Abstract
The loss of neurons due to injury and disease results in a wide spectrum of highly disabling neurological and neurodegenerative conditions, given the apparent limited capacity of endogenous repair of the adult central nervous system (CNS). Therefore, it is important to develop technologies that can promote de novo neural stem cell and neuron generation. Current insights in CNS development and cellular reprogramming have provided the knowledge to finely modulate lineage-restricted transcription factors and microRNAs (miRNA) to elicit correct neurogenesis. Here, we discuss the current knowledge on the direct reprogramming of somatic non-neuronal cells into neural stem cells or subtype specific neurons in vitro and in vivo focusing on miRNA driven reprogramming. miRNA can allow rapid and efficient direct phenotype conversion by modulating gene networks active during development, which promote global shifts in the epigenetic landscape pivoting cell fate decisions. Furthermore, we critically present state-of-the-art and recent advances on miRNA therapeutics that can be applied to the diseased CNS. Together, the advances in our understanding of miRNA role in CNS development and disease, recent progress in miRNA-based therapeutic strategies, and innovative drug delivery methods create novel perspectives for meaningful therapies for neurodegenerative disorders.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
The loss of neuronal cell populations is a key feature that underlies different neurological and neurodegenerative diseases, which severely affect the life of many patients [1, 2]. The vast majority of these conditions still lack effective therapies. Since the disability is due to the critical loss of neurons, a rational approach aims to therapeutically induce neurogenesis compensating for the amount of dead cells [3,4,5]. The adult CNS is apparently incapable of major repair capacity given its inability to effectively replace neuronal circuitries and damaged tissues. The reasons of this defect are largely undetermined and it occurs despite the presence in the CNS of specific areas in which are located progenitor cells, which hold a certain degree of regenerative ability [6, 7]. However, the demonstration of self-renewing stem/progenitor cell populations in the adult CNS has raised the hypothesis to artificially manipulate their potential for an effective endogenous CNS regeneration after injuries [3,4,5].
On the other hand, the regenerative efficacy of transplanted neuronal stem and progenitor cells has been increasingly analyzed, also in clinical trials, but this approach is still in its infancy and likely requires invasive cell administration to the CNS [8]. As alternative, in vivo direct reprogramming of somatic CNS cells into neural stem cells (NSCs) or directly into specific neuronal subtype has been suggested as a possible approach for tissue repairing, overcoming the limits related to invasive cell transplantation. Many of the experimental efforts focus on converting glial cells into stem cell, progenitor or fully differentiated neurons. Glial cells are the most abundant cells in the adult brain and thus could represent a suitable target [1, 9].
In 2006, Takahashi and Yamanaka modified the paradigm of immutable terminal cell lineage commitment, demonstrating the capacity of a combination of defined factors central for pluripotency preservation (i.e., Oct4, Sox2, Klf4, and c-Myc), to reprogram terminally differentiated skin fibroblasts into embryonic stem cells [10]. The so-called induced pluripotent stem cells (iPSCs) opened a new field in the regenerative medicine [10, 11]. These data demonstrated not only that the cell fate is not absolutely determined but also that a combination of transcription factors is sufficient to reprogram cells.
In analogy to the iPSC generation, direct conversion of fibroblasts or astrocytes into induced neurons (iNs) has been obtained by the forced expression of either single or combination of different transcription factors [12]. This strategy is based on the seminal evidence that the combination of only three key neuronal transcription factors, ASCL1/MASH1, BRN2, and MYT1L (the so-called ABM combination), directly reprograms rodent fibroblasts into neuronal cells [13]. Nevertheless, these transcription factors seem to be poorly efficient in the conversion of adult human fibroblasts [14]. Thus, new transcription factors and eventually microRNAs (miRNAs) have been tested in reprogramming experimental protocols.
As abovementioned, non-coding RNAs (ncRNAs), including miRNAs, are known to be critical in regulating epigenetic and gene expression in cells, mediating post-transcriptional processes, and modulating many critical pathways in reprogramming and in maintenance of the reprogrammed cell phenotype [14, 15]. It is worthy to consider that some miRNAs have been demonstrated to be capable to induce cell reprogramming alone without the addition of other factors as elsewhere revised [16].
Here, we will review the data about the miRNA potential role in promoting reprogramming of somatic non-neuronal cells into a neuronal fate for therapeutic purposes.
miRNAs
miRNAs are a class of short, endogenous, non-coding RNAs that represent crucial regulators of the gene expression at the post-transcriptional level [17]. Targeting the majority of protein-coding transcripts, miRNAs play a key role in a wide range of developmental and pathological cellular events. miRNA genes are transcribed by the RNA polymerase II with the creation of a primary (pri)-miRNA [18]. Pri-miRNAs are then transformed into precursor (pre)-miRNAs by the RNA-processing complex that includes the RNase enzyme Drosha and exported from the nucleus by Exportin 5 in a Ran-GTP-dependent way [19]. In the cytosol, pre-miRNAs are subjected to a second modification phase, carried out by the Dicer RNase, which originates the mature miRNA duplex. After maturation, one strand of the miRNA duplex is included into the complex RNA-induced silencing complex (RISC) that uses the miRNA as guide to recognize and negatively regulate the expression of target transcripts. Interestingly, it has been demonstrated that miRNAs are able to cross gap junctional channels and modulate expression in neighboring cells playing a crucial role in regulating intercellular communication and synchronization [20].
miRNA in Cell Reprogramming
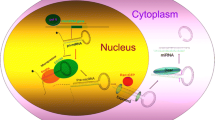
miRNAs are able to negatively modulate the gene expression of various key mRNAs during the reprogramming steps and to ensure the maintenance of the acquired cell phenotype (Fig. 1) [21]. The use of miRNAs in reprogramming methodologies is particularly efficient since even a sole miRNA may exert an effect on numerous pathways concurrently, an event acknowledged as multiplicity of miRNA targets [22]. There is increasing evidence about the potential of different miRNAs (i.e., miR-302-367, miR-290-295, and miR-17-92 clusters and Let7) in playing a significant role in cell reprogramming to a pluripotent state [21, 22]. For instance, miRNA-302 is able to directly inhibit NR2F2 gene expression, a protein that belongs to a nuclear receptor subfamily able to inhibit Oct-4 [23]. This event increases Oct-4 expression, thus promoting reprogramming towards pluripotency.
Schematic representation of miRNA-based cell reprogramming and potential applications. miRNAs can determine an efficient direct phenotype conversion of non-neural cells into neuronal populations by modulating gene networks active during development. Somatic cells like fibroblasts or astrocytes can be completely reprogrammed to a pluripotent cells and then differentiated into neurons or directly converted into neuronal fate. For instance, miRNA-302 is able to directly inhibit NR2F2 gene expression, a protein that belongs to a nuclear receptor subfamily able to inhibit Oct-4. This event increases Oct-4 expression, thus promoting reprogramming towards pluripotency. Mir9 and 124 inhibit the activity of REST/SCP1, some of the major anti-neuronal differentiation factors, thus promoting a neuronal fate acquisition. Derived cells can be exploited to perform in vitro modeling studies and endogenous in situ reprogramming for regenerative purposes. Moreover, the delivery of specific identified miRNA could be crucial in modulating altered pathways in human diseases
Another important step during the change of cell lineage is represented by the modification of the DNA methylation pattern, in particular in the promoter and regulatory sequences of master transcription factor genes [24]. In this context, miRNAs have been shown to target several transcription factors and epigenetic regulators, thus modulating proteins important for DNA methylation and eventually changing gene expression towards the effective conversion of the cell fate. In the majority of cases, miRNAs do not act alone, but in synergy with different master transcription factors and additional controller elements that create the precise network necessary for the reprogramming. For example, Lin28, a relevant pluripotency regulator protein, promotes the cell reprogramming by inhibiting the Let7 miRNA [25]. Hence, beside miRNA and transcription factors, other molecules appear to take part in the reprogramming biological process. Therefore, further investigations are needed to recognize novel key controllers and targets of miRNAs, which will provide a deeper knowledge of the events responsible for miRNA-driven cell fate conversion.
miRNAs in Neuronal Reprogramming
The increased knowledge in the molecular mechanisms determining CNS development in combination with the recent progresses in the stem cell field has contributed to investigate miRNA involvement in neurogenesis. Complementarily, this knowledge could be applied to direct cell reprogramming [25, 26]. Here, we present and discuss the experiments that used miRNAs to promote the reprogramming of somatic non-neuronal cells towards a neuronal fate (Table 1).
miRNA-200 family, which plays a key role in cancer initiation and metastasis [33], has been described as crucial for the development and survival of ventral neurons in the mouse midbrain, including dopaminergic neurons, and hindbrain. These miRNAs are able to target specific SOX2, a well-known pluripotency factor, and E2F3, which regulates the cell cycle [34]. Finally, they can prompt the cell to exit the cell cycle and differentiate towards a neuronal fate.
miR-124 and miR-9 are two of the most important miRNA in neurogenesis, widely applied in reprogramming strategies. They are the most represented miRNAs in the mammalian CNS [35, 36] and play important roles in controlling neuronal fate and function. miR-124 (also called miR-124a) is markedly upregulated during neuronal differentiation and in mature neurons [37, 38], while it is not expressed in non-neuronal cells, such as glia [39]. miR-9 is encoded by three genes (miR-9-1, miR-9-2, and miR-9-3) sited in different chromosomes in mammals. After processing, each of the miR-9 genes originates two mature miRNAs, miR-9 (that is the 5′ strand, miR-9-5p) and miR-9* (that is the 3′ strand, miR-9-3p) [37]. miR-9 and miR-124 recognize distinct molecular targets even if partially overlapping and act synergistically. They are able to promote the development and acquisition of a neuronal fate in progenitor cells [40,41,42]. A single lentiviral vector encoding both precursors of miR-9/9* and miR-124 could prompt the conversion of human fibroblasts into iNs in a short period of time; this process was significantly enhanced by the addition of the neurogenic transcription factor NeuroD2 in a synergistic way [15]. However, the spectrum of the acquired neuronal phenotype was quite limited and Yoo’s group tested in a second work the hypothesis that other lineage-specific factors could be necessary to finely tune the reprogramming into specific neuron subtypes [27]. Thus, they co-expressed miR-9/9* and miR-124 with transcription factors present during the striatum development (BCL11B, DLX1, DLX2, and MYT1L). This combination was able to drive the conversion of human postnatal and adult fibroblasts into iNs that resembled striatal medium spiny neurons (iMSNs), a neuronal subpopulation that has a crucial role in motor control and harbors selective susceptibility to cell death in Huntington’s disease.
In line with these experiments, Sheng Ding’s group demonstrated that the transfer of miR-124 with a couple of transcription factors (Myt1l plus Brn2, instead of Ascl1) was sufficient to reprogram both postnatal and adult human fibroblasts into functional iNs, with a conversion rate similar to that described by Yoo’s group [43]. Remarkably, they demonstrated for the first time that, in the presence of miR-124, ASCL1/MASH1 was dispensable for direct neuronal conversion. The obtained iNs presented a typical neuronal shape and appropriate neuronal markers positivity. Furthermore, these cells were functionally active as demonstrated by the production of action potential and synapses [37]. The effective functional maturation was attributed to miR-124 action. This protocol generated in a prevalent way GABA and glutamatergic neurons, while other phenotypes, like dopaminergic and serotoninergic neurons, were scarce or absent, supporting the abovementioned hypothesis that other master transcription factors may be need to modulate the reprogramming to specific neuronal cell phenotypes. Thus, miRNAs synergize with transcription factors and have a pivotal role in neuronal maturation in terms of morphology and function.
Regarding miRNA mechanisms of action during reprogramming, one possible hypothesis is that miRNAs inhibit a subset of genes that need to be repressed in the neuronal phenotype, while the transcription factors activate key neuronal genes, leading to an additive, synergistic effect. miR-9/9* and miR-124 play a key role in the epigenetic modulation, leading to diffuse chromatin remodeling also by acting on chromatin complexes like BAF. They modulate the activity of some of the major anti-neuronal differentiation factors, including RE1-silencing transcription factor (REST)/SCP1 [37], and promote the expression of neural genes by inhibition of REST, which represses a large array of neuronal-specific genes in non-neuronal cells. Moreover, miR-124 and miR-9 target many genes, estimated as more than 1000, the majority of which are downregulated during neuronal differentiation [44]. Among them, PTBP1 is a known repressor of splicing of neuron-specific alternative exons highly expressed in non-neural cells [45]. Remarkably, it has been described that the repression of PTBP1 protein, which is normally mediated by miR-124, is sufficient to induce the conversion of fibroblasts into functional neurons [46].
Parmar’s group described that the exogenous transcription factors responsible for neuronal reprogramming of fibroblasts in iN could be switched off once the cell had reached a stable neuronal fate, under the control of miR-124 [47]. In this study, iNs were obtained through the use of lentiviral vectors encoding for the conversion factors Ascl1, Brn2, and Myt1L (ABM) followed and regulated by four copies of perfect matching miR-124-target sequences (miR-124. T regulation). Thus, when endogenous miR-124 is not expressed (i.e., in fibroblasts), the ABM complex is not repressed leading to high levels of reprogramming factors. Instead, when a stable neuronal phenotype has been reached, the hiNs activate their endogenous miR-124 that binding to the miR-target sequence in the vector-derived mRNA powerfully represses the expression of the ABM factors. This finely modulated strategy can be associated with integrase-deficient vectors, offering a non-integrating and self-regulated reprogramming method that overcomes issues related to the risk of integration of viral transgenes into the human genome. This strategy is potentially applicable in clinic and thus constitutes a major achievement towards the use of iNs for therapeutics.
Moreover, miRNA-9/124 can synergize with other miRNAs relevant for the achievement and maintenance of pluripotent state as it has been shown for the miRNA-302/367 cluster. Zhou et al. recently reported that the expression of miRNA-302/367 cluster (that is known to be crucial in the maintenance and self-renewal of embryonic stem cells) in combination with miRNA-9/9* and miRNA-124 could effectively transform fibroblasts into neuronal cells [29]. In this study, the efficient reprogramming could be achieved only with the combination of all miRNAs, but without the need of other transcription factors.
In another study focused on in vivo reprogramming, viral vectors encoding for the same cluster (miR-302/367) and the gene reporter GFP were injected into the striatal regions of the mice brain parenchyma to promote transdifferentiation of endogenous non-neuronal cells, mainly astrocytes, into neuroblasts [30]. The combination of miR-302/367 and valproic acid (VPA), a demethylating agent, resulted in the presence of GFP-positive cells into the brain of the treated mice, expressing doublecortin, a neuroblast marker normally not expressed in the adult CNS. Since the vast majority of GFP cells in the first days after injection were astrocytes, this finding supports the conversion of astrocytes into neuroblasts. The addition of VPA increased the efficiency of the reprogramming by modifying the epigenetic regulation of the transcription factor Oct4. After the achievement of the neuronal phenotype, the expression of the pluripotency marker was not detected, suggesting that the cells have been directly converted. It is worth noticing that the cells surrounding the injection site wound appeared to be more prone to the change of fate. A previous work suggested that the reprogramming of NG2+ glial cells into neurons within the rodent cortex was unachievable in the absence of a stab wound [48]. This suggests that reactive cells surrounding the injury are probably exposed to a higher concentration of neurogenic factors, which facilitate the reprogramming. In a clinical perspective, this selectivity could be crucial in order to obtain a conversion of reactive astrocytes surrounding an injury (i.e., an infarcted area) into relevant neuronal cells. In neurodegenerative disorders, there is no acute damage, but several models of different diseases have shown a variable quote of reactive gliosis and neuroinflammation, which could in turn favor optimal conditions for reprogramming and regeneration.
A different group published interesting results on in vivo reprogramming of adult oligodendrocytes in mature neurons [32]. The authors exploited a miRNA-GFP construct able to reduce the expression of polypyrimidine tract-binding protein and delivered it to the rat striatum with an oligodendrocyte targeting adeno-associated virus vector (AAV). Six weeks after the treatment, the majority of GFP cells displayed a proper neuronal morphology and electrophysiological features of mature neurons. It is worth noticing that in a future clinical setting, some problems could arise from the loss of myelinated areas that followed the transition to a neuronal phenotype. This problem could be overcome by the development of viral vectors targeting oligodendrocyte progenitors instead of mature myelinating oligodendrocytes. Overall, the generation of effective vectors targeting specific cell populations for the reprogramming appears to be a promising path towards the development of effective replacement therapies for neuronal loss due to injury or degeneration.
Recent studies investigated the reprogramming of retina glial cells, the so-called Muller glia. Interestingly, Muller glia are positive not only for astrocytic antigens but also for some neural progenitor markers, an aspect that suggests their potential role as neuronal precursors. In non-mammalian animals, retina damage is a signal for Muller glia to re-enter the cell cycle, de-differentiate and originate novel neuronal cells. In mammals, these events are not present or barely detectable, supporting the idea that mammalian Muller glia is not or scarcely able to substitute neuronal loss in the retina. Thus, artificially harnessing the potential repair of Muller glia could hold a therapeutic potential.
Wohl and Reh [31] investigated the reprogramming of murine Müller glia cultures with the combination of miR-124 and miRNA-9/9* in association with the transcription factors Ascl1 and Mash1. Reprogrammed Muller glial cells presented a decrease in Ctdsp1 and PTBP1 proteins, demonstrating a key role of the silencing of REST pathway in the re-activation of neuronal genes. These findings imply that both miR-124-9-9* and the REST complex are key elements in the reprogramming of Müller glia to neuronal progenitors towards retina regeneration. The generation of neuronal progenitor from Muller glia relies on global demethylation, followed by a new methylation pattern, the activation of Lin28/Let-7 miRNA loop, and Sonic Hedgehog/Wnt-β-catenin signalling. Yao et al. [49] described that the modulation of Wnt/Lin28/Let-7 miRNAs promotes Muller glia proliferation also in the absence of the signal generated by a retinal injury. Genetic transfer of β-catenin promotes proliferation of Muller glia by binding to the Lin28 promoter and increasing its transcription that in turn suppresses Let7. Interestingly, a subset of cell cycle-reactivated Muller glia was positive for markers typical of amacrine cells, a subtype of retinal interneurons. These data support the role of Lin28/Let-7 miRNA in regulating proliferation and neurogenic potential. Whether this pathway can have a similar equivalent in CNS astrocytes has to be investigated.
miRNA-Directed Neuronal Reprogramming Therapeutic Perspectives
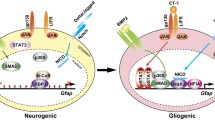
Patients’ derived cells can be directly reprogrammed into neurons with the aid of miRNAs and exploited for disease modeling and drug discovery. Directly reprogrammed cells into neural stem cells and neurons may also be used as a source for transplantation in cell-mediated therapy for neurodegenerative diseases including Parkinson’s disease and amyotrophic lateral sclerosis (ALS). Furthermore, the modulation of miRNA expression in vivo can lead to in situ cell reprogramming of non-neuronal cells, like astrocytes, into NSCs and neurons for tissue repair in neurological disorders (Fig. 2). In general, as miRNAs are capable to target numerous genes within the same pathway or even numerous pathways simultaneously, they might represent encouraging therapeutic tools for human diseases and regenerative medicine. miRNAs act as either neurogenesis enhancers or cell proliferation inhibitors [50]. These events can be modulated through two different strategies of miRNA-based therapeutics: (1) repress the expression of specific miRNAs (i.e., in the case of Let7) or (2) increase miRNA expression (i.e., for miR-9/9* and miR-124). It is possible to inhibit miRNA expression by administering synthetic miRNAs that function with the principles of RNA interference (RNAi), degrading the target miRNA or blocking its binding to the mRNA [51]. However, many aspects need to be considered before applying the silencing approach: how to precisely control the possible widespread effect of the miRNA reduction, the feasibility of RNAi, and a suitable way of in vivo delivery. Small miRNAs, synthetic miRNAs, and antisense oligonucleotides are the most widely employed tools for silencing miRNA [52]. miRNA expression may be modulated by the use of synthetic antisense single-stranded RNA or DNA oligonucleotides, called antagomirs or antimiRs, complementary to mature endogenous miRNAs that can bind and then silence their targets miRNA. Phosphorothioate oligonucleotides are the most common exploited first-generation antisense oligonucleotides. They can be synthesized with a range of different chemical features to increase their stability and efficacy, but these modifications do not confer cell or organ specificity. miRNA sponge and miRNA masking represent some of the latest biotechnological strategies available. The miRNA sponges harbor multiple binding sites for their target miRNAs, competitively antagonizing the binding to the mRNA, thus interfering with miRNA function [53]. Instead, miRNA masking holds a complimentary miRNA binding site in the 3′ UTR of the target mRNA to inhibit competitively and decreases the activity of endogenous miRNAs [54]. On the other hand, the overexpression of the target miRNAs can be obtained by the delivery of synthetic miRNA (miRNA mimics) that possess sequences identic to the miRNA of interest. MRX34, a mimic of miR-34 that acts as tumor suppressor inhibiting multiple oncogenic pathways and enhancing anti-tumor immune response, is now being tested in a phase I clinical trials in patients with advanced hepatocarcinoma [55].
Therapeutic perspectives of miRNA-mediated reprogramming. miRNAs can be exploited in vitro to reprogram cells and obtain a source for cell transplantation therapies. A complementary strategy involves miRNA delivery for in situ cell reprogramming of non-neuronal cells, like astrocytes, into NSCs and neurons for tissue repair in neurological disorders
miRNA-mediated reprogramming in vivo requires innovative administration strategies in the CNS that should be non-invasive and organ specific. The blood brain barrier poses a significant limit for the use of systemic intravenous delivery. Moreover, systemic injection lacks organ specificity, leading to possible side off-target effects. Intrathecal delivery warrants exploration in the miRNA field since it can allow organ selectivity, has minimal invasiveness, and can partially circumvent the presence of the blood brain barrier. The implementation of a broad spectrum of chemical variants to increase tissue and cell penetration of synthetic miRNA mimics/antagomirs is ongoing [56]. Furthermore, innovative drug delivery systems, comprising liposomes, polymeric micelles and vesicles, nanoparticles, and dendrimers are currently under development [57]. One strategy to target a specific cell type (i.e., only the astrocytes) is the so-called epitope targeting drug delivery. It consists into binding the synthetic miRNA mimic or antagomir with peptides that recognize and enter only in certain cells [58]. Another promising novel delivery tool is represented by the AAVs that can allow the effective delivery of vector encoding miRNA in the CNS [59]. A new developed AAV vector targeting oligodendrocytes has been used to deliver miRNAs into the rat striatum [32]. There are several available viral vectors and among them, the AAVs represent a promising tool for therapeutic applications due to their lack of pathogenicity, their ability to persist within the cell, and the many existing serotypes. They are able to enter and integrate in the nucleus of nondividing cells and they do not elicit a significant immunological response [60].
Taking this into consideration, gene transfer will become permanent and the potential side effect of miRNA overexpression has to be carefully considered. A possible strategy to reduce the risk could comprehend the selection of miRNAs that are already highly expressed and proven to be well tolerated in normal tissues.
Conclusion and Future Perspectives
miRNA can regulate the expression of a wide range of target genes by multiple mechanisms well beyond RNAi alone, both by directly interacting with the gene promoter and by epigenetic action through the modification of the DNA methylation [61].
The understanding of miRNA role in neurogenesis and reprogramming is rapidly evolving with the potential to significantly modify in the near future the methodologies of direct cell somatic conversion in vitro and in vivo. At the present, the most frequently employed miRNAs are that with a clear role in neurogenesis, like miRNA-124 and miRNA-9, but novel combination can be explored and applied at different time points of cell fate conversion.
miRNA-based strategies allow a rapid and efficient cell reprogramming due to miRNA broad impact on the cell gene expression pattern. miRNAs can limit the need of transcription factors, and thanks to the ongoing technological advances, this may lead to the realization of proficient non-viral, non-integrating direct reprogramming strategies in vitro and in vivo for therapeutic purpose. However, given the complexity of the reprogramming process and the potential broad effect of miRNAs, a complete knowledge of miRNA mechanisms and effects is needed for their effective and safe application in the clinical setting.
In conclusion, miRNA-mediated reprogramming may represent a promising tool to generate novel neuronal cells for the development of therapeutics for neurological diseases.
AAVs, adeno-associated viruses; ABM, Ascl1, Brn2, and Myt1L; CNS, central nervous system; iMSNs, induced medium spiny neurons; iNs, induced neurons; iPSCs, induced pluripotent stem cells; miRNA, microRNA; ncRNAs, non-coding RNAs; NSCs, neural stem cells; RISC, RNA-induced silencing complex; RNAi, RNA interference
References
Dametti S, Faravelli I, Ruggieri M et al (2015) Experimental advances towards neural regeneration from induced stem cells to direct in vivo reprogramming. Mol Neurobiol. doi:10.1007/s12035-015-9181-7
Sandoe J, Eggan K (2013) Opportunities and challenges of pluripotent stem cell neurodegenerative disease models. Nat Neurosci 16:780–789. doi:10.1038/nn.3425
Buffo A, Vosko MR, Ertürk D et al (2005) Expression pattern of the transcription factor Olig2 in response to brain injuries: implications for neuronal repair. Proc Natl Acad Sci U S A 102:18183–18188. doi:10.1073/pnas.0506535102
Jones KS, Connor B (2016) Adult neurogenesis and in vivo reprogramming: combining strategies for endogenous brain repair. Neural Regen Res 11:1748–1749. doi:10.4103/1673–5374.194712
Li H, Chen G (2016) In vivo reprogramming for CNS repair: regenerating neurons from endogenous glial cells. Neuron 91:728–738. doi:10.1016/j.neuron.2016.08.004
Merkle FT, Tramontin AD, García-Verdugo JM, Alvarez-Buylla A (2004) Radial glia give rise to adult neural stem cells in the subventricular zone. Proc Natl Acad Sci U S A 101:17528–17532. doi:10.1073/pnas.0407893101
Mu L, Berti L, Masserdotti G et al (2012) SoxC transcription factors are required for neuronal differentiation in adult hippocampal neurogenesis. J Neurosci 32:3067–3080. doi:10.1523/JNEUROSCI.4679-11.2012
Faravelli I, Riboldi G, Nizzardo M et al (2014) Stem cell transplantation for amyotrophic lateral sclerosis: therapeutic potential and perspectives on clinical translation. Cell Mol Life Sci CMLS 71:3257–3268. doi:10.1007/s00018-014-1613-4
Arlotta P, Berninger B (2014) Brains in metamorphosis: reprogramming cell identity within the central nervous system. Curr Opin Neurobiol 27:208–214. doi:10.1016/j.conb.2014.04.007
Takahashi K, Yamanaka S (2006) Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126:663–676. doi:10.1016/j.cell.2006.07.024
Peng J, Zeng X (2011) The role of induced pluripotent stem cells in regenerative medicine: neurodegenerative diseases. Stem Cell Res Ther 2:32. doi:10.1186/scrt73
Ruggieri M, Riboldi G, Brajkovic S et al (2014) Induced neural stem cells: methods of reprogramming and potential therapeutic applications. Prog Neurobiol 114:15–24. doi:10.1016/j.pneurobio.2013.11.001
Vierbuchen T, Ostermeier A, Pang ZP et al (2010) Direct conversion of fibroblasts to functional neurons by defined factors. Nature 463:1035–1041. doi:10.1038/nature08797
Pang ZP, Yang N, Vierbuchen T et al (2011) Induction of human neuronal cells by defined transcription factors. Nature 476:220–223. doi:10.1038/nature10202
Yoo AS, Sun AX, Li L et al (2011) MicroRNA-mediated conversion of human fibroblasts to neurons. Nature 476:228–231. doi:10.1038/nature10323
Srivastava D, DeWitt N (2016) In vivo cellular reprogramming: the next generation. Cell 166:1386–1396. doi:10.1016/j.cell.2016.08.055
Ambros V (2001) microRNAs: tiny regulators with great potential. Cell 107:823–826
Lee Y, Kim M, Han J et al (2004) MicroRNA genes are transcribed by RNA polymerase II. EMBO J 23:4051–4060. doi:10.1038/sj.emboj.7600385
Ha M, Kim VN (2014) Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol 15:509–524. doi:10.1038/nrm3838
Zong L, Zhu Y, Liang R, Zhao H-B (2016) Gap junction mediated miRNA intercellular transfer and gene regulation: a novel mechanism for intercellular genetic communication. Sci Rep 6:19884. doi:10.1038/srep19884
Kuo C-H, Ying S-Y (2012) Advances in microRNA-mediated reprogramming technology. Stem Cells Int 2012:823709. doi:10.1155/2012/823709
Yang H, Zhang L, An J et al (2016) MicroRNA-mediated reprogramming of somatic cells into neural stem cells or neurons. Mol Neurobiol. doi:10.1007/s12035-016-0115-9
Hu S, Wilson KD, Ghosh Z et al (2013) MicroRNA-302 increases reprogramming efficiency via repression of NR2F2. Stem Cells Dayt Ohio 31:259–268. doi:10.1002/stem.1278
Gruber AJ, Zavolan M (2013) Modulation of epigenetic regulators and cell fate decisions by miRNAs. Epigenomics 5:671–683. doi:10.2217/epi.13.65
Nam Y, Chen C, Gregory RI et al (2011) Molecular basis for interaction of let-7 microRNAs with Lin28. Cell 147:1080–1091. doi:10.1016/j.cell.2011.10.020
Stevanato L, Sinden JD (2014) The effects of microRNAs on human neural stem cell differentiation in two- and three-dimensional cultures. Stem Cell Res Ther 5:49. doi:10.1186/scrt437
Victor MB, Richner M, Hermanstyne TO et al (2014) Generation of human striatal neurons by microRNA-dependent direct conversion of fibroblasts. Neuron 84:311–323. doi:10.1016/j.neuron.2014.10.016
Richner M, Victor MB, Liu Y et al (2015) MicroRNA-based conversion of human fibroblasts into striatal medium spiny neurons. Nat Protoc 10:1543–1555. doi:10.1038/nprot.2015.102
Zhou C, Gu H, Fan R et al (2015) MicroRNA 302/367 cluster effectively facilitates direct reprogramming from human fibroblasts into functional neurons. Stem Cells Dev 24:2746–2755. doi:10.1089/scd.2015.0123
Ghasemi-Kasman M, Hajikaram M, Baharvand H, Javan M (2015) MicroRNA-mediated in vitro and in vivo direct conversion of astrocytes to neuroblasts. PLoS One 10:e0127878. doi:10.1371/journal.pone.0127878
Wohl SG, Reh TA (2016) miR-124-9-9* potentiates Ascl1-induced reprogramming of cultured Müller glia. Glia 64:743–762. doi:10.1002/glia.22958
Weinberg MS, Criswell HE, Powell SK et al (2017) Viral vector reprogramming of adult resident striatal oligodendrocytes into functional neurons. Mol Ther J Am Soc Gene Ther 25:928–934. doi:10.1016/j.ymthe.2017.01.016
Humphries B, Yang C (2015) The microRNA-200 family: small molecules with novel roles in cancer development, progression and therapy. Oncotarget 6:6472–6498. doi:10.18632/oncotarget.3052
Peng C, Li N, Ng Y-K et al (2012) A unilateral negative feedback loop between miR-200 microRNAs and Sox2/E2F3 controls neural progenitor cell-cycle exit and differentiation. J Neurosci 32:13292–13308. doi:10.1523/JNEUROSCI.2124-12.2012
Lagos-Quintana M, Rauhut R, Yalcin A et al (2002) Identification of tissue-specific microRNAs from mouse. Curr Biol 12:735–739
Landgraf P, Rusu M, Sheridan R et al (2007) A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 129:1401–1414. doi:10.1016/j.cell.2007.04.040
Conaco C, Otto S, Han J-J, Mandel G (2006) Reciprocal actions of REST and a microRNA promote neuronal identity. Proc Natl Acad Sci U S A 103:2422–2427. doi:10.1073/pnas.0511041103
Deo M, Yu J-Y, Chung K-H et al (2006) Detection of mammalian microRNA expression by in situ hybridization with RNA oligonucleotides. Dev Dyn Off Publ Am Assoc Anat 235:2538–2548. doi:10.1002/dvdy.20847
Cheng L-C, Pastrana E, Tavazoie M, Doetsch F (2009) miR-124 regulates adult neurogenesis in the subventricular zone stem cell niche. Nat Neurosci 12:399–408. doi:10.1038/nn.2294
Gu X, Fu C, Lin L et al (2017) miR-124 and miR-9 mediated downregulation of HDAC5 promotes neurite development through activating MEF2C-GPM6A pathway. J Cell Physiol. doi:10.1002/jcp.25927
Xue Q, Yu C, Wang Y et al (2016) miR-9 and miR-124 synergistically affect regulation of dendritic branching via the AKT/GSK3β pathway by targeting Rap2a. Sci Rep 6:26781. doi:10.1038/srep26781
Yoo AS, Staahl BT, Chen L, Crabtree GR (2009) MicroRNA-mediated switching of chromatin-remodelling complexes in neural development. Nature 460:642–646. doi:10.1038/nature08139
Ambasudhan R, Talantova M, Coleman R et al (2011) Direct reprogramming of adult human fibroblasts to functional neurons under defined conditions. Cell Stem Cell 9:113–118. doi:10.1016/j.stem.2011.07.002
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120:15–20. doi:10.1016/j.cell.2004.12.035
Makeyev EV, Zhang J, Carrasco MA, Maniatis T (2007) The MicroRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Mol Cell 27:435–448. doi:10.1016/j.molcel.2007.07.015
Xue Y, Ouyang K, Huang J et al (2013) Direct conversion of fibroblasts to neurons by reprogramming PTB-regulated microRNA circuits. Cell 152:82–96. doi:10.1016/j.cell.2012.11.045
Lau S, Rylander Ottosson D, Jakobsson J, Parmar M (2014) Direct neural conversion from human fibroblasts using self-regulating and nonintegrating viral vectors. Cell Rep 9:1673–1680. doi:10.1016/j.celrep.2014.11.017
Heinrich C, Bergami M, Gascón S et al (2014) Sox2-mediated conversion of NG2 glia into induced neurons in the injured adult cerebral cortex. Stem Cell Rep 3:1000–1014. doi:10.1016/j.stemcr.2014.10.007
Yao K, Qiu S, Tian L et al (2016) Wnt regulates proliferation and neurogenic potential of Müller glial cells via a Lin28/let-7 miRNA-dependent pathway in adult mammalian retinas. Cell Rep 17:165–178. doi:10.1016/j.celrep.2016.08.078
Adlakha YK, Seth P (2017) The expanding horizon of MicroRNAs in cellular reprogramming. Prog Neurobiol 148:21–39. doi:10.1016/j.pneurobio.2016.11.003
Takeshita F, Patrawala L, Osaki M et al (2010) Systemic delivery of synthetic microRNA-16 inhibits the growth of metastatic prostate tumors via downregulation of multiple cell-cycle genes. Mol Ther J Am Soc Gene Ther 18:181–187. doi:10.1038/mt.2009.207
Fanini F, Fabbri M (2016) MicroRNAs and cancer resistance: A new molecular plot. Clin Pharmacol Ther 99:485–493. doi:10.1002/cpt.353
Liu D, Li Y, Luo G et al (2017) LncRNA SPRY4-IT1 sponges miR-101-3p to promote proliferation and metastasis of bladder cancer cells through up-regulating EZH2. Cancer Lett 388:281–291. doi:10.1016/j.canlet.2016.12.005
Murakami K, Miyagishi M (2014) Tiny masking locked nucleic acids effectively bind to mRNA and inhibit binding of microRNAs in relation to thermodynamic stability. Biomed Rep 2:509–512. doi:10.3892/br.2014.260
Beg MS, Brenner AJ, Sachdev J et al (2017) Phase I study of MRX34, a liposomal miR-34a mimic, administered twice weekly in patients with advanced solid tumors. Investig New Drugs 35:180–188. doi:10.1007/s10637-016-0407-y
Krützfeldt J (2016) Strategies to use microRNAs as therapeutic targets. Best Pract Res Clin Endocrinol Metab 30:551–561. doi:10.1016/j.beem.2016.07.004
Liu J, Meng T, Yuan M et al (2016) MicroRNA-200c delivered by solid lipid nanoparticles enhances the effect of paclitaxel on breast cancer stem cell. Int J Nanomedicine 11:6713–6725. doi:10.2147/IJN.S111647
Dengl S, Sustmann C, Brinkmann U (2016) Engineered hapten-binding antibody derivatives for modulation of pharmacokinetic properties of small molecules and targeted payload delivery. Immunol Rev 270:165–177. doi:10.1111/imr.12386
Stoica L, Sena-Esteves M (2016) Adeno associated viral vector delivered RNAi for gene therapy of SOD1 amyotrophic lateral sclerosis. Front Mol Neurosci 9:56. doi:10.3389/fnmol.2016.00056
Daya S, Berns KI (2008) Gene therapy using adeno-associated virus vectors. Clin Microbiol Rev 21:583–593. doi:10.1128/CMR.00008-08
Zhou X, Yang P-C (2012) MicroRNA: a small molecule with a big biological impact. MicroRNA Shariqah United Arab Emir 1:1
Acknowledgements
The authors wish to thank the Associazione Centro Dino Ferrari for their support.
Author information
Authors and Affiliations
Contributions
IF and SC conceived the idea, revised all the literature, and contributed to all parts. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interests
The authors declare that they have no conflict of interest.
Funding
AriSLA provided financial support to SC through a research grant (AriSLAsmallRNALS). AriSLA had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript: and in the decision to publish the results.
Rights and permissions
About this article
Cite this article
Faravelli, I., Corti, S. MicroRNA-Directed Neuronal Reprogramming as a Therapeutic Strategy for Neurological Diseases. Mol Neurobiol 55, 4428–4436 (2018). https://doi.org/10.1007/s12035-017-0671-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-017-0671-7