Abstract
The apolipoprotein E ε4 (APOE ε4) allele is the most important genetic risk factor for Alzheimer’s disease (AD); however, the underlying mechanisms responsible for it remain controversial. We used the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database to examine the influence of APOE ε4 dose on clinical and neuroimaging biomarkers across the AD spectrum (from cognitive normal to AD patients with severe cognitive impairment). A total of 1718 participants from the ADNI cohort were selected, and we evaluated the impact of ε4 dose on cerebrospinal fluid (CSF) levels’ Abeta1-42 (Aβ1-42), tau, and phosphorylated-tau (p-tau); cortical amyloid deposition (Florbetapir-PET-AV45); brain atrophy (MRI); brain metabolism (FDG-PET); hippocampal metabolism; and cognitive declines, through different cognitive subgroups. We found that (1) ε4 was associated with decreased CSF beta-amyloid (Aβ1-42) and increased cerebral Aβ deposition across the AD spectrum; (2) increased CSF tau, P-tau and cerebral hypometabolism, hippocampal atrophy, and cognition decline were all associated with APOE ε4 in prodromal AD stage; (3) increased CSF tau, P-tau and cerebral hypometabolism appear to begin earlier than hippocampal atrophy and cognitive decline. We hypothesized that APOE ε4 increases cerebral amyloid-β (Aβ) deposition in all the stages of AD development, and also influences Aβ-initiated cascade of downstream neurodegenerative effects, thereby increasing the risk of AD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Apolipoprotein E (APOE) has been established unequivocally as the most important susceptibility gene for late-onset Alzheimer’s disease (LOAD). The ε4 polymorphism of APOE is the most common genetic risk factor for LOAD [1]. Thus far, APOE is the primary target of numerous studies investigating the disease’s underlying molecular neuropathology, pharmacological therapy, clinical progression, diagnosis, prevention, and treatment response.
Though the mechanism underlying APOE ε4 allele linked modulation of Alzheimer’s disease (AD) development is still not completely understood, emerging data suggest that APOE contributes to AD pathogenesis through a wide range of biological functions, including amyloid beta (Aβ)-dependent pathway and Aβ-independent pathway [2–4]. Some hypotheses suggest that APOE genotype is associated with AD biomarkers, with higher levels of Aβ deposition [5–7], higher degree and faster rate of neurodegeneration [8, 9], changes in the brain volume [10], function and glucose metabolism [11, 12], influence of the cerebrospinal fluid (CSF) measures of amyloid and tau [13, 14], and more severe impaired cognition [15–18], However, some other studies have yielded controversial results as to these biomarkers investigations associated with APOE [19, 20]. Furthermore, although there have been previous investigations of cognitive and neuroimaging differences between mild cognitive impairment (MCI) or AD patients who are APOE ε4 carriers vs. noncarriers, no prior work has brought these lines of research together towards normal cognition (NC), early mild cognitive impairment (E-MCI), late mild cognitive impairment (L-MCI), and AD to identify the underlying neuroanatomical basis of this genetically influenced gene. Therefore, the goal of this study is to evaluate the effect of APOE ε4 allele on neurodegeneration, cognition, CSF, and neuroimaging biomarkers across the AD spectrum.
Materials and Methods
Alzheimer’s Disease Neuroimaging Initiative
Data used in the preparation of this paper were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (www.loni.ucla.edu/ADNI). ADNI was launched in 2003 by the National Institute on Aging, the National Institute of Biomedical Imaging and Bioengineering, the Food and Drug Administration, private pharmaceutical companies, and non-profit organizations as a $60 million, 5-year public-private partnership [21]. Subjects have been recruited from over 50 sites across the USA and Canada. To date, these three protocols have recruited over 1500 adults, ages 55 to 90, to participate in the research, consisting of cognitively normal older individuals, early or late MCI, and early AD. Further information can be found at www.adni-info.org for up-to-date information and previous reports [22, 23]. Data for this present analysis were downloaded from the ADNI web site in March 2014. This study was approved by institutional review boards of all participating institutions and written informed consent was obtained from all participants or authorized representatives.
Participants
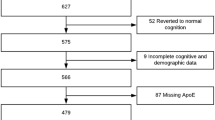
To examine the APOE genetic influence across different clinical and cognitive status further, according to the Clinical Dementia Rating Scale (Sum of Boxes score, CDR-SB), we divided the AD group to the mild AD (M-AD, CDR-SB < 4.5) and severe AD (S-AD, CDR-SB ≥ 4.5) [24]. In this paper, a total of 1718 (S-AD = 180, M-AD = 156, E-MCI = 305, L-MCI = 561, NC = 516) individuals from the ADNI cohort (ADNI1, ADNI2, and ADNI-Go) whose data met all quality control (QC) criteria were included. Detailed quality control steps for CSF [25] and genotype data [26] have been previously reported. Table 1 lists the detailed demographics of all these subjects. Of the 1718 participants, our study included 461 subjects (including 49 AD, 263 MCI, and 149 NC) in MRI analysis, 1288 subjects (including 239 AD, 664 MCI, and 385 NC) in PET analysis, and 1037 subjects (including 343 NC, 301 E-MCI, 257 L-MCI, and 136 AD) in F18-PET-AV45 analysis.
Genotyping, Clinical and Neuropsychological Assessments
APOE genotyping was described in http://www.adni-info.org in detail. The APOE gene is polymorphic with the following three major isoforms: APOE ε2, APOE ε3, APOE ε4. In our analyses that controlled for APOE status (carrier of zero, one, or two APOE ε4 alleles), APOE ε4 carriers were coded as 0, 1, and 2, respectively.
All clinical and neuropsychological test performance data for included participants were downloaded from the ADNI clinical data repository on the Laboratory of Neuro-Imaging (LONI) site. Participants underwent a comprehensive battery of neuropsychological tests, but we only evaluated participant performance on the Mini-Mental State Exam (MMSE), CDR-SB, Alzheimer’s disease Assessment Scale (ADAS), Rey Auditory Verbal Learning Test (RAVLT, total), and Functional Activities Questionnaire (FAQ). Three participants were missing ADAS11 and ADAS13 data and additional five participants were missing FAQ data. Thus, the final samples for neuropsychological testing included 1715 in the ADAS11 and ADAS13 score analysis, and 1713 in the FAQ score analysis.
Cerebrospinal Fluid Data
Levels of Aβ1-42, tau, and phosphorylated-tau (p-tau) were measured from all available CSF samples as described previously [25, 27]. CSF data was downloaded from the LONI site and extracted for all included participants. Of the 1718 included participants, there were only 804 participants with detailed CSF tau, p-tau, and Aβ1-42 data. The final samples for CSF analyses included 221 NC, 192 E-MCI, 264 L-MCI, 62 M-AD, and 65 S-AD participants.
Neuroimaging Data
The neuroimaging data, such as regional volume on MRI, cerebral metabolic rate for glucose (CMRgl) on FDG-PET, and florbetapir cortical standardized uptake values ratios (SUVR) via F18-PET-AV45, were all downloaded from the ADNI dataset. The neuroimaging methods utilized by ADNI have been described in detail previously utilizing the calibration techniques to maintain the consistent protocols across scanners and sites [28].
In our study, we used the regions of interest (ROIs) analysis to calculate differences between APOE genotypes across the AD spectrum. Based on the revised guidelines [29] and our previous meta-analysis [18], we adopted the hippocampal volume and CMRgl in the regional volume to analyze the hippocampal neurodegeneration between the APOE genotypes and AD spectrum. Furthermore, we compared the changed values on hippocampal volume (percent atrophy of hippocampus from baseline) in the follow-up study of 2 years.
Five individual important hypometabolic ROI, including bilateral posterior cingular, left angular gyrus, right angular gyrus, left inferior temporal gyrus, and right inferior temporal gyrus, are downloaded from LONI [30]. These five predefined regions of interest (MetaROIs) are selected based on coordinates cited frequently in other FDG studies comparing AD, MCI, and NCs [31]. We extracted data of florbetapir means of F18-PET-AV45 from four regions (frontal, anterior/posterior cingulate, lateral parietal, and lateral temporal) and global florbetapir SUVR to calculate the amyloid burden.
Statistical Analysis
With each biomarker treated as a continuous scale, differences were calculated by one-way analysis of variance (ANOVA); for categorical marker’s data, differences were tested by Spearman’s correlation analysis. In order to examine the correlation between clinical disease severity and clinical biomarkers, we separate the AD group to the mild AD (M-AD, CDR-SB < 4.5) and severe AD (S-AD, CDR-SB ≥ 4.5) subgroups by their CDR-SB scores [32]. We completed statistics across five subgroups (NC, E-MCI, L-MCI, M-AD, and S-AD) to test the association of APOE ε4 genotypes with the biomarkers in subgroups containing no less than 10 individuals. Furthermore, a Multiple Linear Regression model which considered age, gender, and education as covariates in total sample and subgroups was used to estimate coefficients and the 95 % confidence interval (CI) for testing possible correlation between three APOE ε4 genotypes in these five cognitive groups. To explore this association further, we conducted a multiple linear regression analysis which consider age, gender, and education as covariates in total sample, and calculated the 95 % CI. A Bonferroni-corrected P value (P c) of 0.05 was considered significant, after adjusted for age, sex, and education. We also calculated the 24-month percent changes of cognitive scores, and 12-month and 24-month percent volumetric changes of hippocampus for longitudinal analysis. Sample of CSF and other neuroimaging biomarkers in 24 months was not sufficient for analysis. All statistical analyses were performed by SPSS 19.0 statistics for IBM.
Results
Clinical and Neuropsychometric Findings
The 1718 subjects’ baseline demographic characteristics, neuropsychometric scores, and APOE gene ε4 doses are described in Table 1. As expected, significant effects of diagnosis on the neuropsychometric scores CDR-SB, ADAS11, ADAS13, MMSE, RAVLT, and FAQ were observed (P c < 0.001): AD participants showed a greater CDR-SB, ADAS11, ADAS13, FAQ, as well as lower MMSE and RAVLT scores. Also as expected, the AD and MCI groups had significantly higher proportion of subjects with one or two copies of the APOE ε4 allele (P c < 0.01) than the NC group, with more than 60 % participants (223 out of 336) in AD compared to 28.5 % in NC.
The results of six neuropsychometric scores across AD spectrum are displayed in Supplementary Table S1. In E-MCI, heterozygotes and homozygotes showed significant higher ADAS11 scores than noncarriers (corrected P c = 0.015); in L-MCI, the ADAS11, ADAS13, and RAVLT scores were considerably different across the three genotype groups (P c < 0.01) (Supplementary Table S1 and Fig. S1). APOE ε4 carriers had significantly higher scores on ADAS11 and ADAS13 measures, as well as significantly lower scores on RAVLT measures. MMSE scores and FAQ scores did not show significant difference among the three ε4 allele groups.
CSF Biomarkers
APOE ε4 was significantly associated with lower levels of CSF Aβ1-42 throughout the five diagnostic groups, even after adjusting for age, gender, and education (noncarriers > heterozygotes > homozygotes, Bonferroni-corrected P < 0.01, Fig. 2a, Supplementary Table S2). CSF p-tau and tau protein levels showed remarkable difference in E-MCI and L-MCI subgroups (tau: E-MCI P c < 0.01, L-MCI P c < 0.01; P-tau: E-MCI P c < 0.01, L-MCI P c < 0.01), with the CSF p-tau and tau protein level in noncarriers lower than the others.
Hippocampal Neurodegeneration
As expected, hippocampal volume was indicated to be linearly correlated with the cognitive level (P < 0.001; Table 1). APOE ε4 allele was significantly associated with the hippocampal volumes in the L-MCI and M-AD groups (L-MCI, P c < 0.001; M-AD, P c = 0.015), with both heterozygotes and homozygotes showing more hippocampal atrophy than noncarriers. Besides, the subcortical volume of the left hippocampus was marked associated with the APOE ε4 allele in L-MCI participants (L-MCI, P c < 0.001): the more APOE ε4 dose the more hippocampal atrophy in the L-MCI participants (Supplementary Table S3). In L-MCI and AD groups, APOE ε4 carriers showed more right hippocampal atrophy than noncarriers (L-MCI, P c < 0.001; AD, P c = 0.036). However, no significant difference was shown between APOE ε4 genotypes and hippocampal CMRgl in either cognitive groups.
Neuroimaging biomarkers analysis
In our analysis of ROIs via MRI in baseline, APOE ε4 was significantly associated with several regions across the AD spectrum (see Table 2). In L-MCI, when adjusted for age, gender, and education, the difference remained in several regions: subcortical volume of the right thalamus, subcortical volume of the right ventral DC, subcortical volume of the left amygdala, subcortical volume of the left accumbens area, subcortical volume of the left hippocampus, subcortical volume of the right amygdala, and subcortical volume of the right hippocampus (P c < 0.05, Table 2). Patients with L-MCI who are APOE ε4 carriers exhibit greater atrophy in the right thalamus, right ventral DC, amygdala, left accumbens, and hippocampus.
In the analysis of CMRgl on FDG-PET, only E-MCI group exhibited positive results. In E-MCI, the heterozygotes and homozygotes showed significantly lower CMRgl than the noncarriers on the four regions (right angular gyrus, left angular gyrus, left inferior temporal gyrus, and right inferior temporal gyrus; Supplementary Table S4). Besides, we did not observe significant difference on other groups.
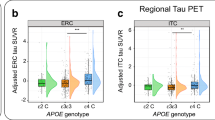
Interestingly, the summary cortical florbetapir SUVRs were significantly associated with the APOE ε4 across the AD spectrum, with the carriers (one or two alleles) showing higher amyloid deposition than noncarriers (P c < 0.05, Fig. 1b). The mean cortical florbetapir SUVR was also associated with the APOE ε4 carriers across the AD spectrum (Supplementary Fig. S2). In the normal and mild cognitive groups (NC, E-MCI, and L-MCI), the highest SUVR (in frontal, cingulate, lateral parietal, and lateral temporal) in subjects with two alleles of APOE ε4 is increasing as the number of alleles increases (P c < 0.01, Fig. 1b).
CSF Aβ1-42 and summary florbetapir cortical SUVR by PET in APOE ε4-negative, homozygous, and heterozygous AD dementia patients. a CSF Aβ1-42 in APOE ε4-negative, homozygous, and heterozygous AD dementia patients. b Summary florbetapir cortical SUVR by PET in APOE ε4-negative, homozygous, and heterozygous AD dementia patients. Multivariate analysis of variance, with APOE dose as independent variable and age, gender, and education as covariates, showed a main effect for APOE dose (all P c < 0.01, except for the Global SUVR in M-AD P c = 0.075). AD = Alzheimer’s disease; M-AD = mild AD patients with high CDR-SB scores; S-AD = AD patients with low CDR-SB scores; SUVR = standardized uptake value ratio
Follow-up Researches
Our results revealed that the APOE ε4 was significantly associated with cognitive decline in L-MCI. Remarkably, APOE ε4 carriers showed significant cognitive decline than noncarriers (P < 0.05, Fig. 2a). APOE ε4 was also significantly associated with the hippocampus volume, with carriers showing more atrophy than the noncarriers (P < 0.05, Fig. 2b). In addition, we detected strong associations on the hippocampus volume percent changes over 24 months from the longitudinal analysis (noncarriers < heterozygotes < homozygotes, L, P < 0.001; R, P < 0.001).
Change from baseline in late mild cognitive impairment (L-MCI). a Change from baseline on cognitive assessment measures in late mild cognitive impairment (L-MCI). Adjusted P value: CDR-SB P = 0.005; MMSE P = 0.13; RAVLT P = 0.002; FAQ P = 0.008. b Percent atrophy of the hippocampus from baseline in late mild cognitive impairment (L-MCI). L m12, left hippocampus volume, 12 months; R m12, right hippocampus volume, 12 months; L m24, left hippocampus volume, 24 months; R m24, right hippocampus volume, 24 months. Adjusted P value: L m12, P < 0.001; R m12, P = 0.004; L m24, P < 0.001, R m24, P < 0.001
Discussion
This study provides a comprehensive evaluation of the impact of APOE ε4 status on CSF Aβ, tau, and p-tau levels; cognitive performance; cerebral atrophy; and brain metabolism across AD spectrum. In our study, we have three major findings: (1) APOE ε4 allele dosage was significantly associated with decreased CSF Aβ1-42 and increased cerebral amyloid deposition across the AD spectrum; (2) The APOE ε4 was significantly associated with increased CSF tau and p-tau in E-MCI and L-MCI subgroups. The ε4 carriers showed significantly cerebral hypometabolism than ε4 noncarriers only in E-MCI; (3) Hippocampal atrophy was associated with APOE ε4 allele in L-MCI and M-AD subgroups. In L-MCI, the APOE ε4 was significantly associated with atrophy of several cerebral regions, as well as cognitive decline manifested by higher ADAS11, ADAS13 and lower RAVLT scores.
In the present study, we confirm that APOE ε4 has a powerful dose-dependent effect on cerebral Aβ deposition. Previous researches had provided evidence on the important role of APOE genotypes in Aβ metabolism [33, 34]. Our findings were consistent with the previous reports that participants (including NC, MCI, and AD) carrying APOE ε4 alleles had lower CSF Aβ1-42 levels than those without an ε4 allele [35, 27, 6] and in ADNI 1 cohort [25]. Current study suggests that APOE genotype strongly affect deposition of Aβ in the brain. Cross-sectional studies by PiB-PET or Florbetapir-PET have consistently reported that cortical Aβ levels were increased in APOE ε4 carriers of E-MCI, L-MCI, and AD [25, 6, 7, 36]. Interestingly, the associations between APOE and amyloid-PET had mainly been seen in NC subjects, but less observed in AD dementia [37]. Although several studies in AD patients have produced paradoxical results against our positive results [38], it should be noted that these studies have focused on clinically manifest AD or severe AD, which may affect the pathologic status significantly.
Nevertheless, the accordance of the both measures of Aβ, Aβ imaging and direct measuring of CSF Aβ concentrations, indicates that cerebral Aβ deposition is the major pathobiological phenotype of APOE ε4 genotype, which would persist during the dementia development. APOE ε4-associated cerebral Aβ deposition could be interpreted by the hypothesized effect of APOE on Aβ clearance and aggregation. Biochemical evidence had shown that the APOE genotype could affect Aβ clearance rate, with the APOE ε4 isoform showing significantly slower clearance [39, 40]. Neuropathological evidence also suggested that APOE ε4 dosage is associated with increased Aβ, Aβ oligomers, and plaque accumulation in the brain [41].
With the analysis across the spectrum of AD, our data indicate that APOE ε4 affects other biomarkers mainly in prodromal stages (E-MCI and L-MCI) of AD. It seems to affect CSF tau, p-tau and regional FDG metabolism in E-MCI stage, and affect the pattern of regional brain atrophy and hippocampal atrophy in L-MCI. These relationships all direct to the APOE-related neurodegenerative function [42].
Our evidence of significant brain Aβ deposition, either from amyloid imaging or CSF Aβ1-42 concentrations, precedes the cerebral neurodegeneration and clinical cognitive changes. Interestingly, the results presented herein are completely consistent with the hypothetical model of dynamic biomarkers for the progression of neuropathology associated with AD (Fig. 3) [42, 27]. These support the hypothesis that the APOE ε4 has a leftward (during the disease progression) shift function of both the Aβ and neurodegenerative biomarkers cascades.
Modulator of APOE ε4 on hypothetical model of biomarkers change across the cascade of Alzheimer’s disease (AD) pathologic progression. Aβ is identified by CSF Aβ42 or PET amyloid imaging. Tau-mediated neuronal injury and dysfunction is identified by CSF tau or fluorodeoxyglucose-PET. Brain structure is measured by use of structural MRI. Aβ = β-amyloid. MCI = mild cognitive impairment
The strength of this study is that it shows relative changes in AD processes across the spectrum of AD. However, interpretations of these results are not certain because the current analyses are based on cross-sectional data, which do not represent individual longitudinal changes. Although relatively large for this study, one limitation of this report is the special ADNI cohort, which is not a population-based cohort. Nevertheless, studies including larger samples are needed to further examine the effect of APOE functions. In addition, not all known biomarkers of AD, including PET-PIB and advanced MRI techniques, were included in this study because of the small sample.
On the basis of our results, we propose that cerebral Aβ deposition is highly associated with APOE genotype, which initiate the pathological cascade of preclinical AD. These were majorly consistent with a recently proposed biomarker model [42], which was hypothesized that accumulation of amyloid-β initiates a cascade of downstream effects, such as neuronal dysfunction and neurodegeneration, which would be amplified by the gene mutation. Therefore, these findings underline the importance of APOE for the disease conversion and progression, which are mediated by its effects on Aβ deposition and a cascade of downstream effects. Elucidating the contribution of APOE ε4 to the neuropathology of AD is a considerable challenge for us, but it provides great support for combating AD.
References
Poirier J, Davignon J, Bouthillier D, Kogan S, Bertrand P, Gauthier S (1993) Apolipoprotein E polymorphism and Alzheimer’s disease. Lancet 342(8873):697–699
Holtzman DM, Herz J, Bu G (2012) Apolipoprotein E and apolipoprotein E receptors: normal biology and roles in Alzheimer disease. Cold Spring Harbor Perspectives Med 2(3):a006312. doi:10.1101/cshperspect.a006312
Liu CC, Kanekiyo T, Xu H, Bu G (2013) Apolipoprotein E and Alzheimer disease: risk, mechanisms and therapy. Nat Rev Neurol 9(2):106–118. doi:10.1038/nrneurol.2012.263
Yu JT, Tan L, Hardy J (2014) Apolipoprotein E in Alzheimer’s disease: an update. Annu Rev Neurosci. doi:10.1146/annurev-neuro-071013-014300
Drzezga A, Grimmer T, Henriksen G, Muhlau M, Perneczky R, Miederer I, Praus C, Sorg C et al (2009) Effect of APOE genotype on amyloid plaque load and gray matter volume in Alzheimer disease. Neurology 72(17):1487–1494. doi:10.1212/WNL.0b013e3181a2e8d0
Morris JC, Roe CM, Xiong C, Fagan AM, Goate AM, Holtzman DM, Mintun MA (2010) APOE predicts amyloid-beta but not tau Alzheimer pathology in cognitively normal aging. Ann Neurol 67(1):122–131. doi:10.1002/ana.21843
Fleisher AS, Chen K, Liu X, Roontiva A, Thiyyagura P, Ayutyanont N, Joshi AD, Clark CM et al (2011) Using positron emission tomography and florbetapir F18 to image cortical amyloid in patients with mild cognitive impairment or dementia due to Alzheimer disease. Arch Neurol 68(11):1404–1411. doi:10.1001/archneurol.2011.150
Caroli A, Frisoni GB, Alzheimer’s Disease Neuroimaging I (2010) The dynamics of Alzheimer’s disease biomarkers in the Alzheimer’s Disease Neuroimaging Initiative cohort. Neurobiol Aging 31(8):1263–1274. doi:10.1016/j.neurobiolaging.2010.04.024
Bookheimer SY, Strojwas MH, Cohen MS, Saunders AM, Pericak-Vance MA, Mazziotta JC, Small GW (2000) Patterns of brain activation in people at risk for Alzheimer’s disease. N Engl J Med 343(7):450–456. doi:10.1056/NEJM200008173430701
Moffat SD, Szekely CA, Zonderman AB, Kabani NJ, Resnick SM (2000) Longitudinal change in hippocampal volume as a function of apolipoprotein E genotype. Neurology 55(1):134–136
Bondi MW, Houston WS, Eyler LT, Brown GG (2005) fMRI evidence of compensatory mechanisms in older adults at genetic risk for Alzheimer disease. Neurology 64(3):501–508. doi:10.1212/01.WNL.0000150885.00929.7E
Langbaum JB, Chen K, Lee W, Reschke C, Bandy D, Fleisher AS, Alexander GE, Foster NL et al (2009) Categorical and correlational analyses of baseline fluorodeoxyglucose positron emission tomography images from the Alzheimer’s Disease Neuroimaging Initiative (ADNI). NeuroImage 45(4):1107–1116. doi:10.1016/j.neuroimage.2008.12.072
Vemuri P, Wiste HJ, Weigand SD, Knopman DS, Shaw LM, Trojanowski JQ, Aisen PS, Weiner M et al (2010) Effect of apolipoprotein E on biomarkers of amyloid load and neuronal pathology in Alzheimer disease. Ann Neurol 67(3):308–316. doi:10.1002/ana.21953
Tosun D, Schuff N, Shaw LM, Trojanowski JQ, Weiner MW, Alzheimer’s Disease NeuroImaging I (2011) Relationship between CSF biomarkers of Alzheimer’s disease and rates of regional cortical thinning in ADNI data. J Alzheimer’s Dis JAD 26(Suppl 3):77–90. doi:10.3233/JAD-2011-0006
Mayeux R, Small SA, Tang M, Tycko B, Stern Y (2001) Memory performance in healthy elderly without Alzheimer’s disease: effects of time and apolipoprotein-E. Neurobiol Aging 22(4):683–689
Farlow MR, He Y, Tekin S, Xu J, Lane R, Charles HC (2004) Impact of APOE in mild cognitive impairment. Neurology 63(10):1898–1901
Caselli RJ, Dueck AC, Locke DE, Hoffman-Snyder CR, Woodruff BK, Rapcsak SZ, Reiman EM (2011) Longitudinal modeling of frontal cognition in APOE epsilon4 homozygotes, heterozygotes, and noncarriers. Neurology 76(16):1383–1388. doi:10.1212/WNL.0b013e3182167147
Liu Y, Yu JT, Wang HF, Han PR, Tan CC, Wang C, Meng XF, Risacher SL et al (2014) APOE genotype and neuroimaging markers of Alzheimer’s disease: systematic review and meta-analysis. J Neurol Neurosurg Psychiatry. doi:10.1136/jnnp-2014-307719
Motter R, Vigo-Pelfrey C, Kholodenko D, Barbour R, Johnson-Wood K, Galasko D, Chang L, Miller B et al (1995) Reduction of beta-amyloid peptide42 in the cerebrospinal fluid of patients with Alzheimer’s disease. Ann Neurol 38(4):643–648. doi:10.1002/ana.410380413
Barber R, Gholkar A, Scheltens P, Ballard C, McKeith IG, Morris CM, O’Brien JT (1999) Apolipoprotein E epsilon4 allele, temporal lobe atrophy, and white matter lesions in late-life dementias. Arch Neurol 56(8):961–965
Mueller SG, Weiner MW, Thal LJ, Petersen RC, Jack CR, Jagust W, Trojanowski JQ, Toga AW et al (2005) Ways toward an early diagnosis in Alzheimer’s disease: the Alzheimer’s Disease Neuroimaging Initiative (ADNI). Alzheimer’s & Dementia J Alzheimer’s Assoc 1(1):55–66. doi:10.1016/j.jalz.2005.06.003
Jack CR Jr, Bernstein MA, Borowski BJ, Gunter JL, Fox NC, Thompson PM, Schuff N, Krueger G et al (2010) Update on the magnetic resonance imaging core of the Alzheimer’s disease neuroimaging initiative. Alzheimer’s Dementia J Alzheimer’s Assoc 6(3):212–220. doi:10.1016/j.jalz.2010.03.004
Petersen RC, Aisen PS, Beckett LA, Donohue MC, Gamst AC, Harvey DJ, Jack CR Jr, Jagust WJ et al (2010) Alzheimer’s Disease Neuroimaging Initiative (ADNI): clinical characterization. Neurology 74(3):201–209. doi:10.1212/WNL.0b013e3181cb3e25
O’Bryant SE, Waring SC, Cullum CM, Hall J, Lacritz L, Massman PJ, Lupo PJ, Reisch JS et al (2008) Staging dementia using Clinical Dementia Rating Scale Sum of Boxes scores: a Texas Alzheimer’s research consortium study. Arch Neurol 65(8):1091–1095. doi:10.1001/archneur.65.8.1091
Shaw LM, Vanderstichele H, Knapik-Czajka M, Clark CM, Aisen PS, Petersen RC, Blennow K, Soares H et al (2009) Cerebrospinal fluid biomarker signature in Alzheimer’s disease neuroimaging initiative subjects. Ann Neurol 65(4):403–413. doi:10.1002/ana.21610
Shen L, Kim S, Risacher SL, Nho K, Swaminathan S, West JD, Foroud T, Pankratz N et al (2010) Whole genome association study of brain-wide imaging phenotypes for identifying quantitative trait loci in MCI and AD: a study of the ADNI cohort. NeuroImage 53(3):1051–1063. doi:10.1016/j.neuroimage.2010.01.042
Risacher SL, Saykin AJ (2013) Neuroimaging and other biomarkers for Alzheimer’s disease: the changing landscape of early detection. Annu Rev Clin Psychol 9:621–648. doi:10.1146/annurev-clinpsy-050212-185535
Zhang D, Wang Y, Zhou L, Yuan H, Shen D (2011) Multimodal classification of Alzheimer’s disease and mild cognitive impairment. NeuroImage 55(3):856–867. doi:10.1016/j.neuroimage.2011.01.008
McKhann GM, Knopman DS, Chertkow H, Hyman BT, Jack CR Jr, Kawas CH, Klunk WE, Koroshetz WJ et al (2011) The diagnosis of dementia due to Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimer’s & dementia J Alzheimer’s Assoc 7(3):263–269. doi:10.1016/j.jalz.2011.03.005
Landau SM, Harvey D, Madison CM, Koeppe RA, Reiman EM, Foster NL, Weiner MW, Jagust WJ et al (2011) Associations between cognitive, functional, and FDG-PET measures of decline in AD and MCI. Neurobiol Aging 32(7):1207–1218. doi:10.1016/j.neurobiolaging.2009.07.002
Landau SM, Lal R, O’Neil JP, Baker S, Jagust WJ (2009) Striatal dopamine and working memory. Cereb Cortex 19(2):445–454. doi:10.1093/cercor/bhn095
O’Bryant SE, Lacritz LH, Hall J, Waring SC, Chan W, Khodr ZG, Massman PJ, Hobson V et al (2010) Validation of the new interpretive guidelines for the clinical dementia rating scale sum of boxes score in the national Alzheimer’s coordinating center database. Arch Neurol 67(6):746–749. doi:10.1001/archneurol.2010.115
Kok E, Haikonen S, Luoto T, Huhtala H, Goebeler S, Haapasalo H, Karhunen PJ (2009) Apolipoprotein E-dependent accumulation of Alzheimer disease-related lesions begins in middle age. Ann Neurol 65(6):650–657. doi:10.1002/ana.21696
Klunk WE, Engler H, Nordberg A, Wang Y, Blomqvist G, Holt DP, Bergstrom M, Savitcheva I et al (2004) Imaging brain amyloid in Alzheimer’s disease with Pittsburgh Compound-B. Ann Neurol 55(3):306–319. doi:10.1002/ana.20009
Tapiola T, Pirttila T, Mehta PD, Alafuzofff I, Lehtovirta M, Soininen H (2000) Relationship between apoE genotype and CSF beta-amyloid (1-42) and tau in patients with probable and definite Alzheimer’s disease. Neurobiol Aging 21(5):735–740
Reiman EM, Chen K, Liu X, Bandy D, Yu M, Lee W, Ayutyanont N, Keppler J et al (2009) Fibrillar amyloid-beta burden in cognitively normal people at 3 levels of genetic risk for Alzheimer’s disease. Proc Natl Acad Sci U S A 106(16):6820–6825. doi:10.1073/pnas.0900345106
Villemagne VL, Pike KE, Chetelat G, Ellis KA, Mulligan RS, Bourgeat P, Ackermann U, Jones G et al (2011) Longitudinal assessment of Abeta and cognition in aging and Alzheimer disease. Ann Neurol 69(1):181–192. doi:10.1002/ana.22248
Ossenkoppele R, van der Flier WM, Zwan MD, Adriaanse SF, Boellaard R, Windhorst AD, Barkhof F, Lammertsma AA et al (2013) Differential effect of APOE genotype on amyloid load and glucose metabolism in AD dementia. Neurology 80(4):359–365. doi:10.1212/WNL.0b013e31827f0889
Deane R, Sagare A, Hamm K, Parisi M, Lane S, Finn MB, Holtzman DM, Zlokovic BV (2008) apoE isoform-specific disruption of amyloid beta peptide clearance from mouse brain. J Clin Invest 118(12):4002–4013. doi:10.1172/JCI36663
Castellano JM, Kim J, Stewart FR, Jiang H, DeMattos RB, Patterson BW, Fagan AM, Morris JC et al (2011) Human apoE isoforms differentially regulate brain amyloid-beta peptide clearance. Sci Transl Med 3(89):89ra57. doi:10.1126/scitranslmed.3002156
Hashimoto T, Serrano-Pozo A, Hori Y, Adams KW, Takeda S, Banerji AO, Mitani A, Joyner D et al (2012) Apolipoprotein E, especially apolipoprotein E4, increases the oligomerization of amyloid beta peptide. J Neurosci Off J Soc Neurosci 32(43):15181–15192. doi:10.1523/JNEUROSCI.1542-12.2012
Jack CR Jr, Knopman DS, Jagust WJ, Shaw LM, Aisen PS, Weiner MW, Petersen RC, Trojanowski JQ (2010) Hypothetical model of dynamic biomarkers of the Alzheimer’s pathological cascade. Lancet Neurol 9(1):119–128. doi:10.1016/S1474-4422(09)70299-6
Acknowledgments
Data collection and sharing was funded by ADNI (National Institutes of Health U01 AG024904). ADNI is funded by the National Institute on Aging; the National Institute of Biomedical Imaging and Bioengineering; the Alzheimer’s Association; the Alzheimer’s Drug Discovery Foundation; BioClinica, Inc; Biogen Idec Inc; Bristol-Myers Squibb Co, F. Hoffmann-LaRoche Ltd and Genetech, Inc; GE Healthcare; Innogenetics, NV; IXICO Ltd; Janssen Alzheimer Immunotherapy Research & Development LLC; Medpace, Inc; Merck & Co, Inc; Meso Scale Diagnostics, LLC; NeuroRx Research; Novartis Pharmaceuticals, Co, Pfizer, Inc; Piramal Imaging; Servier; Synarc Inc; and Takeda Pharmaceutical Co. The Canadian Institutes of Health Research is providing funds to support ADNI clinical sites in Canada. Private sector contributions are facilitated by the Foundation for the National Institutes of Health (www.fnih.org). The grantee organization was the Northern California Institute for Research and Education, and the study was coordinated by the Alzheimer’s Disease Cooperative Study at the University of California, San Diego. ADNI data are disseminated by the Laboratory for Neuro Imaging at the University of California, Los Angeles. This research was also supported by grants from the National Natural Science Foundation of China (81471309, 81371406, 81171209), the Shandong Provincial Outstanding Medical Academic Professional Program, Qingdao Key Health Discipline Development Fund, and Qingdao Outstanding Health Professional Development Fund.
Conflict of Interest
The authors have no conflicts of interest to report.
Author information
Authors and Affiliations
Consortia
Corresponding authors
Additional information
Data used in preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in analysis or writing of this report. A complete listing of ADNI investigators can be found at: http://adni.loni.usc.edu/wp-content/uploads/how_to_apply/ADNI_Acknowledgement_List.pdf
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table S1
(DOCX 17 kb)
Supplementary Table S2
(DOCX 17 kb)
Supplementary Table S3
(DOCX 16 kb)
Supplementary Table S4
(DOCX 18 kb)
Fig. s1
Cross-sectional analyses of six psychological scores across Alzheimer’s disease spectrum. Significant results were marked red star in below. MMSE = Mini-Mental State Exam, CDR-SB = Clinical Dementia Rating Scale Sum of Boxes score, ADAS = Alzheimer’s disease Assessment Scale, RAVLT = Rey Auditory Verbal Learning Test, FAQ = Functional Activities Questionnaire. (GIF 91 kb)
Fig. s2
Cross-sectional analyses of florbetapir cortical retention via PET AV45 across Alzheimer’s disease spectrum. Significant P values after Bonferroni-corrected were indicated using bold font. (GIF 99 kb)
Rights and permissions
About this article
Cite this article
Liu, Y., Tan, L., Wang, HF. et al. Multiple Effect of APOE Genotype on Clinical and Neuroimaging Biomarkers Across Alzheimer’s Disease Spectrum. Mol Neurobiol 53, 4539–4547 (2016). https://doi.org/10.1007/s12035-015-9388-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-015-9388-7