Abstract
Schizophrenia (SCZ) is a mental disorder arising from a complex interaction of genetic and environmental factors. It has been suggested that treatment-resistant schizophrenia (TRS) is a distinct, more severe, and homogenous subgroup of schizophrenia that could present specific biological markers. Our aim was to characterize expression of target genes in blood of TRS patients compared with non-TRS (NTRS) patients and healthy controls (HC). TRS has been defined using failure to respond to two previous antipsychotic trials. We hypothesized that genes involved in neurodevelopment, myelination, neuroplasticity, neurotransmission, and miRNA processing could be involved in treatment resistance; then, we investigated 13 genes related to those processes in 256 subjects, being 94 healthy controls and 162 schizophrenia patients treated with antipsychotics. Of those, 78 were TRS patients and 84 were NTRS patients. Peripheral blood samples were collected from all subjects and RNA was isolated. Gene expression analysis was performed using the TaqMan low-density array (TLDA) technology. To verify the influence of expression quantitative trait loci (eQTLs), we evaluated single-nucleotide polymorphism (SNP) of all genes using data from GTEx Project. SNP genotypes were obtained from HumanOmniExpress BeadChip. We did not detect gene expression differences between TRS and NTRS subjects, indicating candidate genes specific to treatment resistance. We detected an upregulation of CNR1 and UFD1L gene expression in patients (TRS and NTRS groups) when compared to controls, that may be associated with the release of neurotransmitters, which can influence neuronal plasticity, or with a stress response-activating protein degradation. DICER1 and AKT1 expression increased slightly across the groups and could differentiate only the extreme opposite groups, HC and TRS. Both genes act in heterogeneous pathways, such as cell signaling and miRNA processing, and seem to have an increased demand in the TRS group. We did not detect any eQTLs in our sample that could explain differences in mRNA levels, suggesting a possible regulation by other mechanism, not driven by genotypes. Our data strengthen the importance of several biological pathways involved in the schizophrenia refractoriness and severity, adding knowledge to develop more effective treatments in the future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Schizophrenia (SCZ) is a complex disease with a heterogeneous psychopathology. Several hypotheses have been raised about the etiology of SCZ. In neurodevelopment hypothesis, it is proposed that the disease has its origins in disturbed development of the nervous system prior to the onset of clinical symptoms, due to a combination of genetic and environmental factors [1].
Although genetic factors are considered the strongest risk factor for SCZ [2], finding the genetic etiology of the disease is still a challenge due to the complex combination of many common genetic variants of small effect (predominantly single-nucleotide polymorphisms—SNPs), rare variants with high predictive value (such as copy number variations—CNVs), and epigenetics [3]. Furthermore, complex diseases such as SCZ tend to result not only from alterations in individual genes, but from changes in molecular networks [4].
Antipsychotic drugs in current use can be classified into two categories: typical antipsychotics and atypical antipsychotics. The two classes have similar efficacy in the control of positive and negative symptoms, but differ in the propensity to trigger extrapyramidal symptoms [5]. Antipsychotic therapy is an important approach in SCZ treatment, but psychotic relapse rates remain high. It is estimated that 20–30% of patients with schizophrenia do not respond to treatment with conventional antipsychotics [6, 7].
The treatment-resistant schizophrenia (TRS) is defined by an inadequate response to a sequence of treatments [8]. An inadequate response needs not to be restricted to the persistence of positive symptoms, but this is the most common definition. The concept of treatment resistance should not be confused with “chronic schizophrenia” as chronicity and “resistance” are different concepts. Chronic patients can respond to standard treatments and treatment resistance may be as high as 15% even in first-episode patients [9]. Studies have shown that individuals with a family history of psychosis are more likely to poorly respond to antipsychotic drugs [10, 11], suggesting a genetic influence in the development of TRS. As TRS seem to be a distinct and homogenous subgroup of schizophrenia [12, 13], studies have focused on finding biological markers that could predict the response to antipsychotics and consequently improve the prognosis of the patients. Our main aim was to compare expression of target genes in blood among TRS patients, non-TRS (NTRS) patients, and healthy controls (HC) and to find eQTLs that could have an influence on the expression of those genes.
Materials and Methods
Study Population
SCZ group was comprised of chronically outpatients recruited from Schizophrenia Program (PROESQ) of UNIFESP, treated with diverse types of typical and atypical antipsychotics. The diagnosis of SCZ was established according to the criteria of the Diagnostic and Statistical Manual of Mental Disorders (DSM-IV) using the Structured Clinical Interview (SCID-I). Treatment-resistant (TR) status was defined using modified criteria of The International Psychopharmacology Algorithm Project (IPAP)—schizophrenia algorithm. Patients were regarded as TRS in the presence of (a) no response to at least two antipsychotics used in monotherapy for a minimum period of 4 to 6 weeks with appropriate doses of the drug and (b) presence of moderate to severe psychopathology, especially positive symptoms [14]. Antipsychotic types, doses, and number of medications were registered and compared between the TRS and NTRS groups. To evaluate the effect of dosages of different medications in mRNA expression, chlorpromazine (CPZ) equivalent doses were estimated using regression equations according to Andreasen et al. (2010) [15] and were compared between the groups. In patients using more than one type of medication, CPZ doses were calculated, summing equivalent doses of each antipsychotic. Patients were also evaluated using standardized instruments, including the Positive and Negative Syndrome Scale (PANSS) and Global Assessment of Functioning (GAF). The HC group comprised volunteers with no current or previous psychiatric diagnoses or first-degree family history of psychotic disorders. The Research Ethics Committee of UNIFESP approved the research protocol, and all participants and family members provided written informed consent prior to enrolment in the study (CAAE 06191612.7.0000.5505).
Genetic Analyses
A total of 5 mL of whole blood was collected in PAXgene® RNA tubes (PreAnalytix, Hombrechtikon, Switzerland) and RNA was isolated using the PAXgene® Blood RNA kit (Qiagen, Germantown, MD), according to the manufacturer’s instructions. RNA quality and integrity was evaluated using electrophoresis on a 1.0%-agarose gel and NanoDrop® ND-1000 spectrophotometer (NanoDrop, Wilmington, DE), respectively.
Approximately 400 ng of each RNA sample was reverse-transcribed using the High-Capacity cDNA Reverse Transcription Kit (Life Technologies, Foster City, CA). Then, 20 to 100 ng of cDNA was diluted in H2O, mixed with the TaqMan® Universal PCR Master Mix (Life Technologies, Foster City, EUA), and loaded on TaqMan low-density array (TLDA) microfluidic cards (Life Technologies, Foster City, EUA). Probes and primers of 13 target genes, 2 housekeeping genes (HKGs: ACTB and GAPDH), and 1 positive control for the reaction (18S) were preloaded in the 384 wells of each TLDA card (Life Technologies). The HKGs were selected based on the detection of a strong correlation of relative threshold method (Crt) values between ACTB and GAPDH genes in blood, no association of both gene Crt values with SCZ diagnosis and psychopathology symptoms, and the detection of smallest stability values using NormFinder [16]. We selected the candidate genes based on a review of the literature considering the following aspects: (i) at least one report on association with SCZ, (ii) plausibility of their involvement in SCZ pathophysiology, and (iii) expression in blood according to information in the Anatomy tool of GENEVESTIGATOR [17]. Genes that presented medium to high mRNA levels in blood in this tool were defined as expressed in whole blood. We selected genes related to neurotransmission (COMT and CNR1), inflammation and the immune system (TNF), neurodevelopment (DISC1, PAFAH1B1, and NDEL1), myelination (MBP), cell signaling (AKT1), the microRNA machinery (DGCR8, DICER1, and DROSHA), protein degradation (UFD1L), and adhesion (DGCR2). Some genes were selected mainly because of being located in the genomic region known as the most important genetic risk factor for psychotic disorders: the 22q11.2 region (COMT, DGCR2, DGCR8, and UFD1L) [18]. The experiments were performed in accordance with the manufacturer’s instructions using the ViiA™ 7 Real-Time PCR System (Life Technologies, Foster City, EUA).
Gene expression was quantified using the Crt, discounting the ACTB and GAPDH geometric mean (GM) as endogenous controls (ΔCrt = Crttarget gene − CrtGM). To each sample, a ΔCrt was calculated and a fold change (FC) was calculated using 2-ΔΔCrt, always considering the HC group as reference. Gene expression and ΔCrt values are negatively correlated; therefore, higher ΔCrt values mean lower gene expression.
Polymorphisms located within those 13 target genes were extracted from the Genotyping Array using PLINK software version 2.0, and the correlations between their genotype and tissue-specific gene expression levels described in this manuscript were obtained from the Single-Tissue eQTLs and/or Gene eQTL Visualizer on GTEx Portal on dbGaP Accession phs000424.v6.p1.
Statistical Analysis
The statistical analyses were performed in the Statistical Package for the Social Sciences (SPSS), version 15.0 dataset. Gender differences between the SCZ and HC groups were verified using the chi-squared test. Age, disease, and treatment duration were compared among the three groups (TRS, NTRS, and HC) using analysis of variance (ANOVA) test and between two groups (TRS and NTRS) using Student’s t test. ΔCrt values were compared among each group using the ANOVA, and post hoc comparisons were carried out using the Bonferroni test. Pearson correlation was used to test correlations between mRNA levels and PANSS scores, GAF scores, and CPZ equivalent doses.
Results
Samples
We recruited a total of 94 healthy controls and 162 SCZ patients, 78 of whom were TRS patients and the other 84 were NTRS patients. The characteristics of samples from SCZ patients (TRS and NTRS) and controls are described in Table 1. Sex (chi-squared = 1.160; p = 0.560) and age (F(253,2) = 2.278; p = 0.105) did not differ among the groups.
The number of antipsychotics being used between the TRS and NTRS groups were different, with a smaller number of TRS subjects being treated in monotherapy (N = 49; 43.4%) in comparison to that of NTRS subjects (N = 64; 56.6%), and a higher number of TRS patients (N = 26; 70.3%) being treated with two or more antipsychotics than that of NTRS patients (N = 11; 29.7%) (chi-squared = 9.288; p = 0.002). In monotherapy, olanzapine and risperidone accounts for the treatment of 50 and 18.8% of the 64 NTRS subjects, respectively. In the TRS group, clozapine was the antipsychotic used by 77.6%, followed by olanzapine, used by 8.2% of the 49 patients. We observed differences in CPZ equivalent doses of antipsychotics between the groups, with TRS patients receiving higher doses of medication (t − 6.421; df 142; p < 0.001) than NTRS patients (Table 2).
No differences in disease or treatment duration between the TRS and NTRS groups were detected. We observed that TRS patients are affected earlier than NTRS patients, with mean age of onset smaller than NTRS patients (t 2.142; df 160; p = 0.034). As expected, total PANSS (t − 4.116; df 153; p < 0.001) and GAF (t 3.924; df 107; p < 0.001) scores were higher in TRS patients in comparison to NTRS patients (Table 2).
Gene Expression and eQTL Analyses
Regarding treatment resistance, we did not observe gene expression differences between the TRS and NTRS subjects. We also did not detect gene expression differences between patients in use of clozapine compared to other antipsychotic drugs. Although differences between CPZ equivalent doses were detected between the groups, no correlation was observed between CPZ equivalent doses and mRNA levels of the 13 genes analyzed (p > 0.05).
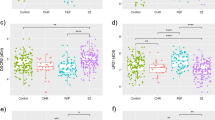
We detected differences in CNR1 (F(174,2) = 7.00; p = 0.001) and UFD1L (F(237,2) = 12.69; p < 0.001) gene expression among the HC, TRS, and NTRS groups. The expression of CNR1 and UFD1L increases proportionately with the severity of the disease, being upregulated in patients (NTRS and TRS) in relation to HC (Table 3). However, there was no correlation between CNR1 and UFD1L mRNA levels and psychopathology, measured by PANSS or GAF (p > 0.05). Fold change and ANOVA p values, after Bonferroni post hoc test, of CNR1 (NTRS: FC = 1.91, p = 0.004; and TRS: FC = 1.96, p = 0.002) and UFD1L (NTRS: FC = 1.18, p = 0.01; and TRS: FC = 1.31, p < 0.001) are presented in Fig. 1(a, b).
Comparison of CNR1 (a), UFD1L (b), DICER1 (c), and UFD1L (d) mRNA levels among groups. ΔCrt and gene expression are inverse correlated; therefore, TRS group exhibits the highest expression levels of all genes. ∆Crt delta relative threshold obtained in real-time PCR, HC healthy controls, TRS treatment-resistant schizophrenia, NTRS non-treatment-resistant schizophrenia, p significance level for ANOVA test after post hoc comparison between NTRS and HC groups, p* significance level for ANOVA test after post hoc comparison between TRS and HC groups, FC mRNA expression fold change of NTRS group using HC as reference, FC* mRNA expression fold change of TRS group using HC as reference
We observed differences in mRNA levels of AKT1 (F(240,2) = 4.47; p = 0.01) and DICER1 (F(241,2) = 3.81; p = 0.02) between the HC and TRS group. Both genes were upregulated in the TRS group in comparison to the HC group. Fold change and ANOVA p values, after Bonferroni post hoc test, of AKT1 (FC = 1.11, p = 0.01) and DICER1 (FC = 1.13, p = 0.03) are presented in Fig. 1(c, d).
According to the GTEx database, the 13 genes studied present 25 eQTLs of brain and blood tissues. Of those, we had genotype information from 19 SNPs; however, none of them were situated in genes differently expressed among the groups.
Discussion
Although TRS patients present characteristics that support a categorically different illness subtype [19], such as gray matter alterations [20], glutamatergic abnormalities [21], and a lack of dopaminergic abnormalities [22, 23], in our study, we did not observe differences in whole blood gene expression that could discriminate TRS and NTRS patients in a very specific manner. However, we found two genes probably associated with disease pathophysiology, CNR1 and UFD1L, that were differentially expressed in the blood of HC subjects, when compared to TRS and NTRS patients, and other two genes, AKT1 and DICER1, that were differentially expressed between the extreme groups HC and TRS. CNR1 gene is located on 6q14–15, a region of replicated linkage for SCZ, and encodes type-1 cannabinoid receptors (CB1) that are activated by endocannabinoids neurotransmitters, by cannabinoids such as the compound delta-9-tetrahydrocannabinol (THC) and by synthetic analogues of THC [24].
CNR1 is proposed as a candidate gene to SCZ etiology since the endocannabinoid system has emerged as a possible link between cannabis and psychosis, especially after the comprehension that the psychogenic properties of cannabis are mainly mediated by THC stimulation of CB1 [25]. Some SNPs of this gene have been associated with central nervous system effects [24, 26, 27] with SCZ subtypes [28,29,30] and treatment refractoriness [31]. However, it is not clear how the genetic variants could influence in disease development, since most of them are not related to amino acid substitution and protein structure changes [24]. Stress induced by maternal diet deprivation seems to trigger executive function deficits associated to an overexpression of Cnr1, in mouse prefrontal cortex (PFC) [32]. These executive function deficits, although unspecific, are associated with different psychiatric diagnoses [33,34,35]. Increased expression of Cnr1 gene was also reported in the frontal cortex of male rats after maternal diet deprivation, associated with depressive-like and psychotic-like symptoms [36]. Upregulation of CNR1 was also described in peripheral blood mononuclear cells (PBMCs) of SCZ patients in comparison to HC, but not in subjects with bipolar disorder types I and II, or major depressive disorder (MDD) compared to controls [37]. Supporting this upregulation, decreased promotor DNA methylation was observed in CNR1 of those SCZ patients. Similarly, to what is observed in humans, increased expression of Cnr1 was detected in the PFC of rats exposed to methylazoxymethanol acetate (MAM), a developmental animal model of SCZ, compared to a control group of animals. Less DNA methylation at promoter region was also reported in Cnr1 gene of this animal model of SCZ. It is important to highlight that these changes in mRNA expression and DNA methylation were investigated in other genes which are part of endocannabinoid system, such as endocannabinoid receptor 2 (CB2) among others, but only CNR1 was significantly changed [37]. CB1 receptors regulate the release of inhibitory and excitatory neurotransmitters such as glutamate, dopamine, and gamma-aminobutyric acid (GABA) that can influence neuronal plasticity [38] and may be important to SCZ development.
UFD1L gene is located at chromosome 22q11.2, whose deletion is responsible for the DiGeorge syndrome, considered one of the main genetic risk factors for SCZ [39]; hence, genes located in this region may contribute to SCZ susceptibility [40]. UFD1L gene encodes for the human homolog of yeast ubiquitin-fusion-degradation 1 protein, expressed from fetal to post-natal period [41], possibly acting in neurodevelopment. It is suggested that UFD1L is directly associated with stress response in endoplasmatic reticulum (ER)-associated degradation (ERAD) [42], since this protein, in complex with others, extracts misfolded proteins from ER to be degraded in cytosol by the proteasome [43]. Polymorphism of UFD1L was previously associated with the disease [40, 44], with age of onset [40], and cognitive deficits in SCZ [45], but not to treatment resistance. In other study of our group, increased UFD1L expression was described in early stage of psychosis, comparing gene expression between individuals in ultra-high risk (UHR), those with first episode of psychosis (FEP), and HC. Higher levels of UFD1L mRNA was observed in those in UHR when compared to other groups, and pointed this gene as candidate as biomarker of prodromal phases of SCZ. Similarly to the findings in the UHR study, the upregulation of UFD1L observed in our patients could be associated with an increased demand of UFD1L protein in response to cellular stress, via ERAD pathways. Although reinforcing this gene importance in SCZ etiology, it is not clear how UFD1L mRNA expression varies across SCZ stages.
Beyond genes possibly related to the disease, we also detected alterations in mRNA levels of DICER1 and AKT1 genes in the blood of HC when compared to TRS. DICER1 and AKT1 expressions increased slightly across groups, and could differentiate only the extreme opposite groups, HC and TRS, not being associated with treatment resistance itself. Both genes were upregulated in first episode of psychosis patients who met bipolar disorder (with psychotic symptoms) diagnostic criteria in comparison to first episode of psychosis patients with SCZ diagnosis and HC [46]. Since AKT1 and DICER1 act in heterogeneous processes, it is not clear if the absence of differences between TRS and NTRS was due to some variable not evaluated in the TRS group, such as presence of other comorbidities and use of other pharmacological drugs, among others.
DICER1 is a ribonuclease that is central in the biogenesis of miRNAs, being responsible for the processing of the pre-miRNA hairpin structure to form mature miRNAs. Upregulation of DICER1 was already shown in dorsolateral prefrontal cortex [47, 48] and lymphoblastoid cell lines of SCZ patients [49]. Upregulation of several specific miRNAs was described both in the brain and blood of SCZ subjects [50], and in plasma of TRS individuals compared to NTRS individuals [51]. The upregulation of DICER1 levels observed in our TRS patients could be associated with an increased demand of enzymes that process miRNAs throughout the disease evolution, being more pronounced in the TRS group.
AKT1 is a member of the serine-threonine protein kinase involved in cell signaling and neural development [52]. Increased activity of AKT genes has been associated with several human diseases, including different types of cancer [53, 54]. Impairment of AKT signaling pathway was described in a number of brain disorders, including SCZ [54, 55], in which decreased protein and mRNA levels of AKT1 were reported in the blood and brain [56, 57]. Activation of AKT1 was described in the brain of mice after antipsychotic treatment [58], reaffirming a role of AKT1 in SCZ development. In the present study, different than others [58,59,60], we detected an increase of AKT1 mRNA levels in SCZ patients, specifically TRS patients, compared only to HC.
We did not detect any eQTLs in our sample that might explain differences in mRNA levels, suggesting a possible regulation by other mechanism, not driven by genotypes. Following this line, D’Addario and co-workers presented evidences of epigenetic regulation of CNR1 expression in humans and animal model [37]. Other agents, like virus and vitamins, can also change the expression of DICER1 [61, 62].
Our sample of TRS patients was affected earlier by SCZ in comparison to NTRS patients, and presented more severe psychopathology, in line to other cases described in literature, and reinforcing a possible pronounced genetic basis of the TR outcome [63,64,65].
In the present study, we detected differences of CNR1 and UFD1L expressions in the blood of healthy subjects and SCZ patients, independently of treatment response, highlighting the importance of these genes to SCZ etiological mechanisms. Besides, we also detected differences of AKT1 and DICER1 mRNA expressions between TRS patients and HC, showing these genes might be related to the severity rather than to the disease itself. All genes were upregulated in TRS compared to HC and seem to present a slightly increased expression throughout the disease progression.
Although these findings should be interpreted in light of some limitations, like the heterogeneity of disease and treatment resistance, and the absence of data related to possible confounders (like the use of medications not related to antipsychotic, BMI index, and the presence of other comorbidities), our results can provide clues of biological pathways involved in TRS that could contribute to better understand treatment response and to develop more effective treatments in the future.
References
Owen MJ, O'Donovan MC, Thapar A, Craddock N (2011) Neurodevelopmental hypothesis of schizophrenia. Br J Psychiatry 198(3):173–175. https://doi.org/10.1192/bjp.bp.110.084384
Henriksen MG, Nordgaard J, Jansson LB (2017) Genetics of schizophrenia: overview of methods, findings and limitations. Front Hum Neurosci 11:322. https://doi.org/10.3389/fnhum.2017.00322
Gejman PV, Sanders AR, Kendler KS (2011) Genetics of schizophrenia: new findings and challenges. Annu Rev Genomics Hum Genet 12(1):121–144. https://doi.org/10.1146/annurev-genom-082410-101459
Boyle EA, Li YI, Pritchard JK (2017) An expanded view of complex traits: from polygenic to omnigenic. Cell 169(7):1177–1186. https://doi.org/10.1016/j.cell.2017.05.038
Leucht S, Pitschel-Walz G, Abraham D, Kissling W (1999) Efficacy and extrapyramidal side-effects of the new antipsychotics olanzapine, quetiapine, risperidone, and sertindole compared to conventional antipsychotics and placebo. A meta-analysis of randomized controlled trials. Schizophr Res 35(1):51–68. https://doi.org/10.1016/S0920-9964(98)00105-4
Kapur S, Seeman P (2001) Does fast dissociation from the dopamine d(2) receptor explain the action of atypical antipsychotics?: a new hypothesis. Am J Psychiatry 158(3):360–369. https://doi.org/10.1176/appi.ajp.158.3.360
Kane JM, Honigfeld G, Singer J, Meltzer H (1988) Clozapine in treatment-resistant schizophrenics. Psychopharmacol Bull 24(1):62–67
Taylor DM, Duncan-McConnell D (2000) Refractory schizophrenia and atypical antipsychotics. J Psychopharmacol 14(4):409–418. https://doi.org/10.1177/026988110001400411
McGorry P, Killackey E, Elkins K, Lambert M, Lambert T (2003) Summary Australian and New Zealand clinical practice guideline for the treatment of schizophrenia. Australasian Psychiatry 2(11):136–147
Crespo-Facorro B, de la Foz VO, Ayesa-Arriola R, Perez-Iglesias R, Mata I, Suarez-Pinilla P et al (2013) Prediction of acute clinical response following a first episode of non affective psychosis: results of a cohort of 375 patients from the Spanish PAFIP study. Prog Neuro-Psychopharmacol Biol Psychiatry 44:162–167. https://doi.org/10.1016/j.pnpbp.2013.02.009
Huber CG, Naber D, Lambert M (2008) Incomplete remission and treatment resistance in first-episode psychosis: definition, prevalence and predictors. Expert Opin Pharmacother 9(12):2027–2038. https://doi.org/10.1517/14656566.9.12.2027
Meltzer HY (1999) The role of serotonin in antipsychotic drug action. Neuropsychopharmacology 21(2 Suppl):106S–115S. https://doi.org/10.1016/S0893-133X(99)00046-9
Meltzer HY (1990) Defining treatment refractoriness in schizophrenia. Schizophr Bull 16(4):563–565. https://doi.org/10.1093/schbul/16.4.563
IPAP. The International Psychopharmacology Algorithm Project. 2009 [cited 2017 21/12/2017]; Available from: http://www.ipap.org/schiz/index.php.
Andreasen NC, Pressler M, Nopoulos P, Miller D, Ho BC (2010 Feb 1) Antipsychotic dose equivalents and dose-years: a standardized method for comparing exposure to different drugs. Biol Psychiatry 67(3):255–262. https://doi.org/10.1016/j.biopsych.2009.08.040
Spindola LM, Pan PM, Moretti PN, Ota VK, Santoro ML, Cogo-Moreira H, Gadelha A, Salum G et al (2017) Gene expression in blood of children and adolescents: mediation between childhood maltreatment and major depressive disorder. J Psychiatr Res 92:24–30. https://doi.org/10.1016/j.jpsychires.2017.03.015
Hruz T, Laule O, Szabo G, Wessendorp F, Bleuler S, Oertle L et al (2008) Genevestigator v3: a reference expression database for the meta-analysis of transcriptomes. Adv Bioinforma 2008:420747
Christofolini DM, Bellucco FT, Ota VK, Belangero SI, Cernach MC, Gadelha A et al (2011) Assessment of 22q11.2 copy number variations in a sample of Brazilian schizophrenia patients. Schizophr Res 132(1):99–100. https://doi.org/10.1016/j.schres.2011.07.007
Gillespie AL, Samanaite R, Mill J, Egerton A, MacCabe JH (2017) Is treatment-resistant schizophrenia categorically distinct from treatment-responsive schizophrenia? A systematic review. BMC Psychiatry 17(1):12. https://doi.org/10.1186/s12888-016-1177-y
Anderson VM, Goldstein ME, Kydd RR, Russell BR (2015) Extensive gray matter volume reduction in treatment-resistant schizophrenia. Int J Neuropsychopharmacol 18(7):pyv016
Demjaha A, Egerton A, Murray RM, Kapur S, Howes OD, Stone JM, McGuire PK (2014) Antipsychotic treatment resistance in schizophrenia associated with elevated glutamate levels but normal dopamine function. Biol Psychiatry 75(5):e11–e13. https://doi.org/10.1016/j.biopsych.2013.06.011
Demjaha A, Murray RM, McGuire PK, Kapur S, Howes OD (2012) Dopamine synthesis capacity in patients with treatment-resistant schizophrenia. Am J Psychiatry 169(11):1203–1210. https://doi.org/10.1176/appi.ajp.2012.12010144
Krebs MO, Sautel F, Bourdel MC, Sokoloff P, Schwartz JC, Olie JP et al (1998) Dopamine D3 receptor gene variants and substance abuse in schizophrenia. Mol Psychiatry 3(4):337–341. https://doi.org/10.1038/sj.mp.4000411
Gouvea ES, Santos AFF, Ota VK, Mrad V, Gadelha A, Bressan RA et al (2017) The role of the CNR1 gene in schizophrenia: a systematic review including unpublished data. Rev Bras Psiquiatr 39(2):160–171. https://doi.org/10.1590/1516-4446-2016-1969
D'Souza DC, Abi-Saab WM, Madonick S, Forselius-Bielen K, Doersch A, Braley G et al (2005) Delta-9-tetrahydrocannabinol effects in schizophrenia: implications for cognition, psychosis, and addiction. Biol Psychiatry 57(6):594–608. https://doi.org/10.1016/j.biopsych.2004.12.006
Dinu IR, Popa S, Bicu M, Mota E, Mota M (2009) The implication of CNR1 gene’s polymorphisms in the modulation of endocannabinoid system effects. Rom J Intern Med 47(1):9–18
Okahisa Y, Kodama M, Takaki M, Inada T, Uchimura N, Yamada M, Iwata N, Iyo M et al (2011 Mar) Association study of two cannabinoid receptor genes, CNR1 and CNR2, with methamphetamine dependence. Curr Neuropharmacol 9(1):183–189. https://doi.org/10.2174/157015911795017191
Ujike H, Takaki M, Nakata K, Tanaka Y, Takeda T, Kodama M, Fujiwara Y, Sakai A et al (2002) CNR1, central cannabinoid receptor gene, associated with susceptibility to hebephrenic schizophrenia. Mol Psychiatry 7(5):515–518. https://doi.org/10.1038/sj.mp.4001029
Chavarria-Siles I, Contreras-Rojas J, Hare E, Walss-Bass C, Quezada P, Dassori A et al (2008) Cannabinoid receptor 1 gene (CNR1) and susceptibility to a quantitative phenotype for hebephrenic schizophrenia. Am J Med Genet B Neuropsychiatr Genet 147(3):279–284. https://doi.org/10.1002/ajmg.b.30592
Martinez-Gras I, Hoenicka J, Ponce G, Rodriguez-Jimenez R, Jimenez-Arriero MA, Perez-Hernandez E et al (2006) (AAT)n repeat in the cannabinoid receptor gene, CNR1: association with schizophrenia in a Spanish population. Eur Arch Psychiatry Clin Neurosci 256(7):437–441. https://doi.org/10.1007/s00406-006-0665-3
Hamdani N, Tabeze JP, Ramoz N, Ades J, Hamon M, Sarfati Y, Boni C, Gorwood P (2008) The CNR1 gene as a pharmacogenetic factor for antipsychotics rather than a susceptibility gene for schizophrenia. Eur Neuropsychopharmacol 18(1):34–40. https://doi.org/10.1016/j.euroneuro.2007.05.005
Grissom NM, Herdt CT, Desilets J, Lidsky-Everson J, Reyes TM (2015) Dissociable deficits of executive function caused by gestational adversity are linked to specific transcriptional changes in the prefrontal cortex. Neuropsychopharmacology 40(6):1353–1363. https://doi.org/10.1038/npp.2014.313
Kuntsi J, Frazier-Wood AC, Banaschewski T, Gill M, Miranda A, Oades RD, Roeyers H, Rothenberger A et al (2013) Genetic analysis of reaction time variability: room for improvement? Psychol Med 43(6):1323–1333. https://doi.org/10.1017/S0033291712002061
Johnson KA, Robertson IH, Kelly SP, Silk TJ, Barry E, Daibhis A et al (2007) Dissociation in performance of children with ADHD and high-functioning autism on a task of sustained attention. Neuropsychologia 45(10):2234–2245. https://doi.org/10.1016/j.neuropsychologia.2007.02.019
Ben Shalom D, Ronel Z, Faran Y, Meiri G, Gabis L, Kerns KA (2017) A double dissociation between inattentive and impulsive traits, on tasks of visual processing and emotion regulation. J Atten Disord 21(7):543–553. https://doi.org/10.1177/1087054713510351
Marco EM, Echeverry-Alzate V, Lopez-Moreno JA, Gine E, Penasco S, Viveros MP (2014) Consequences of early life stress on the expression of endocannabinoid-related genes in the rat brain. Behav Pharmacol 25(5–6):547–556. https://doi.org/10.1097/FBP.0000000000000068
D'Addario C, Micale V, Di Bartolomeo M, Stark T, Pucci M, Sulcova A, Palazzo M, Babinska Z et al (2017) A preliminary study of endocannabinoid system regulation in psychosis: distinct alterations of CNR1 promoter DNA methylation in patients with schizophrenia. Schizophr Res 188:132–140. https://doi.org/10.1016/j.schres.2017.01.022
Wilson RI, Nicoll RA (2002) Endocannabinoid signaling in the brain. Science 296(5568):678–682. https://doi.org/10.1126/science.1063545
Bassett AS, Chow EW, Husted J, Weksberg R, Caluseriu O, Webb GD, Gatzoulis MA (2005) Clinical features of 78 adults with 22q11 deletion syndrome. Am J Med Genet A 138(4):307–313. https://doi.org/10.1002/ajmg.a.30984
Ota VK, Belangero SI, Gadelha A, Bellucco FT, Christofolini DM, Mancini TI, Ribeiro-dos-Santos ÂK, Santos SE et al (2010) The UFD1L rs5992403 polymorphism is associated with age at onset of schizophrenia. J Psychiatr Res 44(15):1113–1115. https://doi.org/10.1016/j.jpsychires.2010.04.008
Novelli G, Mari A, Amati F, Colosimo A, Sangiuolo F, Bengala M, Conti E, Ratti A et al (1998) Structure and expression of the human ubiquitin fusion-degradation gene (UFD1L). Biochim Biophys Acta 1396(2):158–162. https://doi.org/10.1016/S0167-4781(97)00211-X
Chen M, Gutierrez GJ, Ronai ZA (2011) Ubiquitin-recognition protein Ufd1 couples the endoplasmic reticulum (ER) stress response to cell cycle control. Proc Natl Acad Sci U S A 108(22):9119–9124. https://doi.org/10.1073/pnas.1100028108
Ye Y, Meyer HH, Rapoport TA (2001) The AAA ATPase Cdc48/p97 and its partners transport proteins from the ER into the cytosol. Nature. [Research Support, Non-U.S. Gov't Research Support, U.S. Gov't, P.H.S.] 414(6864):652–656
De Luca A, Pasini A, Amati F, Botta A, Spalletta G, Alimenti S et al (2001) Association study of a promoter polymorphism of UFD1L gene with schizophrenia. Am J medical Genetics. [Multicenter Study Research Support, Non-U.S. Gov't] 105(6):529–533
Ota VK, Berberian AA, Gadelha A, Santoro ML, Ottoni GL, Matsuzaka CT, Mari JJ, Melaragno MI et al (2013 Aug 30) Polymorphisms in schizophrenia candidate gene UFD1L may contribute to cognitive deficits. Psychiatry Res 209(1):110–113. https://doi.org/10.1016/j.psychres.2013.03.035
Gouvea ES, Ota VK, Noto C, Santoro ML, Spindola LM, Moretti PN, Carvalho CM, Xavier G et al (2016 Oct 04) Gene expression alterations related to mania and psychosis in peripheral blood of patients with a first episode of psychosis. Transl Psychiatry 6(10):e908. https://doi.org/10.1038/tp.2016.159
Santarelli DM, Beveridge NJ, Tooney PA, Cairns MJ (2011) Upregulation of dicer and microRNA expression in the dorsolateral prefrontal cortex Brodmann area 46 in schizophrenia. Biol Psychiatry 69(2):180–187. https://doi.org/10.1016/j.biopsych.2010.09.030
Beveridge NJ, Gardiner E, Carroll AP, Tooney PA, Cairns MJ (2010) Schizophrenia is associated with an increase in cortical microRNA biogenesis. Mol Psychiatry 15(12):1176–1189. https://doi.org/10.1038/mp.2009.84
Sanders AR, Goring HH, Duan J, Drigalenko EI, Moy W, Freda J et al (2013) Transcriptome study of differential expression in schizophrenia. Hum Mol Genet 22(24):5001–5014. https://doi.org/10.1093/hmg/ddt350
Camkurt MA, Karababa F, Erdal ME, Bayazit H, Kandemir SB, Ay ME et al (2016) Investigation of dysregulation of several microRNAs in peripheral blood of schizophrenia patients. Clin Psychopharmacol Neurosci 14(3):256–260. https://doi.org/10.9758/cpn.2016.14.3.256
Alacam H, Akgun S, Akca H, Ozturk O, Kabukcu BB, Herken H (2016) miR-181b-5p, miR-195-5p and miR-301a-3p are related with treatment resistance in schizophrenia. Psychiatry Res 245:200–206. https://doi.org/10.1016/j.psychres.2016.08.037
Balu DT, Carlson GC, Talbot K, Kazi H, Hill-Smith TE, Easton RM, Birnbaum MJ, Lucki I (2012 Feb) Akt1 deficiency in schizophrenia and impairment of hippocampal plasticity and function. Hippocampus 22(2):230–240. https://doi.org/10.1002/hipo.20887
Woodgett JR (2005) Recent advances in the protein kinase B signaling pathway. Curr Opin Cell Biol 17(2):150–157. https://doi.org/10.1016/j.ceb.2005.02.010
Arguello PA, Gogos JA (2008) A signaling pathway AKTing up in schizophrenia. J Clin Invest 118(6):2018–2021. https://doi.org/10.1172/JCI35931
Emamian ES (2012) AKT/GSK3 signaling pathway and schizophrenia. Front Mol Neurosci 5:33
Aubry JM, Schwald M, Ballmann E, Karege F (2009) Early effects of mood stabilizers on the Akt/GSK-3beta signaling pathway and on cell survival and proliferation. Psychopharmacology 205(3):419–429. https://doi.org/10.1007/s00213-009-1551-2
Tan HY, Nicodemus KK, Chen Q, Li Z, Brooke JK, Honea R, Kolachana BS, Straub RE et al (2008) Genetic variation in AKT1 is linked to dopamine-associated prefrontal cortical structure and function in humans. J Clin Invest 118(6):2200–2208. https://doi.org/10.1172/JCI34725
Emamian ES, Hall D, Birnbaum MJ, Karayiorgou M, Gogos JA (2004) Convergent evidence for impaired AKT1-GSK3beta signaling in schizophrenia. Nat Genet 36(2):131–137. https://doi.org/10.1038/ng1296
Karege F, Perroud N, Schurhoff F, Meary A, Marillier G, Burkhardt S et al (2010) Association of AKT1 gene variants and protein expression in both schizophrenia and bipolar disorder. Genes Brain Behav 9(5):503–511. https://doi.org/10.1111/j.1601-183X.2010.00578.x
van Beveren NJ, Buitendijk GH, Swagemakers S, Krab LC, Roder C, de Haan L et al (2012) Marked reduction of AKT1 expression and deregulation of AKT1-associated pathways in peripheral blood mononuclear cells of schizophrenia patients. PLoS One 7(2):e32618. https://doi.org/10.1371/journal.pone.0032618
Casseb SM, Simith DB, Melo KF, Mendonca MH, Santos AC, Carvalho VL, Cruz ACR, Vasconcelos PFC (2016) Drosha, DGCR8, and Dicer mRNAs are down-regulated in human cells infected with dengue virus 4, and play a role in viral pathogenesis. Genet Mol Res 15(2). https://doi.org/10.4238/gmr.15027891
Dambal S, Giangreco AA, Acosta AM, Fairchild A, Richards Z, Deaton R, Wagner D, Vieth R et al (2017) microRNAs and DICER1 are regulated by 1,25-dihydroxyvitamin D in prostate stroma. J Steroid Biochem Mol Biol 167:192–202. https://doi.org/10.1016/j.jsbmb.2017.01.004
Meltzer HY, Rabinowitz J, Lee MA, Cola PA, Ranjan R, Findling RL, Thompson PA (1997 Apr) Age at onset and gender of schizophrenic patients in relation to neuroleptic resistance. Am J Psychiatry 154(4):475–482. https://doi.org/10.1176/ajp.154.4.475
Lieberman JA, Safferman AZ, Pollack S, Szymanski S, Johns C, Howard A, Kronig M, Bookstein P et al (1994) Clinical effects of clozapine in chronic schizophrenia: response to treatment and predictors of outcome. Am J Psychiatry 151(12):1744–1752. https://doi.org/10.1176/ajp.151.12.1744
Quintero J, Barbudo del Cura E, Lopez-Ibor MI, Lopez-Ibor JJ (2011) The evolving concept of treatment-resistant schizophrenia. Actas Esp Psiquiatr 39(4):236–250
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The Research Ethics Committee of UNIFESP approved the study, and all the participants and family members provided written informed consent prior to enrolment in the study (CAAE 06191612.7.0000.5505).
Rights and permissions
About this article
Cite this article
Moretti, P.N., Ota, V.K., Gouvea, E.S. et al. Accessing Gene Expression in Treatment-Resistant Schizophrenia. Mol Neurobiol 55, 7000–7008 (2018). https://doi.org/10.1007/s12035-018-0876-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-018-0876-4