PEG indicating a decrease in the polarity around the probe, which also validates the volumetric and compressibility measurements. The fluorescence intensity of ANS bound to CHT is found to decrease along with a small red-shift with the addition of PEG (inset of figure 4b), which suggests that ANS binding site is now experiencing higher polarity and hence greater contribution from bulk-type water compared to that in the absence of PEG. The presence of faster decay components at the blue end and a rise in component at the red wavelength is consistent with the picture of solvation dynamics (inset of figure 4c). A representative TRES for ANS-CHT in the buffer presented in figure 4c shows a significant dynamic fluorescence Stokes shift of 1256 cm−1 in 8 ns. The faster hydration dynamics (figure 4d) leading to enhanced tumbling motion of enzyme (inset of figure 4d) at the higher concentration of PEG suggests thinning of the hydration water shell around the enzyme, which leads to decrease in enzymatic activity of CHT.
(a) KM (filled squares) and kcat (open squares) for the catalytic activity of CHT on the substrate AMC as a function of PEG concentrations. The plot of ln \( \frac{{K_{M}^{O} }}{{K_{M}^{\pi } }} \)(filled squares) and ln\( \frac{{V_{M}^{O} }}{{V_{M}^{\pi } }} \) (open squares) against Δπ are shown in the inset. (b) Emission spectra of proflavin in buffer, and CHT with various wt% PEG (λex = 409 nm). Emission spectra of free ANS in buffer, and CHT with various wt% PEG (λex = 375 nm) are shown in inset. (c) Time-resolved emission spectra (TRES) for ANS–CHT complex in buffer. Picosecond-resolved fluorescence transients of ANS-CHT adduct in buffer are shown in the inset (c). Solvation correlation function, C(t), of ANS-CHT adduct in buffer and 5 and 30 wt% PEG (d). Time-resolved anisotropy decay, r(t), of ANS-CHT in buffer and 30 wt% PEG is shown in the inset (d).
Second, we systematically brief the role of molecular crowding (polyethylene glycol, MW 3350) on dynamics of endonuclease glycoprotein known as bovine pancreatic deoxyribonuclease I (DNase I), which leads to differential activity of the protein (Singh et al. 2017a). The kinetics of hydrolysis of the substrate dsDNA-EB by DNase I has been measured by steady-state fluorescence technique, which indicates that the rate of exclusion of the EB from DNA upon hydrolysis by DNase I increases gradually as the concentration of PEG increases in the solution. The calculated Michaelis-Menten constant (Km) and maximum velocity (Vmax) for the different PEG concentrations are presented in figure 5a and its inset, which clearly indicates that with the increase in PEG concentration, Vmax increases, however, Km for the enzymatic activities of DNase I for substrate DNA decreases. These kinetic parameters reveal that molecular crowding influences the hydrolytic activity of the DNase I by affecting catalytic activity as well as its binding affinity towards the substrate DNA (Wenner and Bloomfield 1999; Zimmerman and Harrison 1987; Morán-Zorzano et al. 2007; Su et al. 2013). Far UV CD corroborates that the secondary structures of the enzyme (Ajtai and Venyaminov 1983) is insignificantly perturbed in the presence of PEG, indicating the altered enzymatic activity induced by PEG are not accompanied by alterations in the secondary and tertiary structures of DNase I (Singh et al. 2017a). In order to investigate the effect of PEG in hydrolysis of DNA, picosecond-resolved transients of EB intercalated to DNA upon interaction with DNase I are studied in presence of 15 wt% PEG (figure 5b). It is observed that the maximum population of the dye intercalated to the DNA ~ 21 ns (54%) decreases upon hydrolysis by DNase I (~21 ns (18%)), which is found to be extensively lower than that of DNA hydrolysis in buffer solution (~21 ns (29%)), corroborates the higher cleavage in presence of PEG (inset of figure 5b). Further, the dynamics and binding affinity of DNase I with DNA has been monitored by using a biologically relevant probe 8-anilino-1-naphthalenesulfonic acid ammonium salt (ANS) (figure 5c). The enhanced dynamical flexibility of protein in presence of PEG as revealed from polarization gated anisotropy of ANS (figure 5d) has been correlated with the stronger DNA binding leading to higher nuclease activity as the efficiency of energy transfer is enhanced in the presence of PEG.
(a) KM for the catalytic activity of DNase I on the substrate DNA as a function of PEG concentrations. The plot of Vmax against PEG concentration is shown in the inset. Picosecond-resolved transient of free EB, EB in DNA and in DNA–DNase I complex in presence of (b) 15 wt% PEG and 0 wt% PEG are shown in inset. Picosecond-resolved transients of the donor (ANS-DNase I-DNA complex) in the absence and presence of the acceptor (EB) in DNA in (c) 15 wt% PEG and 0 wt% PEG are shown in inset. Time-resolved fluorescence anisotropy of ANS bound to DNase I–DNA complex in presence of (d) 15 wt% PEG and 0 wt% PEG are shown in inset.
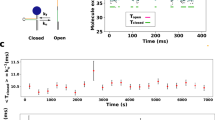
Lastly, in this section, the picosecond-resolved anisotropy was used to explore the dynamics of the serine residue in the active site of CHT, which correlates the observed dynamics with the temperature-dependent catalytic efficiency of the enzyme (Banerjee and Pal 2008). The temperature-dependent hydrolysis of the substrate peptide Ala-Ala-Phe-7-amido-4-methyl coumarin (AMC) by CHT reveals maximum catalytic efficiency at 37 °C coinciding with the normal body temperature of homeothermal animals (figure 6a). The fall in the turnover rate (kcat) and the increase in association (KM) possibility of the enzyme–substrate complex at 50 °C suggest that, at the higher temperatures, even when an enzyme–substrate complex is formed favorably, product formation is hindered. To explore the globular tertiary structures of the protein at different temperatures, DLS and CD techniques were used. While CD measurement (Banerjee and Pal 2008) didn’t disclose any major structural perturbations at a higher temperature, DLS measurements reveals decease in hydrodynamic diameter of CHT from ~6 nm to ~1 nm above 50 °C (inset of figure 6a). However, such fragments do not appear for lower concentration (<50 μM) of the protein at any temperature, indicates that the fall in enzymatic activity of the protein at high temperatures is neither due to autocatalysis nor due to the massive unfolding of the enzyme. The decrease in hydrodynamic diameter of proteins at the high temperature can be associated with the thinning of the hydration shell, which is also corroborated by an increase in adiabatic compressibility (φk) with increase in the temperature (Banerjee and Pal 2008). In order to explore the local dynamics of proteins, picosecond-gated fluorescence spectroscopy was used by covalently labelling the protein using fluorophores anthraniloyl group (attached to the serine-195 residue) and proflavin (PF) (attached to the active site of the protein). Figure 6(b) and its inset reflects the change in the fast and slow components of the rotational relaxation of PF at the protein active site. While slow components of PF represent microviscosity of the active site becoming buffer-like at the higher temperature, fast components of PF reflects the subslip rotational motion of the probe (Inamdar et al. 2006; Narayanan et al. 2008), which decreases monotonically with temperature (table 1). The faster time constants ~30 ps associated with the anthraniloyl group remains constant in the temperature range 10–40 °C. However, at 50 °C, this motion of the serine residue becomes slower (figure 6c), indicating a lower frequency of vibrations. The loss in catalytic activity at this temperature, coinciding with the slowing down of this component, suggests that the fast dynamics of the serine residue is associated with an enzymatic activity. Further, the global motion and the normal mode dynamics of the protein having a time constant of 12 ns becomes faster (9 ns) at high temperatures and vanishes at 30 °C (where the protein activity is near maximum). The absence of this component at higher temperatures suggests that the active site conformation, suitable for catalysis, is preachieved by the protein at that temperature. The consequences of the conformational dynamics to the active site structure at different temperatures have been further monitored through fluorescence-detected circular dichroism (FDCD) studies by using anthraniloyl and the well-known CHT inhibitor PF. The monotonic decreases in the FDCD signal with increasing temperatures suggest the progressive decrease in the induced chirality of the bound fluorophore (figure 6d). This loss in induced chirality of the fluorophore with increasing temperature suggests greater conformational flexibility of the protein at higher temperatures, which leads to increase in the catalytic activity up to 40 °C. The fall of catalytic activity at 50 °C could occur due to least rigid binding of the substrate mimic/inhibitor because of the higher conformational flexibility of the protein, which suggests that the existence of a critical conformation is required for enhanced catalysis.
(a) The variation of the catalytic efficiency of CHT with temperature. The solid line is a Gaussian fit. Hydrodynamic of CHT at different temperatures are shown in inset. The rotational correlation times of fluorescence anisotropy of PF in CHT (b) and inset of (d) and anthraniloyl-CHT (ANT-CHT) (c) and inset of (c). The solid lines are the fit of the Stokes–Einstein–Debye equation; the broken lines are a guide for the eye. The FDCD spectra of CHT using (d) PF-CHT and ANT as fluorescent probes are shown in inset.
2.3 Photoinduced ultrafast dynamics-mediated molecular recognition
Herein, we report the photo-control dynamical alteration of protein and liposome by using a new class of photochromic ligand, dihydroindolizine (DHI). The enzymatic activity by photochromic DHI has been explored in a light-responsive manner. The photo-controlled alteration of liposome (L-α-phosphatidylcholin) dynamics and morphology via the incorporation of a new class of synthesized photochromic material, dihydroindolizine (DHI), has also been explored to investigate its efficacy for controlled drug delivery (Bagchi et al. 2016; Singh et al. 2017b). First, we demonstrate herein that the molecular recognition of a photochromic ligand, dihydroindolizine (DHI), by serine protease α-chymotrypsin (CHT) leads to the photo-control of enzymatic activity (Bagchi et al. 2016). The basis of the photochromic behavior of DHI is light-induced reversible pyrroline ring opening, which transforms the molecule from a light yellow-coloured form (cis) to a red-coloured betaine form (trans) (Fernando et al. 2015). Betaines undergo a thermal back-reaction to their corresponding DHI cis form by 1,5-electrocyclization (Fernando et al. 2016). The optical absorption spectrum of the cis-isomer in acetonitril shows a peak at 390 nm, which reduces in intensity when exposed to UV light, yielding a subsequent peak at 520 nm (Bagchi et al. 2016). To monitor the kinetic of isomerization, cis to trans conversion has been monitored by measuring the increase in absorbance at 520 nm, whereas trans to cis conversion is followed by a decrease in absorbance at 520 nm. The decreased isomerization of DHI upon interaction with CHT corroborates the stability of interaction of hydrophobic DHI isomer with various hydrophobic binding sites present in the CHT (figure 7a and inset of figure 7a). CD experiments of CHT and CHT-DHI (in dark and light conditions) reveal no significant alteration in the structure of CHT and hence are not responsible for the change in enzymatic activity (Bagchi et al. 2016). After confirming the successful interaction between CHT and DHI in the dark and in the presence of UV light, the corresponding enzymatic activities were explored, which was found to be more hindered in dark compared to the light-irradiated condition, indicating that the cis-isomer is more capable of inhibiting CHT activity compared with the trans-isomer (figure 7b). Inset of figure 7b clearly depicts that the decay constant is much slower in dark compared to presence of light (table 2). In order to explore the site of CHT, which interacts with different isomers of DHI leading to its altered enzymatic activity, FRET techniques were used by labelling the CHT using fluorescent probes NPA and PF separately. While NPA was used as a FRET pair for cis isomer of DHI, location of trans isomer was confirmed by FRET pair PF (figure 7c). A shorter excited-state lifetime of the CHT-NPA in the presence of the DHI clearly reveals the energy transfer from NPA to cis-isomer of DHI, whose efficiency is calculated to be 79% (figure 7c). This observation confirms that the DHI cis-isomer attachment site is located at a distance of 2.5 nm from the NPA binding site (table 3), which subsequently proved that the DHI cis-isomer binds at a site other than the S1 pocket of CHT. Further location of trans-isomer of DHI in CHT were employed by the simultaneous binding of PF and DHI in CHT, enabling the possibility of energy transfer from CHT-PF (donor) to the DHI trans-isomer (i.e. when illuminated with UV light). The quenching in the excited-state lifetime of the CHT-PF in presence of DHI (trans) indicates the energy transfer process whose efficiency is calculated to be 61% and hence the distance between the donor CHT-PF and acceptor DHI trans-isomer is determined to be 33±2.5 Å nm (inset of figure 7c). Figure 7d depicts distribution of the donor–acceptor distances in the CHT-DHI (trans)-PF, revealing an internal fluctuation with a full-width half-maximum (FWHM) of 2.5 Å. Since enzymatic inhibition does not result due to competitive interaction of DHI at the Ser-195-His-57 active site, inhibition must result due to its interactions with a control or allosteric site. This allosteric site may be hydrophobic in nature, and thus the hydrophobic inhibitor may bind via electrostatic and hydrophobic interactions and leads to unfavourable conformational change at some different sites on the enzyme, significantly hampering its enzymatic activity (Smith and Hansch 1973).
(a) Kinetics of the trans to cis conversion reaction of CHT-DHI in phosphate buffer. Inset shows the corresponding cis to trans conversion rate. (b) Enzymatic activity of CHT-DHI in absence and presence of UV light. Photo-control of enzymatic activity of CHT-DHI are shown in inset. (c) The fluorescence transients of CHT-DHI-PF (excitation at 375 nm) in cis and in trans forms collected at 520 nm are shown. The fluorescence transients of CHT-NPA (excitation at 375 nm) in the absence and in the presence of DHI cis collected at 450 nm are shown in inset. (d) Distribution of donor–acceptor distances between CHT-NPA-DHI cis-isomer and the CHT-PF-DHI (trans).
Second, the efficacy of photoresponsive destabilization to phosphatidylcholine liposome, which is used as potential drug delivery vehicles, was investigated by using a synthesized photochromic dye dihydroindolizine (DHI) (Singh et al. 2017b). This structural conversion of DHI from closed to open isomer can fluctuate or defect the liposomal membrane by mechanical stress and leads photoresponsive destabilization to liposome. The consequence of different isomerization of DHI on liposome stability was monitored by steady-state, time-resolved fluorescence and polarisation-gated fluorescence spectroscopy by labelling the liposome with fluorescent probe ANS. The small red-shift (~5 nm) in emission maximum of ANS-PC-DHI upon UVA irradiation with decrease in the fluorescence intensity corroborates that ANS is now experiencing higher polarity upon closed-to-open transition of DHI (inset of figure 8a). Further, the faster fluorescence decay of ANS-PC-DHI upon UVA irradiation leading to the enhanced internal rotation of the fluorophore relative to the liposome also indicates a progressive release of restriction on the probe might be due to increase in the mobility of solvating species (figure 8b). To understand the fusion phenomenon, Förster resonance energy transfer (FRET) techniques were used where ANS was encapsulated in a group of liposomes, and doxorubicin (DOX) was encapsulated in another group of liposomes both having DHI. The decrease in average lifetime of ANS-PC-DHI upon UVA-irradiation due to energy transfer from ANS to DOX, which otherwise does not depicts any quenching when simple DOX is diluted in the medium containing ANS-PC-DHI, indicates that UVA-irradiated DHI-sensitized liposome could not lead to rearrangement of bilayer and total membrane perturbation; rather, it leads to fusion of liposome (figure 8c). Based on the photoresponsive properties and microstructural change investigated earlier, DHI-liposome could be considered as photoresponsive drug delivery system. Inset of figure 8d corroborates a burst release occurred upon UVA irradiation at the 30 min, followed by a slower sustained release up to 1 h, whereas spontaneous release of DOX was observed in the group without UVA irradiation. The therapeutic efficacy of the drug-loaded liposome was further evaluated against cervical cancer cell line HeLa by exposing the cells directly to the PC-DHI-DOX in presence or absence of UVA (figure 8d). MTT assay studies reveal an enhanced cellular uptake of DOX leading to significant reduction in cell viability (~40%) of HeLa, followed by photoresponsive destabilization of liposome. The results presented in this study indicated that DHI-encapsulated liposome could serve as a safe and promising drug delivery vehicle.
(a) Time-resolved transients of ANS bound to PC and ANS bound to PC-DHI in the presence and absence of UVA light. Corresponding steady-state emission spectra of ANS in different systems are shown in inset. (b) Time-resolved anisotropy of ANS bound to PC-DHI in presence of UVA light and dark are shown in inset. Picosecond-resolved transients of the donor-acceptor in the absence and presence of UVA light, (c) donor is (ANS-PC-DHI) and acceptor is (DOX-PC-DHI) and donor is (ANS-PC-DHI) and acceptor is free DOX are shown in inset. (d) Cytotoxicity assay in HeLa cells with PC-DHI, PC-DOX and PC-DHI-DOX with MTT as an indicator dye in the presence and absence of UV light and in vitro release profile of DOX from PC-DHI in absence and presence of UVA light are shown in inset.
3 Conclusion
The review provides substantial insight into the ultrafast dynamical studies of protein which were found to be essential for biomolecular recognition. In one of the exemplary studies, an attempt was made to correlate the ultrafast dynamical changes in the model protein dimers GalR with different operator DNA sequences OE and OI, consequently leading to protein–protein interaction (tetramerisation) to form a DNA loop encompassing the promoter segment. FRET from the single tryptophan residue to a covalently attached probe IAEDANS at a cysteine residue in the C-terminal domain of the protein revealed that intra-protein fluctuation, which was found to be slower upon binding with OI than OE. However, for FRET between IAEDANS and another extrinsic probe FITC in the operator DNA, the faster fluctuation was observed for the GalR–OI complex. The flexibility of the C-terminal domain of the protein was measured by picosecond-resolved polarisation-gated fluorescence spectroscopy, which corroborates the faster IAEDANS motions upon binding to OI than to OE. Thus, differential flexibility at the active site of the protein upon recognition with two different DNA operator sequences were shown to be crucial for dimer–dimer interaction through C terminal domains (active sites).
Ultrafast dynamics were also shown to be important for the operation of the genetic switch in the life cycle of the λ-phase virus, where the interaction of λ-repressor protein with different operator DNAs played a crucial role. In case of λ-Repressor protein–operator DNA sites interaction with OR1 and OR2 DNAs, the FRET from the dansyl (bound to the C-terminal domain of the protein) to the EtBr (intercalated in the operator DNA) directed a more compact structure of the protein in OR2 complex than in the OR1 complex. Picosecond-resolved fluorescence anisotropy revealed enhanced flexibility of the C-terminal domain of the repressor at fast timescales after complex formation with OR1. In contrast, OR2-bound repressor has revealed no significant enhancement of protein dynamics at these timescales. These differences are shown to be important for correct protein–protein interactions.
The ultrafast dynamics of water molecules in the close vicinity of protein (hydration) were shown to play an important role in the physical functionalities of the proteins. Here we discussed the role of hydration dynamics on the enzymatic activity by using crowding agent/osmolyte polyethylene glycol (PEG). The enzymatic activity of CHT was found to be decreased upon addition of osmotic/crowding agent, PEG. A detailed, energetic calculation showed that the stabilization of the entrance path (decrease in KM) essentially destabilized the exit channel (decrease in kcat) with increase in osmotic stress. The enhanced rotational as well as faster hydration dynamics at the surface of an enzyme was found to be responsible for the decrease in the enzymatic activity due to dehydration of the biological water. It has to be noted that the effect of osmolyte on proteins may not be considered as monotonously general as it involves crucial dynamical events in and around the proteins under investigation. In another study involving DNase I, the osmolytic stress essentially increased the enzymatic activity. The reduced water activity at the enzyme surface due to osmotic stress of the molecular crowding agent enhancing the dynamical flexibility of the enzyme was responsible to increase the DNA binding, which eventually accelerates the hydrolysis reaction of DNase I. Other than ultrafast domain motion of proteins, there are evidences that ultrafast dynamics of a specific residue of a protein plays a crucial role in the molecular recognition, leading to overall enzymatic turnover in physiological condition. In this direction, the dynamics of a serine residue in the active site of the proteolytic enzyme α-chymotrypsin (CHT) was correlated with its temperature-dependent catalytic efficiency. The catalytic efficiency kcat/KM of the enzyme at different temperatures showed maxima at 37 °C, coinciding with the normal body temperature of homeothermals. We have also discussed the case where ultrafast dynamics (cis–trans isomerization) of small ligands dihydroindolizine (DHI) dictates the molecular recognition by the protein CHT. The recognition of photochromic DHI by CHT altered the enzymatic activity of CHT in a light-responsive manner. The relative orientations and variations in the interactions of different isomer (cis–trans) of DHI within the protein cavity have been responsible for alteration of enzymatic activity. Further, the photo-controlled modulation of liposome (L-phosphatidylcholin) dynamics and morphology via the incorporation of dihydroindolizine (DHI) demonstrated that structural conversion of DHI from closed to open isomer could fluctuate or defect the liposomal membrane by mechanical stress and hence responsible for the fabrication of light-triggered drug delivery systems. To our understanding, the relevance of ultrafast processes in the relatively slower molecular recognition may find impact in the knowledge of molecular recognition in atomic resolution.
References
Ajtai K and Venyaminov SY 1983 CD study of the actin DNase I complex. FEBS Lett. 151 94–96
Andreatta D, Pérez Lustres JL, Kovalenko SA, Ernsting NP, Murphy CJ, Coleman RS and Berg MA 2005 Power–law solvation dynamics in DNA over six decades in time. J. Am. Chem. Soc. 127 7270–7271
Bagchi D, Ghosh A, Singh P, Dutta S, Polley N, Althagafi II, Jassas RS, Ahmed SA and Pal SK 2016 Allosteric inhibitory molecular recognition of a photochromic dye by a digestive enzyme: dihydroindolizine makes α-chymotrypsin photo-responsive. Sci. Rep. 6
Banerjee D and Pal SK 2008 Conformational dynamics at the active site of α-chymotrypsin and enzymatic activity. Langmuir 24 8163–8168
Brauns EB, Madaras ML, Coleman RS, Murphy CJ and Berg MA 1999 Measurement of local DNA reorganization on the picosecond and nanosecond time scales. J. Am. Chem. Soc. 121 11644–11649
Brauns EB, Madaras ML, Coleman RS, Murphy CJ and Berg MA 2002 Complex local dynamics in DNA on the picosecond and nanosecond time scales. Phys. Rev. Lett. 88 158101
Changeux J-P and Edelstein SJ 2005 Allosteric mechanisms of signal transduction. Science 308 1424–1428
Choudhury S, Naiya G, Singh P, Lemmens P, Roy S and Pal SK 2016 Modulation of ultrafast conformational dynamics in allosteric interaction of gal repressor protein with different operator DNA sequences. ChemBioChem 17 605–613
del Sol A, Tsai C-J, Ma B and Nussinov R 2009 The origin of allosteric functional modulation: multiple pre-existing pathways. Structure 17 1042–1050
Dosztanyi Z, Chen J, Dunker AK, Simon I and Tompa P 2006 Disorder and sequence repeats in hub proteins and their implications for network evolution. J. Proteome Res. 5 2985–2995
Dunker AK, Romero P, Obradovic Z, Garner EC and Brown CJ 2000 Intrinsic protein disorder in complete genomes. Genomics Inform 11 161–171
Dyson HJ and Wright PE 2005 Intrinsically unstructured proteins and their functions. Nat. Rev. Mol. Cell Biol. 6 197–208
Ekman D, Light S, Björklund ÅK and Elofsson A 2006 What properties characterize the hub proteins of the protein–protein interaction network of Saccharomyces cerevisiae? Genome Biol. 7 R45
Fenimore PW, Frauenfelder H, McMahon BH and Parak FG 2002 Slaving: solvent fluctuations dominate protein dynamics and functions. Proc. Natl. Acad. Sci. USA 99 16047–16051
Fernando A, Malalasekera AP, Yu J, Shrestha TB, McLaurin EJ, Bossmann SH and Aikens CM 2015 Refined insights in the photochromic spiro-dihydroindolizine/betaine system. J. Phys. Chem. A 119 9621–9629
Fernando A, Shrestha TB, Liu Y, Malalasekera AP, Yu J, McLaurin EJ, Turro C, Bossmann SH and Aikens CM 2016 Insights from theory and experiment on the photochromic spiro-dihydropyrrolo–pyridazine/betaine system. J. Phys. Chem. A 120 875–883
Frederick KK, Marlow MS, Valentine KG and Wand AJ 2007 Conformational entropy in molecular recognition by proteins. Nature 448 325–329
Fuxreiter M, Tompa P, Simon I, Uversky VN, Hansen JC and Asturias FJ 2008 Malleable machines take shape in eukaryotic transcriptional regulation. Nat. Chem. Biol. 4 728–737
Goodey NM and Benkovic SJ 2008 Allosteric regulation and catalysis emerge via a common route. Nat. Chem. Biol. 4 474–482
Grünberg R, Nilges M and Leckner J 2006 Flexibility and conformational entropy in protein–protein binding. Structure 14 683–693
Inamdar S, Mannekutla J, Mulimani B and Savadatti M 2006 Rotational dynamics of nonpolar laser dyes. Chem. Phys. Lett. 429 141–146
Kuriyan J and Eisenberg D 2007 The origin of protein interactions and allostery in colocalization. Nature 450 983–990
Lee J, Natarajan M, Nashine VC, Socolich M, Vo T, Russ WP, Benkovic SJ and Ranganathan R 2008 Surface sites for engineering allosteric control in proteins. Science 322 438–442
Luby-Phelps K, Lanni F and Taylor DL 1988 The submicroscopic properties of cytoplasm as a determinant of cellular function. Annu. Rev. Biophys. Biophys. Chem. 17 369–396
Mittag T, Kay LE and Forman-Kay JD 2010 Protein dynamics and conformational disorder in molecular recognition. J. Mol. Recognit. 23 105–116
Mondol T, Batabyal S, Mazumder A, Roy S and Pal SK 2012 Recognition of different DNA sequences by a DNA binding protein alters protein dynamics differentially. FEBS Lett. 586 258–262
Morán-Zorzano MT, Viale AM, Muñoz FJ, Alonso-Casajús N, Eydallín GG, Zugasti B, Baroja-Fernández E and Pozueta-Romero J 2007 Escherichia coli AspP activity is enhanced by macromolecular crowding and by both glucose-1, 6-bisphosphate and nucleotide-sugars. FEBS Lett. 581 1035–1040
Nag S, Sarkar B, Chandrakesan M, Abhyanakar R, Bhowmik D, Kombrabail M, Dandekar S, Lerner E, Haas E and Maiti S 2013 A folding transition underlies the emergence of membrane affinity in amyloid-β. Phys. Chem. Chem. Phys. 15 19129–19133
Narayanan SS, Sarkar R, Sinha SS, Dias F, Monkman A and Pal SK 2008 Luminescence depolarization dynamics of quantum dots: Is it hydrodynamic rotation or exciton migration? J. Phys. Chem. C 112 3423–3428
Rupley JA and Careri G 1991 Protein hydration and function. Adv. Protein Chem. 41 37–172
Singh P, Choudhury S, Dutta S, Adhikari A, Bhattacharya S, Pal D and Pal SK 2017a Ultrafast spectroscopy on DNA-cleavage by endonuclease in molecular crowding. Int. J. Biol. Macromol. 103 395–402
Singh P, Choudhury S, Kulanthaivel S, Bagchi D, Banerjee I, Ahmed SA and Pal SK 2017b Photo-triggered destabilization of nanoscopic vehicles by dihydroindolizine for enhanced anticancer drug delivery in cervical carcinoma. Colloids Surf. B
Smith RN and Hansch C 1973 Hydrophobic interaction of small molecules with alpha-chymotrypsin. Biochemistry 12 4924–4937
Smock RG and Gierasch LM 2009 Sending signals dynamically. Science 324 198–203
Soranno A, Holla A, Dingfelder F, Nettels D, Makarov DE and Schuler B 2017 Integrated view of internal friction in unfolded proteins from single-molecule FRET, contact quenching, theory, and simulations. Proc. Natl. Acad. Sci. USA 114 E1833–E1839
Su X, Zhang C, Zhu X, Fang S, Weng R, Xiao X and Zhao M 2013 Simultaneous fluorescence imaging of the activities of DNases and 3′ exonucleases in living cells with chimeric oligonucleotide probes. Anal. Chem. 85 9939–9946
Swenson J, Jansson H, Hedström J and Bergman R 2007 Properties of hydration water and its role in protein dynamics. J. Phys. Condens. Matter 19 205109
Tarek M and Tobias D 2002 Role of protein–water hydrogen bond dynamics in the protein dynamical transition. Phys. Rev. Lett. 88 138101
Tompa P, Szasz C and Buday L 2005 Structural disorder throws new light on moonlighting. Trends Biochem. Sci. 30 484–489
Tzeng S-R and Kalodimos CG 2009 Dynamic activation of an allosteric regulatory protein. Nature 462 368–372
Verma PK, Rakshit S, Mitra RK and Pal SK 2011 Role of hydration on the functionality of a proteolytic enzyme α-chymotrypsin under crowded environment. Biochimie 93 1424–1433
Ward JJ, Sodhi JS, McGuffin LJ, Buxton BF and Jones DT 2004 Prediction and functional analysis of native disorder in proteins from the three kingdoms of life. J. Mol. Cell Biol. 337 635–645
Wenner JR and Bloomfield VA 1999 Crowding effects on EcoRV kinetics and binding. Biophys. J. 77 3234–3241
Williams RJ 1989 NMR studies of mobility within protein structure. FEBS J. 183 479–497
Zimmerman SB and Harrison B 1987 Macromolecular crowding increases binding of DNA polymerase to DNA: an adaptive effect. Proc. Natl. Acad. Sci. USA 84 1871–1875
Acknowledgements
DB thanks INSPIRE (DST) for the research fellowship. We thank DBT (India) and DST (India) for financial Grants (BT/PR11534/NNT/28/766/2014) and (EMR/2016/004698). We thank colleagues in our laboratory at S. N. Bose National Centre for Basic Sciences, whose contribution over the years, acknowledged in the references, have been priceless in the successful evolution of work in this area. In particularly, we thank Mr Susobhan Choudhury, Ms Gitashri Naiya, Dr Tanumoy Mondol, Dr Subrata Batabyal, Dr Pramod Kumar Verma and Dr Debapriya Banerjee. We thank Prof Siddhartha Roy, Dr Saleh A Ahmed, Dr Indranil Banerjee, Dr Siddhartha Bhattacharya, Dr Debasish Pal, Dr Rajib Kumar Mitra and Prof Peter Lemmens for the collaboration work.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Singh, P., Bagchi, D. & Pal, S.K. Ultrafast dynamics-driven biomolecular recognition where fast activities dictate slow events. J Biosci 43, 485–498 (2018). https://doi.org/10.1007/s12038-018-9776-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12038-018-9776-z