Abstract
While prion diseases have been described in numerous species, some, including those of the Canidae family, appear to show resistance or reduced susceptibility. A better understanding of the factors underlying prion susceptibility is crucial for the development of effective treatment and control measures. We recently demonstrated resistance to prion infection in mice overexpressing a mutated prion protein (PrP) carrying a specific amino acid substitution characteristic of canids. Here, we show that coexpression of this mutated PrP and wild-type mouse PrP in transgenic mice inoculated with different mouse-adapted prion strains (22 L, ME7, RML, and 301C) significantly increases survival times (by 45 to 113%). These data indicate that this amino acid substitution confers a dominant-negative effect on PrP, attenuating the conversion of PrPC to PrPSc and delaying disease onset without altering the neuropathological properties of the prion strains. Taken together, these findings have important implications for the development of new treatment approaches for prion diseases based on dominant-negative proteins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Transmissible spongiform encephalopathies (TSEs), or prion diseases, are a group of neurodegenerative diseases of animals and humans that can be sporadic (putatively spontaneous), genetic, or acquired by infection [1]. TSEs are caused by the accumulation of a misfolded protein, the scrapie-associated prion protein (PrPSc), which is produced by posttranslational conversion of the physiologically expressed cellular prion protein (PrPC) via an unknown mechanism. This abnormal form of the protein is protease resistant and is composed almost entirely of β-sheet structures [2,3,4,5]. PrPSc deposition results in spongiosis, vacuolation, neuronal death, and glial reactions in the central nervous system of affected individuals [3, 6,7,8].
TSEs naturally affect a wide variety of mammalian species and include Creutzfeldt–Jakob disease (CJD) in humans, scrapie in sheep and goats, bovine spongiform encephalopathy (BSE) in cattle, and chronic wasting disease (CWD) in cervids [9]. Since the emergence of BSE and its association with variant CJD (vCJD) in humans [10, 11], transmission of prion diseases between species has become a major public health concern. Spongiform encephalopathies have been identified in numerous ruminant, feline, and primate species, all of which had consumed cattle meat or feed containing ruminant meat and bone meal, or were in close proximity to infected animals [12,13,14]. However, the absence of prion diseases in other mammals exposed to contaminated food, including rabbits, equids, and canids, suggested the existence of prion-resistant species [15]. This was further supported by unsuccessful attempts to overcome TSE transmission barriers in those species, which contributed to preserve for decades the concept of prion-resistant mammals [16]. Of all putative prion-resistant species, rabbits are the most extensively studied. In vitro studies using protein misfolding cyclic amplification (PMCA) and subsequent in vivo experiments have shown that rabbits are not disease-resistant per se, but are poorly susceptible to prion diseases [17]. Equids represent a very interesting group of mammals that, while not completely resistant to TSEs, show a very peculiar susceptibility displaying an unusual replicative phenomenon termed nonadaptative prion amplification (NAPA), described in transgenic mice expressing horse PrP [18]. Moreover, the use of recombinant proteins in the presence of chaotropic agents [19] and in vitro prion amplification techniques [20] to study the propensity of prion protein misfolding in different mammalian species suggest that susceptibility to prion diseases is lowest in canids. To identify the specific features of canine PrPC that account for its strong resistance to misfolding, we previously generated a transgenic mouse model expressing a PrP variant (N158D PrP), containing a single specific amino acid substitution, characteristic of the dog PrPC [21]. We found that this model was completely resistant to intracerebral infection with several mouse-adapted prion strains, indicating that a single amino acid substitution is sufficient to inhibit the misfolding of the mutated protein.
In the present study, we investigated whether this mutant could act as a dominant-negative protein and prevent PrPSc formation when coexpressed with wild-type (wt) PrPC. To this end, we created a new mouse model coexpressing wt mouse PrPC and the aforementioned mutant PrP variant carrying the critical dog amino acid substitution. These mice were intracerebrally inoculated with different mouse-adapted prion strains and the results of the in vivo challenge compared with those obtained in mice expressing comparable levels of wt mouse prion protein. Surprisingly, coexpression of the mutated protein significantly delayed the onset of disease induced by all prion strains studied. Survival periods were increased by 45% to 113% with respect to mice expressing wt protein alone, thereby demonstrating the dominant-negative effect of the mutant protein. Our findings show that this specific dog amino acid substitution confers the protein the ability to interfere with the propagation of wt prions in transgenic mice. These findings have important implications for the development of therapeutic strategies against prion diseases.
Materials and Methods
Generation and Inoculation of Transgenic Mouse Models
Three different transgenic mouse models were used in the present study: (1) Tga20xTga20 mice (hereafter referred to as Tga20 mice) expressing mouse PrPC at a levels ∼ 8-fold higher than those observed in the mouse brain [22]; (2) Tga20xPrnp 0/0 [23] (hereafter referred to as Tga20xKO mice) mice expressing mouse PrPC at a levels ∼ 4-fold higher than those observed in mouse brain; and (3) Tga20xTgN158D mice (hereafter referred to as Tga20xN158D mice) [21]expressing mouse PrPC at levels ∼ 4-fold higher and N158D mouse PrPC at levels ∼ 2-fold higher than those observed in mouse brain. The murine PRNP promoter was used for N158D mouse PrPC expression.
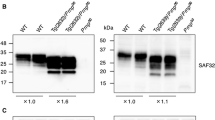
PrP expression levels from Tga20, Tga20xKO, and Tga20xN158D mice were analyzed by Western blot using SAF83 (1:400) and 5C6 (1:2000) monoclonal antibodies and compared with those obtained in TgN158DxTgN158D mice (hereafter referred to as TgN158D mice), expressing only N158D mouse PrPC [21]. 5C6 antibody (PRC5 antibody) was kindly provided by Dr. Glenn Telling (Prion Research Center, Colorado State University). This antibody requires asparagine at mouse PrP residue 158 [24] and therefore does not detect N158D PrP, whereas SAF83 antibody recognizes both wt and N158D mouse PrPs (Supplementary Fig. 1).
Mice aged 6 to 8 weeks were anesthetized with isoflurane and intracerebrally inoculated (left cerebral hemisphere) with mouse-adapted prion strains 22L, RML, ME7, or 301C using 20 μl of a 10% brain homogenate. Injections were administered using a 50-μl syringe and a 25-G needle. Analgesia was achieved by subcutaneous injection of buprenorphine (0.3 mg/kg). Animals were subsequently housed in filtered cages and monitored three times per week for neurologic dysfunction. Mice were euthanized by cervical dislocation upon detection of clinical signs of terminal disease (severe ataxia, inability to stand and poor body condition).
All experimental procedures were approved by the Ethics Committee for Animal Experiments of the University of Zaragoza (permit number PI32/13) and performed in accordance with the recommendations for the care and use of experimental animals and in agreement with Spanish law (R.D. 1201/05).
Sample Processing and Histopathological Evaluation
After euthanasia, brains were removed, and transversal sections from the frontal cortex and medulla oblongata were separated and frozen at − 80 °C for subsequent biochemical analyses. The remaining tissue was fixed in 10% formalin for neuropathological studies. After fixation, brains were cut at four standard levels for the histological evaluation of the following nine brain regions: frontal cortex (Fc), septal area (Sa), thalamic cortex (Tc), hippocampus (Hc), thalamus (T), hypothalamus (Ht), mesencephalon (Mes), cerebellum (Cbl), and medulla oblongata (Mo) [25]. Tissues were embedded in paraffin, cut into 4-μm-thick sections on a microtome, and mounted on glass slides for staining with hematoxylin and eosin. Sections were examined using an optical microscope (Zeiss Axioskop 40), and the extent of vacuolation and spongiosis in each area was blindly evaluated and semi-quantitatively scored on a scale of 0 (absence of lesions) to 5 (high intensity lesions).
Analysis of PrPSc Deposition
The intensity and distribution of PrPSc deposition was evaluated using the paraffin-embedded tissue (PET) blot method, as previously described [26]. Sections from paraffin-embedded brains (4 μm thick) were collected on a nitrocellulose membrane (0.45-μm pore size; Bio-Rad, Richmond, CA) and dried at 55 °C for 24 h. After deparaffination and rehydration, sections were digested for 2 h at 56 °C with 250 μg/ml proteinase-K (PK) (Applied Biosystems) in PK digestion buffer containing TBS (Tris-buffered saline) and 0.1% Brij 35P (Sigma-Aldrich). After washing with TBST (Tris buffered saline; 0.05% Tween 20), membrane-attached proteins were denatured in 3 M guanidine thiocyanate (Sigma-Aldrich). Sections were then blocked with 1% casein in TBST and incubated with Sha31 primary monoclonal antibody (1:8000; SPI-Bio). After incubation with an alkaline phosphatase-coupled goat anti-mouse antibody (DAKO), immunostaining was visualized using NBT/BCIP (Nitro blue tetrazolium/5-bromo-4-chloro-3-indolyl-phosphate; Sigma-Aldich). PrPSc deposits were evaluated semi-quantitatively, as described for spongiform lesions, using a Zeiss Stemi DV4 stereomicroscope.
Histological Analysis of PrPC Distribution
The localization and distribution of PrPC in the brains of Tga20xN158D mice was analyzed by immunohistochemistry. Brains from TgN158D mice were used as controls. Serial paraffin-embedded sections were incubated with a peroxidase blocking reagent (Dako) for 20 min followed by hydrated autoclaving at 100 °C in citrate buffer for 30 min. Immunodetection was performed overnight at 4 °C using SAF32 (1:1000; SPI-Bio) and 5C6 (1:1000) anti-PrP monoclonal antibodies. The anti-mouse Envision polymer (Dako) was used as the visualization system and DAB (diaminobenzidine, Dako) as the chromogen.
The localization of N158D PrP was analyzed using immunofluorescence and confocal imaging. Immunofluorescence staining was performed as described previously [27], with specific modifications to adapt the protocol to paraffin-embedded samples. Paraffin-embedded tissue sections from TgN158D mice were deparaffinated and rehydrated and then blocked with 1% H2O2 for 30 min. After pretreatment with 0.1% Triton X-100 for 3 h at room temperature, samples were subjected to hydrated autoclaving and incubated with SAF32 antibody (1:100) followed by a goat anti-mouse IgG biotin conjugate (1:100; Invitrogen) and an Alexa fluor 594 streptavidin conjugate (1:1000; Invitrogen). Sections were analyzed using a Zeiss laser-scanning confocal microscope LSM 510 (Carl Zeiss MicroImaging).
Biochemical Analysis
Frozen brain sections stored for biochemical analysis, as described above, were homogenized in 1% (w/v) in PBS (phosphate-buffered saline) using a ribolyzer. The resulting samples were digested with Protease K (PK) for 1 h at 42 °C and used for Western blot. Immunodetection was performed using SAF83 (1: 400; SPI-Bio) and 5C6 (1:2000) primary antibodies.
Data Analysis
Survival times were analyzed by Kaplan-Meier survival analysis, and the resulting survival curves were compared using the log rank test (α = 0.050). Differences in spongiform lesions (distribution and intensity) and PrPSc deposition profiles between different transgenic mouse models were evaluated using the nonparametric Mann-Whitney U test and considered significant at p < 0.05. GraphPad Prism version 6.0 (GraphPad Software, La Jolla, CA, USA) was used to perform all statistical analyses, generate Kaplan Meier curves, and to graph histopathology results.
Results
Coexpression of the N158D PrP Substitution Greatly Increases Survival Time in Inoculated Mice
Three groups of mice expressing different levels of wt protein, either alone or together with N158D PrP [(Tga20 (8x), Tga20xKO (4x+0x), and Tga20xN158D (4x+2x)] (Supplementary Fig. 1) were challenged by intracerebral inoculation with the 22L, RML, 301C, or ME7 mouse-adapted prion strains. Owing to its high levels of PrPC expression in the brain (∼ 8-fold higher than wt mice), the TSE incubation period in the Tga20 mouse is relatively short, making it a useful model for prion research. Moreover, the histopathological and biochemical features of several mouse-adapted TSE strains, including those used in the present study, are well defined in this transgenic model [22, 28]. Tga20xN158D mice were used to achieve coexpression of the wt and N158D PrP. Tga20xKO mice were selected as controls, given that their wt PrPC expression level is identical to that of Tga20xN158D mice. No significant differences were observed in electrophoretic migration patterns between the different mouse lines (Supplementary Fig. 1). In addition, both wt and N158D PrPs are present in high amounts and are distributed normally in the brain of transgenic mice (Supplementary Fig. 2).
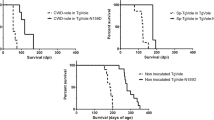
Tga20xN158D mice inoculated with the 22L, RML, 301C, or ME7 prion strains showed increases in survival times of 113, 45, 71, and 49%, respectively, as compared with Tga20xKO mice, which express the same amount of wt PrPC (Table 1). Significant differences were observed between genotypes for all inocula (Fig. 1). In the case of the 301C strain, survival times in Tga20xN158D mice were not as homogeneous as those observed for the other strains, as evidenced by less steep decline in the Kaplan-Meier curve (Fig. 1).
Survival curves for Tga20, Tga20xKO, and Tga20xN158D mice challenged with different mouse-adapted prion strains. Comparison of Tga20xN158D curves with Tga20xKO curves using the log rank test (α = 0.050) revealed very significant differences for the 22L, RML, and ME7 (p < 0.0001) and the 301C (p < 0.0033) inoculation groups. Survival curves for Tga20 mice inoculated with the corresponding strains are also shown. Tga20xN158D mice infected with the 22L, RML, 301C, or ME7 prion strains showed relative increases in survival times of 113, 45, 71, and 49%, respectively, when compared with those of Tga20xKO mice
Although survival time was significantly increased in Tga20xN158D mice, the clinical presentation in these mice was indistinguishable from that of mice expressing only the wt protein. Mice developed clinical signs typical of TSEs in rodents, including poor hair coat condition and kyphosis in the early stages of the disease, followed by proprioceptive deficits, head twitching, and progressive ataxia, which became severe in terminal stages. However, certain clinical signs were more evident in animals infected with a given inoculum (e.g., the opisthotonos observed in all three genotypes of mice inoculated with the 22L strain, a sign that may be due to cerebellar lesions).
Expression of the Dominant-Negative Protein Did Not Alter the Neuropathological Features of the Disease
Despite the substantial prolongation of survival time in mice coexpressing the N158D PrP substitution, an exhaustive comparison of the two models expressing equivalent levels of wt PrPC (Tga20xKO and Tga20xN158D mice) revealed no significant differences in terms of the neuropathological characteristics of the disease. Lesion profiles and prion protein deposition patterns, evaluated semi-quantitatively on a scale of 0 to 5, were very similar between different genotypes inoculated with the same strain (Figs. 2 and 3). Our results are consistent with those of another study in which Tga20 mice were infected with all the same inocula [28], indicating that the mouse-adapted strains used in the present study retained their characteristic histopathological features and PrPSc deposition profiles. All mice infected with the 22L strain developed particularly severe spongiform lesions and showed marked PrPSc deposition in the T, Ht, Mes, and Mo (Figs. 2 and 3a). Inoculation with the RML strain resulted in intense histopathological changes and PrPSc deposition predominantly in the T, Mes, and Mo (Figs. 2 and 3a), with low vacuolation scores observed in the Cbl. The spongiform lesions caused by the 301C strain were mainly located in the T, Mes, and Mo (Fig. 2). Compared with the other strains used, this mouse-adapted BSE strain produced slightly less intense PrPSc deposition throughout the brain (Fig. 3a), as described previously in wt mice [29]. Finally, inoculation with ME7 resulted in severe spongiosis and vacuolation in the Sa, T, Ht, and brainstem (Fig. 2) and very intense PrPSc deposition in the Hc, T, and Sa in all genotypes (Fig. 3a), indicating that the main features of the ME7 strain, as previously described in Tga20 mice [28], were preserved (Fig. 3b).
Brain lesion profiles of Tga20, Tga20xKO, and Tga20xN158D mice inoculated with different mouse-adapted prion strains. Spongiosis and vacuolation were evaluated semiquantitatively on a scale of 0 (absence of lesions) to 5 (high intensity lesions) in the following nine brain areas: frontal cortex (Fc), septal area (Sa), thalamic cortex (Tc), hippocampus (Hc), thalamus (T), hypothalamus (Ht), mesencephalon (Mes), cerebellum (Cbl), and medulla oblongata (Mo). Comparison of the lesion profiles of Tga20xKO and Tga20xN158D mice revealed a very similar lesion distribution (*p < 0.05, Mann-Whitney U test)
a PrPSc deposition profiles in the brains of Tga20, Tga20xKO, and Tga20xN158D mice inoculated with 22L, RML, 301C, or ME7 prion strains. PrPSc deposition was evaluated semiquantitatively on a scale of 0 (absence of deposits) to 5 (high intensity deposition) in the following nine brain areas: frontal cortex (Fc), septal area (Sa), thalamic cortex (Tc), hippocampus (Hc), thalamus (T), hypothalamus (Ht), mesencephalon (Mes), cerebellum (Cbl), and medulla oblongata (Mo). Comparison of the PrPSc deposition profiles of Tga20xKO and Tga20xN158D mice revealed almost identical PrPSc profiles (*p < 0.05, Mann-Whitney U test). b PET blot images of coronal sections of the mesencephalon from Tga20xKO and Tga20xN158D mice inoculated with the RML or ME7 strain. Note that the PrPSc deposition profile of mice expressing the mutant PrP is almost identical to that of Tga20xKO mice. Moreover, the characteristic deposition patterns of the inoculated strains are retained: note the marked deposition in the hippocampus in ME7-inoculated mice (arrows), a feature not observed in RML-inoculated mice
The results of biochemical analyses were consistent with the histopathological findings. No significant differences in PrP glycosylation and electrophoretic mobility patterns between Tga20, Tga20xKO, and Tga20xN158D mice were observed for any of the strains inoculated (Fig. 4). We further investigated whether the similarities observed between Tga20xKO and Tga20xN158D mice regarding the histopathological and biochemical features of the disease could be related to an exclusive conversion of wt PrPC. Serial dilutions of brain homogenates from 22L infected Tga20xKO and Tga20xN158D mice were analyzed for PrPres using two different antibodies: 5C6, which is unable to detect N158D PrP since it requires the presence of asparagine at codon 158 [24], and SAF83, which detects both wt and N158D PrPs. No differences were observed in the amount of PrPres detected by these antibodies in Tga20xN158D mice, indicating that only wt PrPC was converted (Supplementary Fig. 3).
PrPres detection from 22L, ME7, 301C, and RML inoculated Tga20, Tga20xKO, and Tga20xN158D mouse brains. Ten percent brain homogenates from 22L, ME7, 301C, and RML inoculated Tga20, Tga20xKO, and Tga20xN158D mice were digested with 80 μg/ml of Protease K (PK). Digested samples were analyzed by Western blot using SAF83 (1:400). No significant differences are observed between any of the Tga20, Tga20xKO, and Tga20xN158D brain homogenates suggesting that N158D PrPC did not alter the major biochemical characteristics of any of the four prion strains. Control: undigested Tga20xKO brain homogenate. Mw molecular weight
Discussion
Certain PrP polymorphisms are strongly linked to susceptibility/resistance to prion diseases. This relationship has been well documented in sheep, leading to the establishment of five haplotypic PrP gene variants associated with scrapie susceptibility [30]. Among the three main polymorphisms of ovine PRNP, variations at codon 171 appear to be the principal determinants of resistance to classical scrapie; sheep with arginine at this specific residue are resistant to natural [31, 32] and experimental [33] scrapie infection. Heterozygosity at certain PrP positions also exerts protective effects against human prion diseases [34].
The ability of certain variant proteins to interfere with coexpressed wt PrP and block prion replication is known as a dominant-negative effect. This has been experimentally reproduced in cells and in transgenic mice and may have implications for the development of therapeutic strategies for prion diseases [35,36,37,38]. The use of PMCA in in vitro studies has proved an efficient means of testing a wide variety of PrPs with different substitutions in order to identify the most appropriate dominant-negative changes [16].
In the search for PrPs that exert a consistent and potent inhibitory effect on in vivo prion propagation, it seems reasonable to begin with PrPs from species with demonstrated low susceptibility to prion diseases. For the purposes of this study, we selected dog prion protein, in which low susceptibility has been proven [19, 20]. Using cell and brain-based PMCA, we previously demonstrated that the substitution of asparagine with aspartic or glutamic acid at codon 163, a distinctive substitution from the Canidae family [39], strongly inhibits prion replication in vitro. Moreover, we found that when transgenic mice overexpressing a PrP variant carrying this substitution were challenged with several mouse-adapted prion strains, they were completely resistant to prion infection [21]. Based on these findings, we investigated whether the coexpression of this mutant PrP together with wt mouse PrP could interfere with prion propagation, thereby preventing or delaying the onset of the disease in vivo.
Coexpression of both proteins dramatically increased survival times after inoculation with any of the four mouse-adapted prion strains tested (22L, RML, 301C, and ME7). Furthermore, survival times in Tga20 and Tga20xKO mice differed significantly. This was not unexpected since PrPC expression levels dramatically influence the incubation time in prion diseases, and expression levels of PrPC are inversely proportional to the duration of the survival period [40, 41]. However, it is important to note that, in our study, the appropriate comparison of survival period is with that of mice expressing an equivalent amount of wt PrP (i.e., Tga20xKO vs. Tga20xN158D mice; Table 1).
The elongation of the survival times produced by the coexpression of an exogenous protein can be the result of several processes. We observed that, when PrPres levels from Tga20xKO and Tga20xN158D mice culled at different days postinoculation (dpi) were compared, Tga20xKO mice showed higher amounts of PrPres, even at shorter incubation periods than Tga20xN158D mice (Supplementary Fig. 4). Thus, we can suggest that the longer survival times observed in Tga20xN158D mice may be due to a slower rate of misfolding of the wt PrP, therefore producing a delayed accumulation of PrPSc. However, the molecular mechanisms by which N158D PrP delays prion propagation remain unclear. Several theories, most of them developed using scrapie-infected cell models, have been proposed to explain how dominant-negative proteins inhibit prion propagation. Although differing only at one position from the wt PrP, dominant-negative proteins could obstruct the interactions between similar PrP monomers [40, 42,43,44] since the difference between mutant and wt PrP could make them structurally incompatible [45]. This dissimilarity could interfere with the rate of formation [38] and the stability of PrPSc polymers [42, 46]. In addition, it has been also proposed that dominant-negative proteins may compete with wt PrPC for binding to newly formed PrPSc molecules [46, 47]. Thus, the prolongation in survival times observed in the present study might also be the result of a greater affinity of N158D PrP for interacting with PrPSc than that of wt PrP. Due to the apparent resistance of N158D PrP to misfold (Supplementary Fig. 3), a competition of this mutant and wt PrP for the same binding site in PrPSc would explain the delay of the disease observed in Tga20xN158D mice, as previously reported [46, 47].
Although survival times were significantly increased in Tga20xN158D mice inoculated with all experimental strains, this effect was not homogeneous for all strains. The greatest increase was observed in mice inoculated with the 22L strain: survival time in mice carrying the N158D PrP variant was 113% longer than that of controls. The smallest increase in survival times was observed in RML-inoculated Tga20xN158D mice (45% increase). It is well demonstrated that when propagated in vivo, distinct mouse-adapted prion strains differ in terms of incubation period, as well as their biochemical and neuropathological features [48,49,50,51]. Strains can also show biophysical, molecular, and, as in the case of the strains used in the present study, ultrastructural differences [52, 53]. These findings could explain that different tertiary and/or quaternary structures were also differentially affected by the blockade of a dominant-negative protein. Our findings suggest that the dominant-negative effect of this mutant protein is stronger with certain strains (22L and 301C) than with others (RML and ME7). Other dominant-negative proteins have been also reported to interfere with the generation of PrPSc in a strain-specific manner. As an example, Q218K PrP strongly inhibits the misfolding of coexpressed wt PrP in Chandler-infected cells but produces a much weaker inhibition with 22L strain. This distinct effect was attributed to the structural differences, determined by IR spectroscopy, between Chandler and 22L strains [54]. As aforementioned, we cannot know for certain what precise molecular mechanisms are involved in the partial dominance exerted by N158D PrP. However, if the dominant-negative protein blocks fibril growth, ultrastructural differences between strains could account for the differential effect (45–113% increase) of the dominant-negative protein on the survival period.
The dominant-negative effect of certain mutant PrPs on PrPSc formation has been already demonstrated in vivo in transgenic mice coexpressing wt PrP [37]. In that study, mice expressing PrPs containing ovine and human TSE resistance-associated substitutions were not completely resistant to prion formation when mutant and wt mouse PrPs were coexpressed. Our findings are in agreement with those results and demonstrate that minimal amino acid changes can produce highly efficient dominant-negative variants able to double the survival period when coexpressed with wt mouse PrPC. Extrapolating these findings to humans, in which the incubation period of prion diseases can last for decades, it seems possible that affected individuals may never develop clinical signs. The ability shown by certain PrP molecules with single residue substitutions to interfere with the misfolding of the endogenous PrPC has already been demonstrated. However, most of the approaches have been performed using cell cultures [35, 42, 54], whereas in vivo studies are limited [37, 55]. In addition, the dominant-negative effects described for this type of molecules, albeit potent, have been demonstrated against a limited number of strains [37, 42, 54]. Herein, we show that N158D PrP produces a dominant-negative inhibition in the propagation of a variety of prion strains, of both scrapie (22L, RML, and ME7) and BSE (301C) origins. The delay of the disease was not homogeneous among the strains despite showing inhibition against the propagation of all of them. When describing dominant-negative PrPs, it is important to check their ability to interfere with the propagation of prions from different origins and characteristics. Other naturally occurring amino acid variants of PrPC, such as sheep Q171R, have demonstrated a strong dominant-negative inhibition in the propagation of scrapie strains [35,36,37, 46]. However, it has been shown that sheep with Q171R are susceptible to atypical scrapie [56] as well as BSE [57]. Thus, our study suggests that N158D PrP, a substitution found in canids in which no natural prion diseases have been reported, may be a dominant-negative protein with a broader inhibitory effect.
The prolongation of the incubation period seen in the present study was less dramatic than that reported by Perrier and coworkers in RML-inoculated mice coexpressing equal amounts of wt PrP and dominant-negative PrP. However, wt PrP expression levels in our Tga20xN158D mice are four times higher than those of wt mice, making it more difficult to fully block prion formation. It cannot be ruled out that if an equimolecular amount of dominant-negative PrP and wt PrP is required for the complete blockade of prion propagation, we would need to double the amount of N158D PrP. The dose-dependent, dominant-negative inhibition by other similar molecules has already been demonstrated [37, 42, 46], showing that certain dominant-negative proteins need to be present in high amounts to inhibit endogenous PrPC conversion [37, 46]. We have observed that N158D PrP, even being expressed at lower levels than wt PrP, is able to significantly extend the survival period in Tga20xN158D mice.
The neuropathological changes seen in our Tga20xN158D mice were very similar to those observed in mice expressing only wt PrP, with few significant differences observed in terms of lesion and PrPSc deposition profiles (Figs. 2 and 3). These findings, coupled with the complete resistance to intracerebral challenge seen in mice expressing N158D mutant protein only [21], could lead us to think that the pathological form detected, and therefore, the neuropathological hallmarks observed in Tga20xN158D mice are due only to the conversion of the mouse wt protein. Fortunately, the expression of aspartic acid at 158 residue of mouse N158D PrPC impedes the epitope recognition of 5C6 antibody [24], and therefore, it allows discrimination between wt and N158D PrPC. Our results indicate that only mouse wt PrPC was converted in Tga20xN158D mice (Supplementary Fig. 3). Accordingly with this suggestion, most of the pathological features previously described in Tga20 mice inoculated with the strains used in the present study [28] were reproduced in Tga20xN158D mice. All of the prion strains tested produced marked spongiosis and PrPSc deposition in both the thalamus and brainstem of Tga20xN158D mice (Figs. 2 and 3), regions previously proposed as clinical target areas of these strains in Tga20 mice [28]. In mice coexpressing N158D PrP, these different prion strains retained their specific pathological characteristics, as evidenced by the marked PrPSc deposition in the hippocampus of ME7-inoculated mice (Fig. 3b) [28] and the characteristic affectation of the cerebellum in those inoculated with the 22L strain [58]. Expression of the dominant-negative protein therefore appears not to have affected the characteristic pathological hallmarks of these strains, indicating that the increase in survival times observed in Tga20xN158D mice is not due to strain modifications caused by the amino acid substitution of the dominant-negative protein.
Based on our findings, we conclude that N158D PrP acts as a dominant-negative protein to partially block the conversion of PrPC to PrPSc and is thus a promising candidate for gene therapy strategies for the treatment of TSEs.
References
Prusiner SB (1998) The prion diseases. Brain Pathol 8(3):499–513
Prusiner SB (1998) Prions. Proc Natl Acad Sci U S A 95(23):13363–13383. https://doi.org/10.1073/pnas.95.23.13363
Prusiner SB (1982) Novel proteinaceous infectious particles cause scrapie. Science 216(4542):136–144. https://doi.org/10.1126/science.6801762
Smirnovas V, Baron GS, Offerdahl DK, Raymond GJ, Caughey B, Surewicz WK (2011) Structural organization of brain-derived mammalian prions examined by hydrogen-deuterium exchange. Nat Struct Mol Biol 18(4):504–506. https://doi.org/10.1038/nsmb.2035
Vazquez-Fernandez E, Vos MR, Afanasyev P, Cebey L, Sevillano AM, Vidal E, Rosa I, Renault L et al (2016) The structural architecture of an infectious mammalian prion using electron cryomicroscopy. PLoS Pathog 12(9):e1005835. https://doi.org/10.1371/journal.ppat.1005835
Fraser H (1976) The pathology of a natural and experimental scrapie. Front Biol 44:267–305
Budka H, Aguzzi A, Brown P, Brucher JM, Bugiani O, Gullotta F, Haltia M, Hauw JJ et al (1995) Neuropathological diagnostic criteria for Creutzfeldt-Jakob disease (CJD) and other human spongiform encephalopathies (prion diseases). Brain Pathol 5(4):459–466. https://doi.org/10.1111/j.1750-3639.1995.tb00625.x
Vidal E, Acin C, Foradada L, Monzon M, Marquez M, Monleon E, Pumarola M, Badiola JJ et al (2009) Immunohistochemical characterisation of classical scrapie neuropathology in sheep. J Comp Pathol 141(2–3):135–146. https://doi.org/10.1016/j.jcpa.2009.04.002
Collins SJ, Lawson VA, Masters CL (2004) Transmissible spongiform encephalopathies. Lancet 363(9402):51–61. https://doi.org/10.1016/S0140-6736(03)15171-9
Will RG, Ironside JW, Zeidler M, Cousens SN, Estibeiro K, Alperovitch A, Poser S, Pocchiari M et al (1996) A new variant of Creutzfeldt-Jakob disease in the UK. Lancet 347(9006):921–925. https://doi.org/10.1016/S0140-6736(96)91412-9
Bruce ME, Will RG, Ironside JW, McConnell I, Drummond D, Suttie A, McCardle L, Chree A et al (1997) Transmissions to mice indicate that ‘new variant’ CJD is caused by the BSE agent. Nature 389(6650):498–501. https://doi.org/10.1038/39057
Kirkwood JK, Cunningham AA (1994) Epidemiological observations on spongiform encephalopathies in captive wild animals in the British Isles. Vet Rec 135(13):296–303. https://doi.org/10.1136/vr.135.13.296
Kirkwood JK, Cunningham AA, Wells GA, Wilesmith JW, Barnett JE (1993) Spongiform encephalopathy in a herd of greater kudu (Tragelaphus strepsiceros): epidemiological observations. Vet Rec 133(15):360–364. https://doi.org/10.1136/vr.133.15.360
Sigurdson CJ, Miller MW (2003) Other animal prion diseases. Br Med Bull 66(1):199–212. https://doi.org/10.1093/bmb/66.1.199
Fernandez-Borges N, Chianini F, Eraña H, Vidal E, Eaton SL, Pintado B, Finlayson J, Dagleish MP et al (2012) Naturally prion resistant mammals: a utopia? Prion 6(5):425–429. https://doi.org/10.4161/pri.22057
Fernandez-Borges N, de Castro J, Castilla J (2009) In vitro studies of the transmission barrier. Prion 3(4):220–223. https://doi.org/10.4161/pri.3.4.10500
Chianini F, Fernandez-Borges N, Vidal E, Gibbard L, Pintado B, de Castro J, Priola SA, Hamilton S et al (2012) Rabbits are not resistant to prion infection. Proc Natl Acad Sci U S A 109(13):5080–5085. https://doi.org/10.1073/pnas.1120076109
Bian J, Khaychuk V, Angers RC, Fernandez-Borges N, Vidal E, Meyerett-Reid C, Kim S, Calvi CL et al (2017) Prion replication without host adaptation during interspecies transmissions. Proc Natl Acad Sci U S A 114(5):1141–1146. https://doi.org/10.1073/pnas.1611891114
Khan MQ, Sweeting B, Mulligan VK, Arslan PE, Cashman NR, Pai EF, Chakrabartty A (2010) Prion disease susceptibility is affected by beta-structure folding propensity and local side-chain interactions in PrP. Proc Natl Acad Sci U S A 107(46):19808–19813. https://doi.org/10.1073/pnas.1005267107
Vidal E, Fernandez-Borges N, Pintado B, Ordoñez M, Marquez M, Fondevila D, Torres JM, Pumarola M et al (2013) Bovine spongiform encephalopathy induces misfolding of alleged prion-resistant species cellular prion protein without altering its pathobiological features. J Neurosci 33(18):7778–7786. https://doi.org/10.1523/JNEUROSCI.0244-13.2013
Fernández-Borges N, Parra B, Vidal E, Eraña H, Sánchez-Martín MA, de Castro J, Elezgarai SR, Pumarola M, Mayoral T, Castilla J (2017) Unraveling the key to the resistance of canids to prion diseases. PLoS Pathog 13 (11):e1006716. https://doi.org/10.1371/journal.ppat.1006716
Fischer M, Rulicke T, Raeber A, Sailer A, Moser M, Oesch B, Brandner S, Aguzzi A et al (1996) Prion protein (PrP) with amino-proximal deletions restoring susceptibility of PrP knockout mice to scrapie. EMBO J 15(6):1255–1264
Manson JC, Clarke AR, Hooper ML, Aitchison L, McConnell I, Hope J (1994) 129/Ola mice carrying a null mutation in PrP that abolishes mRNA production are developmentally normal. Mol Neurobiol 8(2–3):121–127. https://doi.org/10.1007/BF02780662
Kang HE, Weng CC, Saijo E, Saylor V, Bian J, Kim S, Ramos L, Angers R et al (2012) Characterization of conformation-dependent prion protein epitopes. J Biol Chem 287(44):37219–37232. https://doi.org/10.1074/jbc.M112.395921
Fraser H, Dickinson AG (1968) The sequential development of the brain lesion of scrapie in three strains of mice. J Comp Pathol 78(3):301–311. https://doi.org/10.1016/0021-9975(68)90006-6
Schulz-Schaeffer WJ, Tschoke S, Kranefuss N, Drose W, Hause-Reitner D, Giese A, Groschup MH, Kretzschmar HA (2000) The paraffin-embedded tissue blot detects PrP(Sc) early in the incubation time in prion diseases. Am J Pathol 156(1):51–56. https://doi.org/10.1016/S0002-9440(10)64705-0
Sarasa R, Martinez A, Monleon E, Bolea R, Vargas A, Badiola JJ, Monzon M (2012) Involvement of astrocytes in transmissible spongiform encephalopathies: a confocal microscopy study. Cell Tissue Res 350(1):127–134. https://doi.org/10.1007/s00441-012-1461-1
Karapetyan YE, Saa P, Mahal SP, Sferrazza GF, Sherman A, Sales N, Weissmann C, Lasmezas CI (2009) Prion strain discrimination based on rapid in vivo amplification and analysis by the cell panel assay. PLoS One 4(5):e5730. https://doi.org/10.1371/journal.pone.0005730
Tuzi NL, Cancellotti E, Baybutt H, Blackford L, Bradford B, Plinston C, Coghill A, Hart P et al (2008) Host PrP glycosylation: a major factor determining the outcome of prion infection. PLoS Biol 6(4):e100. https://doi.org/10.1371/journal.pbio.0060100
Belt PB, Muileman IH, Schreuder BE, Bos-de Ruijter J, Gielkens AL, Smits MA (1995) Identification of five allelic variants of the sheep PrP gene and their association with natural scrapie. J Gen Virol 76(Pt 3):509–517. https://doi.org/10.1099/0022-1317-76-3-509
Westaway D, Zuliani V, Cooper CM, Da Costa M, Neuman S, Jenny AL, Detwiler L, Prusiner SB (1994) Homozygosity for prion protein alleles encoding glutamine-171 renders sheep susceptible to natural scrapie. Genes Dev 8(8):959–969. https://doi.org/10.1101/gad.8.8.959
Clouscard C, Beaudry P, Elsen JM, Milan D, Dussaucy M, Bounneau C, Schelcher F, Chatelain J et al (1995) Different allelic effects of the codons 136 and 171 of the prion protein gene in sheep with natural scrapie. J Gen Virol 76(Pt 8):2097–2101. https://doi.org/10.1099/0022-1317-76-8-2097
Hunter N, Cairns D, Foster JD, Smith G, Goldmann W, Donnelly K (1997) Is scrapie solely a genetic disease? Nature 386(6621):137. https://doi.org/10.1038/386137a0
Shibuya S, Higuchi J, Shin RW, Tateishi J, Kitamoto T (1998) Codon 219 Lys allele of PRNP is not found in sporadic Creutzfeldt-Jakob disease. Ann Neurol 43(6):826–828. https://doi.org/10.1002/ana.410430618
Kaneko K, Zulianello L, Scott M, Cooper CM, Wallace AC, James TL, Cohen FE, Prusiner SB (1997) Evidence for protein X binding to a discontinuous epitope on the cellular prion protein during scrapie prion propagation. Proc Natl Acad Sci U S A 94(19):10069–10074. https://doi.org/10.1073/pnas.94.19.10069
Zulianello L, Kaneko K, Scott M, Erpel S, Han D, Cohen FE, Prusiner SB (2000) Dominant-negative inhibition of prion formation diminished by deletion mutagenesis of the prion protein. J Virol 74(9):4351–4360. https://doi.org/10.1128/JVI.74.9.4351-4360.2000
Perrier V, Kaneko K, Safar J, Vergara J, Tremblay P, DeArmond SJ, Cohen FE, Prusiner SB et al (2002) Dominant-negative inhibition of prion replication in transgenic mice. Proc Natl Acad Sci U S A 99(20):13079–13084. https://doi.org/10.1073/pnas.182425299
Lee CI, Yang Q, Perrier V, Baskakov IV (2007) The dominant-negative effect of the Q218K variant of the prion protein does not require protein X. Protein Sci 16(10):2166–2173. https://doi.org/10.1110/ps.072954607
Stewart P, Campbell L, Skogtvedt S, Griffin KA, Arnemo JM, Tryland M, Girling S, Miller MW et al (2012) Genetic predictions of prion disease susceptibility in carnivore species based on variability of the prion gene coding region. PLoS One 7(12):e50623. https://doi.org/10.1371/journal.pone.0050623
Prusiner SB, Scott M, Foster D, Pan KM, Groth D, Mirenda C, Torchia M, Yang SL et al (1990) Transgenetic studies implicate interactions between homologous PrP isoforms in scrapie prion replication. Cell 63(4):673–686. https://doi.org/10.1016/0092-8674(90)90134-Z
Bueler H, Raeber A, Sailer A, Fischer M, Aguzzi A, Weissmann C (1994) High prion and PrPSc levels but delayed onset of disease in scrapie-inoculated mice heterozygous for a disrupted PrP gene. Mol Med 1(1):19–30
Priola SA, Caughey B, Race RE, Chesebro B (1994) Heterologous PrP molecules interfere with accumulation of protease-resistant PrP in scrapie-infected murine neuroblastoma cells. J Virol 68(8):4873–4878
Bolton DC, Bendheim PE (1988) A modified host protein model of scrapie. CIBA Found Symp 135:164–181
Hope J, Morton LJ, Farquhar CF, Multhaup G, Beyreuther K, Kimberlin RH (1986) The major polypeptide of scrapie-associated fibrils (SAF) has the same size, charge distribution and N-terminal protein sequence as predicted for the normal brain protein (PrP). EMBO J 5(10):2591–2597
Jahandideh S, Jamalan M, Faridounnia M (2015) Molecular dynamics study of the dominant-negative E219K polymorphism in human prion protein. J Biomol Struct Dyn 33(6):1315–1325. https://doi.org/10.1080/07391102.2014.945486
Geoghegan JC, Miller MB, Kwak AH, Harris BT, Supattapone S (2009) Trans-dominant inhibition of prion propagation in vitro is not mediated by an accessory cofactor. PLoS Pathog 5(7):e1000535. https://doi.org/10.1371/journal.ppat.1000535
Yuan J, Zhan YA, Abskharon R, Xiao X, Martinez MC, Zhou X, Kneale G, Mikol J et al (2013) Recombinant human prion protein inhibits prion propagation in vitro. Sci Rep 3(1):2911. https://doi.org/10.1038/srep02911
Fraser H, Dickinson AG (1973) Scrapie in mice. Agent-strain differences in the distribution and intensity of grey matter vacuolation. J Comp Pathol 83(1):29–40. https://doi.org/10.1016/0021-9975(73)90024-8
Bruce ME, McConnell I, Fraser H, Dickinson AG (1991) The disease characteristics of different strains of scrapie in Sinc congenic mouse lines: implications for the nature of the agent and host control of pathogenesis. J Gen Virol 72(Pt 3):595–603. https://doi.org/10.1099/0022-1317-72-3-595
Bruce ME (1993) Scrapie strain variation and mutation. Br Med Bull 49(4):822–838. https://doi.org/10.1093/oxfordjournals.bmb.a072649
Baron T, Crozet C, Biacabe AG, Philippe S, Verchere J, Bencsik A, Madec JY, Calavas D et al (2004) Molecular analysis of the protease-resistant prion protein in scrapie and bovine spongiform encephalopathy transmitted to ovine transgenic and wild-type mice. J Virol 78(12):6243–6251. https://doi.org/10.1128/JVI.78.12.6243-6251.2004
Kascsak RJ, Rubenstein R, Merz PA, Carp RI, Wisniewski HM, Diringer H (1985) Biochemical differences among scrapie-associated fibrils support the biological diversity of scrapie agents. J Gen Virol 66(Pt 8):1715–1722. https://doi.org/10.1099/0022-1317-66-8-1715
Sim VL, Caughey B (2009) Ultrastructures and strain comparison of under-glycosylated scrapie prion fibrils. Neurobiol Aging 30(12):2031–2042. https://doi.org/10.1016/j.neurobiolaging.2008.02.016
Atarashi R, Sim VL, Nishida N, Caughey B, Katamine S (2006) Prion strain-dependent differences in conversion of mutant prion proteins in cell culture. J Virol 80(16):7854–7862. https://doi.org/10.1128/JVI.00424-06
Telling GC, Scott M, Mastrianni J, Gabizon R, Torchia M, Cohen FE, DeArmond SJ, Prusiner SB (1995) Prion propagation in mice expressing human and chimeric PrP transgenes implicates the interaction of cellular PrP with another protein. Cell 83(1):79–90. https://doi.org/10.1016/0092-8674(95)90236-8
Buschmann A, Biacabe AG, Ziegler U, Bencsik A, Madec JY, Erhardt G, Luhken G, Baron T et al (2004) Atypical scrapie cases in Germany and France are identified by discrepant reaction patterns in BSE rapid tests. J Virol Methods 117(1):27–36. https://doi.org/10.1016/j.jviromet.2003.11.017
Houston F, Goldmann W, Chong A, Jeffrey M, Gonzalez L, Foster J, Parnham D, Hunter N (2003) Prion diseases: BSE in sheep bred for resistance to infection. Nature 423(6939):498. https://doi.org/10.1038/423498a
Fraser H (1979) Neuropathology of scrapie: the precision of the lesions and their diversity. In: Prusiner SB, Hadlow WJ (eds) Slow transmissible diseases of the nervous system, vol 1. Academic Press, New York, pp. 387–406
Acknowledgments
We thank MINECO for the Severo Ochoa Excellence Accreditation (SEV-2016-0644). The authors would like to thank the following for their support: the IKERBasque Foundation, the staff at the CIC bioGUNE animal facility, Dr. Belén Pintado for the Tga20 mouse embryo rederivation, Dr. Glenn Telling for kindly providing the 5C6 antibody, and Patricia Piñeiro, Silvia Ruiz, and Sonia Gómez for their technical assistance. The authors would also like to acknowledge Sara Gutiérrez for the image editing. Alicia Otero was supported by a research grant from the Government of Aragón (C020/2014) cofinanced by the European Social Fund.
Funding
This work was supported financially by grants from the Spanish (AGL2015-65046-C2-1-R, AGL2015-65560-R, and PCIN-2013-065) and Basque (2014111157) governments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This study was approved by the Ethics Committee for Animal Experiments of the University of Zaragoza (permit number PI32/13) and was performed in accordance with the recommendations for the care and use of experimental animals and with Spanish national law (R.D. 1201/05).
Conflict of Interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(PDF 985 kb)
Rights and permissions
About this article
Cite this article
Otero, A., Bolea, R., Hedman, C. et al. An Amino Acid Substitution Found in Animals with Low Susceptibility to Prion Diseases Confers a Protective Dominant-Negative Effect in Prion-Infected Transgenic Mice. Mol Neurobiol 55, 6182–6192 (2018). https://doi.org/10.1007/s12035-017-0832-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-017-0832-8