Abstract
Nonamyloidogenic processing of amyloid precursor protein (APP) by augmenting ADAM10 is a promising therapeutic strategy for Alzheimer’s disease (AD). Therefore identification of molecular pathways that regulate ADAM10 expression is crucial. Autophagy is strongly dysregulated in AD, and TFEB was recently shown to be a master regulator of autophagy-lysosome pathway (ALP). Here, we report that TFEB expression in HeLa cells increased ADAM10 mature form by 72% (p < 0.01, n = 4), while TFEB knockdown by CRISPR strategy reduced ADAM10 mature form by 36% (p < 0.05, n = 4). Autophagy inhibition by 3-methyladenine (3-MA), but not bafilomycin A1 (BAF1), reduced ADAM10 mature form by 49% (p < 0.05, n = 4) in the TFEB expressing HeLa cells. Autophagy activation by 3 h of starvation increased ADAM10 to 91% (p < 0.001, n = 6) relative to 51% (p < 0.01, n = 6) in the nutrient-fed cells. Further, siRNAs targeted against PPARα in HeLa cells decreased ADAM10 levels by 28% (p < 0.05, n = 6) relative to the cells treated with scrambled siRNAs. Further, incubation of EGFP-TFEB expressing HeLa cells with PPARα antagonist, but not PPARβ or PPARγ antagonists, prevented TFEB-induced increase in ADAM10 levels. Importantly, flag-TFEB expression in the brain also increased ADAM10 by 60% (p < 0.05, n = 3) in the cortical and 34% (p < 0.001, n = 3) in the hippocampal homogenates. ADAM10 activity also increased by 57% (p < 0.01, n = 3) in the HeLa cells. Finally, TFEB-induced ADAM10 potentiation led to increased secretion of sAPPα by 154% (p < 0.001, n = 3) in the cortex and 62% (p < 0.001, n = 3) in the hippocampus. Thus, TFEB expression enhances nonamyloidogenic processing of APP. In conclusion, TFEB expression induces ADAM10 in an autophagy-dependent manner through PPARα.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ever since Andrea Ballabio’s group described the pivotal role of the transcription factor EB (TFEB) in the autophagy-lysosome pathway (ALP) [1, 2], numerous studies have further documented the beneficial roles of TFEB in several models of neurological diseases. TFEB controls ALP by regulating the expression of more than 35 CLEAR network genes [1, 3]. Thus, viral transduction of TFEB has demonstrated therapeutic benefits in mouse models of Alzheimer’s disease (AD) [4,5,6], tauopathy [7, 8], Parkinson’s disease (PD) [9, 10], and Huntington’s disease (HD) [11, 12]. Indeed, by generating flag-TFEB expressing transgenic mice for the first time, we also recently demonstrated that TFEB overexpression in the P301S model of tauopathy markedly cleared PHF-tau levels which led to the recovery of cognitive deficits [13]. Importantly, TFEB expression significantly reduced the age-associated lipofuscin pigments which were so far considered to be nondegradable [13]. An expanding role of TFEB in various other non-neuronal diseases is also becoming clear especially in the lysosomal storage diseases (LSDs), diet-induced obesity, multiple sulfatase deficiency, and α-1-antitrypsin deficiency [14]. The beneficial role of TFEB-mediated clearance of toxic protein aggregates in multiple neurodegenerative diseases is not surprising given that TFEB protein levels are reduced in many patient brains with neurological diseases including AD, amyotrophic lateral sclerosis (ALS) [15, 16], PD [17], and mouse models of SODG93A, HD, PD, and polyQ AR spinobulbar muscular atrophy [18,19,20,21]. Because cytoplasm to nuclear translocation of TFEB is critical for its transcriptional regulation of ALP, replenishing the reduced nuclear TFEB levels by overexpression studies is a logical step towards therapeutic benefits in various neurological models. Thus, since enhancing the TFEB pathway is expected to have a broad impact on human health, it has triggered intense research efforts in understanding the molecular basis for many of these beneficial properties of TFEB. Yet, how TFEB expression decreases Aβ and hyperphosphorylated tau and also increases longevity [22, 23] remains poorly understood.
ADAM10 (a disintegrin and metalloproteinase 10) has recently emerged as the major α-secretase responsible for amyloid precursor protein (APP) processing [24, 25]. Multiple pieces of evidence, including A673T mutation in the APP which decreases Aβ levels leading to protection against AD [26], K16N mutation at the APP α-secretase cleavage site [27], and two (Q170H and R181G) mutations within ADAM10 [28], which also increases Aβ levels and leads to late-onset Alzheimer’s disease (LOAD), all provide solid evidence that inadequate ADAM10 activity is likely the cause of not only familial but also sporadic cases of AD. Moreover, ADAM10 has been shown to play a pivotal role in the proliferation of neural precursor cells (NPCs) and neurogenesis in the adult brain [29, 30]. Equally important, ADAM10 has demonstrated neuroprotection against Aβ and excitotoxicity [31]. Additionally, ADAM10 plays a crucial role in axon guidance [32], spine density regulation, and in decreasing neuroinflammation [33, 34]. Further, ADAM10 expression correlates with mini-mental state of AD patients [35] and alleviates long-term potentiation (LTP) and cognitive deficits in an APP transgenic mouse model [31, 34]; its impaired trafficking to the synapse generates a model of the sporadic AD [36], and there is now definitive evidence that reduction of ADAM10 activity can cause AD [30], and finally, ADAM10 is particularly high in the subventricular zone (SVZ) suggesting a correlation between this pathway and the adult neurogenesis [37]. Although this overwhelming evidence suggests the ADAM10 pathway is the right target to combat AD, so far only limited studies have identified genetic regulation of ADAM10 activity.
Here, we unexpectedly discovered that TFEB increases the expression of mature ADAM10 in HeLa cells. Further, using pharmacological and genetic approaches, we identified PPARα as the mediator of increased ADAM10 in TFEB expressing cells. Finally, using our recently generated flag-TFEB transgenic mice, we confirmed the increased levels of mature ADAM10 protein and enzyme activity in both the cortex and hippocampus of mouse brains. This remarkable finding further reinforces TFEB as a reliable candidate therapeutic target for AD and possibly several other neurological disorders.
Results
TFEB Increases ADAM10 Protein Levels in HeLa Cells
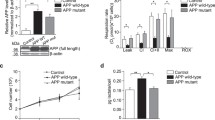
Several studies have demonstrated that TFEB expression using adenoviral vectors reduces Aβ generation in cells as well as in the brain [4, 5], but the mechanism by which TFEB reduces Aβ is not known. Since ADAM10 is the major α-secretase known to reduce Aβ generation through the nonamyloidogenic pathway, in this study, we wanted to measure if there are any alterations in ADAM10 protein levels and activity following TFEB expression. HeLa cells stably expressing EGFP-TFEB were successfully generated and confirmed the expression of EGFP-tagged TFEB using both TFEB-specific and GFP-specific antibodies. Quantitation of actin normalized levels of ADAM10, and analysis by t test revealed significant differences. ADAM10 mature form was increased by 72% (p < 0.01, n = 4) in the EGFP-TFEB expressing HeLa cells compared to HeLa control cells. Similarly, ADAM10 proform was increased by 118% (p < 0.001, n = 4) compared to HeLa control cells (Fig. 1 a and b). Thus, TFEB expression leads to a robust increase in both proform and mature forms of ADAM10 in HeLa cells. We next used CRISPR strategy to knockdown endogenous TFEB to verify whether reduced TFEB expression decreases ADAM10 protein levels. HeLa cells stably expressing CRISPR construct targeted against TFEB were generated, and ADAM10 levels were compared between HeLa, HeLa-EGFP-TFEB cells, and HeLa-TFEB-CRISPR cells. Statistical tests by analysis of variance (ANOVA) followed by Dunnett’s test showed a significant effect. In the HeLa-CRISPR stable cells, the endogenous TFEB levels were only 12% (p < 0.05, n = 4) of the HeLa control cells (Fig. 1 c and d), suggesting that the used CRISPR construct correctly targeted the endogenous TFEB. Results showed that mature ADAM10 protein level was decreased by 36% (p < 0.05, n = 4) in the HeLa-CRISPR cells while increased by 46% (p < 0.01, n = 4) in the HeLa-EGFP-TFEB cells compared to HeLa control cells (Fig. 1e). ADAM10 proform was also increased by 160% (p < 0.01, n = 4) in the HeLa-EGFP-TFEB cells but decreased by 30% in the HeLa-CRISPR cells (Fig. 1f) which were statistically significant.
Effect of TFEB on ADAM10 protein levels in the HeLa cells. a HeLa cells stably expressing EGFP-TFEB and control HeLa cells were plated in the 6-well plates in duplicates, and next day cells were lysed, and the lysates were separated by SDS-PAGE and transferred to nitrocellulose membranes. Representative images are shown for the HeLa control and HeLa-EGFP cells in duplicate wells. b Quantitation of ADAM10 proform and mature forms normalized to actin levels showed significant differences by t test. In the EGFP-TFEB expressing cells, ADAM10 mature form was increased by 72% (**, p < 0.01, n = 4), and the ADAM10 proform was increased by 118% (***, p < 0.001, n = 4) compared to HeLa control cells. c In addition to HeLa cells and HeLa-EGFP cells, another HeLa cells line stably expressing CRISPR construct targeted against TFEB were plated in duplicate wells, and the lysates were subjected to immunoblotting. TFEB antibody detected endogenous TFEB protein (panel 3), while GFP antibody detected EGFP-TFEB protein (panel 5). d Bar diagrams show that in the HeLa-CRISPR stable cells, the endogenous TFEB levels were only 12% (*, p < 0.05, n = 4) of the HeLa control cells which was statistically significant as analyzed by analysis of variance (ANOVA) followed by Dunnett’s test. e ADAM10 mature form was decreased by 36% (*, p < 0.05, n = 4) in the HeLa-CRISPR cells while increased by 46% (**, p < 0.01, n = 4) in the HeLa-EGFP-TFEB cells compared to HeLa control cells. ADAM10 proform was also increased by 160% (**, p < 0.01, n = 4) in the HeLa-EGFP-TFEB cells, but the 30% decrease in the HeLa-CRISPR cells were not statistically significant
Increased ADAM10 by TFEB Is Autophagy-Dependent in HeLa Cells
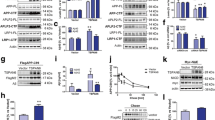
Having confirmed the increased ADAM10 protein levels, we wanted to investigate whether this effect of TFEB on ADAM10 is autophagy-dependent since TFEB is the master regulator of the ALP [1, 2]. To address this point, we utilized both autophagy inhibition approach using pharmacological inhibitors and autophagy activation through starvation in HeLa control cells as well as HeLa cells under TFEB overexpression conditions (Fig. 2a). To increase rigor in the study, we used both 3-methyladenine (3-MA) which inhibits autophagy by blocking autophagosome formation by inhibiting type III phosphatidylinositol 3-kinases (PI-3K) [38] and bafilomycin A1 (BAF1) which is a blocker of autophagy flux by disrupting the autophagosome-lysosome (A-L) fusion, thereby inhibiting ALP pathway (39). As expected mature ADAM10 protein levels normalized to actin were increased in untreated HeLa-EGFP-TFEB cells compared to untreated HeLa control cells by 67% (p < 0.05, n = 4) which was statistically significant by t test (Fig. 2b). Autophagy inhibition per se either by BAF1 or 3-MA did not alter ADAM10 protein levels as seen in the HeLa cells. However, comparison among all the groups that include HeLa cells and HeLa-EGFP-TFEB cells treated or untreated with BAF1 or 3-MA showed significant differences. Quantitation and analysis of mature ADAM10 by ANOVA followed by Dunnett’s test revealed a 67% (p < 0.01, n = 6) increase in the HeLa-EGFP-TFEB cells and a 54% (*, p < 0.05, n = 6) increase in the BAF1-treated cells when compared to HeLa control cells (Fig. 2c), suggesting that autophagy inhibition by 3-MA but not BAF1 can prevent TFEB-induced increased ADAM10 protein levels. This implies that the TFEB-mediated effect on ADAM10 is autophagy-dependent. Although BAF1 is known to inhibit autophagy, LC3-II levels were significantly increased (100%, *, p < 0.05) in the HeLa control cells (Fig. 2c). Also in the HeLa-EGFP-TFEB cells, LC3-II levels showed 147% (**, p < 0.01, n = 6) increase in the BAF1-treated cells and 145% (p < 0.01, n = 6) increase in the 3-MA-treated cells when compared to HeLa control cells (Fig. 2c).
Effect of autophagy inhibition on ADAM10, LC3-II, and p62 protein levels in the HeLa cells and HeLa-EGFP-TFEB cells. a Both HeLa control cells and HeLa-EGFP-TFEB stable cells were plated on to 6-well plates in duplicates and were treated with either vehicle as control, bafilomycin a1 (BAF1) at 100 nM final concentration or 3-methyladenine (3-MA) at 5 mM final concentration. After 4 h, cell lysates were subjected to SDS-PAGE and transferred to nitrocellulose membranes. Representative images are shown for several proteins and actin. b As analyzed by t test, mature ADAM10 protein levels were significantly increased by 67% (*, p < 0.05, n = 6) when HeLa control cells were compared to HeLa-EGFP-TFEB cells. However, mature ADAM10 levels were not altered when comparisons were made between HeLa-EGFP-TFEB cells and the same cells treated with either BAF1 or 3-MA. c Quantitation of mature ADAM10 by ANOVA followed by Dunnett’s test revealed a 67% (**, p < 0.01, n = 6) increase in the HeLa-EGFP-TFEB cells, a 54% (*, p < 0.05, n = 6) increase in the BAF1-treated cells when compared to HeLa control cells. Also in the HeLa-EGFP-TFEB cells, LC3-II levels showed 147% (**, p < 0.01, n = 6) increase in the BAF1-treated cells and 145% (p < 0.01, n = 6) increase in the 3-MA-treated cells when compared to HeLa control cells. BAF1 treatment also increased LC3-II levels by) 100% (*, p < 0.05) in the HeLa control cells
Starvation is a well-characterized method to induce autophagy which is also known to activate TFEB [1, 2]. Therefore to induce autophagy, HeLa control cells and HeLa-EGFP-TFEB cells were exposed to either DMEM medium with 10% fetal bovine serum (FBS) as well-fed controls or induced starvation incubating cells with HBSS medium without fetal bovine serum (FBS) for 1 h or 3 h (Fig. 3a). Autophagy activation per se, similar to autophagy inhibition, did not alter ADAM10 protein levels in the HeLa control cells (Fig. 3b). Consistent with results in Figs. 1 and 2, compared to control HeLa cells, mature ADAM10 protein levels were increased in the HeLa-EGFP-TFEB cells by 51% (p < 0.01, n = 4). Autophagy activation by starvation further increased ADAM10 mature form by 63% (p < 0.001, n = 4) after 1 h and 91% (p < 0.001, n = 4) after 3 h which were all highly significant by ANOVA followed by Student-Newman-Keuls multiple comparisons test (Fig. 3b). When the comparison is made between the fully fed HeLa-EGFP-TFEB cells and after starvation for 3 h, there was a further 40% ($$, p < 0.01) increase in mature ADAM10 levels (Fig. 3b), suggesting that under TFEB expression conditions, starvation-induced autophagy activation enhances ADAM10 protein levels. A similar trend of ADAM10 proform was also noted, i.e., no changes in the HeLa cells subjected to starvation, but increased in the HeLa-EGFP-TFEB cells by 41% (p < 0.001, n = 4) when compared to HeLa control cells, by 72% (p < 0.001, n = 6) after 1 h starvation and by 111% (p < 0.001, n = 4) after 3 h starvation (Fig. 3b). Activation of autophagy by TFEB expression is indicated by LC3-II levels which were increased by 62% (p < 0.05) in the HeLa-EGFP-TFEB cells when compared to HeLa cells. However, it is intriguing that starvation-induced autophagy activation failed to increase the levels of either LC3-I, LC3-II, or P62 levels, although an increasing trend is noted in the HeLa-EGFP-TFEB cells subjected to starvation (Fig. 3b). Although we did not quantify, starvation is known to dephosphorylate TFEB which is apparent in this study (Fig. 3a) indicating autophagy activation. Overall, results from both autophagy inhibition and starvation-induced autophagy activation studies suggest that the TFEB-mediated increase in ADAM10 protein levels is autophagy-dependent.
Effect of starvation-induced autophagy activation on ADAM10 and markers of autophagy in the HeLa cells and HeLa-EGFP-TFEB cells. a Cells were exposed to either DMEM medium with 10% fetal bovine serum (FBS) as well-fed control or induced starvation with HBSS medium without FBS for 1 h or 3 h, followed by SDS-PAGE analysis of cell lysates. Representative images of blots from different experiments are shown for several proteins along with actin for each of these proteins. b Compared to control HeLa cells, mature ADAM10 protein levels were increased in the HeLa-EGFP-TFEB cells by 51% (**, p < 0.01, n = 4) or 63% (***, p < 0.001, n = 4) after 1 h starvation and 91% (***, p < 0.001, n = 4) after 3 h starvation as analyzed by ANOVA followed by Student-Newman-Keuls multiple comparisons test. Compared to HeLa-EGFP-TFEB cells, there was a 41% ($$, p < 0.01) further increase in mature ADAM10 levels after 3 h starvation. ADAM10 proform levels were also increased in HeLa-EGFP-TFEB cells by 41% (***, p < 0.001, n = 4), after 1 h starvation by 72% ((***, p < 0.001, n = 4) and after 3 h starvation by 111% (***, p < 0.001, n = 4). LC3-II levels were increased by 62% (*, p < 0.05) only when HeLa cells were compared to HeLa-EGFP-TFEB cells. However, LC3-I levels were not statistically significant
Genetic and Pharmacological Evidence that TFEB Increases ADAM10 Through PPARα Signaling
Both TFEB [40] and peroxisome proliferator-activated receptor α (PPARα) [41] regulate peroxisome biogenesis. Both these transcription factors are also known to regulate fatty acid metabolism and lysosome biogenesis in the same pathway. Moreover, like TFEB [4, 5], PPARα activation also decreases Aβ generation likely through the activation of the nonamyloidogenic pathway [42]. Accordingly, we suspected that PPARα signaling might be responsible for TFEB-induced ADAM10 potentiation. Therefore, we used PPARα-specific synthetic siRNAs or scrambled control siRNAs which were transiently transfected into the HeLa-EGFP-TFEB cells and compared with those of HeLa control cells. There was a 57% (p < 0.01, n = 6) increase of normalized mature ADAM10 in the HeLa-EGFP-TFEB cells transfected with control siRNA and only 29% (p < 0.05, n = 6) increase in the cells treated with PPAR-α-specific siRNA compared to HeLa cells (Fig. 4a). Thus, a 28% reduction in ADAM10 protein levels may be due to a reduction of PPAR-α levels by 28% ($$, p < 0.01) when compared to EGFP-TFEB cells transfected with control siRNAs (Fig. 4a). These results suggest that a decrease in PPARα leads to a reduction in the amount of ADAM10 mature protein in HeLa cells. To confirm these results, we also used a pharmacological approach. There was a 33% (p < 0.01, n = 4) increase in ADAM10 protein levels when HeLa-EGFP-TFEB cells were compared to HeLa control cells (Fig. 4b). When HeLa-EGFP-TFEB cells were treated with GSK0660, a PPAR-β antagonist, there was 40% (p < 0.01, n = 4) increase in ADAM10 protein levels, with GW9662, a PPAR-γ antagonist, there was 28% (p < 0.05, n = 4) increase, which was all statistically significant as analyzed by ANOVA and Dunnett’s test (Fig. 4b). However, when HeLa-EGFP-TFEB cells were treated with GW6471, a PPAR-α antagonist, there was no significant change in the ADAM10 protein levels and was similar to HeLa control cells. Similarly, ADAM10 proform increased in the EGFP-TFEB cells treated with the vehicle by 69% (p < 0.05); the increase was 114% (p < 0.01) after GSK0660 treatment, 86% (p < 0.01) after GW6471 treatment, and by 75% (*, p < 0.05) in the GW9662-treated cells (Fig. 4b). These results from pharmacological inhibition studies of PPAR isoforms are consistent with the results from PPAR-α targeted siRNA experiments. Taken together, these results suggest that TFEB-induced increased mature ADAM10 protein levels is mediated through PPAR-α.
PPAR-α is responsible for TFEB-mediated increased mature ADAM10 levels in HeLa cells. a HeLa cells and HeLa-EGFP-TFEB cells were plated in 6-well plates in duplicate wells, and HeLa-EGFP-TFEB cells were transiently transfected twice at 24 h interval with either control scrambled siRNA or PPAR-α-specific siRNA. Twenty-four hours after the second transfection, cells were lysed and subjected to SDS-PAGE. Blots shown are from different experiments. Quantitation and analysis by ANOVA followed by Student-Newman-Keuls multiple comparisons test showed a 57% (**, p < 0.01, n = 6) increase of mature ADAM10 in the HeLa-EGFP-TFEB cells transfected with control siRNA and only 29% (*, p < 0.05, n = 6) increase in the cells treated with PPAR-α-specific siRNA compared to HeLa cells. PPAR-α levels were increased to 17% (*, p < 0.05, n = 6) in the HeLa-EGFP-TFEB cells but reduced to 89% ($$, p < 0.01, n = 6) after transfection with PPAR-α-specific siRNA. b HeLa-EGFP-TFEB cells were treated with either vehicle control or GSK0660 (PPAR-β antagonist), GW6471 (PPAR-α antagonist), or GW9662 (PPAR-g antagonist) for 24 h, and then lysates were immunoblotted. Mature ADAM10 levels increased in the EGFP-TFEB cells treated with the vehicle by 33% (**, p < 0.01, n = 4), in the GSK0660-treated cells by 40% (**, p < 0.01, n = 4), and in the GW9662-treated cells by 28% (*, p < 0.05, n = 4) but was not altered in the GW6471-treated cells. Similarly, ADAM10 proform increased in the EGFP-TFEB cells treated with the vehicle by 69% (*, p < 0.05, n = 4), in the GSK0660-treated cells by 114% (**, p < 0.01, n = 4), in the GW6471-treated cells by 86% (**, p < 0.01, n = 4), and by 75% (*, p < 0.05, n = 4) in the GW9662-treated cells
TFEB Expression Increases ADAM10 Protein Levels In Vivo in the Transgenic Mouse Brain Expressing Flag-TFEB
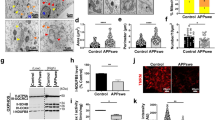
We next examined whether TFEB expression can increase ADAM10 in vivo in the mouse brains. To investigate this, we used our flag-TFEB transgenic mice [13]. We found a significant increase of ADAM10 protein levels by 60% (p < 0.05, n = 3) in the cortex and by 34% (p < 0.001, n = 3) in the hippocampal homogenates of flag-TFEB mice compared to wild-type (WT) mice and as analyzed by t test (Fig. 5a). Since ADAM10 is known to be localized to synaptic terminals [43, 44], we also wanted to assess whether TFEB has any differential effects on the cellular localization of ADAM10. Therefore, we also quantified ADAM10 protein levels in the synaptosomes. We used our optimized protocol for the separation of synaptosomes, the purity of which has been confirmed by both biochemical methods and transmission electron microscopy [45, 46]. Similar to results in the homogenates, quantitation of mature ADAM10 protein levels normalized to actin in the synaptosomes fractions revealed an increase of 56% (p < 0.01, n = 3) in the cortex and 64% (p < 0.01, n = 3) in the hippocampal synaptosomes which were also statistically significant by t test (Fig. 5b). This in vivo evidence of an increased mature form of ADAM10 protein levels by TFEB is consistent with the results from HeLa cells.
TFEB expression increases mature ADAM10 levels in the mouse brain. a Cortical and hippocampal brain homogenates were prepared from wild-type (WT) and flag-TFEB mice and immunoblotted showing ADAM10, flag-TFEB, and actin protein blots. Quantitation of mature ADAM10 protein levels revealed an increase of 60% (*, p < 0.05, n = 3) in the cortex and 34% (***, p < 0.001, n = 3) in the hippocampal homogenates. b Cortical and hippocampal synaptosomes were prepared from the WT and flag-TFEB mice and immunoblotted. Quantitation of mature ADAM10 protein levels revealed an increase of 56% (**, p < 0.01, n = 3) in the cortex and 64% (**, p < 0.01, n = 3) in the hippocampal homogenates which were statistically significant by t test
TFEB Expression Increases ADAM10 Enzyme Activity in HeLa Cells and In Vivo in the Mouse Brain
To ascertain that increased ADAM10 protein levels by TFEB indeed result in increased ADAM10 enzyme activity, we initially prepared cell lysates from HeLa control cells and HeLa cells stably expressing EGFP-TFEB. Results showed that compared to HeLa control cells, HeLa-EGFP cells showed an increased ADAM10 enzyme activity by 57% (p < 0.01, n = 3) (Fig. 6a). Similarly, cortical and hippocampal homogenate fractions derived from WT and flag-TFEB mice were subjected to a fluorogenic method of ADAM10 enzyme activity assay. A 37% (p < 0.01, n = 3) and 50% (p < 0.05, n = 3) increase in ADAM10 enzyme activity was found in the cortex and hippocampus, respectively, as revealed by t test (Fig. 6b). Thus, TFEB is a bona fide activator of ADAM10 activity with enormous therapeutic potential.
TFEB expression increases ADAM10 enzyme activity in HeLa cells and in the mouse brain. a ADAM10 activity in cell lysates showed an increase by 57% (p < 0.01, n = 3) in the HeLa-EGFP stable cells compared to HeLa control cells. b Cortical and hippocampal brain homogenates were prepared from the WT and flag-TFEB mice, and the measurement of ADAM10 activity showed a statistically significant increase of 37% (**, p < 0.01, n = 3) in the cortex and 50% (*, p < 0.05, n = 3) in the hippocampus as revealed by t test
Increased ADAM10 Activity by TFEB Results in Increased sAPPα Levels in the Mouse Brain
Since the major α-secretase in the brain is ADAM10 [47, 48], its enhanced cleavage of amyloid precursor protein (APP) should invariably increase the generation of soluble-APP-α (sAPPα). To prove that the increased ADAM10 protein levels and enzyme activity by TFEB are physiologically relevant, we measured the quantity of secreted sAPPα levels in the brain lysates from WT and flag-TFEB mice. Using mouse sAPPα-specific antibody M3.2, we found an increase of sAPPα levels normalized to actin by 154% (***, p < 0.001, n = 3) in the cortex and by 62% (***, p < 0.001, n = 3) in the hippocampus (Fig. 7 a and b) which were highly significant by t test. To further confirm these results, based on our previous experience, we also used a 6E10 antibody which detects mouse sAPPα also, although 6E10 antibody was raised against the human protein. With 6E10 antibody, we found 153% (p < 0.001, n = 3) and 176% (p < 0.01, n = 3) increase of sAPPα levels in the cortex and hippocampus, respectively (Fig. 7 a and b). There was no apparent change in the levels of APP holoprotein either in the cortex or hippocampus of flag-TFEB mice compared to WT mice. Additionally, we found decreased c-terminal fragment (CTF)-β levels by 46% (p < 0.001) in the cortex, but not in the hippocampus (Fig. 7 a and b). Increased nonamyloidogenic processing of APP is expected to decrease CTF-β levels. Therefore, the observed decreased CTF-β levels are consistent with the increased sAPPα levels.
Increased ADAM10 activity by TFEB increases sAPPα levels and reduces c-terminal fragment-β (CTF-β) levels in the brain. Secreted proteins such as sAPPα were extracted using tris buffer, while the other cellular proteins such as CTF-β, amyloid precursor protein (APP), and flag-TFEB were extracted using a detergent containing buffer and immunoblotted as shown. In the cortex, sAPPα levels increased by 154% (***, p < 0.001, n = 3) as detected by M3.2 antibody or by 153% (***, p < 0.001, n = 3) as detected by 6E10 antibody and analyzed by t test. In the hippocampus, sAPPα levels increased by 62% (***, p < 0.001, n = 3) as detected by the M3.2 antibody or by 176% (**, p < 0.01, n = 3) as detected by 6E10 antibody and analyzed by t test. Increased sAPPα levels were confirmed using two antibodies, M3.2 and 6E10. CTF-β levels were significantly reduced by 46% (***, p < 0.001, n = 3) in the cortex, but in the hippocampus, it was insignificant. Similar levels of APP, as detected by anti-APP antibody, were seen in both WT and flag-TFEB brains. As expected flag antibody detected flag-TFEB expression only from flag-TFEB mice and not from WT mice
Discussion
A promising alternative approach for the therapeutic benefit of AD patients is the activation or potentiation of the nonamyloidogenic processing of APP [49,50,51]. Activation of the nonamyloidogenic ADAM10 pathway not only precludes Aβ generation but most importantly increases sAPPα which is known to exert potent neuroprotective properties as well as regulate synaptic plasticity and learning and memory [52]. The reduced ADAM10 trafficking [53], protein expression [54], and enzyme activity [55] in AD patients underscore the significance of ADAM10 in AD. Therefore, it is crucial to identify and understand the mechanistic regulation of ADAM10.
Here, we found that TFEB overexpression robustly increased actin normalized levels of mature ADAM10 protein in the HeLa cells. Importantly, CRISPR-based stable knockdown of TFEB in the same HeLa cells, as confirmed by a marked reduction in the levels of endogenous TFEB, significantly reduced mature ADAM10 protein levels. This suggests that TFEB indeed has a pertinent role in regulating ADAM10 protein levels in the HeLa cells. From this study, two strong pieces of evidence also suggest that TFEB may increase ADAM10 mature form through the autophagy pathway. First, autophagy inhibition by 3-MA prevented TFEB-induced increased ADAM10 mature form in the HeLa-EGFP-TFEB cells. However, mature ADAM10 protein levels in the BAF1-treated HeLa-EGFP-TFEB cells were not significantly reduced compared to HeLa-EGFP-TFEB cells. 3-MA inhibits autophagosome initiation by inhibiting type III phosphatidylinositol 3-kinases (PI-3K) of class III PI3K [38, 56]; thus, 3-MA is an early-stage autophagy-lysosome pathway inhibitor. But BAF1inhibits the late stage of the ALP pathway during autophagosome-lysosome fusion [39]. These observations imply that TFEB-induced increased ADAM10 might require activation of the entire autophagy-lysosome pathway including the initiation of autophagosome formation. Second, after the activation of autophagy by starvation, especially after 3 h, the increase in ADAM10 mature protein levels was significantly more than the completely nutrient-fed HeLa-EGFP-TFEB cells. Interestingly, starvation alone, in the absence of TFEB expression, failed to increase ADAM10 protein levels in the HeLa control cells, although an insignificant increasing trend was noted. Thus although endogenous TFEB in HeLa cells as well as exogenously expressed EGFP-TFEB both underwent dephosphorylation, a necessary step for TFEB-mediated activation of the autophagy-lysosome pathway, ADAM10 levels were increased only after TFEB overexpression. This evidence together with the results from autophagy inhibition studies strongly suggests that TFEB influences ADAM10 levels through the autophagy-lysosome pathway. The role of the autophagy-lysosome pathway in ADAM10 regulation has not been thoroughly investigated previously except a recent demonstration that knockdown of ATG16L1, a key player in basic autophagy mechanism, decreased levels of ADAM10 mature form [57].
Intriguingly, in the present study, autophagy marker LC3-II was not significantly altered following starvation-induced autophagy, but as expected, we observed increased LC3-II levels only following TFEB overexpression during nutrient-rich conditions, which is consistent with the previous reports [58, 59]. Surprisingly, under nutrient-rich conditions, BAF1 rather increased LC3-II levels, but when TFEB was overexpressed, both BAF1 and 3-MA increased LC3-II levels. Increased LC3-II activity of 3-MA may be due to its differential temporal effects on class I and class III PI3K. In our study, although both inhibitors were expected to decrease LC3-II levels during autophagy inhibition exposed to 4 h, a previous study also found increased LC3-II levels after exposure of several cell types to 3-MA for 9 h [60]. BAF1 also has been shown to increase LC3-II levels [61]. Increased LC3-II may simply reflect the accumulation of autophagosomes, rather than increased autophagy flux or may even be due to blockage of fusion of autophagosomes with lysosomes as has been shown previously by others [62,63,64]. Alternatively, TFEB expression and autophagy activation may be dominant over autophagy inhibition by 3-MA and BAF1. We also did not find any changes in the levels of P62 following autophagy inhibition with 3-MA or BAF1 or during starvation-induced autophagy activation. This may likely be due to our short time exposures ranging from 1 to 4 h. In HeLa cells, the half-life of P62 is about 6 h [65]. So, detectable differences in P62 protein levels may not be apparent by 4 h.
An important finding from the present study is the demonstration that PPARα is the mediator of TFEB-induced increased ADAM10 mature form in HeLa cells by using PPARα-specific siRNA-based knockdown method. Further, using isoform-specific inhibitors, our pharmacological studies confirmed that PPARα, but not either PPARβ or PPARγ, is responsible for TFEB-induced increased ADAM10 protein levels. Interestingly, PPARα is one of the major genes that is significantly enhanced by increased TFEB activity under starvation-induced stress [66]; thus, PPARα is downstream to TFEB and can mediate this function. Additionally, the ADAM10 promoter harbors PPAR response elements (PPRE) [67], and among several isoforms, an inhibitor of PPARα but not PPARβ, PPAR δ, or PPARγ decreased the ADAM10 protein levels in primary hippocampal neurons [67]. Thus our identification of PPARα as the mediator of increased ADAM10 is consistent with these reported observations. However, it should also be noted that the mouse TFEB promoter harbors a PPRE, and PPARα can also regulate transcriptional activation of TFEB [66, 68]. Thus TFEB can also function downstream of PPARα as confirmed by gemfibrozil, a PPARα agonist [68]. Indeed, PPARα stimulation has been shown to regulate ADAM10 protein expression [67]. On the contrary, PPARα deficiency in brains and neurons results in impaired ADAM10 expression [67]. These multiple pieces of evidence support our finding that PPARα can mediate TFEB-induced ADAM10 protein maturation.
While these observations in HeLa cells are important especially for identifying the mechanism, it is even more crucial to verify whether TFEB expression can also increase ADAM10 mature form in vivo in the brain. Our flag-TFEB mice [13] became an invaluable tool to address this issue. Indeed, similar to results from HeLa cells, flag-TFEB expression significantly increased ADAM10 mature form in the whole homogenate fractions of both the cortex and hippocampus. This observation in the brain confirms that TFEB-induced increased ADAM10 mature protein occurs in vivo and is consistent with increased ADAM10 enzyme activity observed in the same brain regions as well as HeLa cells. Since ADAM10 may also play a critical role at synapses as reflected by its colocalization with the presynaptic protein synaptophysin and postsynaptic protein SAP-97 [43], and in dendritic spine generation [33], we quantified ADAM10 protein levels in the purified synaptosomes and found a robust increase from both hippocampal and cortical synaptosomes. As both ADAM10 deficiency [33] and Alzheimer’s disease [69, 70] result in fewer and abnormally shaped spines, TFEB-induced enhanced ADAM10 expression can be expected to rescue such synaptic abnormalities which is the best pathological correlate of cognitive deficits in AD. Thus, cognitive deficits that correlate with reduced ADAM10 levels seen in AD patient platelets [35, 71] may also be rescued with the TFEB-induced ADAM10 expression, thereby may reverse cognitive deficits.
Another important proof for the physiological significance of TFEB-induced increased ADAM10 mature protein and enzyme activity comes from the observed more sAPPα in the cortex and hippocampus of flag-TFEB mice than the WT mice. ADAM10 being the major α-secretase in the brain for cleaving APP at the α-secretase cleavage site [24, 25], increased sAPPα level is another solid evidence in support of increased ADAM10 by TFEB. Overexpression of ADAM10 that results in only 30% more enzymes over the endogenous levels leading to increased sAPPα levels was shown to almost completely prevent plaque formation and learning and memory deficits in the APP/PS1 transgenic mice [34]. In this study, we found a more than 30% increase in the mature ADAM10 protein levels, as well as ADAM10 enzyme activity suggesting that the extent of ADAM10 induction in the brain regions by TFEB is sufficient for significantly reducing, amyloid plaques, synaptic deficits, and the associated cognitive deficits. Pieces of evidence such as a robust elevation in plaque area, number, and size in the brain following PPARα deficiency in the 5XFAD mice [67] and increased ADAM10 protein both proform and mature form as well as sAPPα levels after PPARα agonist, WY14643 stimulation [67], together with the present results imply that TFEB-induced enhanced ADAM10 and sAPPα occur through PPARα pathway. Although the mature form of ADAM10 has been recently shown to rapidly undergo autoproteolytic degradation in the normal lysis buffers [72], it is important to note that we have used the same lysis buffer and similar lysis conditions in both the control and experimental conditions, and therefore, if at all there was any degradation, it is expected to occur in both the groups. Moreover, increased ADAM10 enzyme activity as well as sAPPα levels in the brain validated the increased levels of mature form of ADAM10 by TFEB.
The current study has several limitations. First, we have demonstrated ADAM10 potentiation and PPARα mechanism only in HeLa cells, and therefore, it is important to investigate whether the observed effects also occur in other cell types. Autophagy inhibition by BAF1 and 3-MA may not be selective on autophagy, and other pathways may be affected. The doses of BAF1 and 3-MA were selected based on literature reports. Future studies should look at a range of doses and multiple time points so that autophagy markers should reflect the correct stage of the autophagy-lysosome pathway. Finally, future studies could use cells derived from PPARα deficient mice to provide confirmatory evidence that PPARα mediates TFEB-induced increased ADAM10. Nevertheless, the present data, especially from the mouse brain, provides solid data for our conclusion.
In summary, we have demonstrated that TFEB activates ADAM10 through PPARα pathway in HeLa cells and that such ADAM10 potentiation resulting in nonamyloidogenic processing of APP and sAPPα generation also occurs in vivo in the brain. Since there is an age-associated decline in nonamyloidogenic processing of APP [73] and since sAPPα is also known to directly bind to BACE1, thereby inhibiting BACE1 and reducing Aβ [74, 75], both TFEB and PPARα are excellent targets to augment nonamyloidogenic pathway as a therapeutic strategy for AD.
Materials and Methods
Chemicals and Antibodies
GW7647 (cat # G6793), GSK0660 (cat # G5797), GW6471 (cat # G5045), protease inhibitor cocktail (cat # P8340), dithiothreitol (cat # D9779), sodium orthovanadate (cat # 450243), and G418 disulfate (cat # A1729) were all purchased from Sigma-Aldrich (St. Louis, MO, USA). Puromycin dihydrochloride (cat # 540222) and microcystin-LR (cat# 475815) were purchased from Calbiochem-Millipore (Temecula, CA, USA). Nonidet-P40 substitute (cat # M158) was obtained from AMRESCO (Solon, OH, USA). Fetal bovine serum (FBS) was purchased from BioFluid Technologies (cat # SKU: 200-500-Q). Polyclonal TFEB antibody (cat # 4240) was purchased from Cell Signaling (Danvers, MA, USA.). Polyclonal rabbit anti-ADAM10 antibody (cat # A10348) was from ABclonal. Monoclonal mouse anti-PPARα antibody (cat # sc-398,394) was from Santa Cruz Biotechnology (Dallas, TX, USA). Monoclonal flag antibody (M2, cat # F1804) and mouse monoclonal anti-GFP antibody (cat # SAB5300167) were purchased from Sigma-Aldrich (St. Louis, MO, USA). For the detection of sAPPα from soluble fractions, 6E10 antibody (cat # SIG-39320) and M3.2 antibody (cat # SIG-39155) were purchased from Covance. Anti-actin antibody (cat # JLA20) deposited by Jim Jung-Ching Lin was purchased from Developmental Studies Hybridoma Bank (DSHB), University of Iowa (Iowa City, IA, USA). Mouse monoclonal antibody against the alpha-tubulin (cat # A01410) was purchased from GenScript (Piscataway, NJ, USA). Secondary antibodies such as peroxidase-conjugated AffiniPure goat anti-mouse (code # 115-035-146) and goat anti-rabbit (code # 111-035-144) IgG (H + L) were purchased from Jackson ImmunoResearch Laboratories (West Grove, PA, USA). All primary antibodies for immunoblot analysis were diluted in 5% nonfat milk in tris-buffered saline with 0.1% Tween-20 (TBS-T) buffer, while the secondary antibodies were diluted in the TBS-T buffer.
Generation of Stable HeLa Cells Expressing EGFP-TFEB or mCherry-TFEB sgRNA
The plasmid pEGFP-N1-TFEB expressing human TFEB, deposited by Shawn Michael Ferguson, Department of Cell Biology, Yale School of Medicine, was obtained from Addgene (cat # 38119). HeLa cells (cat # CCL-2) derived from epithelial cells of human origin were purchased from the American Type Culture Collection (Manassas, VA, USA). To generate HeLa stable clones, cells were plated in a 10-cm plate and after 24 h were transiently transfected with the pEGFP-N1-TFEB plasmid using lipofectamine 2000 according to the manufacturer’s instructions. Following 24 h of transfection, the medium was replaced with DMEM medium containing the antibiotic G418 at 600 μg/mL concentration. Cells were cultured under the G418 selection-medium for approximately 1 month, which resulted in the growth of about 20 individual colonies. Each colony was further expanded and evaluated for the expression levels of EGFP-TFEB using a fluorescence microscope. Out of 20 clones, 3 were narrowed down, and one clone expressing highest levels of EGFP-TFEB was selected and maintained in the culture medium containing 300 μg/mL of G418. This cell line was designated as HeLa-EGFP-TFEB and was used in the present experiments. Similarly to generate TFEB knockdown, we purchased a lentiviral sgRNA clone for the human TFEB gene (cat # HCP219038-LvSG03-1-B) with the targeting sequence “GCCACCATGGCGTCACGCAT” against human TFEB in the dual-use vector pCRISPR-LvSG03 from GeneCopoeia (Rockville, MD, USA). After transient transfection of the construct with lipofectamine 2000, stable clones resistant to puromycin were selected and expanded. The stable cells were used only after the successful knockdown of TFEB was confirmed by immunoblotting.
Generation of Transgenic Mice with Neuronal Expression of Flag-TFEB Using Mouse Thy1 Promoter
All animal studies were approved by the institutional animal care and use committee (IACUC) and followed all the NIH ethical guidelines. The detailed methods of flag-TFEB transgenic mice generation are available in our previous publication [13]. Briefly, using pLenti6.2/V5-TFEB construct as the source of human TFEB cDNA, the p3XFlag-TFEB-P2A-EYFP construct was made by the combination of both restriction digestion and polymerase chain reaction (PCR). A single XhoI site within the TFEB cDNA was removed by site-directed mutagenesis, and the entire 3XFlag-TFEB-P2A-EYFP sequence was shuttled and fused with mouse thy-1 promoter at the XhoI restriction site in the pTSC21K plasmid. The use of thy1 promoter was successful in restricting flag-TFEB expression only in the neurons of the postnatal brain.
Isolation of Synaptosomes from the Mouse Brains
We used Syn-PER synaptic protein extraction reagent (cat # 87793, Thermo Scientific, Rockland, IL, USA) to prepare synaptic proteins that have been previously shown to retain phosphoprotein integrity. We followed the manufacturer’s instructions with a slight modification concerning the speed and duration of centrifugation steps. Briefly, freshly dissected cortical and hippocampal brain tissues isolated from the wild-type (WT) and flag-TFEB transgenic mice were mixed with Syn-PER reagent at 100 mg/mL and hand homogenized using Teflon pestle glass homogenizer. Tissue debris was removed by centrifuging the homogenate at 3000×g for 10 min, followed by another centrifugation at 15,000×g. The supernatant was used as the homogenate fraction, and the pellet was dissolved in Syn-PER reagent at 1.0 mL per 100 mg initial brain tissue and used as synaptosome fractions. We previously confirmed the integrity of synaptosomes isolated by this method by enriched detection of synaptic protein markers by western blots and the presence of postsynaptic density by transmission electron microscopy.
Quantitation of Protein Levels by Immunoblotting
HeLa cells stably expressing EGFP-TFEB and control HeLa cells were plated into 6-well plates at about 70% confluency. The next day EGFP-TFEB cells were treated with either the vehicle control or GSK0660 (PPARβ antagonist), GW6471 (PPARα antagonist), or GW9662 (PPARγ antagonist), all at 1.0 μM final concentration, and after 24 h, the cells were lysed using lysis buffer (1% NP40) with complete protease inhibitor mix (Sigma) supplemented with sodium vanadate and microcystin. To quantitate protein levels in the brain, WT and flag-TFEB mice were euthanized with isoflurane, decapitated immediately, and cortical and hippocampal brain regions were rapidly separated into Syn-PER reagent containing the complete protease inhibitor cocktail plus the microcystin and sodium vanadate. For synaptic protein quantitation, synaptosomes isolated as described above were used. Lysate samples were loaded into each well and subjected to SDS-PAGE electrophoresis. The proteins were then transferred onto PVDF membranes, blocked with 5% milk, and incubated overnight with primary antibodies followed by 1-h incubation with HRP-conjugated secondary antibodies. The protein signals were detected using super signal west pico chemiluminescent substrate (Pierce, USA). Quantification of Western blot signals was done using ImageJ software.
Quantitation of ADAM10 Enzyme Activity
To quantitate ADAM10 activity, we used a fluorometric SensoLyte® 520 ADAM10 activity assay kit from AnaSpec (cat # AS-72226) and followed their recommended protocol. Briefly, working solutions of ADAM10 substrate, recombinant ADAM10 enzyme, and inhibitors were prepared as per their recommended protocol. Lysates from HeLa and HeLa-EGFP cells were prepared using assay buffer with additional protease inhibitor cocktail. Similarly, mice were euthanized, and brain regions such as cortex and hippocampus were rapidly removed and homogenized using cold assay buffer, centrifuged for 15 min at 10,000×g at 4 °C, and 25 uL of supernatants was added to the plate in a total volume of 100 μL reaction. For endpoint reading, the reactions were incubated at 37 °C for 1 h and read at excitation 490 nm and emission at 520 nm. The provided 5-FAM fluorescence reference standard was used to plot the standard curve as relative fluorescence units (RFU), and the concentration of the final product generated from the brain samples was deduced from the standard graph.
Statistical Analysis
ImageJ software was used to quantify the protein bands from the immunoblots. Actin levels were used for normalization of proteins of interest to reflect true changes. Statistical analysis was performed by using Instat3 software (GraphPad Software, San Diego, CA, USA). In those experiments where only two groups were involved (ADAM10 protein levels in the cortex and hippocampal brain regions, HeLa cells), the student’s t test was used assuming values are sampled from Gaussian distribution and using two-tail p value. In experiments involving more than two groups (HeLa cells exposed to PPAR antagonists), either ordinary ANOVA or repeated measures one-way analysis of variance (ANOVA) as standard parametric methods was used followed by post hoc Dunnett’s multiple comparison test or Student-Newman-Keuls multiple comparisons test for comparisons among the different types of PPAR antagonists. The data presented are the mean + standard deviation (SD). The data were considered significant only if p < 0.05. * indicates p < 0.05, ** p < 0.01, and *** p < 0.001.
Data Availability
All data are contained within the manuscript.
Abbreviations
- AD:
-
Alzheimer’s disease
- ADAM10:
-
ADAM metallopeptidase domain 10
- ALP:
-
Autophagy-lysosome pathway
- APP:
-
Amyloid precursor protein
- BAF1:
-
Bafilomycin A1
- CRISPR:
-
Clustered regularly interspaced short palindromic repeats
- LC3:
-
Microtubule-associated protein 1A/1B-light chain 3
- 3-MA:
-
3-Methyl adenine
- PPARα:
-
Peroxisome proliferator-activated receptor-alpha
- PPRE:
-
Peroxisome proliferator response element
- SAP97:
-
Synapse-associated protein 97
- sAPPα:
-
Soluble amyloid precursor protein alpha
- TFEB:
-
Transcription factor EB
- UPS:
-
Ubiquitin-proteasome system
References
Sardiello M, Palmieri M, di Ronza A, Medina DL, Valenza M, Gennarino VA, Di Malt AC, Donaudy F et al (2009) A gene network regulating lysosomal biogenesis and function. Science 325:473–477
Settembre C, Di Malta C, Polito VA, Garcia Arencibia M, Vetrini F, Erdin S, Erdin SU, Huynh T et al (2011) TFEB links autophagy to lysosomal biogenesis. Science 332:1429–1433
Palmieri M, Impey S, Kang H, di Ronza A, Pelz C, Sardiello M, Ballabio A (2011) Characterization of the CLEAR network reveals an integrated control of cellular clearance pathways. Hum Mol Genet 20:3852–3866
Xiao Q, Yan P, Ma X, Liu H, Perez R, Zhu A, Gonzales E, Tripoli DL et al (2015) Neuronal-targeted TFEB accelerates lysosomal degradation of APP, reducing Aβ generation and amyloid plaque pathogenesis. J Neurosci 35:12137–12151
Bao J, Zheng L, Zhang Q, Li X, Zhang X, Li Z, Bai X, Zhang Z et al (2014) Deacetylation of TFEB promotes fibrillar Aβ degradation by upregulating lysosomal biogenesis in microglia. Protein Cell 7:417–433
Zhang YD, Zhao JJ (2015) TFEB participates in the Aβ-induced pathogenesis of Alzheimer's disease by regulating the autophagy-lysosome pathway. DNA Cell Biol 34:661–668
Polito VA, Li H, Martini-Stoica H, Wang B, Yang L, Xu Y, Swartzlander DB, Palmieri M et al (2014) Selective clearance of aberrant tau proteins and rescue of neurotoxicity by transcription factor EB. EMBO Mol Med 6:1142–1160
Kim S, Choi KJ, Cho SJ, Yun SM, Jeon JP, Koh YH, Song J, Johnson GV et al (2016) Fisetin stimulates autophagic degradation of phosphorylated tau via the activation of TFEB and Nrf2 transcription factors. Sci Rep 6:24933
Kilpatrick K, Zeng Y, Hancock T, Segatori L (2015) Genetic and chemical activation of TFEB mediates clearance of aggregated α-synuclein. PLoS One 10:e0120819
Ebrahimi-Fakhari D, Wahlster L (2013) Restoring impaired protein metabolism in Parkinson's disease--TFEB-mediated autophagy as a novel therapeutic target. Mov Disord 28:1346
Tsunemi T et al (2012) PGC-1alpha rescues Huntington's disease proteotoxicity by preventing oxidative stress and promoting TFEB function. Sci Transl Med 4:142ra197
Vodicka P, Chase K, Iuliano M, Tousley A, Valentine DT, Sapp E, Kegel-Gleason KB, Sena-Esteves M et al (2016) Autophagy activation by transcription factor EB (TFEB) in striatum of HDQ175/Q7 mice. J Huntington’s Dis 5:249–260
Wang H, Wang R, Carrera I, Xu S, Lakshmana MK (2016) TFEB Overexpression in the P301S model of tauopathy mitigates increased PHF1 levels and lipofuscin puncta and rescues memory deficits. eNeuro 23:ENEURO.0042–16.2016
Ballabio A (2016) The awesome lysosome. EMBO Mol Med 8:73–76
Wang H, Wang R, Xu S, Lakshmana MK (2016) Transcription factor EB is selectively reduced in the nuclear fractions of Alzheimer's and amyotrophic lateral sclerosis brains. Neurosci J 2016:4732837
Tiribuzi R, Crispoltoni L, Porcellati S et al (2014) MiR128 up-regulation correlates with impaired amyloid β(1-42) degradation in monocytes from patients with sporadic Alzheimer's disease. Neurobiol Aging 35:345–356
Decressac M et al (2013) TFEB-mediated autophagy rescues midbrain dopamine neurons from alpha-synuclein toxicity. Proc Natl Acad Sci U S A 110:E1817–E1826
Chen Y, Liu H, Guan Y, Wang Q, Zhou F, Jie L, Ju J, Pu L et al (2015) The altered autophagy mediated by TFEB in animal and cell models of amyotrophic lateral sclerosis. Am J Transl Res 7:1574–1587
La Spada AR (2012) PPARGC1A/PGC-1alpha, TFEB and enhanced proteostasis in Huntington disease: defining regulatory linkages between energy production and protein-organelle quality control. Autophagy 8:1845–1847
Tsunemi T et al (2012) PGC-1rescues Huntington’s disease proteotoxicity by preventing oxidative stress and promoting TFEB function. Sci Transl Med 4:142–197
Cortes CJ et al (2014) Polyglutamine-expanded androgen receptor interferes with TFEB to elicit autophagy defects in SBMA. Nat Neurosci 17:1180–1189
Lapierre LR, De Magalhaes Filho CD, McQuary PR, Chu CC, Visvikis O, Chang JT, Gelino S, Ong B et al (2013) The TFEB orthologue HLH-30 regulates autophagy and modulates longevity in Caenorhabditis elegans. Nat Commun 4:2267
Nakamura S, Karalay Ö, Jäger PS, Horikawa M, Klein C, Nakamura K, Latza C, Templer SE et al (2016) Mondo complexes regulate TFEB via TOR inhibition to promote longevity in response to gonadal signals. Nat Commun 7:10944
Kuhn PH, Wang H, Dislich B, Colombo A, Zeitschel U, Ellwart JW, Kremmer E, Rossner S et al (2010) ADAM10 is the physiologically relevant, constitutive alpha-secretase of the amyloid precursor protein in primary neurons. EMBO J 29:3020–3032
Lichtenthaler SF (2011) α-Secretase in Alzheimer's disease: molecular identity, regulation, and therapeutic potential. J Neurochem 116:10–21
Jonsson T, Atwal JK, Steinberg S, Snaedal J, Jonsson PV, Bjornsson S, Stefansson H, Sulem P et al (2012) A mutation in APP protects against Alzheimer’s disease and age-related cognitive decline. Nature 488:96–99
Kaden D, Harmeier A, Weise C, Munter LM, Althoff V, Rost BR, Hildebrand PW, Schmitz D et al (2012) Novel APP/Aβ mutation K16N produces highly toxic heteromeric Ab oligomers. EMBO Mol Med 4:647–659
Suh J, Choi SH, Romano DM, Gannon MA, Lesinski AN, Kim DY, Tanzi RE (2013) ADAM10 missense mutations potentiate β-amyloid accumulation by impairing the prodomain chaperone function. Neuron 80:385–401
Jorissen E, Prox J, Bernreuther C, Weber S, Schwanbeck R, Serneels L, Snellinx A, Craessaerts K et al (2010) The disintegrin/metalloproteinase ADAM10 is essential for the establishment of the brain cortex. J Neurosci 30:4833–4844
Suh J, Choi SH, Romano DM, Gannon MA, Lesinski AN, Kim DY, Tanzi RE (2013) ADAM10 missense mutations potentiate beta-amyloid accumulation by impairing the prodomain chaperone function. Neuron 80:385–401
Clement AB, Hanstein R, Schröder A, Nagel H, Endres K, Fahrenholz F, Behl C (2008) Effects of neuron-specific ADAM10 modulation in an in vivo model of acute excitotoxic stress. Neuroscience. 152:459–468
Chen YY, Hehr CL, Atkinson-Leadbeater K, Hocking JC, McFarlane S (2007) Targeting of retinal axons requires the metalloproteinase ADAM10. J Neurosci 27:8448–8456
Prox J, Bernreuther C, Altmeppen H, Grendel J, Glatzel M, D’Hooge R, Stroobants S, Ahmed T et al (2013) Postnatal disruption of the disintegrin/metalloproteinase ADAM10 in brain causes epileptic seizures, learning deficits, altered spine morphology, and defective synaptic functions. J Neurosci 33:12915–12928
Postina R, Schroeder A, Dewachter I, Bohl J, Schmitt U, Kojro E, Prinzen C, Endres K et al (2004) A disintegrin-metalloproteinase prevents amyloid plaque formation and hippocampal defects in an Alzheimer disease mouse model. J Clin Invest 113:1456–1464
Manzine PR, Barham EJ, Vale Fde A, Selistre-de-Araújo HS, Iost Pavarini SC, Cominetti MR (2013) Correlation between mini-mental state examination and platelet ADAM10 expression in Alzheimer's disease. J Alzheimers Dis 36:253–260
Epis R et al (2010) Blocking ADAM10 synaptic trafficking generates a model of sporadic Alzheimer’s disease. Brain 133:3323–3335
Marcello E, Borroni B, Pelucchi S, Gardoni F, Di Luca M (2017) ADAM10 as a therapeutic target for brain diseases: From developmental disorders to Alzheimer's disease. Expert Opin Ther Targets 21:1017–1026
Blommaart EFC, Krause U, Schellens JPM, Vreeling-Sindelárová H, Meijer AJ (1997) The phosphatidylinositol 3-kinase inhibitors wortmannin and LY294002 inhibit autophagy in isolated rat hepatocytes. Eur J Biochem 243:240–246
Mauvezin C, Neufeld TP (2015) Bafilomycin A1 disrupts autophagic flux by inhibiting both V-ATPase-dependent acidification and Ca-P60A/SERCAdependentautophagosome-lysosome fusion. Autophagy 11:1437–1438
Eun SY, Lee JN, Nam IK, Liu ZQ, So HS, Choe SK, Park R (2018) PEX5 regulates autophagy via the mTORC1-TFEB axis during starvation. Exp Mol Med 50:4
Issemann I, Green S (1990) Activation of a member of the steroid hormone receptor superfamily by peroxisome proliferators. Nature 347:645–650
Luo R, Su LY, Li G, Yang J, Liu Q, Yang LX, Zhang DF, Zhou H et al (2020) Activation of PPARA-mediated autophagy reduces Alzheimer disease-like pathology and cognitive decline in a murine model. Autophagy. 16:52–69
Lundgren JL, Ahmed S, Schedin-Weiss S, Gouras GK, Winblad B, Tjernberg LO, Frykman S (2015) ADAM10 and BACE1 are localized to synaptic vesicles. J Neurochem 135:606–615
Malinverno M, Cart AM, Epis R, Marcello E, Verpelli C, Cattabeni F, Sala C, Mulle C et al (2010) Synaptic localization and activity of ADAM10 regulate excitatory synapses through N-cadherin cleavage. J Neurosci 30:16343–16355
Wang H, Lewsadder M, Dorn E, Xu S, Lakshmana MK (2014) RanBP9 overexpression reduces dendritic arbor and spine density. Neuroscience. 265:253–262
Palavicini JP, Wang H, Bianchi E, Xu S, Rao JS, Kang DE, Lakshman MK (2013) RanBP9 aggravates synaptic damage in the mouse brain and is inversely correlated to spinophilin levels in Alzheimer's brain synaptosomes. Cell Death Dis 4:e667
Postina R, Schroeder A, Dewachter I et al (2004) A disintegrinmetalloproteinase prevents amyloid plaque formation and hippocampal defects in an Alzheimer disease mouse model. J Clin Invest 113:1456–1464
Kuhn P-H, Wang H, Dislich B, Colombo A, Zeitschel U, Ellwart JW, Kremmer E, Rossner S et al (2010) ADAM10 is the physiologically relevant, constitutive alpha secretase of the amyloid precursor protein in primary neurons. EMBO J 29:3020–3032
Fahrenholz F (2007) Alpha-secretase as a therapeutic target. Curr Alzheimer Res 4:412–417
Lichtenthaler SF (2011) Alpha-secretase in Alzheimer's disease: molecular identity, regulation and therapeutic potential. J Neurochem 116:10–21
Bandyopadhyay S, Goldstein LE, Lahiri DK, Rogers JT (2007) Role of the APP non-amyloidogenic signaling pathway and targeting alpha-secretase as an alternative drug target for treatment of Alzheimer's disease. Curr Med Chem 14:2848–2864
Kögel D, Deller T, Behl C (2012) Roles of amyloid precursor protein family members in neuroprotection, stress signaling and aging. Exp Brain Res 217:471–479
Colciaghi F, Marcello E, Borroni B, Zimmermann M, Caltagirone C, Cattabeni F, Padovani A, Di Luca M (2004) Platelet APP, ADAM 10 and BACE alterations in the early stages of Alzheimer disease. Neurology. 62:498–501
Marcello E, Saraceno C, Musardo S, Vara H, de la Fuente AG, Pelucchi S, Di Marino D, Borroni B et al (2013) Endocytosis of synaptic ADAM10 in neuronal plasticity and Alzheimer’s disease. J Clin Invest 123:2523–2538
Colciaghi F, Borroni B, Pastorino L, Marcello E, Zimmermann M, Cattabeni F, Padovani A, Di Luca M (2002) [alpha]-Secretase ADAM10 as well as [alpha]APPs is reduced in platelets and CSF of Alzheimer disease patients. Mol Med 8:67–74
Petiot A, Ogier-Denis E, Blommaart EF, Meijer AJ, Codogno P (2000) J Biol Chem 275:992–998
Keller MD, Ching KL, Liang FX, Dhabaria A, Tam K, Ueberheide BM, Unutmaz D, Torres VJ et al (2020) Decoy exosomes provide protection against bacterial toxins. Nature 579:260–264
Zheng G, Zhan Y, Li X, Pan Z, Zheng F, Zhang Z, Zhou Y, Wu Y et al (2018) TFEB, a potential therapeutic target for osteoarthritis via autophagy regulation. Cell Death Dis 9:858
Pan B, Zhang H, Cui T, Wang X (2017) TFEB activation protects against cardiac proteotoxicity via increasing autophagic flux. J Mol Cell Cardiol 113:51–62
Wu YT, Tan HL, Shui G, Bauvy C, Huang Q, Wenk MR, Ong CN, Codogno P et al (2010) Dual role of 3-methyladenine in modulation of autophagy via different temporal patterns of inhibition on class I and III phosphoinositide 3-kinase. J Biol Chem 285:10850–10861
Séité S, Pioche T, Ory N, Plagnes-Juan E, Panserat S, Seiliez I (2019) The autophagic flux inhibitor bafilomycine A1 affects the expression of intermediary metabolism-related genes in trout hepatocytes. Front Physiol 10:263
Klionsky DJ, Abdelmohsen K, Abe A, Abedin MJ, Abeliovich H, Acevedo Arozena A et al (2016) Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 12:1–222
Mizushima N (2004) Methods for monitoring autophagy. Int J Biochem Cell Biol 36:2491–2502
Mizushima N, Yoshimori T (2007) How to interpret LC3 immunoblotting. Autophagy 3:542–545
Bjørkøy G, Lamark T, Brech A, Outzen H, Perander M, Overvatn A, Stenmark H, Johansen T (2005) p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J Cell Biol 171:603–614
Settembre C, De Cegli R, Mansueto G, Saha PK, Vetrini F, Visvikis O, Huynh T, Carissimo A et al (2013) Tfeb controls cellular lipid metabolism through a starvation-induced autoregulatory loop. Nat Cell Biol 15:647–658
Corbett GT, Gonzalez FJ, Pahan K (2015) Activation of peroxisome proliferator-activated receptor α stimulates ADAM10-mediated proteolysis of APP. Proc Natl Acad Sci U S A 112:8445–8450
Ghosh A, Jana M, Modi K, Gonzalez FJ, Sims KB, Berry-Kravis E, Pahan K (2015) Activation of peroxisome proliferator-activated receptor alpha induces lysosomal biogenesis in brain cells: Implications for lysosomal storage disorders. J Biol Chem 290:10309–10324
Terry RD, Masliah E, Salmon DP, Butters N, DeTeresa R, Hill R et al (1991) Physical basis of cognitive alterations in Alzheimer's disease: synapse loss is the major correlate of cognitive impairment. Ann Neurol 30:572–580
DeKosky S, Scheff S (1990) Synapse loss in frontal cortex biopsies in Alzheimer's disease: Correlation with cognitive severity. Ann Neurol 27:457–464
Manzine PR, Barham EJ, Vale FA, Selistre-De-Araujo HS, Pavarini SC, Cominetti MR (2014) Platelet a disintegrin and metallopeptidase 10 expression correlates with clock drawing test scores in Alzheimer’s disease. Int J Geriatr Psychiatry 29:414–420
Brummer T, Pigoni M, Rossello A et al (2018) The metalloprotease ADAM10 (a disintegrin and metalloprotease 10) undergoes rapid, postlysis autocatalytic degradation. FASEB J 32(7):3560–3573
Kern A, Roempp B, Prager K, Walter J, Behl C (2006) Down-regulation of endogenous amyloid precursor protein processing due to cellular aging. J Biol Chem 281:2405–2413
Peters-Libeu C, Jesus C, Mitsumori M, Poksay K, Spilman P et al (2015) sAßPPα is a potent endogenous inhibitor of BACE 1. J Alzheimer’s Dis 47:545–555
Obregon D, Hou H, Deng J, Giunta B, Tian J, Darlington D, Shahaduzzaman M, Zhu Y et al (2012) Soluble amyloid precursor protein-alpha modulates beta-secretase activity and amyloid-beta generation. Nat Commun 3:777
Acknowledgments
We want to thank the staff at the Florida International University (FIU) vivarium for taking care of the mouse colonies.
Funding
This work was supported by the National Institute of Aging (NIA), National Institute of Health (NIH) to MKL through grant # 1R21AG060299.
Author information
Authors and Affiliations
Contributions
Lakshmana MK and Nair M designed the research; Wang H. Lakshmana MK and Mohan Kumar MK formed the research and analyzed the data; Nair M contributed to reagents, and Lakshmana MK and Nair M wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflicts of interest.
Ethics Statement
All animal procedures were approved by an Institutional Animal Care and Use Committee (IACUC) of Florida International University (FIU).
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 146 kb)
Rights and permissions
About this article
Cite this article
Wang, H., Muthu Karuppan, M.K., Nair, M. et al. Autophagy-Dependent Increased ADAM10 Mature Protein Induced by TFEB Overexpression Is Mediated Through PPARα. Mol Neurobiol 58, 2269–2283 (2021). https://doi.org/10.1007/s12035-020-02230-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-020-02230-8