Abstract
Transforming growth factor beta (TGF-β) is suggestive of a molecular target for cancer therapy due to its involvement in cell cycle, differentiation, and morphogenesis. Meanwhile, survivin is identified as an apoptosis inhibitor and involved in tumorgenesis. Here, we aimed to investigate the potential associations between TGF-β and survivin in glioblastoma U87 cell line. Survivin small interfering RNA (siRNA), Western blotting, and cell cycle analysis were introduced to detect relevant proteins in TGF-β pathways. In this study, we observed a concentration- and time-dependent increase of survivin expression after treatment with TGF-β1. However, the kinase inhibitors U0126 and LY294002 inhibited the upregulation of survivin in comparison with DMSO. In addition, survivin siRNA effectively abrogated survivin expression in U87 cells, therefore affected cells’ entry into the S phase of cell cycle, and then repressed the expression of epidermal growth factor receptor (EGFR) and matrix metalloproteinase 9 (MMP9) in comparison with non-transfection. In conclusion, the present study shows that TGF-β upregulates survivin expression via ERK and PI3K/AKT pathway, leading to glioblastoma cell cycle progression. Thus, the blockade of survivin will allow for the treatment of glioblastoma, partially attributing to the inhibition of EGFR and MMP9 expression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glioblastoma ranks among the most prevalent brain tumors, with a poor median survival of less than 1 year [1–3]. Poor survival status contributed to promoting survival-prolonging therapies [4–6]. A number of recent advances have shown prospects for combating resistance to chemotherapy, and then multimodal regimen including radiotherapy and chemotherapy has succeed in modestly improving survival [7]. As a result of little improvement in the patient survival, there have been intensified investigation efforts on the molecule-based therapy.
Transforming growth factor beta (TGF-β) has been reported as a multifunctional cytokine with three isoforms: TGF-β1, TGF-β2, and TGF-β3, among which TGF-β1 is the most abundant in the human body [8]. TGF-β is synthesized as an inactive form by various cells, such as epithelial, hematopoietic, and neuronal cells [9]. TGF-β super family signaling mediators are important regulators of diverse physiological and pathological events [10–12]. Several pathways, involving many downstream proteins, mediate the effects of TGF-β1. Many critical steps in intracellular TGF-β signaling are mediated by Smad proteins. However, Smad-independent signaling transduction pathways are also involved in the biological activities of TGF-β [13, 14]. As the Smad pathway principally regulates gene expression, it was originally thought that non-Smad effectors mediate the rapid or direct effects of TGF-β on tumor invasiveness.
In addition, survivin is a member of the inhibitor of apoptosis protein family, which is highly expressed in most cancer tissues and induces resistance to radiation therapy. It was reported survivin expression is regulated by pre-transcriptional and post-translational factors. Transcription factors dominating survivin gene mainly include hypoxia inducible factor 1α [15], specificity protein 1, STAT3, and Notch [16, 17]. Up to date, the association between TGF-β and survivin has not been clear. In this work, we used survivin small interfering RNA (siRNA), Western blotting, and cell cycle analysis to determine whether a cell survives when survivin levels were depleted. These results provided a novel regulation mechanism of TGF-β1, which can benefit glioblastoma patients.
Materials and Methods
Cell Culture and Reagents
Human glioblastoma cell line U87 was purchased from American Type Culture Collection (ATCC, Manassas, VA) and was cultured in DMEM (Gibco) supplemented with 10 % fetal bovine serum (FBS; HyClone) and 100 U/ml penicillin/streptomycin (Gibco) and were maintained in a humidified atmosphere with 5 % CO2 at 37 °C. Recombinant human TGF-β1 was purchased from Sigma (St. Louis, USA). Specific inhibitors of MEK (U0126) and PI3K (LY294002) were purchased from Calbiochem (La Jolla, CA, USA). Antibodies were purchased from the same resources: anti-survivin, anti-ERK, anti-AKT, anti-cyclin D1, anti-CDK2, anti-epidermal growth factor receptor (EGFR), anti-matrix metalloproteinase 9 (MMP9), and anti-β-actin antibody (Santa Cruz Biotechnology, Santa Cruz, USA). To test the effects of inhibitors on particular signaling molecules, the inhibitors were added to cells 1 h prior to TGF-β1 treatment. All experiments were carried out in the absence of FBS.
Transfection of Survivin siRNA
Cell lines were seeded in a 6-cm dish at density of 5 × 105 cells/dish and incubated overnight. U87 was prepared for transfection of survivin siRNA or control siRNA (generously provided by Dr. Wang, Chinese Academy of Medical Sciences). siRNA was added to Opti-MEM with Lipofectamine 2000 (Invitrogen) for transfection, according to the manufacturer’s instructions. Twelve hours following incubation, the medium was changed into a fresh medium containing 10 % FBS. Cells were harvested at 72 h following transfection of survivin siRNA. Efficiencies of survivin siRNA and non-silencing control siRNA were tested using Western blot.
Flow Cytometry Analyses
Cell cycle analysis was performed on the flow cytometer, after propidium iodide (PI) staining. U87 cells, in the logarithmic phase of growth, were seeded in six-well plates and incubated overnight. The cells, except in control group, were transfected with survivin siRNA or control siRNA. After 24 h, the cells were collected, washed with Dulbecco’s phosphate-buffered saline (DPBS; Genview, El Monte, USA), fixed in 70 % ethanol, and incubated overnight at 4 °C. Ethanol was removed by centrifugation. The cell pellets were washed with DPBS followed by incubation with 300 μL PI solutions, for 30 min in the dark at 37 °C. Cells were then analyzed by flow cytometry.
Western Blotting
Total protein from cell lines was extracted in cell lysis buffer (PIERCE, Rockford, IL) and quantified using the BSA method. A 10 % sodium dodecyl sulfate (SDS)-PAGE was performed, and 30 μg of protein of each sample were analyzed. Proteins in the SDS gels were transferred to a polyvinylidene difluoride membrane by an electroblot apparatus. Membranes were incubated with primary antibodies. Antibody recognition was detected with either anti-mouse IgG or anti-rabbit IgG antibody linked to horseradish peroxidase (Sigma). Immunocomplexes were visualized by ECL (Amersham Pharmacia Biotech).
Statistical Analysis
The data was presented as the mean ± SEM. The analysis of difference was performed using one-way ANOVA with the LSD post hoc test for multiple comparisons with SPSS version 13.0 software. The p < 0.05 was considered as significant difference.
Results
TGF-β1 Induces Survivin Expression in A Dose- and Time-Dependent Manner
To figure out the effects of TGF-β1 on survivin expression, firstly, we treated cells with serum-starved medium for 30 min, and then cells were subjected to different TGF-β1 treatment and Western blotting analysis. After TGF-β1 gradient induction, Western blotting analysis showed that the expression of survivin protein increased in a concentration-dependent fashion. At the same time, we also found 20 ng/ml of TGF-β1 can induce the highest protein expression level (Fig. 1a).
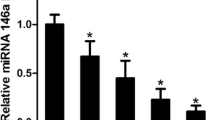
Survivin is regulated in response to TGF-β1 treatment. a Cells were pretreated with serum-starved medium for 30 min, and then treated with 0, 10, 20, and 50 ng/ml of TGF-β1. Subsequently, survivin was detected using Western blotting, and then measured with Image J. β-actin was used as a normalization control. b After that, U87 cells were cultured in the absence or presence of 20 ng/ml of TGF-β1 for different times. Cell lysates were prepared and the level of survivin protein was analyzed by Western blotting with a specific antibody. Each bar represents the mean ± SEM of three independent experiments; *p < 0.001, compared with control, one-way ANOVA with the LSD post hoc test
Subsequently, we performed Western blot analyses on U87 cells treated with 20 ng/ml of TGF-β1 and harvested at 0, 2, 6, 12, 36, and 48 h post-treatment, respectively. Results from densitometry measurements revealed that survivin protein levels also increased in TGF-β1-treated cells in a time-dependent fashion, and differences from control were significant (p < 0.001) (Fig. 1b). This increase started at the 2nd hour following TGF-β1 treatment and peaked at the 12th hour.
Upregulation of Survivin Induced by TGF-β1 Is Reduced by Inhibition of Both ERK and PI3K/AKT Pathways
Recent reports suggested that the activation of conventional ERK and PI3K/AKT signaling pathways were involved in the TGF-β1 pathway, which may be required for the upregulation of survivin. To investigate the role of the key signaling mediators in the upregulation of survivin, we used kinase inhibitors to individually block each signaling pathway in U87 cells, and then treated with 20 ng/ml of TGF-β1, and then examined the level of survivin expression. Both U0126 and LY294002 significantly reduced the survivin expression level in U87 cells, compared to U87 cells treated with DMSO alone (Fig. 2). These data suggests that ERK and PI3K/AKT signaling are important for the upregulation of survivin in response to TGF-β1.
TGF-β1-induced upregulation of survivin is attributed to the activation of ERK and PI3K/AKT pathways. Inhibition of the MEK/ERK and PI3K/AKT signaling pathways by chemical inhibitors MAP/ERK kinase inhibitor U0126 or 40 mM of PI3K inhibitor LY294002, respectively. Serum-starved U87 cells were pretreated for 1 h with DMSO alone or 20 mM of U0126 or 40 mM of LY294002, and then cultured in the presence of 20 ng/ml TGF-β1 for 12 h. Cell lysates were prepared and the level of survivin protein was analyzed by Western blotting. Each bar represents the mean ± SEM of three independent experiments; *p < 0.001, compared with control, one-way ANOVA with the LSD post hoc test
Cell Cycle Progression Is Regulated by Survivin in U87 Cells
To further investigate the role of survivin induced by TGF-β1 in cell cycle progression of U87 cells, we efficiently inhibited the expression of endogenous survivin using transfection with survivin siRNA and detected cell cycle progression of U87 cells treated with TGF-β1. We found that survivin siRNA significantly inhibited the protein level of survivin, compared with control siRNA (Fig. 3a). We further assessed the role of survivin in cell cycle using flow cytometry. The proportion of cells in different phases of the cell cycle was determined by flow cytometry. In comparison with TGF-β1 alone or control siRNA treatment, survivin inhibition by survivin siRNA did significantly decrease the S phase of U87 cells (p < 0.001). This indicates that survivin indeed affected cell cycle progression at the G1/S phase (Fig. 3b).
Survivin is critical for TGF-β1-mediated cell cycle progression. a After transfection, the expression of survivin protein was evaluated using Western blot and compared to untransfected cells. Cells transfected with survivin siRNA completely inhibited survivin expression compared with si-control. b Samples were taken after 24 h of TGF-β1 treatment, and DNA content was assayed by flow cytometry and PI staining. The gates and percentages of cells in the G1, S, and G2 phases are indicated. c, d Serum-starved U87 cells were cultured in the presence of 20 ng/ml TGF-β1 for 24 h. Western blotting analysis of expression of Cyclin D1 and CDK2 was conducted in cell lysates of U87 cells. β-actin served as the loading control. Each bar represents the mean ± SEM of three independent experiments; *p < 0.001, compared with control, one-way ANOVA with the LSD post hoc test
As reported, Cyclin D1 and CDK2 were recommended as the important biomarker during the cell cycle; we also tested the expression status of Cyclin D1 and CDK2. Our data revealed that survivin knockdown can inhibit the expression of Cyclin D1 and CDK2 compared with non-transfection and control (Fig. 3c, d), indicating that cell cycle progression was affected by survivin.
TGF-β1 Regulates the EGFR and MMP9 expression via Survivin
Glioblastoma is characterized by local invasiveness and frequent recurrence. The surrounding stroma, composed of different cell types and extracellular matrix, may influence glioblastoma invasion. Furthermore, tumor and stromal cells secrete matrix metalloproteases, which, in turn, can modulate the matrix and promote the release of growth factors. Among these growth factors, EGF and its receptor EGFR have already been shown to stimulate MMP9 synthesis to contribute to the tumor invasiveness. To examine whether the TGF-β1-induced survivin affected the expression of EGFR and MMP9, we cultured U87 with transfection of survivin siRNA or control siRNA, and then exposed to 20 ng/ml of TGF-β1. In the present study, the expression of EGFR and MMP9 protein was tested through Western blotting in different times. As shown in Fig. 4, Western blot analysis revealed that EGFR and MMP9 protein were significantly decreased in a time-dependent manner in response to survivin siRNA. In contrast, control siRNA efficaciously promoted EGFR and MMP9 expression. Taken together, these results suggested that the TGF-β1-induced survivin upregulation regulated the expression of EGFR and MMP9.
Expression of EGFR and MMP9 protein is regulated by survivin in U87. U87 was transfected as described above. Serum-starved U87 cells were treated with 20 ng/ml of TGF-β1 for different times, and then cell lysates were prepared for Western blotting with a specific antibody. Western blotting analysis of EGFR and MMP9 expression results were normalized with the level of β-actin. The relative intensity was expressed as a fraction of β-actin. Data represent the mean ± SEM of at least three independent experiments. *p < 0.001, vs control siRNA
Discussion
Survivin is widely implicated in tumor development and progression due to its ability to inhibit apoptosis, promote cell cycle progression, facilitate metastasis, and enhance angiogenesis [16]. TGF-β is also a multifunctional cytokine and a key regulator of cell fate, such as epithelial-to-mesenchymal transition, which is involved in cancer invasion and progression. Once binding to its type I and type II serine/threonine kinase receptors, TGF-β conveys its signals to the Smad2 and Smad3 signaling protein, and then the phosphorylation of Smad2 and Smad3 promotes their association with Smad4, which regulates the expression of targets genes, such as Smad7, p21, and c-Jun. While the connection between survivin and TGF-β has not been extensively documented, we provided evidence to highlight a role of survivin in the cell cycle via ERK and PI3K pathway in U87 cells.
It should be noted that the non-Smad pathways include various branches of MAPK pathways and PI3K/AKT pathways. At the same time, pharmacological modulation of survivin was tagged with its evolving functional complexity associated with various cell signaling cascades including PI3K/AKT, ERK, p53, EGFR, etc. so we hypothesized that ERK and PI3K/AKT was involved in the activation of survivin. In the present study, we used specific kinase inhibitors to individually block each signaling pathway in U87 cells treated with TGF-β1, and then examined the level of survivin expression. Both U0126 and LY294002 significantly reduced the survivin induction in U87 cell line, compared to the control cells treated with DMSO alone. These data suggest that ERK and PI3K/AKT signaling is required for the upregulation of survivin in response to TGF-β1.
It is widely accepted that during the late stage of tumor progression, inhibitory effects of TGF-β on cell proliferation are lost. The prominent mechanisms of TGF-β on tumor progression are epithelial-to-mesenchymal transition, tumor-stroma interaction, and microenvironment [18], and TGF-β was suggested as a tumor promoter [19]. In this work, we investigated whether TGF-β promoted cell cycle via survivin. In comparison with TGF-β1 alone or control siRNA treatment, survivin inhibition by survivin siRNA significantly inhibited the entry into S phase of U87 cells. This indicates that survivin indeed affected cell cycle progression.
Cyclin D1 is a putative regulator of cell cycle, regulating the G1/S transition [20–22]. Cyclin D1 expression is regulated by mitogenic signaling through the Ras-signaling pathway involving Ras/Raf/mitogen-activated protein [23]. Cyclin D1 levels are also post-translationally regulated by its degradation through the following ubiquitin-proteasome pathway. In addition, CDK2 binds to cyclin E or cyclin A and exclusively promotes the G1/S phase transition and that Cdc2/cyclin B complexes play a major role in mitosis [24–26]. In this study, we showed that TGF-β1 increased the active level of CDK2 and Cyclin D1. Conversely, this level was affected by depletion of survivin, suggesting that TGF-β1 induces cell cycle progression by regulating the survivin activity.
At last, we studied the effects of survivin on TGF-β1-induced EGFR and MMP9. Recent data suggest that increased expression of EGF and its receptor EGFR in gastric mucosa may induce changes in gastric epithelial cells leading to tumorigenesis [27, 28]. EGF, through interaction with its receptor, stimulates the cell proliferation and migration and triggers epithelial cell signaling [29, 30]. Besides, matrix metalloproteinases are secreted during the growth, invasion, metastases, and angiogenesis of tumors and can affect the surrounding microenvironment, causing dynamic changes of biological behaviors of the tumor [31]. However, the precise molecular mechanisms between TGF-β, EGFR, and MMP9 in the cellular malignant and invasive phenotypes are not fully understood. We next identified the downstream signaling events responsible for the upregulation of survivin in response to TGF-β1. EGFR and MMP-9 protein were significantly decreased in a time-dependent manner in response to survivin siRNA. In contrast, control siRNA efficaciously promoted EGFR and MMP9 expression. Taken together, these results suggested that the TGF-β1-induced upregulation of survivin regulated the expression of EGFR and MMP9.
In summary, our data demonstrated that survivin serves as a regulator of TGF-β1-induced cell cycle. TGF-β1 can upregulate survivin expression via the ERK and PI3K pathway, and this increased level of survivin promotes cell cycle progression and the expression of EGFR and MMP9. Thus, the blockade of survivin will allow for the treatment of glioblastoma, partially attributing to the inhibition of EGFR and MMP9 expression.
References
Buckner JC (2003) Factors influencing survival in high-grade gliomas. Semin Oncol 30(6 Suppl 19):10–14
Stupp R, Mason WP, van den Bent MJ, Weller M, Fisher B, Taphoorn MJ, Belanger K, Brandes AA et al (2005) Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med 352(10):987–996
Woodworth GF, Dunn GP, Nance EA, Hanes J, Brem H (2014) Emerging insights into barriers to effective brain tumor therapeutics. Front Oncol 4:126
Walker MD, Alexander E Jr, Hunt WE, MacCarty CS, Mahaley MS Jr, Mealey J Jr, Norrell HA, Owens G et al (1978) Evaluation of BCNU and/or radiotherapy in the treatment of anaplastic gliomas. A cooperative clinical trial. J Neurosurg 49(3):333–343
Walker MD, Green SB, Byar DP, Alexander E Jr, Batzdorf U, Brooks WH, Hunt WE, MacCarty CS et al (1980) Randomized comparisons of radiotherapy and nitrosoureas for the treatment of malignant glioma after surgery. N Engl J Med 303(23):1323–1329
Parsa AT, Waldron JS, Panner A, Crane CA, Parney IF, Barry JJ, Cachola KE, Murray JC et al (2007) Loss of tumor suppressor PTEN function increases B7-H1 expression and immunoresistance in glioma. Nat Med 13(1):84–88
Nagasawa DT, Chow F, Yew A, Kim W, Cremer N, Yang I (2012) Temozolomide and other potential agents for the treatment of glioblastoma multiforme. Neurosurg Clin N Am 23(2):307–322
Sporn MB, Roberts AB (1992) Transforming growth factor-beta: recent progress and new challenges. J Cell Biol 119(5):1017–1021
Blobe GC, Schiemann WP, Lodish HF (2000) Role of transforming growth factor beta in human disease. N Engl J Med 342(18):1350–1358
Gao J, Zhu Y, Nilsson M, Sundfeldt K (2014) TGF-β isoforms induce EMT independent migration of ovarian cancer cells. Cancer Cell Int 14(1):72
Gonzalez DM, Medici D (2014) Signaling mechanisms of the epithelial-mesenchymal transition. Sci Signal 7(344):re8
Liu RY, Zeng Y, Lei Z, Wang L, Yang H, Liu Z, Zhao J, Zhang HT (2014) JAK/STAT3 signaling is required for TGF-β-induced epithelial-mesenchymal transition in lung cancer cells. Int J Oncol 44(5):1643–1651
Takai E, Tsukimoto M, Kojima S (2013) TGF-β1 downregulates COX-2 expression leading to decrease of PGE2 production in human lung cancer A549 cells, which is involved in fibrotic response to TGF-β1. PLoS One 8(10):e76346
Cho KH, Jeong KJ, Shin SC, Kang J, Park CG, Lee HY (2013) STAT3 mediates TGF-β1-induced TWIST1 expression and prostate cancer invasion. Cancer Lett 336(1):167–173
Li W, Chen YQ, Shen YB, Shu HM, Wang XJ, Zhao CL, Chen CJ (2013) HIF-1α knockdown by miRNA decreases survivin expression and inhibits A549 cell growth in vitro and in vivo. Int J Mol Med 32(2):271–280
Guha M, Altieri DC (2009) Survivin as a global target of intrinsic tumor suppression networks. Cell Cycle 8(17):2708–2710
Lladser A, Sanhueza C, Kiessling R, Quest AF (2011) Is survivin the potential Achilles’ heel of cancer? Adv Cancer Res 111:1–37
Travis RC, Reeves GK, Green J, Bull D, Tipper SJ, Baker K, Beral V, Peto R et al (2010) Gene-environment interactions in 7610 women with breast cancer: prospective evidence from the Million Women Study. Lancet 375(9732):2143–2151
Sánchez-Elsner T, Botella LM, Velasco B, Corbí A, Attisano L, Bernabéu C (2001) Synergistic cooperation between hypoxia and transforming growth factor-beta pathways on human vascular endothelial growth factor gene expression. J Biol Chem 276(42):38527–38535
Ravikumar G, Ananthamurthy A (2014) Cyclin D1 expression in ductal carcinoma of the breast and its correlation with other prognostic parameters. J Cancer Res Ther 10(3):671–675
Rekhi B, Motghare P (2014) Cyclin D1 and p16INK4 positive endometrial stromal sarcoma: a case report with new insights. Indian J Pathol Microbiol 57(4):606–608
Li T, Zhao X, Mo Z, Huang W, Yan H, Ling Z, Ye Y (2014) Formononetin promotes cell cycle arrest via downregulation of Akt/Cyclin D1/CDK4 in human prostate cancer cells. Cell Physiol Biochem 34(4):1351–1358
Soppa U, Schumacher J, Florencio Ortiz V, Pasqualon T, Tejedor FJ, Becker W (2014) The Down syndrome-related protein kinase DYRK1A phosphorylates p27(Kip1) and Cyclin D1 and induces cell cycle exit and neuronal differentiation. Cell Cycle 13(13):2084–2100
Yang Y, Ma B, Li L, Jin Y, Ben W, Zhang D, Jiang K, Feng S et al (2014) CDK2 and CDK4 play important roles in promoting the proliferation of SKOV3 ovarian carcinoma cells induced by tumor-associated macrophages. Oncol Rep 31(6):2759–2768
Kokontis JM, Lin HP, Jiang SS, Lin CY, Fukuchi J, Hiipakka RA, Chung CJ, Chan TM et al (2014) Androgen suppresses the proliferation of androgen receptor-positive castration-resistant prostate cancer cells via inhibition of Cdk2, CyclinA, and Skp2. PLoS One 9(10):e109170
Ahn SH, Jeong EH, Lee TG, Kim SY, Kim HR, Kim CH (2014) Gefitinib induces cytoplasmic translocation of the CDK inhibitor p27 and its binding to a cleaved intermediate of caspase 8 in non-small cell lung cancer cells. Cell Oncol (Dordr) 37(5):377–386
García I, Vizoso F, Andicoechea A, Fernandez P, Suarez C, García-Muñz JL, Allende MT (2000) C-erbB-2 oncoprotein content in gastric cancer and in adjacent mucosa. Int J Biol Markers 15(3):231–234
Shun CT, Wu MS, Lin JT, Chen SY, Wang HP, Lee WJ, Wang TH, Chuang SM (1997) Relationship of p53 and c-erbB-2 expression to histopathological features, Helicobacter pylori infection and prognosis in gastric cancer. Hepatogastroenterology 44(14):604–609
Dan L, Jian D, Na L, Xiaozhong W (2012) Crosstalk between EGFR and integrin affects invasion and proliferation of gastric cancer cell line, SGC7901. Onco Targets Ther 5:271–277
Sarosiek J, Bilski J, Murty VL, Slomiany A, Slomiany BL (1988) Role of salivary epidermal growth factor in the maintenance of physicochemical characteristics of oral and gastric mucosal mucus coat. Biochem Biophys Res Commun 152(3):1421–1427
Kessenbrock K, Plaks V, Werb Z (2010) Matrix metalloproteinases: regulators of the tumor microenvironment. Cell 141(1):52–67
Acknowledgments
This work is supported by the National Natural Science Foundation of China (grant no. 81402926 to Dr. Wenliang Chen) and Youth Program from Guangzhou City Bureau of Education (no. 2012C204 to Dr. Wen-liang Chen). We gratefully thank other members of Sandy Lab for their critical reading of this paper and valuable suggestions.
Conflicts of interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
This article has been retracted at the request of the Editor-in-Chief and the Publisher per the Committee on Publication Ethics guidelines. There is strong reason to believe that the peer review process was compromised and the authors have plagiarized parts from the following articles:
Anyan Liao, Ranran Shi, Yuliang Jiang, Suqing Tian, Panpan Li, Fuxi Song, Yalan Qu, Jinna Li, Haiqin Yun, Xiangshan Yang, SDF-1/CXCR4 Axis Regulates Cell Cycle Progression and Epithelial-Mesenchymal Transition via Up-regulation of Survivin in Glioblastoma, Molecular Neurobiology, January 2016, Volume 53, Issue 1, pp 210–215, DOI: 10.1007/s12035-014-9006-0
Received: 1 November 2014
Peng Yang, Gang Wang, Hongjun Huo, Qiang Li, Yan Zhao, Yuanhang Liu, SDF-1/CXCR4 signaling up-regulates survivin to regulate human sacral chondrosarcoma cell cycle and epithelial–mesenchymal transition via ERK and PI3K/AKT pathway, Medical Oncology, January 2015, 32:377, DOI: 10.1007/s12032-014-0377-x
Received: 13 November 2014
As such, the validity of the content of this article cannot be verified
An erratum to this article is available at http://dx.doi.org/10.1007/s12035-017-0581-8.
About this article
Cite this article
Chen, W., Zhong, X., Wei, Y. et al. RETRACTED ARTICLE: TGF-β Regulates Survivin to Affect Cell Cycle and the Expression of EGFR and MMP9 in Glioblastoma. Mol Neurobiol 53, 1648–1653 (2016). https://doi.org/10.1007/s12035-015-9121-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-015-9121-6