Abstract
Local control of protein translation is a fundamental process for the regulation of synaptic plasticity. It has been demonstrated that the local protein synthesis occurring in axons and dendrites can be shaped by numerous mechanisms, including miRNA-mediated regulation. However, several aspects underlying this regulatory process have not been elucidated yet. Here, we analyze the differential miRNA profile in cell bodies and neurites of primary hippocampal neurons and find an enrichment of the precursor and mature forms of miR-218 in the neuritic projections. We show that miR-218 abundance is regulated during hippocampal development and by chronic silencing or activation of neuronal network. Overexpression and knockdown of miR-218 demonstrated that miR-218 targets the mRNA encoding the GluA2 subunit of AMPA receptors and modulates its expression. At the functional level, miR-218 overexpression increases glutamatergic synaptic transmission at both single neuron and network levels. Our data demonstrate that miR-218 may play a key role in the regulation of AMPA-mediated excitatory transmission and in the homeostatic regulation of synaptic plasticity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The mammalian brain constantly undergoes structural and functional modifications while interacting with the external environment. The brain architecture can be modified by adjusting the number and the strength of the synapses, a process called synaptic plasticity. Fine-tuning of protein turnover plays an essential role to ensure the steady-state level of specific proteins and to adjust the proteome to external signals [1, 2].
A number of studies have revealed that the control of protein synthesis and degradation is strictly temporally and spatially regulated [3]. Cumulative evidence has shown that mRNAs can be targeted and translated in specific subcellular areas to create functionally distinct domains independent of the perinuclear control of mRNA translation [4,5,6]. mRNA targeting and local translation in dendrites and axons are particularly important for highly polarized cells as neurons, because they avoid the ectopic expression of an active protein while it is transported from the soma to neuronal processes and offer the possibility of a quick “on-demand” local protein synthesis in response to external cues that affect growth, synapse formation, and plasticity [1, 7].
Numerous studies have shown that a large fraction of the total transcriptome can be detected in dendrites, where the machineries for both protein synthesis and protein degradation are also present, including polyribosomes, ribosome binding proteins (RBPs), components of the ubiquitin proteasome system, and regulatory RNAs, such as microRNAs (miRNAs) [8,9,10,11,12,13].
miRNAs are a large family of non-coding small RNAs that have emerged as key posttranscriptional regulators of gene expression [14]. The evolutionarily conserved mechanism which gives rise to mature miRNAs involves two ordered endonucleolytic cleavages by the RNase III enzymes Drosha and Dicer. Following transcription, Drosha processes the primary miRNA transcript (pri-miRNA) into a ~ 60–100 nt hairpin structure termed precursor-miRNA (pre-miRNA) [15, 16]. Following translocation to the cytoplasm, the pre-miRNA is cleaved by Dicer to generate a ~ 22-nt double-stranded miRNA that is incorporated into the miRNA-induced silence complex RISC [17].
A number of studies have revealed that miRNAs are actively involved in the regulation of the expression of synaptic proteins at the cell body level and miRNAs themselves are regulated by synaptic plasticity [18]. Moreover, miRNAs have been shown to regulate local protein synthesis at, or near, synapses [19,20,21]. Interestingly, mature miRNAs, pre-miRNAs, and enzymes for miRNA biogenesis can be localized to dendritic spines and the enrichment of miRNAs in axons, in dendrites, or in synaptosomes isolated from specific brain areas has been reported [22,23,24,25]. However, several aspects of the local regulation of the synaptic proteome and synaptic function by miRNAs have not been elucidated yet.
In this study, we investigated the expression of miRNAs and pre-miRNAs primarily enriched in axons and dendrites and characterized their role in the regulation of local translation. To define the neuritic miRNA profile, we mechanically separated neuronal cell bodies from processes and identified a subset of miRNAs enriched in neuronal processes, including miR-218. By using computational algorithms to predict the physiological synaptic mRNA targets, we found that miR-218, among its several synaptic mRNA targets, is directed to a conserved miRNA response element (MRE) in the 3′UTR of the GRIA2 gene that encodes the GluA2 subunit of AMPA-type glutamate receptors, whose mRNA is targeted to dendrites [26]. Indeed, overexpression of miR-218 in primary neurons was followed by upregulation of GluA2 and a corresponding boosting of excitatory synaptic transmission at both single neuron and network levels. Moreover, in primary neurons, chronic deprivation of electrical activity increased, and chronic hyperactivity decreased, miR-218 expression, revealing a function for miR-218 in homeostatic synaptic responses. miR-218 thus emerges as a novel candidate for miRNA-dependent regulation of synaptic strength and plasticity.
Materials and Methods
Cell Culture Preparations
All experiments were carried out in accordance with the guidelines established by the European Community Council (Directive 2010/63/EU of 22 September 2010) and were approved by the Italian Ministry of Health. Primary hippocampal cultures were prepared from wild-type C57 Bl6/J mice (Charles River). All efforts were made to minimize suffering and to reduce the number of animals used. Mice were sacrificed by CO2 inhalation, and 18-day embryos (E18) were removed immediately by cesarean section. Briefly, enzymatically dissociated hippocampal neurons were plated on 0.4 μm pore PET membrane filter inserts (Corning) or standard glass coated with polyethyleneimine (PEI) (0.1% in 50 mM sodium borate buffer pH 7.4) (Sigma). Cells were plated at a density of 50,000 cells/cm2 in Neurobasal-Media supplemented with 2% B-27 supplement, 2 mM glutamine, and antibiotics (Life Technologies). Cells were allowed to grow at 37 °C, 5% CO2, and 95% humidity for 14 days before harvesting. Neurons were treated with bicuculline (BiC, 30 μM, Tocris), 4-aminopyridine (4-AP, 100 μM Sigma), and tetrodotoxin (TTX, 1 μM Tocris) for 48 h before harvesting.
HEK293T were routinely cultured in IMDM, supplemented with 100 units/ml penicillin, 100 μg/ml streptomycin (Invitrogen), glutamine, and 10% heat-inactivated FCS (Invitrogen). Cells were maintained at 37 °C in a humidified 5% CO2 atmosphere.
Cell Transfection and Transduction
The mouse GluA2 3′UTR was amplified from mouse hippocampal cDNA. The UTR fragment was then inserted into pGL3 promoter vector in which firefly luciferase expression is driven through 3′UTR GluA2 sequence (Promega). Neurons were transfected through electroporation using Mouse Neuron Nucleofector Kit and an Amaxa Nucleofector Device (program 0-003, Lonza) before plating. For luciferase assay experiments, neurons were transfected with 3′UTR GluA2-pGL3 promoter vector together with miR-218, miR-124 mimics, miR-218 inhibitor, and scrambled sequences (indicated in text as Scr) (Ambion) (Supplemental Table S1). The pRL-TK vector expressing Renilla luciferase was co-transfected to normalize for transfection efficiency (Promega). Forty-eight hours after transfection, the luciferase activity was measured using a Dual-Luciferase Assay Kit (Promega) according to the manufacturer’s instructions. Luciferase measurements were performed with a Victor3-V luminometer (PerkinElmer). The results were expressed as the ratio of pGL3 promoter vector over pRL-TK. A version with a mutated miR-218 or mutated miR-124 MRE within the GluA2 3′UTR was generated with the QuikChange site-directed mutagenesis kit (Stratagene) using overlapping PCR (see Supplemental Table S1 for primer sequences). The production of VSV-pseudotyped third-generation lentiviruses was performed as previously described [27]. Lentiviral vectors expressing miR-218, miR-218 sponge (indicated as anti-miR-218), or scrambled sequence were obtained by cloning human U6 promoter and small hairpin RNA that overexpress miR-218 (miR-218, TTGTGCTTGATCTAACCATGT), inhibit miR-218 (ACATGGTTAGATCAAGCACAA), or scrambled sequences (Scr, GCAGTTATCACGTCTATGTTT) into FG12 plasmid (Addgene). The expression levels were assessed by GFP fluorescence. Primary hippocampal neurons were infected at 7 DIV at 10 multiplicity of infection. After 24 h, half of the medium was replaced with fresh medium. All experiments were performed between 14 and 16 DIV.
Sample Collection and RNA Extraction
Media were aspirated and cell processes were washed in PBS and rapidly separated from their cell bodies by scraping the bottom (neuritic fraction) and upper surface (somatic fraction) with a rubber cell scraper in Trizol reagent (Life Technologies). Hippocampal tissues were obtained from C57 Bl6/J mice at the indicated ages and homogenized in Trizol reagent (Life Technologies). Total RNA was extracted according to manufacturer’s instructions.
Taqman Low-Density Array
Taqman low-density array (TLDA) was performed using TaqMan® Array Rodent MicroRNA A + B Cards Set v2.0 (Applied Biosystems, USA). Samples used were somatic and neuritic fractions. Briefly, reverse transcription reaction was performed using rodent Megaplex™ RT primers (Applied Biosystems), which contain a pool of 750 individual miRNA-specific primers, according to the manufacturer’s instructions. Real-time quantitative PCR (RT-qPCR) was then carried out on an ABI 7900HT real-time PCR machine with the microfluidic card block, using predefined TLDA thermal cycling conditions. RT-qPCR data were analyzed using SDS 2.3 and RQ Manager Software (Applied Biosystems). To determine an accurate normalization factor for data analysis, we evaluated the expression stability of various control genes with the GeNorm algorithm available in qBasePlus software (Biogazelle) [28]. miR-877 and miR-719 were selected as reference genes. Heat map was generated using MeV TM4 software V 4.9 (http://www.tm4.org).
Real-Time PCR Analysis
Total RNA was retro-transcribed using the miScript SYBR Green PCR Kit (Qiagen) and the SuperScript III First-Strand Synthesis System for mRNAs (Life Technologies) according to the manufacturer’s instructions. Reactions were run on an ABI 7900HT real-time PCR machine. Relative gene expression was determined using the 2−ΔΔCT method [28]. GAPDH and HPRT1 were used as reference genes. The list of primers used is provided in Supplemental Table 1.
miRNA Target Prediction
Three software packages were applied for miRNA target prediction: Pita [29], TargetScan [30], and PicTar [31]. We considered only targets predicted by all three algorithms. Gene ontology analysis of miR-218 putative target genes was performed using David software (https://david.ncifcrf.gov/).
Western Bloting Analysis
Cells were harvested and lysed in RIPA buffer (50 mM Tris-Cl pH 7.4, 150 mM NaCl, 2 mM EDTA, NP-40 1%, and SDS 0.1%) supplemented with protease inhibitors (complete EDTA-free protease inhibitors, Roche Diagnostic). Equal amounts of proteins were separated by SDS-PAGE and blotted onto nitrocellulose membranes (GE Healthcare). The following primary antibodies were used: polyclonal rabbit anti-GluA2 C-terminal (AGC-005, Alomone); rabbit anti-GAPDH (2118, Cell Signaling); rabbit anti-actin (A2066, Sigma). Secondary antibodies were detected by chemiluminescence with the ECL Western Blotting Detection System (GE Healthcare). Densitometric analysis was performed with ImageJ software.
Immunofluorescence Staining
Hippocampal neurons were fixed in phosphate-buffered saline (PBS)/4% paraformaldehyde for 20 min at RT. Cells were permeabilized with 1% Triton X-100 for 5 min, blocked with 2% bovine serum albumin in PBS/Tween 20 0.05% for 30 min at RT, and incubated with primary antibodies in the same buffer for 45 min. The primary antibodies used were mouse anti-synapsin I (clone 10.22, Millipore) and rabbit anti-GluA2 C-terminal (AGC-005, Alomone). After the primary incubation, neurons were incubated for 45 min with the secondary antibodies in blocking buffer. Fluorescently conjugated secondary antibodies were from Molecular Probes (Thermo-Fisher Scientific; Alexa Fluor 568, A11036; Alexa Fluor 647, A21450). Image acquisitions were performed using a confocal microscope (SP8, Leica Microsystems GmbH, Wetzlar, Germany) at × 63 (1.4 NA) magnification. Z-stacks were acquired every 300 nm; 10 fields/sample. Offline analysis was performed using the ImageJ software and the JACoP plugin for co-localization studies. For each set of experiments, all images were acquired using identical exposure settings. Values were normalized to the relative cell volume calculated on the basis of GFP reporter gene intensity.
Miniature Excitatory Postsynaptic Currents
Whole-cell patch-clamp recordings of AMPA-mediated miniature excitatory postsynaptic currents (mEPSCs) were obtained in the presence of tetrodotoxin (TTX, 1 μM). Patch pipettes, prepared from thin borosilicate glass (Hilgenberg), were pulled and fire-polished to a final resistance of 4–5 MΩ when filled with standard internal solution. Neurons were infected at 7 DIV and recorded at 14–18 DIV. mEPSCs were recorded from cultured pyramidal neurons, morphologically identified by their teardrop-shaped somata and characteristic apical dendrite, using an Axon Multiclamp 700B/Digidata1440A system (Molecular Devices) and an upright BX51WI microscope (Olympus). Cells were maintained in a standard external solution (Tyrode) containing (in mM) 140 NaCl, 2 CaCl2, 1 MgCl2, 4 KCl, 10 glucose, and 10 HEPES (pH 7.3 with NaOH). Unless otherwise indicated, D-(−)-2-amino-5-phosphonopentanoic acid (D-APV; 50 μM; Tocris), bicuculline methiodide (30 μM, Tocris) and ((2S)-3-[[(1S)-1-(3,4-dichlorophenyl) ethyl]amino-2-hydroxypropyl] (phenylmethyl) phosphinic acid hydrochloride; CGP58845 hydrochloride; 5 μM), and TTX (300 nM) were added to the Tyrode external solution to block N-methyl-D-aspartate (NMDA) GABAARs, GABABRs, and voltage-gated Na+ channels, respectively. The standard internal solution contained (in mM) 126 K-gluconate, 4 NaCl, 1 MgSO4, 0.02 CaCl2, 0.1 BAPTA, 15 glucose, 5 HEPES, 3 ATP, and 0.1 GTP (pH 7.2 with KOH). Experiments were performed at 22–24 °C, and miniature EPSCs were acquired at 20 kHz sample frequency and filtered at half the acquisition rate with an 8-pole low-pass Bessel filter. Recordings with leak currents > 100 pA or access resistance > 20 MΩ were discarded. The EPSCs analysis was performed using the Minianalysis program (Synaptosoft, Leonia, NJ) and the Prism software (GraphPad Software, Inc.). The amplitude and frequency of mEPSCs were calculated using a peak detector function with a threshold amplitude set at 4 pA and a threshold area at 50 ms*pA.
Multielectrode Array Electrophysiology
Hippocampal neurons were plated onto 12-well MEA plates (768-GL1-30Au200 from Axion BioSystems). Plates were incubated overnight with poly-L-lysine (0.1 mg/ml; Sigma-Aldrich), washed three times with distilled water, dried, and coated with laminin for 2 h. Dissociated hippocampal neurons were plated at a final density of 80,000 neurons per well and maintained at 37 °C in 5% CO2. Spontaneous network activity was recorded for 15 min at 14 DIV using the Axion Biosystems Maestro MEA at 37 °C at 12.5 kHz sample frequency. A Butterworth band-pass filter (200–3000 Hz) and an adaptive threshold spike detector set at 7× the standard deviation of the noise for each channel were used during recordings. No spike sorting procedure was carried out. Electrodes were considered as active if they had at least 5 spike/min [32]. Data were analyzed using NeuralMetric Tool 2.0.4 (Axion Biosystems). Bursts were detected using an inter-spike interval (ISI) threshold algorithm for each channel [33]. The algorithm relies on defining a burst as a collection of at least 5 spikes, each separated by an ISI ≤ 100 ms (ISI max). Network bursts were identified as a minimum number of 10 spikes on at least 25% of the electrodes with an ISI max of 100 ms were detected. The following parameters were used to describe the electrical activity: firing rate normalized to the number of active electrodes (weighted firing rate), overall bursting rate, bursting rate/electrode, burst duration, network burst rate, and number of spikes/burst. Acute application of 6-cyano-7-nitroquinoxaline-2,3-dione (CNQX, 20 μM Tocris) was performed to assess the specificity of the effects for glutamatergic transmission.
Data Analysis
Results are presented as means ± sem from four independent preparations. Normal distribution of data was assessed using the D’Agostino-Pearson’s normality test. To compare two normally distributed sample groups, the unpaired Student’s t test was used. To compare more than two normally distributed sample groups, one-way ANOVA was used, followed by the post hoc tests (Bonferroni’s test). A value of p < 0.05 was considered significant. Statistical analysis was carried out using SigmaStat 13 (Systat Software).
Results
Physically Separated Cell Bodies and Processes of Cultured Hippocampal Neurons Display a Differential miRNA Profile
To identify miRNAs putatively involved in synaptic function, we isolated miRNAs enriched in neuronal processes, where they could regulate the local translation of crucial mRNA targets. To define the neuritic miRNA profile, we used a mechanical approach to separate neuronal cell bodies from their processes [34, 35]. Dissociated hippocampal neurons were grown for 14 days on polyethylene tetraphthalate (PET) membranes etched with 0.4 μm pores. While neuronal cell bodies were constrained on the top surface of the filters, neuronal processes (including axons and dendrites; hereafter indicated as neuritic fraction) could extend through the pores and grow on the bottom surface of the membrane (Fig. 1a). To evaluate the purity of somatic and neuritic fractions, we analyzed transcripts that are selectively enriched in either compartment by qRT-PCR analysis. Somatically restricted transcripts, such as U6 small nuclear RNA (Rnu6) and glyceraldehyde-3-phosphate dehydrogenase (GAPDH), were enriched in the somatic fraction, but not in the neuritic counterpart [36,37,38]. Conversely, dendrite-specific transcripts, such as microtubule-associated protein 2 (MAP2) and eukaryotic translation elongation factor 1 alpha 1 (EEF1A1), were enriched only in the neuritic fraction, confirming the lack of somatic contamination during sample preparation (Fig. 1b) [39].
Neuronal cell bodies and processes of primary hippocampal neurons display a differential miRNA profile. a Schematic representation of the transwell system used for the mechanical separation of neurites and somatic fractions in primary neuronal cultures. A transwell insert with a base made of microporous membrane (pore size 0.4 μm) separates the well into upper and lower compartments. b Real-time PCR analysis of the transcript levels of U6, GAPDH, MAP2, and EEF1A1 in somatic and neuritic fractions. CaMKIIα was used as reference gene. Graph shows means ± sem (n = 4). c Heatmap shows differentially expressed miRNAs between somatic and neuritic fractions in primary hippocampal neurons grown on porous membranes at 14 DIV. A pseudocolor scale represents the level of miRNA expression (blue, low expression; red, high expression). miR-877 and miR-719 were used as reference. d Real-time PCR analysis of miRNA precursors that are either enriched or depleted in neuritic preparations with respect to somatic fractions from hippocampal neurons at 14 DIV. All results are expressed as expression ratios between neuritic somatic fractions and normalized to the miR-877 and miR-719 expression levels. Data are means ± sem (n = 5). *p < 0.05, **p < 0.01; unpaired Student’s t test
To define the global miRNA profile in somatic and neuritic preparations of primary hippocampal neurons, we analyzed the expression of mature miRNAs using the Taqman low-density quantitative reverse transcription PCR arrays (TLDA RT-qPCR). Of 585 mature miRNAs that were profiled, 35% (216 of 585) were expressed in the processes (cycle threshold, Ct < 32). An unsupervised hierarchical clustering analysis revealed 26 miRNAs differentially regulated between the two fractions (Fig. 1c). Of these, 18 miRNAs were upregulated (at least 1.5-fold, p < 0.05) and 8 downregulated (at least 0.5-fold; p < 0.05) in the neuritic compartment as compared to the somatic counterpart (Fig. 1c and Table S2).
For synaptic compartments located at a distance from the cell body, it may in principle be convenient for a miRNA-mediated regulation to be independent of de novo pri-miRNA synthesis at the cell body level. Such independence can be achieved by using synaptically localized pools of pre-miRNAs that can be rapidly processed on site into mature miRNAs when required [40]. To verify this possibility, we investigated the expression of miRNA precursors (pre-miRs) for the most enriched and depleted miRNAs in the neuronal processes. qRT-PCR analysis showed that nine distinct pre-miRNAs were differently expressed between neuritic and somatic fractions (Fig. 1d). Notably, pre-miR218-1, pre-miR335, and pre-miR376b were highly expressed in the neuritic fraction, suggesting a possible involvement in the regulation of synaptic plasticity (Fig. 1d).
Development and Activity Levels of Hippocampal Networks Affect the Expression of Neurite-Enriched miRNAs
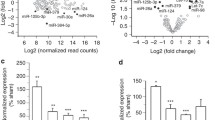
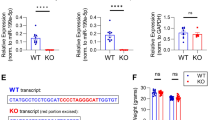
Based on the expression levels of pre-miRNAs, we restricted our analysis to the most significantly upregulated miRNAs. miRNAs involved in the regulation of synaptic transmission and plasticity are likely to be expressed at high levels during synapse maturation. Thus, we assessed the pattern of expression of mature and precursor miR-218, miR-335, and miR-376b by RT-qPCR analysis in the hippocampus at various postnatal ages (P) ranging from 1 to 28 days, a temporal window in which intense synaptogenesis occurs. miR-218 gradually increased with development, whereas miR-335 and miR-376b exhibited the highest expression at P14 and then declined at P28 (Fig. 2a). Of note, we observed a parallel significant increase of pre-miR-218-1 expression at P14 and P28, whereas pre-miR-335 and pre-miR-376b levels were unchanged during postnatal development (Fig. 2a).
Hippocampal development and network activity affect the expression of neurite-enriched miRNAs. a Real-time PCR analysis of mature (solid line) and precursor (dashed line) forms of miR-218, miR-335, and miR-376b in mouse hippocampal tissue at postnatal days 1, 14, and 28. Data are represented as fold changes with respect to the values of postnatal day 1 (means ± sem, n = 5). *p < 0.05, **p < 0.01; one-way ANOVA/Dunnett’s tests versus P1. b Real-time PCR analysis of miR-218, miR-335, and miR-376b in 14 DIV hippocampal neurons grown on standard plates and subjected to either TTX (1 μM) or BIC/4-AP (30 μM and 100 μM, respectively) treatments for 48 h (means ± sem, n = 5). *p < 0.05; one-way ANOVA/Dunnett’s tests versus control (CTR). GAPDH and HPRT1 were used as reference genes. c Venn diagram displaying the computationally predicted miR-218 target genes according to PicTar (green), TargetScan (blue), and Pita (red). d Gene Ontology Biological Processes enriched for miR-218 targets include synaptic transmission, regulation of synaptic plasticity, regulation of synapse maturation, neurotransmitter release, GABA signaling pathway, and regulation of membrane potential. The p values were calculated using Benjamini-corrected modified Fisher’s exact test. e Evolutionarily conserved miR-218 MRE in the 3′UTR of GluA2 in mammals, chicken, and Xenopus. The seed sequence is shown in red
Next, we investigated whether miRNA expression was affected by changes of neural network activity. To test this hypothesis, we applied either the Na+ channel blocker tetrodotoxin (TTX) to chronically silence firing activity or the combination of the GABAA receptor antagonist bicuculline and the K+ channel blocker 4-aminopyridine (BiC/4-AP) to chronically boost neuronal activity. Remarkably, inhibition of spontaneous activity following TTX treatment induced a strong increase in miR-218 expression, whereas miR-335 and miR376b exhibited a slight but not significant induction (Fig. 2b). On the contrary, sustained elevation of synaptic activity with BiC/4-AP decreased miR-218 expression without affecting the level of expression of miR-335 and miR-376b (Fig. 2b). Taken together, these results strongly suggest that miR-218 expression is bidirectionally modulated by the level of network activity. Therefore, we choose to focus our study on miR-218.
To identify protein-coding mRNA transcripts harboring miRNA response elements for miR-218 (miR-218 MRE), three bioinformatic programs for prediction analysis were used (PicTar, TargetScan, and Pita) [29, 30, 41]. All programs identified 159 putative target genes for miR-218 (Fig. 2c). Interestingly, 114 of the 159 genes (∼ 72%) are brain-expressed genes. Among those, gene ontology analysis revealed a significant enrichment of genes related to synaptic function and plasticity (Fig. 2d).
Among these genes, we selected the gene encoding for GluA2 receptor for representing a highly conserved putative miR-218 target across species from Xenopus to human (Fig. 2e) and a key actor in synaptic transmission. GluA2 has a crucial role in fast excitatory transmission. Several studies have shown that GluA2 mRNA is expressed in the dendritic compartment and GluA2 protein is locally synthesized and regulated by miRNAs [21, 26, 42,43,44,45,46,47,48]. Thus, this subunit could be an ideal target for the miRNA-mediated regulation of synaptic plasticity and focused on this target for further analysis.
miR-218 Specifically Targets the GluA2 Subunit of AMPA Glutamate Receptors
To determine whether miR-218 regulates GluA2 mRNA translation, full length 3′UTR of GluA2 was cloned downstream of the firefly luciferase gene in a reporter vector (Fig. 3a). We first expressed it in primary mouse hippocampal neurons together with the miR-218 mimic and measured the resulting luciferase activity using co-transfected Renilla luciferase to monitor transfection efficiency. Notably, miR-218 significantly increased the luciferase signal in a dose-dependent manner as compared to the scrambled control (Scr; Fig. 3b).
GluA2 is a specific target of miR-218 in primary hippocampal neurons. a Schematic of miRNA luciferase reporter plasmid, in which the full length 3′UTR of GluA2 is cloned downstream of the firefly luciferase gene under the control of CMV promoter. miR-124 (position 185–192) and miR-218 (position 476–483) MREs are reported. In all experiments, the relative luciferase activity was measured 48 h after electroporation, expressed as the ratio between firefly and Renilla luciferase signals, and normalized to the expression of control condition (Scr). b Luciferase expression to monitor the dose-dependent effect of miR-218 mimic on 3′UTR of GluA2 in hippocampal neurons as compared to the respective concentrations of scrambled miRNA (Scr). c Effect of the miR-218 inhibitor (anti-miR-218) on the wild-type form of Gria2 3′UTR in the presence of 80 nM miR-218. d Interaction of miR-218 and miR-124 on the wild-type form of Gria2 3′UTR or on its variants bearing mutations in the miR218 (mut218) or miR-124 (mut124) MREs. Graphs in b–d show means ± sem (n = 4). *p < 0.05, **p < 0.01; one-way ANOVA/Bonferroni’s tests. e Real-time PCR analysis of GRIA2 transcript level in hippocampal neurons after lentiviral-mediated expression or inhibition of miR-218 as compared to the scrambled sequence (Scr). GAPDH and HPRT1 were used as reference genes. f Western blotting analysis of the GluA2 subunit in primary hippocampal neurons grown on either standard plates (whole lysate) or porous membranes (soma and neuritic fractions) after lentiviral-mediated expression or inhibition of miR-218 and compared to the scrambled sequence (Scr). GAPDH or actin were used as loading control. Data are means ± sem (n = 4). *p < 0.05, one-way ANOVA/Bonferroni’s tests
Similar results, albeit at lower concentrations, were obtained in cultured HEK293T cells that do not endogenously express GluA2 (Fig. S1A). To verify that the effect was not due to differences in luciferase mRNA stability, we also analyzed luciferase mRNA levels by RT-qPCR and found no changes following miR-218 expression (Fig. S2). Furthermore, overexpression of miR-218 inhibitor (anti-miR-218) resulted in a significant reduction of luciferase signal compared to control conditions (Fig. 3c and Fig. S1B).
Previous studies have demonstrated the inhibitory effect of miR-124 on the GluA2-3′UTR [46,47,48]. We therefore tested whether miR-218 could be able to counteract the effect of miR-124. Consistent with previous reports, miR-124 caused a decrease in the luciferase activity of GluA2-3′UTR as compared to its scrambled control. Interestingly, when both miR-124 and miR-218 were co-expressed in equimolar amounts, we observed an antagonistic effect on GluA2-3′UTR, indicating that the translation rate is strictly dependent on the concentration ratio of the two miRNAs (Fig. 3d).
We next investigated whether the effect of the miR-218 mimic was mediated by the predicted site within the GluA2-3′UTR. Point mutations were introduced in the GluA 3′UTR sequence corresponding to miR-218 MRE (mut-218). When co-expressed with mut-218, miR-218 did not affect the luciferase signal as compared to Scr sample (Fig. 3d and Fig. S2B). Conversely, miR-124 could still provide inhibition of GluA2 translation of mutant miR-218 MRE, even in the presence of the miR-218 mimic (Fig. 3d).
Additionally, we co-expressed a mutated form of the GluA2-3′UTR for miR-124 MRE (mut124) along with miR-218 or miR-124. miR-218 alone or together with miR-124 increased the rate of translation of GluA 3′UTR reporter mutated for miR-124 MRE, whereas no significant differences were observed with the miR-124 mimic as compared to it scrambled control (Scr) (Fig. 3d). Together, these results indicate that miR-218 increases the translation of GluA2 receptor and the effect depends on the direct binding to the 3′UTR.
We finally assessed the ability of miR-218 to modify endogenous GluA2 expression in primary neurons. We induced lentiviral-mediated overexpression or inhibition of miR-218 with a resulting transduction efficiency over 85% as deducted by co-expression of the GFP reporter from the same vector (data not shown). A vector expressing a scrambled sequence was used as a control (Src) to assess the ability of miR-218 to affect the abundance of GRIA2 mRNA. We found no effect of miR-218 on GRIA2 mRNA levels as shown by qRT-PCR analysis, confirming that miR-218-dependent regulation did not change the transcription rate or the GRIA2 transcript stability (Fig. 3e). On the contrary, consistent with the luciferase assay data, the expression levels of GluA2 protein were increased by overexpression and decreased by inhibition of miR-218, as determined by western blotting on whole-cell lysates, soma, and neuritic preparations (Fig. 3f).
miR-218 Increases mEPSC Amplitude and Network Activity in Hippocampal Neurons
To address whether miR-218-mediated regulation of GluA2 has any consequence on AMPA-R function, we monitored GluA2 expression at the synapse. To evaluate synaptic GluA2 following miR-218 overexpression, we performed double-labeling for GluA2 and the presynaptic marker synapsin I. In accordance with the results obtained from western blotting, we confirmed the miR-218-induced upregulation of GluA2 (Fig. 4a). Additionally, in miR-218-overexpressing neurons, the GluR upregulation was associated with an increased expression of synapsin I and an increased co-localization between GluA2 and synapsin I (Fig. 4a).
Activity of excitatory synapses in primary hippocampal neurons after expression of miR-218. aLeft: Representative images of cultured hippocampal neurons double-stained for the presynaptic marker synapsin I (red), GluA2 (blue), and GFP reporter (gray), showing increased synaptic GluA2 after overexpression of miR-218 as compared to the scrambled sequence (Scr). Scale bars 5 μm. Right: Quantification of the intensity of synapsin I and GluA2 immunoreactivities and extent of co-localization of GluA2 and synapsin I after overexpression of miR-218. Data are means ± sem (n = 4). b Representative recordings of miniature excitatory postsynaptic currents (mEPSCs) in neurons expressing miR-218 (red trace) or its scrambled sequence (black trace). c, d Mean (± sem) and cumulative distribution of mEPSC amplitude (c) and mEPSC frequency/inter-event interval (d) in neurons expressing either scrambled (black) or miR-218 (red). e, f Mean (± sem) and cumulative distribution of mEPSC rise (e) and decay (f) times. (n = 5 independent preparations). *p < 0.05, ***p < 0.001; unpaired Student’s t test
To elucidate the physiological role of miR-218 in basal synaptic transmission mediated by AMPA receptors, we recorded miniature excitatory postsynaptic currents (mEPSCs) in DIV14–16 low-density hippocampal neurons overexpressing miR-218 or the scrambled sequence therefrom. In the analysis of mEPSCs, the frequency depends on synaptic density and probability of spontaneous release, while the amplitude reflects the quantal size that predominantly depends on the expression of postsynaptic receptors. In accordance with the increase in GluA2 expression, miR218-infected neurons exhibited a significant increase in mEPSC amplitude as compared to control conditions (Fig. 4a, b), in the absence of significant changes in mEPSC frequency (Fig. 4c) or kinetics (Fig. 4d, e).
To address the impact of synaptic changes on the regulation of the overall network activity, primary neurons overexpressing miR-218 or the scrambled sequence therefrom were plated at high density onto multielectrode array (MEA) chips and monitored at 14 DIV (Fig. 5a, b). Under these conditions, hippocampal network develops electrical activity characterized by random spiking associated with isolated bursts of action potentials that may diffuse to the whole network (network burst), as shown in the raster plots of the spiking activity recorded over 30 s from representative Scr- and miR-218-transduced neurons (Fig. 5c). A significantly increased neuronal activity was present in neuronal networks overexpressing miR-218 in comparison to those transduced with the scrambled control. Indeed, both the firing and the bursting rates resulted significantly increased in miR-218-infected neurons, with a consistently higher number of bursts per electrode and of spikes per burst and no significant change in the burst duration and in the occurrence of network bursts (Fig. 5d). To ascertain whether the increase in network activity was indeed caused by a strengthening of excitatory synapses promoted by miR-218, we challenged it by performing recordings in the presence of the AMPA receptor inhibitor CNQX. Indeed, CNQX prevented the miR-218-induced increases of network activity, which comparable to the levels of Scr-infected neurons (Fig. S3), indicating the synaptic source of the miR-218-induced increased network activity.
Neuronal networks expressing miR-218 display increased excitability. a, b Representative network electrical activity recorded from two adjacent microelectrodes (a) and phase-contrast micrograph of primary neurons on MEA (b). c Raster plots of spiking activity recorded over 30 s from neurons infected at 7 DIV with lentiviral vectors encoding either scrambled (black) or miR-218 (red) and analyzed at 14 DIV. Each bar denotes a spike, each row an electrode. d Analysis of network activity parameters. From left to right: weighted firing rate, overall bursting rate, bursting rate/electrode, burst duration, network burst rate, spikes/burst measured in neuronal networks expressing either scrambled (black bars) or miR-218 (red bars) at 14 DIV. Data are shown as means ± sem (n = 5 independent preparations). *p < 0.05; Student’s t test
Together, our results indicate that miR-218 plays a positive role in the regulation of excitatory synaptic transmission likely mediated by a local induction of GluA2 expression at synaptic level that in turn increases the overall electrical activity of the neuronal network.
Discussion
It is well known that local control of protein synthesis is involved in the expression of several forms of synaptic plasticity and a number of studies have reported the contribution of miRNAs to this process [18]. For instance, miR-132, miR-138, miR-124, and miR-9 regulate dendrite outgrowth and synapse formation; miR-128 and miR-125 contribute to synaptic strength and excitability [18]. In this study, we identified a set of 26 mature miRNAs and 9 precursor miRNAs that are significantly enriched in neuronal processes of primary hippocampal neurons. miR-218 and its precursor pre-miR-218-1 showed a robust increase, confirming previous results obtained in forebrain synaptosomes and hippocampal cultures [20]. Moreover, chronic suppression of neuronal activity increases miR-218 expression and, conversely, chronic hyperactivity reduces miR-218 expression, suggesting a role of miR-218 in the homeostatic regulation of synaptic strength. Interestingly, in silico miR-218 target prediction identified a complex repertoire of mRNAs associated with synaptic transmission and plasticity, including the transcript encoding for the AMPA receptor subunit, GluA2.
AMPA receptors are the principal transducers of fast excitatory transmission [49] and GluA2 is a critical subunit determining AMPA receptor properties, such as calcium permeability, single channel conductance, and rectification [50]. A large body of experimental evidence has shown that GluA2 plays a role in synaptic plasticity, as well as in learning and memory [51,52,53,54]. Interestingly, GluA2 mRNA is not only localized in the soma, but it is also targeted to dendrites, where it can be actively translated into functional receptor subunits [26, 42, 55]. GluA2 subunit is extensively regulated at the transcriptional and translational level [49]. GluA2 3′UTR harbors multiple putative MREs and its translation is downregulated by miRNAs such as miR-181b and miR-124 [43, 47, 48, 56]. Opposite to these miRNAs, we found that miR-218 enhances GluA2 translation by directly interacting with the 3′UTR of GluA2 leading to an increase of the expression of the GluA2 subunit of AMPA receptors. Consistent with this translational effect, we observed that miR-218 levels positively control excitatory synaptic strength by increasing the amplitude of single synaptic currents through an increase of the quantum size. This effect is reflected, at the network level, by an increase in the overall activity of primary hippocampal cultures with an enhancement of their firing and bursting behavior that is directly correlated with the strength and plasticity of excitatory synaptic transmission [57]. Of note, the regulation of miR-218 on GluA2 translation is consistent with previous findings showing that chronic inhibition of neuronal activity enhances, and hyperactivation decreases, GluA2 expression [58,59,60,61,62]. Additionally, several studies have reported that GluA2 receptors increase during postnatal development in hippocampal neurons [63, 64], in parallel with the observed developmental regulation expression of miR-218.
We demonstrated a functional role for miR-218 in upregulating AMPA receptor-mediated transmission in hippocampal cultures. An aspect of the study that needs more clarification is the mechanism by which miR-218 enhances the translation of GluA2. In the general view, binding of a miRNA to its target sequence leads to mRNA destabilization and translational repression by a plethora of mechanisms [65]. Interference with cap recognition, translational initiation, ribosomal subunit joining, and accumulation in decay bodies (P-bodies) have been described as key steps in miRNA-mediated downregulation of translation. However, cumulative evidence has been collected showing that miRNAs are also capable of upregulating gene expression in specific cell types and under particular cellular conditions. The first clue was obtained in liver cells, where miR-122 was able to upregulate hepatitis C virus RNA [66]. Later studies confirmed that miRNA pathways can activate translation in quiescent (G0 phase) mammalian cells and immature Xenopus oocytes [67, 68]. Recent observations have revealed that RISC composition and mRNA sequence features create a permissive or non-permissive environment to stimulate target translation [69]. The presence or absence of specific factors in the RISC complex affects the final output on gene expression. For instance, GW182 is essential for translational repression. Downregulation of GW-182 leads to the incorporation of FXR1 in the miRNP complex followed by miRNA-mediated gene activation [70].
In addition, sequence features of mRNA targets, mainly length of the poly(A) tail and presence of a 5′ terminal oligopyrimidine tract, as well as the ability of miRNAs to compete with repressive pathways have been proposed as alternative mechanisms to explain how a given miRNA can act bidirectionally as positive and negative regulator depending on the specific target [71]. Notably, miR-218 can act also as negative regulator and several mRNAs have been experimentally validated as genuine targets, including the RE-1-silencing transcription factor (REST) [72]. REST is a transcriptional repressor of GluA2 and participates in synaptic homeostasis by reducing the strength of excitatory synapses [73,74,75,76]. Thus, it is possible that miR-218 exerts a concerted action on GluA2 translation and AMPA receptor composition through a direct stimulatory action on the 3′UTR of GluR2 mRNA and by an indirect inhibitory action on the 3′UTR of REST mRNA that in turn relieves the REST-mediated inhibition of Gria2 transcription.
Taken together, our data extend the number of miRNAs involved in the regulation of AMPA receptor-mediated transmission and add complexity to the mechanisms regulating synaptic plasticity. Future studies may clarify the mechanism responsible for miR-218-mediated activation of GluA2 and reveal the effects of miR-218 on additional targets present at the synapse.
References
Cajigas IJ, Will T, Schuman EM (2010) Protein homeostasis and synaptic plasticity. EMBO J 29(16):2746–2752. https://doi.org/10.1038/emboj.2010.173
Yi JJ, Ehlers MD (2005) Ubiquitin and protein turnover in synapse function. Neuron 47(5):629–632. https://doi.org/10.1016/j.neuron.2005.07.008
Holt CE, Schuman EM (2013) The central dogma decentralized: new perspectives on RNA function and local translation in neurons. Neuron 80(3):648–657. https://doi.org/10.1016/j.neuron.2013.10.036
Martin KC, Ephrussi A (2009) mRNA localization: gene expression in the spatial dimension. Cell 136(4):719–730. https://doi.org/10.1016/j.cell.2009.01.044
Richter JD, Klann E (2009) Making synaptic plasticity and memory last: mechanisms of translational regulation. Genes Dev 23(1):1–11. https://doi.org/10.1101/gad.1735809
Sutton MA, Ito HT, Cressy P, Kempf C, Woo JC, Schuman EM (2006) Miniature neurotransmission stabilizes synaptic function via tonic suppression of local dendritic protein synthesis. Cell 125(4):785–799. https://doi.org/10.1016/j.cell.2006.03.040
Cox DJ, Racca C (2013) Differential dendritic targeting of AMPA receptor subunit mRNAs in adult rat hippocampal principal neurons and interneurons. J Comp Neurol 521(9):1954–2007. https://doi.org/10.1002/cne.23292
Ostroff LE, Fiala JC, Allwardt B, Harris KM (2002) Polyribosomes redistribute from dendritic shafts into spines with enlarged synapses during LTP in developing rat hippocampal slices. Neuron 35(3):535–545
Steward O, Levy WB (1982) Preferential localization of polyribosomes under the base of dendritic spines in granule cells of the dentate gyrus. J Neurosci 2(3):284–291
Vessey JP, Vaccani A, Xie Y, Dahm R, Karra D, Kiebler MA, Macchi P (2006) Dendritic localization of the translational repressor Pumilio 2 and its contribution to dendritic stress granules. J Neurosci 26(24):6496–6508. https://doi.org/10.1523/JNEUROSCI.0649-06.2006
Ferrari F, Mercaldo V, Piccoli G, Sala C, Cannata S, Achsel T, Bagni C (2007) The fragile X mental retardation protein-RNP granules show an mGluR-dependent localization in the post-synaptic spines. Mol Cell Neurosci 34(3):343–354. https://doi.org/10.1016/j.mcn.2006.11.015
Lugli G, Larson J, Martone ME, Jones Y, Smalheiser NR (2005) Dicer and eIF2c are enriched at postsynaptic densities in adult mouse brain and are modified by neuronal activity in a calpain-dependent manner. J Neurochem 94(4):896–905. https://doi.org/10.1111/j.1471-4159.2005.03224.x
Rao A, Steward O (1991) Evidence that protein constituents of postsynaptic membrane specializations are locally synthesized: analysis of proteins synthesized within synaptosomes. J Neurosci 11(9):2881–2895
Carthew RW, Sontheimer EJ (2009) Origins and mechanisms of miRNAs and siRNAs. Cell 136(4):642–655. https://doi.org/10.1016/j.cell.2009.01.035
Denli AM, Tops BB, Plasterk RH, Ketting RF, Hannon GJ (2004) Processing of primary microRNAs by the microprocessor complex. Nature 432(7014):231–235. https://doi.org/10.1038/nature03049
Lee Y, Kim M, Han J, Yeom KH, Lee S, Baek SH, Kim VN (2004) MicroRNA genes are transcribed by RNA polymerase II. EMBO J 23(20):4051–4060. https://doi.org/10.1038/sj.emboj.7600385
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116(2):281–297
Ye Y, Xu H, Su X, He X (2016) Role of microRNA in governing synaptic plasticity. Neural Plast 2016:4959523. https://doi.org/10.1155/2016/4959523
Schratt GM, Tuebing F, Nigh EA, Kane CG, Sabatini ME, Kiebler M, Greenberg ME (2006) A brain-specific microRNA regulates dendritic spine development. Nature 439(7074):283–289. https://doi.org/10.1038/nature04367
Siegel G, Obernosterer G, Fiore R, Oehmen M, Bicker S, Christensen M, Khudayberdiev S, Leuschner PF et al (2009) A functional screen implicates microRNA-138-dependent regulation of the depalmitoylation enzyme APT1 in dendritic spine morphogenesis. Nat Cell Biol 11(6):705–716. https://doi.org/10.1038/ncb1876
Fiore R, Rajman M, Schwale C, Bicker S, Antoniou A, Bruehl C, Draguhn A, Schratt G (2014) MiR-134-dependent regulation of Pumilio-2 is necessary for homeostatic synaptic depression. EMBO J 33(19):2231–2246. https://doi.org/10.15252/embj.201487921
Kye MJ, Liu T, Levy SF, Xu NL, Groves BB, Bonneau R, Lao K, Kosik KS (2007) Somatodendritic microRNAs identified by laser capture and multiplex RT-PCR. Rna 13(8):1224–1234. https://doi.org/10.1261/rna.480407
Lugli G, Torvik VI, Larson J, Smalheiser NR (2008) Expression of microRNAs and their precursors in synaptic fractions of adult mouse forebrain. J Neurochem 106(2):650–661. https://doi.org/10.1111/j.1471-4159.2008.05413.x
Natera-Naranjo O, Aschrafi A, Gioio AE, Kaplan BB (2010) Identification and quantitative analyses of microRNAs located in the distal axons of sympathetic neurons. Rna 16(8):1516–1529. https://doi.org/10.1261/rna.1833310
Sasaki Y, Gross C, Xing L, Goshima Y, Bassell GJ (2014) Identification of axon-enriched microRNAs localized to growth cones of cortical neurons. Dev Neurobiol 74(3):397–406. https://doi.org/10.1002/dneu.22113
Grooms SY, Noh KM, Regis R, Bassell GJ, Bryan MK, Carroll RC, Zukin RS (2006) Activity bidirectionally regulates AMPA receptor mRNA abundance in dendrites of hippocampal neurons. J Neurosci 26(32):8339–8351. https://doi.org/10.1523/JNEUROSCI.0472-06.2006
Lignani G, Raimondi A, Ferrea E, Rocchi A, Paonessa F, Cesca F, Orlando M, Tkatch T et al (2013) Epileptogenic Q555X SYN1 mutant triggers imbalances in release dynamics and short-term plasticity. Hum Mol Genet 22(11):2186–2199. https://doi.org/10.1093/hmg/ddt071
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3(7):RESEARCH0034
Kertesz M, Iovino N, Unnerstall U, Gaul U, Segal E (2007) The role of site accessibility in microRNA target recognition. Nat Genet 39(10):1278–1284. https://doi.org/10.1038/ng2135
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120(1):15–20. https://doi.org/10.1016/j.cell.2004.12.035
Lall S, Grun D, Krek A, Chen K, Wang YL, Dewey CN, Sood P, Colombo T et al (2006) A genome-wide map of conserved microRNA targets in C. elegans. Curr Biol 16(5):460–471. https://doi.org/10.1016/j.cub.2006.01.050
McSweeney KM, Gussow AB, Bradrick SS, Dugger SA, Gelfman S, Wang Q, Petrovski S, Frankel WN et al (2016) Inhibition of microRNA 128 promotes excitability of cultured cortical neuronal networks. Genome Res 26(10):1411–1416. https://doi.org/10.1101/gr.199828.115
Chiappalone M, Novellino A, Vajda I, Vato A, Martinoia S, Van Pelt J (2005) Burst detection algorithms for the analysis of spatio-temporal patterns in cortical networks of neurons. Neurocomputing 65-66:653–662
Torre ER, Steward O (1992) Demonstration of local protein synthesis within dendrites using a new cell culture system that permits the isolation of living axons and dendrites from their cell bodies. J Neurosci 12(3):762–772
Poon MM, Choi SH, Jamieson CA, Geschwind DH, Martin KC (2006) Identification of process-localized mRNAs from cultured rodent hippocampal neurons. J Neurosci 26(51):13390–13399. https://doi.org/10.1523/JNEUROSCI.3432-06.2006
Kohler A, Hurt E (2007) Exporting RNA from the nucleus to the cytoplasm. Nat Rev Mol Cell Biol 8(10):761–773. https://doi.org/10.1038/nrm2255
Litman P, Barg J, Ginzburg I (1994) Microtubules are involved in the localization of tau mRNA in primary neuronal cell cultures. Neuron 13(6):1463–1474
Bicker S, Khudayberdiev S, Weiss K, Zocher K, Baumeister S, Schratt G (2013) The DEAH-box helicase DHX36 mediates dendritic localization of the neuronal precursor-microRNA-134. Genes Dev 27(9):991–996. https://doi.org/10.1101/gad.211243.112
Bramham CR, Wells DG (2007) Dendritic mRNA: transport, translation and function. Nat Rev Neurosci 8(10):776–789. https://doi.org/10.1038/nrn2150
Sambandan S, Akbalik G, Kochen L, Rinne J, Kahlstatt J, Glock C, Tushev G, Alvarez-Castelao B et al (2017) Activity-dependent spatially localized miRNA maturation in neuronal dendrites. Science 355(6325):634–637. https://doi.org/10.1126/science.aaf8995
Krek A, Grun D, Poy MN, Wolf R, Rosenberg L, Epstein EJ, MacMenamin P, da Piedade I et al (2005) Combinatorial microRNA target predictions. Nat Genet 37(5):495–500. https://doi.org/10.1038/ng1536
Ju W, Morishita W, Tsui J, Gaietta G, Deerinck TJ, Adams SR, Garner CC, Tsien RY et al (2004) Activity-dependent regulation of dendritic synthesis and trafficking of AMPA receptors. Nat Neurosci 7(3):244–253. https://doi.org/10.1038/nn1189nn1189
Saba R, Storchel PH, Aksoy-Aksel A, Kepura F, Lippi G, Plant TD, Schratt GM (2012) Dopamine-regulated microRNA MiR-181a controls GluA2 surface expression in hippocampal neurons. Mol Cell Biol 32(3):619–632. https://doi.org/10.1128/MCB.05896-11
Harraz MM, Eacker SM, Wang X, Dawson TM, Dawson VL (2012) MicroRNA-223 is neuroprotective by targeting glutamate receptors. Proc Natl Acad Sci U S A 109(46):18962–18967. https://doi.org/10.1073/pnas.1121288109
Dutta R, Chomyk AM, Chang A, Ribaudo MV, Deckard SA, Doud MK, Edberg DD, Bai B et al (2013) Hippocampal demyelination and memory dysfunction are associated with increased levels of the neuronal microRNA miR-124 and reduced AMPA receptors. Ann Neurol 73(5):637–645. https://doi.org/10.1002/ana.23860
Gascon E, Lynch K, Ruan H, Almeida S, Verheyden JM, Seeley WW, Dickson DW, Petrucelli L et al (2014) Alterations in microRNA-124 and AMPA receptors contribute to social behavioral deficits in frontotemporal dementia. Nat Med 20(12):1444–1451. https://doi.org/10.1038/nm.3717
Ho VM, Dallalzadeh LO, Karathanasis N, Keles MF, Vangala S, Grogan T, Poirazi P, Martin KC (2014) GluA2 mRNA distribution and regulation by miR-124 in hippocampal neurons. Mol Cell Neurosci 61:1–12. https://doi.org/10.1016/j.mcn.2014.04.006
Hou Q, Ruan H, Gilbert J, Wang G, Ma Q, Yao WD, Man HY (2015) MicroRNA miR124 is required for the expression of homeostatic synaptic plasticity. Nat Commun 6:10045. https://doi.org/10.1038/ncomms10045
Traynelis SF, Wollmuth LP, McBain CJ, Menniti FS, Vance KM, Ogden KK, Hansen KB, Yuan H et al (2010) Glutamate receptor ion channels: structure, regulation, and function. Pharmacol Rev 62(3):405–496. https://doi.org/10.1124/pr.109.002451
Tanaka H, Grooms SY, Bennett MV, Zukin RS (2000) The AMPAR subunit GluR2: still front and center-stage. Brain Res 886(1–2):190–207
Malinow R, Malenka RC (2002) AMPA receptor trafficking and synaptic plasticity. Annu Rev Neurosci 25:103–126. https://doi.org/10.1146/annurev.neuro.25.112701.142758
Isaac JT, Ashby MC, McBain CJ (2007) The role of the GluR2 subunit in AMPA receptor function and synaptic plasticity. Neuron 54(6):859–871. https://doi.org/10.1016/j.neuron.2007.06.001
Riedel G, Micheau J, Lam AG, Roloff EL, Martin SJ, Bridge H, de Hoz L, Poeschel B et al (1999) Reversible neural inactivation reveals hippocampal participation in several memory processes. Nat Neurosci 2(10):898–905. https://doi.org/10.1038/13202
Wiltgen BJ, Royle GA, Gray EE, Abdipranoto A, Thangthaeng N, Jacobs N, Saab F, Tonegawa S et al (2010) A role for calcium-permeable AMPA receptors in synaptic plasticity and learning. PLoS One 5(9):e12818. https://doi.org/10.1371/journal.pone.0012818
Kacharmina JE, Job C, Crino P, Eberwine J (2000) Stimulation of glutamate receptor protein synthesis and membrane insertion within isolated neuronal dendrites. Proc Natl Acad Sci U S A 97(21):11545–11550. https://doi.org/10.1073/pnas.97.21.1154597/21/11545[pii]
Beveridge NJ, Tooney PA, Carroll AP, Gardiner E, Bowden N, Scott RJ, Tran N, Dedova I et al (2008) Dysregulation of miRNA 181b in the temporal cortex in schizophrenia. Hum Mol Genet 17(8):1156–1168. https://doi.org/10.1093/hmg/ddn005
Lonardoni D, Amin H, Di Marco S, Maccione A, Berdondini L, Nieus T (2017) Recurrently connected and localized neuronal communities initiate coordinated spontaneous activity in neuronal networks. PLoS Comput Biol 13(7):e1005672. https://doi.org/10.1371/journal.pcbi.1005672
Ancona Esselmann SG, Diaz-Alonso J, Levy JM, Bemben MA, Nicoll RA (2017) Synaptic homeostasis requires the membrane-proximal carboxy tail of GluA2. Proc Natl Acad Sci U S A 114(50):13266–13271. https://doi.org/10.1073/pnas.1716022114
Anggono V, Clem RL, Huganir RL (2011) PICK1 loss of function occludes homeostatic synaptic scaling. J Neurosci 31(6):2188–2196. https://doi.org/10.1523/JNEUROSCI.5633-10.2011
Diering GH, Gustina AS, Huganir RL (2014) PKA-GluA1 coupling via AKAP5 controls AMPA receptor phosphorylation and cell-surface targeting during bidirectional homeostatic plasticity. Neuron 84(4):790–805. https://doi.org/10.1016/j.neuron.2014.09.024
Wierenga CJ, Ibata K, Turrigiano GG (2005) Postsynaptic expression of homeostatic plasticity at neocortical synapses. J Neurosci 25(11):2895–2905. https://doi.org/10.1523/JNEUROSCI.5217-04.2005
Lussier MP, Nasu-Nishimura Y, Roche KW (2011) Activity-dependent ubiquitination of the AMPA receptor subunit GluA2. J Neurosci 31(8):3077–3081. https://doi.org/10.1523/JNEUROSCI.5944-10.2011
Jensen V, Kaiser KM, Borchardt T, Adelmann G, Rozov A, Burnashev N, Brix C, Frotscher M et al (2003) A juvenile form of postsynaptic hippocampal long-term potentiation in mice deficient for the AMPA receptor subunit GluR-A. J Physiol 553(Pt 3):843–856. https://doi.org/10.1113/jphysiol.2003.053637
Monyer H, Seeburg PH, Wisden W (1991) Glutamate-operated channels: developmentally early and mature forms arise by alternative splicing. Neuron 6(5):799–810
Jonas S, Izaurralde E (2015) Towards a molecular understanding of microRNA-mediated gene silencing. Nat Rev Genet 16(7):421–433. https://doi.org/10.1038/nrg3965
Jopling CL, Yi M, Lancaster AM, Lemon SM, Sarnow P (2005) Modulation of hepatitis C virus RNA abundance by a liver-specific microRNA. Science 309(5740):1577–1581. https://doi.org/10.1126/science.1113329
Vasudevan S, Tong Y, Steitz JA (2007) Switching from repression to activation: microRNAs can up-regulate translation. Science 318(5858):1931–1934. https://doi.org/10.1126/science.1149460
Mortensen RD, Serra M, Steitz JA, Vasudevan S (2011) Posttranscriptional activation of gene expression in Xenopus laevis oocytes by microRNA-protein complexes (microRNPs). Proc Natl Acad Sci U S A 108(20):8281–8286. https://doi.org/10.1073/pnas.1105401108
Valinezhad Orang A, Safaralizadeh R, Kazemzadeh-Bavili M (2014) Mechanisms of miRNA-mediated gene regulation from common downregulation to mRNA-specific upregulation. Int J Genomics 2014:970607. https://doi.org/10.1155/2014/970607
Bukhari SIA, Truesdell SS, Lee S, Kollu S, Classon A, Boukhali M, Jain E, Mortensen RD et al (2016) A specialized mechanism of translation mediated by FXR1a-associated microRNP in cellular quiescence. Mol Cell 61(5):760–773. https://doi.org/10.1016/j.molcel.2016.02.013
Vasudevan S (2012) Posttranscriptional upregulation by microRNAs. Wiley Interdiscip Rev RNA 3(3):311–330. https://doi.org/10.1002/wrna.121
Liu JJ, Zhao CM, Li ZG, Wang YM, Miao W, Wu XJ, Wang WJ, Liu C et al (2016) miR-218 involvement in cardiomyocyte hypertrophy is likely through targeting REST. Int J Mol Sci 17(6):848. https://doi.org/10.3390/ijms17060848
Myers SJ, Peters J, Huang Y, Comer MB, Barthel F, Dingledine R (1998) Transcriptional regulation of the GluR2 gene: neural-specific expression, multiple promoters, and regulatory elements. J Neurosci 18(17):6723–6739
Calderone A, Jover T, Noh KM, Tanaka H, Yokota H, Lin Y, Grooms SY, Regis R et al (2003) Ischemic insults derepress the gene silencer REST in neurons destined to die. J Neurosci 23(6):2112–2121
Pozzi D, Lignani G, Ferrea E, Contestabile A, Paonessa F, D'Alessandro R, Lippiello P, Boido D et al (2013) REST/NRSF-mediated intrinsic homeostasis protects neuronal networks from hyperexcitability. EMBO J 32(22):2994–3007. https://doi.org/10.1038/emboj.2013.231
Pecoraro-Bisogni F, Lignani G, Contestabile A, Castroflorio E, Pozzi D, Rocchi A, Prestigio C, Orlando M et al (2018) REST-dependent presynaptic homeostasis induced by chronic neuronal hyperactivity. Mol Neurobiol 55(6):4959–4972. https://doi.org/10.1007/s12035-017-0698-9
Acknowledgements
We thank Luigi Naldini for kindly providing lentiviral vectors; Monica Morini, Riccardo Navone (Italian Institute of Technology, Genova, Italy), and Michele Cilli (IRCCS San Martino, Genova, Italy) for help in breeding the mice; and Arta Mehilli (Center for Synaptic Neuroscience, Istituto Italiano di Tecnologia, Genova, Italy) for assistance in the preparation of primary cultures. This study was supported by research grants from Compagnia di San Paolo (2015.0546 to FB and 2017.0589 to PB), EU FP7 Project “Desire” (Grant agreement n. 602531 to FB) and the Italian Ministry of Health Ricerca Finalizzata (GR-2013-02355540 to AR).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Fig. S1
Specificity of the effects of miR-218 on GluA2 3′UTR in HEK293T cells. (A) Luciferase expression was used to monitor the dose-dependent effect of miR-218 mimic on GluA2 3’UTR in HEK293T cells that do not endogenously express GluA2. Scr, scrambled miRNA control. (B) Effect of miR-218 and miR-218 inhibitor (anti-miR-218) on the wild type and mutated forms of Gria2 3′UTR. Data are shown as means ± sem (n = 4). * p < 0.05, ** p < 0.01; unpaired Student’s t test. (PNG 108 kb)
Fig. S2
Luciferase mRNA is not affected by miR-218 overexpression. Real time PCR analysis of luciferase transcript level in hippocampal neurons overexpressing miR-218 mimic as compared to the respective scrambled sequence (Scr). Data are shown as means ± sem (n = 4). (PNG 26 kb)
Fig. S3
The effects of miR-218 on network activity are mediated by an increase in AMPA transmission. (A) Representative raster plots of the network electrical activity recorded in neurons expressing scrambled sequence (black), miR-218 (red) or miR-218 (green) in presence of the specific AMPA glutamate receptor inhibitor CNQX (20 μM) . (B) The overall activity of the network under the various experimental conditions is expressed as the cumulative firing rates recorded over a period of 200 s (Hz) weighted on the number of active electrodes. Data are means ± sem (n = 3 independent preparations). * p < 0.05; ** p < 0.01; one-way ANOVA/Bonferroni’s multiple comparison test. (PNG 153 kb)
ESM 1
(XLSX 11 kb)
ESM 2
(XLSX 10 kb)
Rights and permissions
About this article
Cite this article
Rocchi, A., Moretti, D., Lignani, G. et al. Neurite-Enriched MicroRNA-218 Stimulates Translation of the GluA2 Subunit and Increases Excitatory Synaptic Strength. Mol Neurobiol 56, 5701–5714 (2019). https://doi.org/10.1007/s12035-019-1492-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-019-1492-7