Abstract
To investigate the role of DNA methylation in modulating chronic neuropathic pain (NPP), identify possible target genes of DNA methylation involved in this process, and preliminarily confirm the medicinal value of the DNA methyltransferases (DNMTs) inhibitor 5-azacytidine (5-AZA) in NPP by targeting gene methylation. Two rat NPP models, chronic constriction injury (CCI) and spinal nerve ligation (SNL), were used. The DNA methylation profiles in the lumbar spinal cord were assayed using an Arraystar Rat RefSeq Promoter Array. The underlying genes with differential methylation were then identified and submitted to Gene Ontology and pathway analysis. Methyl-DNA immunoprecipitation quantitative PCR (MeDIP-qPCR) and quantitative reverse transcription-PCR (RT-qPCR) were used to confirm gene methylation and expression. The protective function of 5-AZA in NPP and gene expression were evaluated via behavioral assays and RT-qPCR, respectively. Analysis of the DNA methylation patterns in the lumbar spinal cord indicated that 1205 differentially methylated fragments in CCI rats were located within DNA promoter regions, including 638 hypermethylated fragments and 567 hypomethylated fragments. The methylation levels of Grm4, Htr4, Adrb2, Kcnf1, Gad2, and Pparg, which are associated with long-term potentiation (LTP) and glutamatergic synapse pathways, were increased with a corresponding decrease in their mRNA expression, in the spinal cords of CCI rats. Moreover, we found that the intraperitoneal injection of 5-AZA (4 mg/kg) attenuated CCI- or SNL-induced mechanical allodynia and thermal hyperalgesia. Finally, the mRNA expression of hypermethylated genes such as Grm4, Htr4, Adrb2, Kcnf1, and Gad2 was reversed after 5-AZA treatment. CCI induced widespread methylation changes in the DNA promoter regions in the lumbar spinal cord. Intraperitoneal 5-AZA alleviated hyperalgesia in CCI and SNL rats, an effect accompanied by the reversed expression of hypermethylated genes. Thus, DNA methylation inhibition represents a promising epigenetic strategy for protection against chronic NPP following nerve injury. Our study lays a theoretical foundation for 5-AZA to become a clinical targeted drug.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Neuropathic pain (NPP) is the most common category of pathological pain [1] and is defined as pain brought on by a lesion or illness that affects the nervous system at either the peripheral or central level [2]. It is estimated that NPP has a prevalence of 6–10% in the general population, and it causes difficulties in daily activities and decreases productivity at work [3, 4], resulting in a poorer quality of life. However, current treatments for NPP are poorly effective because of our limited understanding of the molecular mechanisms involved. It has been established that central sensitization in the spinal cord plays a crucial role in the NPP condition [5, 6], and altered gene expression contributes critically to the central sensitization and the initiation and maintenance of chronic NPP [7,8,9]. Besides, long-term NPP in animals also led to alterations in gene expression across the genome, indicating that gene expression alteration in the spinal cord should be noticeable in response to chronic NPP [10,11,12]. Interestingly, therapies that alleviate hyperalgesia are associated with the reversal of NPP-induced regulation of pain-related mediators in the spinal cord [13,14,15].
Without changing the underlying genetic sequence, epigenetic regulation enables the steady control of gene expression in response to experiential and environmental inputs [16]. DNA methylation is an epigenetic process that can be involved in the long-term control of gene expression [17]. DNA methyltransferases (DNMTs), which transfer a methyl group from the methyl donor s-adenosyl methionine to cytosine bases in the DNA, catalyze the methylation of DNA [17]. Typically, DNA methylation in a gene's promoter region causes epigenetic inhibition of gene expression [18]. Numerous studies have shown that chronic pain is associated with changes in DNA methylation throughout cells and tissues involved in pain pathways, including the dorsal root ganglion (DRG) [19, 20], spinal cord [21, 22], and brain [23, 24]. Additionally, artificial enrichment by reducing DNA methylation levels in the spinal cord successfully attenuated pain [25, 26]. As DNMTs inhibitors and regulators, 5-azacytidine (5-AZA) and 5-aza-2′-deoxycytidine (5-AZA-dC) are used to treat a variety of pain-related or pain-comorbid disorders, such as cognitive impairment, depression, osteoarthritis, and cancer pain [27,28,29,30]. However, the specific genes and biological systems associated with effective treatment of NPP have not been well explored. Little is known about the methylated genes that 5-AZA targets in the spinal cord during chronic NPP. The discovery of these systems will shed light on the processes by which the epigenome is altered by chronic NPP and how the epigenome might be therapeutically addressed.
Our previous study found that abnormal DNA methylation is involved in chronic constriction injury (CCI)-induced NPP [21]. In this study, we use an Arraystar Rat RefSeq Promoter Array to analyze changes in DNA methylation at the genome-wide level in the spinal cord caused by peripheral nerve injury. We show that CCI induces widespread lumbar spinal cord methylation changes, and more than 1200 genes are differentially methylated after CCI. Furthermore, the specific hypermethylated genes are located mainly in long-term potentiation (LTP) and glutamatergic synapse pathways. Moreover, intraperitoneal injection of 5-AZA alleviates mechanical allodynia and thermal hyperalgesia and counteracts changes in hypermethylated gene mRNA expression induced by chronic NPP following CCI or spinal nerve ligation (SNL).
Materials and Methods
Animals
IN this study, male Sprague‒Dawley (SD) rats weighing 220–250 g were employed. All animals were kept in a setting with a temperature of 22 ± 1 °C, a relative humidity of 50 ± 1%, and a cycle of light and dark of 12 h. They had access to food and water at all times in their separate cages. All protocols were approved by the ethics committee of the Fujian Cancer Hospital and adhered to the Guide for the Care and Use of Laboratory Animals published by the US National Institutes of Health.
Experimental Paradigm
Behavioral testing was performed before surgery and at 3, 5, 7, 10, and 14 days after surgery in each rat. Under deep anesthesia with intraperitoneal injection (i.p.) of pentobarbital sodium (40 mg/kg), all the rats were killed and their ipsilateral lumbar spinal cords were immediately dissected 14 days after behavioral testing. The first six rats were randomly divided into two groups and subjected to CCI or sham surgery (n = 3 for each group). On day 14 after surgery, ipsilateral lumbar spinal cords in CCI or sham groups were dissected for methyl-DNA immunoprecipitation (MeDIP)-on-chip and microarray data analysis. Then, to verify the chip results and assess the methylation of target genes by MeDIP-qPCR, six rats were randomly divided into CCI and sham groups (n = 3), and their ipsilateral lumbar spinal cords were dissected on day 14 after surgery. Next, twelve rats were randomly divided into CCI and sham groups (n = 6), and their ipsilateral lumbar spinal cords were used to detect the mRNA levels of target genes. Furthermore, to study the effects of 5-AZA on CCI-induced NPP, eighteen rats were randomly divided into three groups (n = 6 for each group): the sham + NS group, CCI + NS group, and CCI + AZA group. CCI + AZA rats were treated with 4 mg/kg (i.p.) 5-AZA at a concentration of 1 mg/ml (Sigma, USA) once a day from days 3 to 14 after CCI. Meanwhile, rats in the sham + NS group and CCI + NS group received the same volume of 0.9% saline. Moreover, to demonstrate the effects of 5-AZA on SNL-induced NPP, eighteen rats were randomly divided into three groups to receive SNL surgery (n = 6 for each group): the Sham + NS group, SNL + NS group, and SNL + AZA group. From days 3 to 14, the SNL + 5-AZA group rats were treated with 4 mg/kg 5-AZA (1 mg/ml, i.p., Sigma, USA) once daily. Meanwhile, sham + NS and SNL + NS rats received the same volume of 0.9% saline. All intraperitoneal injections were administered between 9:00 am and 12:00 am. On day 14 after surgery, the ipsilateral lumbar spinal cords of the last thirty-six rats were dissected to detect the mRNA levels of target genes after behavioral testing.
NPP Model
Chronic Constriction Injury
The CCI procedure was performed as previously described [31]. Briefly, rats were anesthetized with pentobarbital sodium (40 mg/kg, i.p.). The trunk of the right sciatic nerve was freed before it branched. Next, 4–0 chromic catgut was used to mildly ligate the trunk of the nerve with appropriate strength to slightly compress the epineuria and cause a gentle tremor in the associated muscles. Four ligatures were made at intervals of 1 mm in the sciatic nerve trunk. Sham rats underwent the same surgical procedures without ligatures placement.
Spinal nerve ligation
The SNL procedure was performed on the right L5 spinal nerve as previously described [32]. In short, the dorsal vertebral column of L4–L6 was surgically exposed under pentobarbital sodium anesthesia (40 mg/kg, i.p.). At L4–L6, the paraspinal muscles were dissociated from the spinal processes, and the L5 transverse process was delicately removed. The right L5 spinal nerve was exposed and firmly ligated distal to the DRG using 4–0 silk sutures. In the sham-operated group, the same procedure was followed, except that the L5 spinal nerve was not ligated.
Behavioral Testing
The 2390 Electronic von Frey Anesthesiometer (IITC Life Science, USA) and the 37370 Plantar Test Instrument (Ugo Basile, Italy) were used to measure the mechanical withdrawal threshold (MWT) and thermal withdrawal latency (TWL) in all rats before and 3, 5, 7, 10, and 14 days after surgery, respectively, as previously described [25, 26]. Both of the two tests ended with paw removal followed by distinct flinching movements. The intensity was automatically recorded after paw withdrawal. At five-minute intervals, each rat was measured five times. The maximum and minimum values were subtracted before averaging the MWT and TWL. All behavioral measurements were performed between 2:00 pm and 5:00 pm by a researcher who was blinded to the group assignment.
Methyl-DNA Immunoprecipitation-on-Chip
Genomic DNA was extracted from lumbar spinal cords using a DNeasy Blood & Tissue Kit (Qiagen, Germany). Using a Bioruptor sonicator (Diagenode, Belgium), genomic DNA from each sample was sonicated to around 200–1000-bp fragments. One microgram of fragmented genomic DNA was immunoprecipitated with a mouse monoclonal anti-5-methylcytosine antibody (Diagenode, Belgium). MeDIP DNA was purified using Qiagen MinElute columns (Qiagen, Germany). The NimbleGen Dual-Color DNA Labeling Kit (NimbleGen Systems, USA) was used for DNA labeling. One microgram of DNA from each sample was incubated for 10 min at 98 °C with 1 OD of Cy5-9mer primer (IP sample) or Cy3-9mer primer (input sample). The Arraystar Rat RefSeq Promoter Array (Arraystar, USA) was hybridized with labeled DNA. Finally, an Agilent G2505C Scanner (Agilent Technologies, Germany) was used to wash and scan the arrays. The microarray data have been deposited in NCBI Gene Expression Omnibus (GEO) and are accessible through GEO Series accession number GSE219038.
Microarray Data Analysis
From the normalized log2-ratio data, a sliding-window (1500 bp) peak-finding algorithm provided by NimbleScan v2.5 (Roche-NimbleGen, Sweden) was applied to analyze the MeDIP-chip data. A one-sided Kolmogorov‒Smirnov (KS) test was used to determine whether the probes were drawn from a significantly more positive distribution of intensity log2-ratios than the rest of the array. The region was assigned to an enrichment peak (EP), if several adjacent probes were significantly increased above a set threshold. The NimbleScan sliding-window peak-finding algorithm was rerun to identify the differentially enriched peaks (DEPs). The log2-ratio value of all probes in the DEP region was used to make a cluster analysis to directly show the methylation of DEP probes in each sample. Genes containing DEPs in the promoter were categorized as differentially methylated genes (DMGs). Then, the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis was used to analyze these genes. The significance of the pathway correlation to the conditions was denoted by the p value with a cutoff of 0.05. The Gene Ontology project offers a controlled vocabulary to describe gene and gene product properties in any organism (http://www.geneontology.org). Three domains of biological process, cellular component, and molecular function are all covered by Gene Ontology terms. To determine whether there was a greater overlap between the DE list and the GO annotation list than would be predicted by chance, Fisher's exact test was utilized. The significance of GO term enrichment in the DE genes was denoted by the p value with a cutoff of 0.05.
Real-time PCR on MeDIP-Enriched DNA
The MeDIP assay combined with real-time PCR was used to quantitatively assess the methylation status of target genes to verify MeDIP-chip results. DNA extraction and MeDIP were performed as described above. We carried out real-time PCR with immunoprecipitated methylated DNA and input DNA by using the ViiA 7 Real-time PCR System (Applied Biosystems, USA), and 2 × PCR master mix (Arraystar, USA). The methylation levels of Grm4, Htr4, Adrb2, Kcnf1, Gad2, and Pparg were evaluated. The primers used for MeDIP-qPCR are shown in Table 1. 2−△△CT values were used, and the relative enrichment of methylation in each sample was calculated using the signal ratios of immunoprecipitated DNA to input DNA: %MeDIP/Input = 2 (CtInput − CtChIP) × Fd × 100%, Fold Enrichment = [% (MeDIP/Input)]/[% (Negative control/Input)].
Quantitative Reverse Transcription-PCR
The mRNA expression of the genes was determined by quantitative reverse transcription-PCR (RT-qPCR). TRIzol Reagent (Invitrogen, USA) was used to extract total RNA. The cDNA templates were synthesized by reverse transcription using the SuperScript™ III First-Strand Synthesis System for RT-PCR (Invitrogen, USA) and then saved at − 20 °C. The ViiA 7 Real-time PCR System (Applied Biosystems, USA) and 2 × PCR master mix (Arraystar, USA) were used to perform real-time PCR. GAPDH was used as an internal standard, and the 2−ΔΔCT method was used to determine the relative expression levels of the target genes. Primers were produced by Shanghai Lifei Biotechnology Co., Ltd., in China, and are listed in Table 2.
Statistical Analyses
The GraphPad Prism 5.0 software (GraphPad Software, CA) was used to conduct all statistical analyses. Data are presented as the mean ± SD. The unpaired Student’s t-test was used for comparisons of PCR data between two groups. Differences between sham, CCI, or SNL and 5-AZA treatment conditions were analyzed using one-way analysis of variance (ANOVA) followed by Tukey’s post hoc test for multiple comparisons. Behavioral test data were analyzed using two-way repeated-measurement ANOVA followed by Bonferroni’s post hoc test for multiple comparisons. P < 0.05 was considered significant.
Results
DNA Methylation Status in Response to CCI in the Lumbar Spinal Cord in Rats
MWT and TWL in rats were measured using behavioral tests. Our results showed that MWT and TWL significantly decreased in the CCI rats than in the sham rats from 3 to 14 days after surgery (all P < 0.001, Fig. 1A), indicating mechanical allodynia and thermal hyperalgesia in CCI rats were successfully established.
Differential methylation of promoter regions in the lumbar spinal cord of Sham and CCI rats. A Ipsilateral mechanical withdrawal threshold (MWT) and thermal withdrawal latency (TWL) were monitored to identify CCI-induced neuropathic pain (n = 3), values represented as mean ± SD, two-way repeated-measurement ANOVA followed by Bonferroni’s test. *P < 0.05 vs. the Sham group, **P < 0.01 vs. the Sham group, ***P < 0.001 vs. the Sham group. B Clustering of differentially enriched peaks (DEPs) of methylation in the DNA promoter region in the Sham group and CCI group (red: hypermethylation; green: hypomethylation). C Numbers of DEPs of methylation between groups. CCI_Sham represents the DEPs that were hypermethylated in the CCI group compared with the sham group, and Sham_CCI represents the DEPs that were hypermethylated in the sham group compared with the CCI group. D The distribution of the total number of DEPs of methylation in the DNA promoter regions enriched in the microarray analysis in each category. Note: HCP, high CpG-density promoter; ICP, intermediate CpG-density promoter; LCP, low CpG-density promoter
To study whether CCI led to abnormal methylation changes in DNA promoter regions at the genome-wide level, the Arraystar Rat RefSeq Promoter Arrays covering 15,987 gene promoter regions (between approximately − 1300 and + 500 bp of the TSSs) with ~ 180,000 probes separated by around 158 bp were used to detect differentially methylated regions in the DNAs that were extracted from the lumbar spinal cord of the CCI and sham rats on day 14 (Supplementary Table 1). The cluster diagram directly showed the methylation of DEP probes in each sample (Fig. 1B). There were 1205 differentially methylated fragments within DNA promoter regions, including 638 hypermethylated fragments and 567 hypomethylated fragments in CCI rats (Fig. 1C). And the hypermethylated fragments contained 292 (45.77%) high CpG-density promoters (HCP), 169 (26.49%) intermediate CpG-density promoters (ICP), and 177 (27.74%) low CpG-density promoters (LCP). While the hypomethylated fragments contained 267 (47.09%) HCP, 163 (28.75%) ICP, and 137 (24.16%) LCP (Fig. 1D).
Differentially Methylated Functional Domains and Signaling Pathways in CCI Rats
Gene Ontology (GO) analysis was used to identify ontology terms enriched among differentially methylated genes (DMGs) in the lumbar spinal cord of CCI rats. DMGs between hyper- and hypomethylated regions in response to peripheral nerve injury were identified. The top 10 most significant enrichment terms among hyper- and hypomethylated DMGs after CCI are shown in Fig. 2A and B. Organism process, developmental process, anatomical structure development, and ion transport were the top GO terms enriched among hypermethylated genes (Fig. 2A), while cellular component organization or biogenesis, establishment of localization, and transport were the top GO terms enriched among hypomethylated genes (Fig. 2B).
GO and KEGG pathway analysis of differentially methylated genes (DMGs) between the hyper- and hypo-methylation groups in the lumbar spinal cord of CCI rats. A, B Dot graphs of the top ten GO terms based on hypermethylated DMGs (A) and hypomethylated DMGs (B) profiling. The dot plot shows the gene ratio values of the top ten most significant enrichment terms. The gene ratio value equals (the number of DE genes associated with the listed GOID/the total number of DE genes). GO, Gene Ontology; BP, biological process; CC, cellular component; MF, molecular function; GOID, the ID of gene ontology term used in the Gene Ontology Project. C, D Enrichment analysis of KEGG pathways and the top ten activated pathways identified among hypermethylated DMGs (C) and hypomethylated DMGs (D) are shown. The bar plot shows the top ten Enrichment score (-log10(P value)) values of the significant enrichment pathways. The red lines in C and D depict the level of statistical significance (P = 0.01). KEGG, Kyoto Encyclopedia of Genes and Genomes
KEGG pathway analysis was used to identify the signaling pathways among DMGs. The top 10 most significant enrichment pathways among hyper- and hypomethylated DMGs after CCI are shown in Fig. 2C and D. The hypermethylated genes in the CCI group were mainly enriched in long-term potentiation (LTP), glutamatergic synapse, calcium signaling, and cAMP signaling pathways (Fig. 2C). While hypomethylated genes were mainly enriched in the cell cycle, peroxisome, oocyte meiosis, and gastric acid secretion pathways (Fig. 2D). The full list of GO terms and KEGG pathways enriched in the DMGs can be found in Supplementary Tables 2 and 3.
Screening and Verification of DMGs in CCI Rats
In accordance with the specified conditions, some hypermethylated DMGs were identified from the above microarray findings. Table 3 summarizes the six selected hypermethylated genes that predict the chronic NPP-related pathological signature in CCI rats, in order of PeakScore from high to low.
We further validated the methylation status of the six chronic NPP-related genes by MeDIP-qPCR. As shown in Fig. 3, we observed that the methylation levels of Grm4, Htr4, and Kcnf1 were significantly increased in the CCI group compared to the sham group (P < 0.05, n = 3). Meanwhile, CCI-induced gene (Adrb2, Gad2, and Pparg) methylation was increased in comparison with the sham group, albeit without statistical significance (n = 3). These results indicated that the methylation levels of various genes, including Grm4, Htr4, Kcnf1, Adrb2, Gad2, and Pparg, might be enhanced by peripheral nerve injury.
MeDIP-qPCR results of 6 target hypermethylated genes in CCI rats. A DNA methylation of Grm4. B DNA methylation of Htr4. C DNA methylation of Adrb2. D DNA methylation of Kcnf1. E DNA methylation of Gad2. F DNA methylation of Pparg. Note: n = 3; values represented as mean ± SD, unpaired Student’s t-test. *P < 0.05 vs. the sham group
Expression Level of Pain-Related Genes in the Lumbar Spinal Cord After CCI
In line with the alterations in DNA methylation, gene expression detection by RT-qPCR showed a decrease in the mRNA expression of Grm4, Htr4, Adrb2, Kcnf1, and Pparg in the CCI group compared with the sham group (P < 0.05, Fig. 4). And Gad2 mRNA expression was reduced without statistical significance. Based on these findings, peripheral nerve injury could alter the expression and methylation of Grm4, Htr4, Adrb2, Kcnf1, and Pparg genes in rats following the CCI procedure.
5-AZA Alleviated Hyperalgesia and Induced Pain-Related Gene Expression Changes in the Lumbar Spinal Cord After CCI
We examined the MWT and TWL after the intraperitoneal injection of 5-AZA in the CCI-induced NPP model. The MWT and TWL were recorded before CCI (baseline) and on days 3, 5, 7, 10, and 14 after CCI (Fig. 5A). The baseline MWT and TWL in each group showed no significant differences. Compared to those in the sham + NS group, the MWT and TWL were significantly reduced in the CCI + NS and CCI + AZA groups after CCI from day 3 (all P < 0.01, Fig. 5B), indicating the successful establishment of the CCI-induced NPP model. Mechanical allodynia and thermal hyperalgesia were partially improved on day 5 in the CCI + AZA group compared to the CCI + NS group, and this improvement was maintained throughout the study (all P < 0.01, Fig. 5B). These results indicated that the intraperitoneal injection of 5-AZA alleviates the mechanical allodynia and thermal hyperalgesia induced by CCI.
Intraperitoneal injection of 5-AZA alleviated CCI-induced neuropathic pain. A Timescale of the CCI experiment and behavioral test. “i.p.” intraperitoneal injection. B After 5-AZA treatment, mechanical withdrawal threshold (MWT) and thermal withdrawal latency (TWL) were monitored to identify CCI-induced neuropathic pain (n = 6). Data were shown as mean ± SD, two-way repeated-measurement ANOVA followed by Bonferroni’s test. C RT-qPCR of Grm4, Htr4, Adrb2, Kcnf1, Gad2, and Pparg (n = 6). Data were shown as mean ± SD, one-way ANOVA followed by Tukey's test. Note: *P < 0.05 vs. the sham + NS group, **P < 0.01 vs. the sham + NS group, ***P < 0.001 vs. the sham + NS group, #P < 0.05 vs. the CCI + NS group, ##P < 0.01 vs. the CCI + NS group, ###P < 0.001 vs. the CCI + NS group
Based on behavioral experiments, we explored the relationship between 5-AZA and pain-related gene expression using RT-qPCR analysis. All pain-related genes (Grm4, Htr4, Adrb2, Kcnf1, Gad2, and Pparg) were evaluated on day 14 after 5-AZA injection. The results showed that the mRNA expression of Grm4, Htr4, Adrb2, and Kcnf1 was significantly increased in the CCI + 5-AZA group compared to the CCI + NS group (P < 0.05, Fig. 5C).
5-AZA Alleviated Hyperalgesia and Induced Pain-Related Gene Expression Changes in the Lumbar Spinal Cord After SNL
We further examined the MWT and TWL after intraperitoneal injection of 5-AZA in the SNL-induced NPP model. MWT and TWL were recorded before SNL (baseline) and on days 3, 5, 7, 10, and 14 (Fig. 6A). The baselines of MWT and TWL in each group showed no significant differences. Compared to the sham + NS group, the MWT and TWL were significantly reduced in the SNL + NS and SNL + AZA groups from day 3 after SNL (all P < 0.05, Fig. 6B), indicating the successful establishment of the SNL-induced NPP model. Mechanical allodynia and thermal hyperalgesia were partially improved on day 5 in the SNL + AZA group compared to the SNL + NS group and maintained throughout the study (all P < 0.001, Fig. 6B). These results indicated that the intraperitoneal injection of 5-AZA alleviated the mechanical allodynia and thermal hyperalgesia induced by SNL.
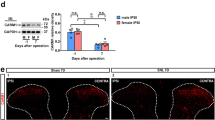
Intraperitoneal injection of 5-AZA alleviated SNL-induced neuropathic pain. A Timescale of SNL experiment and behavioral test, “i.p.” intraperitoneal injection. B After 5-AZA treatment, mechanical withdrawal threshold (MWT) and thermal withdrawal latency (TWL) were monitored to identify SNL-induced neuropathic pain (n = 6). Data were shown as mean ± SD, two-way repeated-measurement ANOVA followed by Bonferroni’s test. C RT-qPCR of Grm4, Htr4, Adrb2, Kcnf1, Gad2 and Pparg (n = 5–6). Data were shown as mean ± SD, one-way ANOVA followed by Tukey’s test. Note:*P < 0.05 vs. the sham + NS group, **P < 0.01 vs. the sham + NS group, ***P < 0.001 vs. the sham + NS group, #P < 0.05 vs. the SNL + NS group, ##P < 0.01 vs. the SNL + NS group, ###P < 0.001 vs. the SNL + NS group
Based on behavioral experiments, we explored the relationship between 5-AZA and pain-related gene expression by RT-qPCR analysis. All pain-related genes (Grm4, Htr4, Adrb2, Kcnf1, Gad2, and Pparg) were evaluated on day 14 after 5-AZA injection. The results showed that the mRNA expression of Grm4, Htr4, Adrb2, Kcnf1, and Gad2 was significantly increased in the SNL + 5-AZA group compared to the SNL + NS group (P < 0.05, Fig. 6C).
Discussion
Chronic NPP is associated with multiple dimensions, including nociception, emotional distress, and cognitive dysfunction. Epigenetic mechanisms are implicated in these conditions. Our results showed that NPP altered DNA methylation profiles and induced a decrease in the mRNA expression of some pain-related hypermethylated genes. Methylation inhibition improved nociception associated with NPP and reversed the decrease in the hypermethylated gene mRNA expression followed by nerve injury.
Over the past few decades, many animal models for nerve injury-induced NPP have been developed. CCI is a widely used model, especially for behavioral studies. CCI simulates post-traumatic peripheral painful neuropathic states in humans and is reliable and simple to create [33]. However, the degree of damage stability in CCI is lower compared with some other models. SNL possesses less experimental variability and is regarded as an excellent model for mechanical allodynia assessment [33]. But the surgical procedure of the SNL model is more challenging, and animals suffer more trauma. Here, CCI and SNL models were both employed in the present study to ensure the reliability of the research.
DNA methylation is critical in pain memory and synaptic plasticity [34]. Previous studies reported abnormal changes in global DNA methylation and the methylation of specific promoters in rodent lumbar spinal cords 1–21 days following CCI/SNL surgery [22, 25, 35]. However, the methylation profiles of the DNA promoter region in the lumbar spinal cord in nerve injury-induced NPP rats have not been well explored. Here, we used the Arraystar Rat DNA Promoter Array, a reliable and efficient platform [36], to identify the methylation profiles at the genome-wide level in the DNA promoter regions of the lumbar spinal cord in rats 14 days following CCI surgery. In this study, more than 1200 fragments were differentially methylated after CCI treatment, including 638 hypermethylated fragments and 567 hypomethylated fragments, which indicated that CCI could be capable of inducing widespread lumbar spinal cord methylation changes.
After identifying differentially methylated genes associated with CCI-induced NPP, GO analysis and KEGG pathway analysis were used to reveal genome-wide methylation patterns. The top GO terms among hypomethylated genes were cellular component organization or biogenesis, establishment of localization, and transport, while the top KEGG pathways were cell cycle, peroxisome, oocyte meiosis, and gastric acid secretion, which seem to be little related to pain. Although studies reported that CCI and SNL induced demethylation and increased expression of receptor tyrosine kinase-like orphan receptor 2 (ROR2) [37] and chemokine receptor CXCR3 [38], the two genes were not observed in our chip results. Besides, our previous study showed that the overall DNA methylation level in the spinal cord increased significantly in CCI rats [25]. Here, we focused on genes whose methylation increased and whose encoded mRNAs were expressed at a lower level in CCI rats.
In the present study, the top GO terms among hypermethylated genes were organism process, developmental process, anatomical structure development, and ion transport, while the top KEGG pathways were long-term potentiation (LTP), glutamatergic synapse, calcium signaling, and cAMP signaling pathways. Among the genes altered by NPP in the promoter region, Grm4, Htr4, Adrb2, Kcnf1, Gad2, and Pparg belong to development processes and ion transport and are involved in LTP and glutamatergic synapse pathways [39,40,41,42,43]. Previous studies have shown that DNA methylation changes in certain genes (such as Htr4, Gad2, Adrb2, and Pparg) could interfere with the expression of their encoded mRNA in some pathological states [30, 44,45,46,47]. Here, our results showed that Grm4, Htr4, and Kcnf1 were significantly hypermethylated and that Adrb2, Gad2, and Pparg were insignificantly hypermethylated in CCI rats. Correspondingly, the mRNA levels of these genes decreased to various degrees, suggesting the negative regulation of DNA methylation in its encoded targets under NPP conditions.
Aberrant DNA methylation in the nervous system is involved in pain conditions. Numerous studies have demonstrated the possibility of attenuating chronic pain by modulating DNA methylation [22, 25, 48, 49]. For example, restoring DNA methylation through the methyl donor s-adenosyl methionine improved cognitive and emotional functions damaged by spared nerve injury [50]. Increased DNMT3a in the DRG led to undesirable opioid analgesic effects, and blocking this increase restored the analgesic effects of morphine or loperamide and attenuated the development of their analgesic tolerance under NPP [51]. Moreover, intrathecal administration of 5-AZA or 5-AZA-dC reversed methylation levels corresponding to the amelioration of pain [25, 26]. The US Food and Drug Administration (FDA) has approved 5-AZA as a chemotherapy agent in the treatment of malignant tumors and myelodysplastic syndromes [52]. However, the operation of intrathecal administration of 5-AZA is complex, so its clinical application is limited. A previous study showed that intraperitoneal administration of 5-AZA-dC provided relief from incisional nociceptive sensitization [49]. Thus, we examined the effect of 5-AZA on CCI/SNL-induced NPP via intraperitoneal injection for the first time.
An increase in DNA methylation in response to nerve injury is well established, and inhibiting this increase resulting in pain relief has been reported. Sun et al. found that DNMT1 expression was upregulated after peripheral nerve injury and that blocking DNMT1 could reverse the abnormal excitability and sensitivity in injured DRG neurons [53]. Liu et al. revealed that blocking the upregulation of DNMTs with RG108 alleviated spared nerve injury-induced pain [54]. Furthermore, nerve injury or incision-related hyperalgesia can be attenuated by 5-AZA-dC, which also induces α5-GABAA receptor or Oprm1 reversion [49, 55]. DNA demethylation in the DRG was proven to alleviate SNL-induced pain hypersensitivities coinciding with rescued expression of μ-opioid receptors and the potassium voltage-gated channel Kv1.2 [56]. Moreover, 5-AZA was reported to reverse the hypermethylation and upregulate the expression of Adrb2 in PM2.5-induced cardiomyocyte apoptosis [47]. Consistent with these studies, our results showed that 5-AZA treatment obviously attenuated CCI/SNL-induced mechanical allodynia and thermal hyperalgesia, coinciding with reversed mRNA expression of the hypermethylated genes; Grm4, Htr4, Adrb2, and Kcnf1 expression was observed in both CCI and SNL rats, and Gad2 expression was observed in SNL rats. We infer that the modulation of DNA methylation in these genes in the spinal cord through 5-AZA treatment might be a promising epigenetic mechanism and therapeutic target for the relief of chronic NPP.
Specifically, Grm4 encodes metabotropic glutamate receptor 4 (mGluR4), which contributes to pain modulation by suppressing glutamatergic hyperactivation [57, 58]; its exogenous activation reversed the sensitivity imbalances of chronic NPP in the dorsal spinal region [59, 60]. Adrb2 encodes the β2-adrenergic receptor (β2-AR) [61], which is widely expressed in the normal and injured central nervous system [41]. Adrb2 activation could alleviate multiple chronic NPPs [62, 63]. Consistent with these studies, our results showed that intraperitoneal 5-AZA reversed Grm4 and Adrb2 mRNA expression, coinciding with attenuated hyperalgesia in both CCI and SNL rats. These results further demonstrate the pivotal role of Grm4 and Adrb2 in NPP. Htr4 encodes 5-hydroxytryptamine receptor 4 (5-HT4R), which has been found to produce antinociceptive or pronociceptive effects in the spinal cord [64,65,66]. We observed that Htr4 mRNA levels recovered after 5-AZA treatment in parallel with pain relief, which indicates that Htr4 may produce antinociceptive effects as an inhibitory receptor subtype of 5-HT in the lumbar spinal cord after nerve injury. Moreover, KCNF1 is a potassium channel gene and encodes Kv5.1, which can form a functional complex with Kv2.1, resulting in physiological effects [67, 68]. The contribution of the Kv5.1 channel to the regulation of NPP has not been investigated, and our results first showed that the decrease in Kv5.1 mRNA was reversed after 5-AZA treatment along with pain relief in CCI/SNL rats, suggesting that Kv5.1 may be involved in the occurrence and development of NPP. Finally, although Gad2 and Pparg have been reported to be downregulated in CCI rats [69, 70], we observed the reduction of Gad2 only after SNL, and that of Pparg only after CCI. More experiments are needed to further investigate the correlations of Gad2 and Pparg with NPP. It is crucial to note that the correlation of Grm4 and Kcnf1 between DNA methylation and NPP was reported here for the first time and might become a new promising direction for pain treatment.
There are several limitations in this study. First, due to the small sample size of 3 to 6 animals, these results should be considered exploratory. Second, DNA methylation assessment was performed at only one time point and therefore cannot depict the temporal dynamics of DNA methylation in response to pain during the acute-to-chronic process. Third, the impact of 5-AZA on the target gene methylation level was not examined. As stated above, chronic systemic administration of 5-AZA relieves NPP along with the restoration of hypermethylated gene expression. Future studies incorporating the modulation of target gene methylation and protein levels are needed to characterize the explicit change during NPP. Finally, the phenomena described here are correlative; further investigation is needed to explore the causal and detailed relationships between spinal cord methylation and chronic NPP. Despite these limitations, these results provide an important insight into the DNA methylation of novel spinal cord target genes associated with the successful treatment of chronic NPP.
Conclusions
In summary, our findings reveal the DNA methylation profiles at the genome-wide level in the DNA promoter regions of the spinal cord in CCI rats and offer important evidence for the reversal of NPP and this decrease in target gene expression following nerve injury by employing an epigenetic regulator to enhance DNA methylation in CCI and SNL rats. The hypermethylation occurred mainly in genes which are associated with development, structure, and ion transport and are involved in LTP and glutamatergic synapse pathways. The methylation and mRNA levels of these target genes (Grm4, Htr4, Adrb2, Kcnf1, Gad2, and Pparg) may contribute to NPP persistence or recovery in the spinal cord after therapeutic interventions with intraperitoneal 5-AZA. These findings provide crucial insight into the relationship and call for increased exploration between chronic NPP and epigenetic interventions, such as intraperitoneal 5-AZA, for the treatment of chronic NPP.
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Gilron I, Baron R, Jensen T (2015) Neuropathic pain: principles of diagnosis and treatment. Mayo Clin Proc 90:532–545. https://doi.org/10.1016/j.mayocp.2015.01.018
Ralf Baron AB, Wasner G (2010) Neuropathic pain: diagnosis, pathophysiological mechanisms, and treatment. Lancet Neurol 9:807–819. https://doi.org/10.1016/S1474-4422(10)70143-5
Bouhassira D (2019) Neuropathic pain: definition, assessment and epidemiology. Revue Neurologique 175:16–25. https://doi.org/10.1016/j.neurol.2018.09.016
Bushnell MC, Ceko M, Low LA (2013) Cognitive and emotional control of pain and its disruption in chronic pain. Nat Rev Neurosci 14:502–511. https://doi.org/10.1038/nrn3516
Tsuda M, Koga K, Chen T, Zhuo M (2017) Neuronal and microglial mechanisms for neuropathic pain in the spinal dorsal horn and anterior cingulate cortex. J Neurochem 141:486–498. https://doi.org/10.1111/jnc.14001
West SJ, Bannister K, Dickenson AH, Bennett DL (2015) Circuitry and plasticity of the dorsal horn–toward a better understanding of neuropathic pain. Neuroscience 300:254–275. https://doi.org/10.1016/j.neuroscience.2015.05.020
Liu Y, Zhou L-J, Wang J, Li D, Ren W-J, Peng J, Wei X, Xu T et al (2017) TNF-α differentially regulates synaptic plasticity in the hippocampus and spinal cord by microglia-dependent mechanisms after peripheral nerve injury. J Neurosci 37:871–881. https://doi.org/10.1523/jneurosci.2235-16.2017
Woolf CJ (2011) Central sensitization: implications for the diagnosis and treatment of pain. Pain 152:S2–S15. https://doi.org/10.1016/j.pain.2010.09.030
Voscopoulos C, Lema M (2010) When does acute pain become chronic? Br J Anaesth 105(Suppl 1):i69-85. https://doi.org/10.1093/bja/aeq323
Parisien M, Samoshkin A, Tansley SN, Piltonen MH, Martin LJ, El-Hachem N, Dagostino C, Allegri M et al (2019) Genetic pathway analysis reveals a major role for extracellular matrix organization in inflammatory and neuropathic pain. Pain 160:932–944. https://doi.org/10.1097/j.pain.0000000000001471
Hozumi T, Sawai S, Jitsuishi T, Kitajo K, Inage K, Eguchi Y, Shiga Y, Narita M et al (2021) Gene expression profiling of the spinal cord at the chronic pain phase identified CDKL5 as a candidate gene for neural remodeling. Neurosci Lett 749:135772. https://doi.org/10.1016/j.neulet.2021.135772
Du H, Liu Z, Tan X, Ma Y, Gong Q (2019) Identification of the genome-wide expression patterns of long non-coding RNAs and mRNAs in mice with streptozotocin-induced diabetic neuropathic pain. Neuroscience 402:90–103. https://doi.org/10.1016/j.neuroscience.2018.12.040
Ghosh K, Pan HL (2022) Epigenetic mechanisms of neural plasticity in chronic neuropathic pain. ACS Chem Neurosci 13:432–441. https://doi.org/10.1021/acschemneuro.1c00841
Hsieh MC, Ho YC, Lai CY, Chou D, Wang HH, Chen GD, Lin TB, Peng HY (2017) Melatonin impedes Tet1-dependent mGluR5 promoter demethylation to relieve pain. J Pineal Res 63. https://doi.org/10.1111/jpi.12436
Hsieh MC, Lai CY, Ho YC, Wang HH, Cheng JK, Chau YP, Peng HY (2016) Tet1-dependent epigenetic modification of BDNF expression in dorsal horn neurons mediates neuropathic pain in rats. Sci Rep 6:37411. https://doi.org/10.1038/srep37411
Topper MJ, Vaz M, Marrone KA, Brahmer JR, Baylin SB (2020) The emerging role of epigenetic therapeutics in immuno-oncology. Nat Rev Clin Oncol 17:75–90. https://doi.org/10.1038/s41571-019-0266-5
Morgan AE, Davies TJ, Mc Auley MT (2018) The role of DNA methylation in ageing and cancer. Proc Nutr Soc 77:412–422. https://doi.org/10.1017/S0029665118000150
Perri F, Longo F, Giuliano M, Sabbatino F, Favia G, Ionna F, Addeo R, Della Vittoria Scarpati G et al (2017) Epigenetic control of gene expression: potential implications for cancer treatment. Crit Rev Oncol Hematol 111:166–172. https://doi.org/10.1016/j.critrevonc.2017.01.020
Garriga J, Laumet G, Chen SR, Zhang Y, Madzo J, Issa JJ, Pan HL, Jelinek J (2018) Nerve injury-induced chronic pain is associated with persistent DNA methylation reprogramming in dorsal root ganglion. J Neurosci 38:6090–6101. https://doi.org/10.1523/JNEUROSCI.2616-17.2018
Pollema-Mays SL, Centeno MV, Apkarian AV, Martina M (2014) Expression of DNA methyltransferases in adult dorsal root ganglia is cell-type specific and up regulated in a rodent model of neuropathic pain. Front Cell Neurosci 8:217. https://doi.org/10.3389/fncel.2014.00217
Wang Y, Lin ZP, Zheng HZ, Zhang S, Zhang ZL, Chen Y, You YS, Yang MH (2016) Abnormal DNA methylation in the lumbar spinal cord following chronic constriction injury in rats. Neurosci Lett 610:1–5. https://doi.org/10.1016/j.neulet.2015.10.048
Shao C, Gao Y, Jin D, Xu X, Tan S, Yu H, Zhao Q, Zhao L et al (2017) DNMT3a methylation in neuropathic pain. J Pain Res 10:2253–2262. https://doi.org/10.2147/JPR.S130654
Gregoire S, Millecamps M, Naso L, Do Carmo S, Cuello AC, Szyf M, Stone LS (2017) Therapeutic benefits of the methyl donor S-adenosylmethionine on nerve injury-induced mechanical hypersensitivity and cognitive impairment in mice. Pain 158:802–810. https://doi.org/10.1097/j.pain.0000000000000811
Topham L, Gregoire S, Kang H, Salmon-Divon M, Lax E, Millecamps M, Szyf M, Stone LS (2020) The transition from acute to chronic pain: dynamic epigenetic reprogramming of the mouse prefrontal cortex up to 1 year after nerve injury. Pain 161:2394–2409. https://doi.org/10.1097/j.pain.0000000000001917
Wang Y, Liu C, Guo QL, Yan JQ, Zhu XY, Huang CS, Zou WY (2011) Intrathecal 5-azacytidine inhibits global DNA methylation and methyl- CpG-binding protein 2 expression and alleviates neuropathic pain in rats following chronic constriction injury. Brain Res 1418:64–69. https://doi.org/10.1016/j.brainres.2011.08.040
Miao XR, Fan LC, Wu S, Mao Q, Li Z, Lutz B, Xu JT, Lu Z et al (2017) DNMT3a contributes to the development and maintenance of bone cancer pain by silencing Kv1.2 expression in spinal cord dorsal horn. Mol Pain 13:1744806917740681. https://doi.org/10.1177/1744806917740681
Xing B, Liu P, Xu WJ, Xu FY, Dang YH (2014) Effect of microinjecting of 5-aza-2-deoxycytidine into ventrolateral orbital cortex on depressive-like behavior in rats. Neurosci Lett 574:11–14. https://doi.org/10.1016/j.neulet.2014.04.050
Yu CC, He C, Du YJ, Gao S, Lin YF, Wang SQ, Wang L, Wang J et al (2021) Preventive electroacupuncture reduces cognitive deficits in a rat model of D-galactose-induced aging. Neural Regen Res 16:916–923. https://doi.org/10.4103/1673-5374.297090
Appel CK, Scheff NN, Viet CT, Schmidt BL, Heegaard AM (2019) Decitabine attenuates nociceptive behavior in a murine model of bone cancer pain. Pain 160:619–631. https://doi.org/10.1097/j.pain.0000000000001442
Zhu X, Chen F, Lu K, Wei A, Jiang Q, Cao W (2019) PPARgamma preservation via promoter demethylation alleviates osteoarthritis in mice. Ann Rheum Dis 78:1420–1429. https://doi.org/10.1136/annrheumdis-2018-214940
Grace PM, Strand KA, Galer EL, Urban DJ, Wang X, Baratta MV, Fabisiak TJ, Anderson ND et al (2016) Morphine paradoxically prolongs neuropathic pain in rats by amplifying spinal NLRP3 inflammasome activation. Proc Natl Acad Sci U S A 113:E3441-3450. https://doi.org/10.1073/pnas.1602070113
Wang D, Chen T, Gao Y, Quirion R, Hong Y (2012) Inhibition of SNL-induced upregulation of CGRP and NPY in the spinal cord and dorsal root ganglia by the 5-HT(2A) receptor antagonist ketanserin in rats. Pharmacol Biochem Behav 101:379–386. https://doi.org/10.1016/j.pbb.2012.02.004
Challa SR (2015) Surgical animal models of neuropathic pain: Pros and Cons. Int J Neurosci 125:170–174. https://doi.org/10.3109/00207454.2014.922559
Moore LD, Le T, Fan G (2013) DNA methylation and its basic function. Neuropsychopharmacology 38:23–38. https://doi.org/10.1038/npp.2012.112
Zhao JY, Liang L, Gu X, Li Z, Wu S, Sun L, Atianjoh FE, Feng J et al (2017) DNA methyltransferase DNMT3a contributes to neuropathic pain by repressing Kcna2 in primary afferent neurons. Nat Commun 8:14712. https://doi.org/10.1038/ncomms14712
Huang CZ, Xu JH, Zhong W, Xia ZS, Wang SY, Cheng D, Li JY, Wu TF et al (2017) Sox9 transcriptionally regulates Wnt signaling in intestinal epithelial stem cells in hypomethylated crypts in the diabetic state. Stem Cell Res Ther 8:60. https://doi.org/10.1186/s13287-017-0507-4
Zhou XL, Zhang CJ, Peng YN, Wang Y, Xu HJ, Liu CM (2019) ROR2 modulates neuropathic pain via phosphorylation of NMDA receptor subunit GluN2B in rats. Br J Anaesth 123:e239–e248. https://doi.org/10.1016/j.bja.2018.08.025
Jiang BC, He LN, Wu XB, Shi H, Zhang WW, Zhang ZJ, Cao DL, Li CH et al (2017) Promoted interaction of C/EBPalpha with demethylated Cxcr3 gene promoter contributes to neuropathic pain in mice. J Neurosci 37:685–700. https://doi.org/10.1523/JNEUROSCI.2262-16.2016
Boccella S, Marabese I, Guida F, Luongo L, Maione S, Palazzo E (2020) The modulation of pain by metabotropic glutamate receptors 7 and 8 in the dorsal striatum. Curr Neuropharmacol 18:34–50. https://doi.org/10.2174/1570159X17666190618121859
McKinnell RM, Armstrong SR, Beattie DT, Choi SK, Fatheree PR, Gendron RA, Goldblum A, Humphrey PP et al (2009) A multivalent approach to the design and discovery of orally efficacious 5-HT4 receptor agonists. J Med Chem 52:5330–5343. https://doi.org/10.1021/jm900881j
Lemmens S, Kusters L, Bronckaers A, Geurts N, Hendrix S (2017) The beta2-adrenoceptor agonist terbutaline stimulates angiogenesis via Akt and ERK signaling. J Cell Physiol 232:298–308. https://doi.org/10.1002/jcp.25483
Smith PA (2020) K+ channels in primary afferents and their role in nerve injury-induced pain. Front Cell Neurosci 14. https://doi.org/10.3389/fncel.2020.566418
Trindade da Silva CA, Clemente-Napimoga JT, Abdalla HB, Basting RT, Napimoga MH (2022) Peroxisome proliferator-activated receptor-gamma (PPARgamma) and its immunomodulation function: current understanding and future therapeutic implications. Expert Rev Clin Pharmacol 15:295–303. https://doi.org/10.1080/17512433.2022.2071697
Hu Z, Ying X, Huang L, Zhao Y, Zhou D, Liu J, Zhong J, Huang T et al (2020) Association of human serotonin receptor 4 promoter methylation with autism spectrum disorder. Medicine 99. https://doi.org/10.1097/md.0000000000018838
Zanni G, Goto S, Fragopoulou AF, Gaudenzi G, Naidoo V, Di Martino E, Levy G, Dominguez CA et al (2021) Lithium treatment reverses irradiation-induced changes in rodent neural progenitors and rescues cognition. Mol Psychiatry 26:322–340. https://doi.org/10.1038/s41380-019-0584-0
Labouesse MA, Dong E, Grayson DR, Guidotti A, Meyer U (2015) Maternal immune activation induces GAD1 and GAD2 promoter remodeling in the offspring prefrontal cortex. Epigenetics 10:1143–1155. https://doi.org/10.1080/15592294.2015.1114202
Yang X, Zhao T, Feng L, Shi Y, Jiang J, Liang S, Sun B, Xu Q et al (2019) PM2.5-induced ADRB2 hypermethylation contributed to cardiac dysfunction through cardiomyocytes apoptosis via PI3K/Akt pathway. Environ Int 127:601–614. https://doi.org/10.1016/j.envint.2019.03.057
Liu L, Xu D, Wang T, Zhang Y, Yang X, Wang X, Tang Y (2020) Epigenetic reduction of miR-214-3p upregulates astrocytic colony-stimulating factor-1 and contributes to neuropathic pain induced by nerve injury. Pain 161:96–108. https://doi.org/10.1097/j.pain.0000000000001681
Taylor B, Sun Y, Sahbaie P, Liang D, Li W, Shi X, Kingery P, Clark JD (2015) DNA methylation modulates nociceptive sensitization after incision. Plos One 10. https://doi.org/10.1371/journal.pone.0142046
Topham L, Gregoire S, Kang H, Salmon-Divon M, Lax E, Millecamps M, Szyf M, Stone L (2021) The methyl donor S-adenosyl methionine reverses the DNA methylation signature of chronic neuropathic pain in mouse frontal cortex. Pain Rep 6:e944. https://doi.org/10.1097/PR9.0000000000000944
Sun L, Zhao JY, Gu X, Liang L, Wu S, Mo K, Feng J, Guo W et al (2017) Nerve injury-induced epigenetic silencing of opioid receptors controlled by DNMT3a in primary afferent neurons. Pain 158:1153–1165. https://doi.org/10.1097/j.pain.0000000000000894
Buckstein R, Yee K, Wells RA, Canadian Consortium on Evidence-based Care in MDS (2011) 5-Azacytidine in myelodysplastic syndromes: a clinical practice guideline. Cancer Treat Rev 37:160–167. https://doi.org/10.1016/j.ctrv.2010.05.006
Sun L, Gu X, Pan Z, Guo X, Liu J, Atianjoh FE, Wu S, Mo K et al (2019) Contribution of DNMT1 to neuropathic pain genesis partially through epigenetically repressing Kcna2 in primary afferent neurons. J Neurosci 39:6595–6607. https://doi.org/10.1523/JNEUROSCI.0695-19.2019
Liu R, Wu XM, He X, Wang RZ, Yin XY, Zhou F, Ji MH, Shen JC (2021) Contribution of DNA methyltransferases to spared nerve injury induced depression partially through epigenetically repressing Bdnf in hippocampus: Reversal by ketamine. Pharmacol Biochem Behav 200:173079. https://doi.org/10.1016/j.pbb.2020.173079
Franco-Enzastiga U, Garcia G, Murbartian J, Gonzalez-Barrios R, Salinas-Abarca AB, Sanchez-Hernandez B, Tavares-Ferreira D, Herrera LA et al (2021) Sex-dependent pronociceptive role of spinal alpha5 -GABAA receptor and its epigenetic regulation in neuropathic rodents. J Neurochem 156:897–916. https://doi.org/10.1111/jnc.15140
Wu Q, Wei G, Ji F, Jia S, Wu S, Guo X, He L, Pan Z et al (2019) TET1 overexpression mitigates neuropathic pain through rescuing the expression of mu-opioid receptor and Kv1.2 in the primary sensory neurons. Neurotherapeutics 16:491–504. https://doi.org/10.1007/s13311-018-00689-x
Zhang HM, Chen SR, Pan HL (2009) Effects of activation of group III metabotropic glutamate receptors on spinal synaptic transmission in a rat model of neuropathic pain. Neuroscience 158:875–884. https://doi.org/10.1016/j.neuroscience.2008.10.042
Vilar B, Busserolles J, Ling B, Laffray S, Ulmann L, Malhaire F, Chapuy E, Aissouni Y et al (2013) Alleviating pain hypersensitivity through activation of type 4 metabotropic glutamate receptor. J Neurosci 33:18951–18965. https://doi.org/10.1523/JNEUROSCI.1221-13.2013
Goudet C, Chapuy E, Alloui A, Acher F, Pin JP, Eschalier A (2008) Group III metabotropic glutamate receptors inhibit hyperalgesia in animal models of inflammation and neuropathic pain. Pain 137:112–124. https://doi.org/10.1016/j.pain.2007.08.020
Wang H, Jiang W, Yang R, Li Y (2011) Spinal metabotropic glutamate receptor 4 is involved in neuropathic pain. NeuroRep 22:244–248. https://doi.org/10.1097/WNR.0b013e3283453843
Nicholson R, Dixon AK, Spanswick D, Lee K (2005) Noradrenergic receptor mRNA expression in adult rat superficial dorsal horn and dorsal root ganglion neurons. Neurosci Lett 380:316–321. https://doi.org/10.1016/j.neulet.2005.01.079
Bohren Y, Tessier LH, Megat S, Petitjean H, Hugel S, Daniel D, Kremer M, Fournel S et al (2013) Antidepressants suppress neuropathic pain by a peripheral beta2-adrenoceptor mediated anti-TNFalpha mechanism. Neurobiol Dis 60:39–50. https://doi.org/10.1016/j.nbd.2013.08.012
Zhang FF, Morioka N, Abe H, Fujii S, Miyauchi K, Nakamura Y, Hisaoka-Nakashima K, Nakata Y (2016) Stimulation of spinal dorsal horn beta2-adrenergic receptor ameliorates neuropathic mechanical hypersensitivity through a reduction of phosphorylation of microglial p38 MAP kinase and astrocytic c-jun N-terminal kinase. Neurochem Int 101:144–155. https://doi.org/10.1016/j.neuint.2016.11.004
Liu QQ, Yao XX, Gao SH, Li R, Li BJ, Yang W, Cui RJ (2020) Role of 5-HT receptors in neuropathic pain: potential therapeutic implications. Pharmacol Res 159:104949. https://doi.org/10.1016/j.phrs.2020.104949
Song Z, Meyerson BA, Linderoth B (2011) Spinal 5-HT receptors that contribute to the pain-relieving effects of spinal cord stimulation in a rat model of neuropathy. Pain 152:1666–1673. https://doi.org/10.1016/j.pain.2011.03.012
Pineda-Farias JB, Barragan-Iglesias P, Valdivieso-Sanchez A, Rodriguez-Silverio J, Flores-Murrieta FJ, Granados-Soto V, Rocha-Gonzalez HI (2017) Spinal 5-HT4 and 5-HT6 receptors contribute to the maintenance of neuropathic pain in rats. Pharmacol Rep 69:916–923. https://doi.org/10.1016/j.pharep.2017.04.001
Bocksteins E (2016) Kv5, Kv6, Kv8, and Kv9 subunits: no simple silent bystanders. J Gen Physiol 147:105–125. https://doi.org/10.1085/jgp.201511507
Stas JI, Bocksteins E, Labro AJ, Snyders DJ (2015) Modulation of Closed-State Inactivation in Kv2.1/Kv6.4 Heterotetramers as Mechanism for 4-AP Induced Potentiation. PLoS One 10:1349. https://doi.org/10.1371/journal.pone.0141349
Jiang P, Jiang Q, Yan Y, Hou Z, Luo D (2021) Propofol ameliorates neuropathic pain and neuroinflammation through PPAR gamma up-regulation to block Wnt/beta-catenin pathway. Neurol Res 43:71–77. https://doi.org/10.1080/01616412.2020.1823107
Farzad B, Rajabi H, Gharakhanlou R, Allison DJ, Hayat P, Jameie SB (2018) Swimming training attenuates allodynia and hyperalgesia induced by peripheral nerve injury in an adult male rat neuropathic model: effects on irisin and GAD65. Pain Med 19:2236–2245. https://doi.org/10.1093/pm/pnx294
Acknowledgements
This study was supported by the Natural Science Foundation of Fujian Province of China (No. 2021J01434), the National Natural Science Foundation of China (No. 81400921), and the Natural Science Foundation of Guangdong Province of China (No. 2018A030313781).
Funding
This study was supported by the Natural Science Foundation of Fujian Province of China (No. 2021J01434), the National Natural Science Foundation of China (No. 81400921), and the Natural Science Foundation of Guangdong Province of China (No. 2018A030313781).
Author information
Authors and Affiliations
Contributions
Ying Wang and Xuan Li designed the study, collated the data, carried out data analyses, and produced the initial draft of the manuscript. DeZhao Liu and ZhiSen Dai provided critical advice and contributed to drafting the manuscript. YiSheng You, Yan Chen, ChenXing Lei, and YouYou Lv performed the experiments. All authors have read and approved the final submitted manuscript.
Corresponding author
Ethics declarations
Ethics Approval
All protocols were approved by the ethics committee of the Fujian Cancer Hospital and adhered to the Guide for the Care and Use of Laboratory Animals published by the US National Institutes of Health.
Consent to Participate
Not applicable.
Consent for Publication
All authors have read and approved the submission of the manuscript.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, X., Liu, D., Dai, Z. et al. Intraperitoneal 5-Azacytidine Alleviates Nerve Injury-Induced Pain in Rats by Modulating DNA Methylation. Mol Neurobiol 60, 2186–2199 (2023). https://doi.org/10.1007/s12035-022-03196-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-022-03196-5