Abstract
Non-small cell lung cancer (NSCLC) is a fatal malignancy all over the world. Emerging studies have shown that curcumin might repress NSCLC progression by regulating ferroptosis, but the underlying mechanism remains unclear. 16HBE, LK-2, and H1650 cell viability was detected using Cell Counting Kit-8 assay. LK-2 and H1650 cell proliferation, apoptosis, and angiopoiesis were measured using 5-ethynyl-2’-deoxyuridine, flow cytometry, and tube formation assay. Superoxide dismutase, Malondialdehyde, Glutathione, and lactate dehydrogenase levels in LK-2 and H1650 cells were examined using special assay kits. Fe+ level was assessed using an iron assay kit. Doublesex and Mab-3 related Transcription Factor 3 (DMRT3) and solute carrier family 7 member 11 (SLC7A11) protein levels were detected using western in NSCLC tissues, adjacent matched normal tissues, 16HBE cells, LK-2 cells, H1650 cells, and xenograft tumor tissues. Glutathione peroxidase 4, Acyl-CoA Synthetase Long Chain Family Member 4, and transferrin receptor 1 protein levels in LK-2 and H1650 cells were examined by western blot assay. DMRT3 and SLC7A11 levels were determined using real-time quantitative polymerase chain reaction. After JASPAR prediction, binding between DMRT3 and SLC7A11 promoter was verified using Chromatin immunoprecipitation and dual-luciferase reporter assays in LK-2 and H1650 cells. Role of curcumin on NSCLC tumor growth was assessed using the xenograft tumor model in vivo. Curcumin blocked NSCLC cell proliferation and angiopoiesis, and induced apoptosis and ferroptosis. DMRT3 or SLC7A11 upregulation partly abolished the suppressive role of curcumin on NSCLC development. In mechanism, DMRT3 was a transcription factor of SLC7A11 and increased the transcription of SLC7A11 via binding to its promoter region. Curcumin inhibited NSCLC growth in vivo by modulating DMRT3. Curcumin might constrain NSCLC cell malignant phenotypes partly through the DMRT3/SLC7A11 axis, providing a promising therapeutic strategy for NSCLC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As a most prevalent and fatal malignant tumor, lung cancer has been recognized as a significant global healthcare problem and an enormous social burden [1, 2]. According to histologic types and prognosis, non-small cell lung cancer (NSCLC) is responsible for over 80% of lung cancer cases, exhibiting a trend of increasing incidence in young adults in recent years [3, 4]. Clinically, standard treatments, targeted therapy, immunotherapy drugs, and combined therapy have made substantial advances, but the overall survival and recovery rates are still unfavorable, which is associated with late diagnosis, drug resistance, side effects, and toxicities [5, 6]. Hence, there is an urgent need to identify more effective therapies for patients with NSCLC, which has important clinical significance.
Currently, increasing attention is focusing on traditional Chinese medicine (TCM), which possesses numerous therapeutic properties with decreased side effects [7]. As a naturally occurring polyphenol compound extracted from the medical plant Curcuma longa (also known as turmeric) or the Indian spice turmeric, curcumin is one of three curcuminoids and possesses various pharmacologic activities, including anti-inflammatory, anti-oxidant, anti-microbial, and anti-cancer [8, 9]. Unfortunately, curcumin is characterized by poor water solubility (0.6 µg/mL), which also means low bioavailability (its level in the serum is up to 60 nM). Currently, many scientists are working on improving the solubility of curcumin in water, such as microencapsulation, nanosuspensions, liposomes, micelles, or complexes with cycloserine [10]. It has been reported that the low solubility of curcumin and its rapid metabolism by intestinal glucuronidation are major factors that contribute to the low bioavailability. In the body, curcumin is metabolized by glucuronidation into dehydrocurcumin, tetrahydrocucumin, and hexahydrocurcumin. Glucuronidation plays a major role in its metabolism since 99% of curcumin is present in the plasma as a glucuronidation conjugate [11]. Furthermore, piperine, a potent inhibitor for UDP-glucuronosyltransferase, was incorporated into the formulations to potentially enhance the bioavailability of curcumin [12]. Of note, the prominent features of curcumin are chemical stability, low toxicity, and wide distribution, thus the application of curcumin in the treatment of diverse cancers is gradually being recognized [13, 14]. Several laboratory works have indicated that curcumin might exert an anti-carcinogenic capability by modulating growth factors, transcription factors, protein kinases, and oncogenes in different cancers [15], including lung cancer [16]. In terms of NSCLC trials, curcumin and its analogs have confirmed control over cell proliferation, ferroptosis, and metastasis via multiple mechanisms [17, 18]. However, the precise molecular mechanisms of curcumin on NSCLC development are far from being addressed.
The Doublesex and Mab-3 related transcription factor (DMRT) is a deeply conserved family of transcription factors with a unique DNA binding motif, which is able to bind to and modulate DNAs [19, 20]. As a member of the DMRT family, DMRT3 has been verified to participate in determining sexual dimorphism and reproducing sexually [21, 22]. Current studies have suggested that DMRT3 is abnormally expressed in many cancers and boosts tumorigenesis and development through different mechanisms [23]. Furthermore, DMRT3, together with TP63/SOX2, is responsible for an oncogenic transcription factor, which co-regulates the genes involved in epithelial cell differentiation during NSCLC process [24]. Nevertheless, it is still unknown whether DMRT3 could participate in curcumin-mediated NSCLC progression.
In the present work, a public prediction server SRAMP initially found that the DMRT3 motif is bound to the promoter regions of solute carrier family 7 member 11 (SLC7A11). It has been reported that SLC7A11 might be modulated at the transcriptional and translational levels [25, 26]. Beyond that, the dysregulation of SLC7A11 might be associated with ferroptosis in lung cancer [27]. Meanwhile, SLC7A11 expression was significantly decreased in curcumin-treated NSCLC cells [17]. Hence, the present work attempted to identify whether DMRT3 might control curcumin-mediated NSCLC process by regulating SLC7A11 expression.
Materials and Methods
Cell Culture and Clinical Samples
Under the standard conditions with 5% CO2 at 37 °C, NSCLC cells (LK-2, CBP60105, Cobioer, Nanjing, China; H1650, CL-0166, Procell, Wuhan, China) and human bronchial epithelial cell line (16HBE, CL-0249, Procell) were routinely grown in RPMI1640 medium (Gibco, Rockville, MD, USA). In addition, the media were supplemented with 10% FBS (Gibco) and 1% penicillin/streptomycin (Invitrogen). For in vitro study, curcumin (purity ≥ 99.5%, dissolved in DMSO, Sigma–Aldrich, St.Louis, MO, USA) was diluted at a dose range of 0 (control), 10, 20, and 40 μM in cell culture medium, followed by incubation with 16HBE cells and NSCLC cell lines (LK-2 and H1650) for 24 h. For function analysis, LK-2 and H1650 cells were cultured in the media containing 20 μM curcumin for 24 h. diluted as needed in cell culture.
Samples of the NSCLC tissues and paired adjacent normal tissues (n = 33) were collected from 33 patients who were diagnosed with NSCLC at the Changzhou Cancer Hospital. Every participant signed the informed consent prior to enrolling in the study. Subsequently, these excised specimens during the surgery were instantly frozen in liquid nitrogen and kept at a stable temperature of -80 °C. For this research, approval was endowed by the Ethics Committee of Changzhou Cancer Hospital.
Counting Kit (CCK-8) Assay
The assessment of cell viability was performed in this experiment. In brief, 5 × 103 cells in 96-well plates were cultured overnight at 37 °C. Then, these cells were exposed to curcumin for 24 h, and 0 μM curcumin was applied as a control. In parallel, a culture system was made by mixing CCK-8 reagent (10 μL, Sigma–Aldrich) with 90 μL fresh complete media, followed by addition to each well. After incubation for 2 h, the absorbance at 450 nm was recorded based on a microplate reader.
5-ethynyl-2′-Deoxyuridine (EdU) Assay
In this experiment, 5 × 104 treated cells in 6-well plates were mixed with 50 μM EdU working solution (RiboBio, Guangzhou, China) for 2 h. After being fixed in 4% formaldehyde for 30 min, cells were subjected to 0.5% Triton-X-100 permeabilization, Apollo reaction cocktail reaction, and DAPI staining (identify the nuclei). Finally, EdU-positive cells were visualized and analyzed based on a fluorescence microscope.
Flow Cytometry for Cell Apoptosis
In short, treated cells were first digested with trypsin (Sigma–Aldrich) to form a single cell, which then was re-suspended in a binding buffer at a density of 5 × 105 cells/well. After being incubated with 5 μL Annexin V-FITC for 10 min, the cells were subjected to 5 μL PI staining for 5 min, followed by the analysis of the apoptotic cells using a FACSCalibur Flow Cytometry instrument.
Tube Formation Assay
In this assay, 300 μL matrigel was first polymerized on 24-well plates. Then, HUVECs (Procell) were incubated with the medium preconditioned with LK-2 and H1650 cells (1 × 105), followed by introduction into the matrigel-coated wells. 18 h later, angiogenic activity was quantified by measuring the length of tube walls formed using ImageJ.
Measurement of Superoxide Dismutase (SOD), Malondialdehyde (MDA), Glutathione (GSH), and Lactate Dehydrogenase (LDH)
These characteristic indicators of ferroptosis were assessed in this experiment. After 20 μM curcumin treatment for 24 h, transfected or un-transfected tumor cells were harvested and lysed. After that, intracellular SOD, MDA, GSH, and LDH levels in the cell extract were examined using commercial SOD kits (S0101S, Beyotime, Shanghai, China; based on the colorimetric reaction of WST-8 for the detection of SOD), MDA kit (ab118970, Abcam, Cambridge, MA, USA; MDA in the sample reacts with thiobarbituric acid (TBA) to generate a MDA-TBA adduct, which was quantified colorimetrically (OD = 532 nm) or fluorometrically (Ex/Em = 532/553 nm), GSH kit (CS0260, Sigma–Aldrich; based on an enzymatic cycling method in the presence of GSH and a chromophore), and LDH kit (C0016, Beyotime; a Diaphorase-based INT chromogenic reaction that detects lactate dehydrogenase activity released in response to cytotoxicity by colorimetry).
Iron Measurement
Briefly, the content of iron ion (Fe2+) in LK-2 and H1650 cells was monitored based on the Iron Colorimetric Assay Kit (ab83366, Abcam; Iron Carrier Protein releases iron into solution as iron/iron ions in an acidic conditions, and free ferrous iron (Fe2 +) reacts with the iron probe to form a stable colored complex with an absorbance of 593 nm), followed by the assessment of the absorbance values using a microplate reader at 593 nm.
Western Blot Assay
In brief, the isolation of total protein from 1 × 106 cells or 100 mg clinical samples was conducted based on RIPA buffer (Keygen, Nanjing, China) and then was quantified by Pierce™ BCA Protein Assay Kit. Subsequently, 30 μg of each protein lysates were resolved by 10% SDS polyacrylamide gel and shifted to nitrocellulose membranes (Millipore, Molsheim, France). After blocking, membranes were labeled at 4 °C with primary antibodies: SLC7A11 (ab216876, 1:1000, Abcam), GPX4 (ab125066, 1:1000, Abcam), ACSL4 (ab155282, 1:10,000, Abcam), TFR1 (ab214039, 1:1000, Abcam), DMRT3 (PA5-116,427, 1:2000, Invitrogen), and β-actin (ab213262, 1:10,000, Abcam) at 4 °C. The next day, the bands were detected by ECL detection system (GE Healthcare, Piscataway, NJ, USA) after incubation with secondary antibody at 37 °C for 2 h.
Real-Time Quantitative Polymerase Chain Reaction (RT-qPCR)
Based on Trizol reagent (Invitrogen), total RNAs from clinical samples and cells were prepared, followed by quantification with the NanoDrop 2000 system. Subsequently, total RNAs were reversely transcribed into cDNA with Prime Script RT Master Mix (Takara, Tokyo, Japan). On the ABI 7500 fast PCR System (Applied Biosystems, Foster City, CA, USA), amplification reaction was implemented according to SYBR Green SuperMix (Roche, Basel, Switzerland). The amplification parameters were: denaturation at 95 °C for 10 min, followed by 40 cycles of denaturation at 95 °C for 30 s, annealing at 60 °C for 30 s and extension at 72 °C for 1 min. Lastly, relative expression from each group was subjected to β-actin normalization and 2–ΔΔCt method analysis. Primers used are exhibited in Table 1.
Cell Transfection
For DMRT3 or SLC7A11 overexpression, the sequences of DMRT3 (NM_021240.4) or SLC7A11 (NM_014331.4) were inserted into pcDNA vector to construct pcDNA- DMRT3 or SLC7A11 (termed DMRT3 or SLC7A11). Then, 6 μg of pcDNA vector specific to DMRT3 or SLC7A11, and pcDNA empty vector (pcDNA) were transfected into LK-2 and H1650 cells according to Lipofectamine 3000 (Invitrogen) for 48 h.
Chromatin Immunoprecipitation (ChIP)
In this assay, SimpleChIP ® Enzymatic Chromatin IP Kit (Cell Signaling Technology, Danvers, MA, USA) was applied to confirm the binding between DMRT3 and SLC7A11. After being fixed in 1% formaldehyde and lysed using lysis buffer from the kit, tumor cells were centrifuged (1000 × g for 5 min at 4 °C) for nuclei pellets. After DNA digestion, the obtained chromatin was added with anti-DMRT3 and anti-IgG and magnetic beads. 16 later, these products were purified and analyzed using RT-qPCR.
Dual-Luciferase Reporter Assay
Furthermore, in order to analyze SLC7A11 promoter activity, we synthesized with SLC7A11 promoter region (-409 to -399 bp) sequences possessing the wild-type (wt) or mutant-type (mut) DMRT3 binding sites (RiboBio), followed by introduction into pmirGLO vector (Promega, Madison, WI, USA). After that, LK-2 and H1650 cells were co-transfected with the acquired SLC7A11-wt/mut vectors and pcDNA or DMRT3 for 48 h. Finally, the luciferase activities in LK-2 and H1650 cell lysates were assessed based on Dual Luciferase Assay Kit (Promega).
Tumor Xenograft Assay
In this animal experiment, male BALB/C (nu/nu) nude mice (4 week-old, Slaike Jingda Laboratory, Hunan, China) under a temperature of 25 °C and relative air humidity between 45 and 50% were randomly assigned to three groups (n = 5 each group): 1) mice were subcutaneously injected with LK-2 cells carrying pcDNA (4 × 106/0.2 mL PBS) and administered 50 mg/kg corn oil (control) orally daily; 2) mice were injected with LK-2 cells carrying pcDNA (4 × 106/0.2 mL PBS) and administered 50 mg/kg curcumin (dissolved in corn oil) orally daily; 3) mice were injected with LK-2 cells carrying DMRT3 (4 × 106/0.2 mL PBS) and given 50 mg/kg curcumin orally. All surgeries were performed in a sterile environment. All mice were allowed free access to drinking water and sterilized standard diet. During tumor growth, the size was measured once a week and calculated. Four weeks later, all mice were euthanized by cervical dislocation under anesthesia, and the dissected tumors were subjected to weight, western blot, and Immunohistochemical (IHC) staining analysis. Besides, all animal processes involving this assay were approved by the Animal Ethics Committee of the Changzhou Cancer Hospital and in accordance with NIH guide for the care and use of Laboratory animals (NIH publications No.85–23, revised 2011).
Statistical Analysis
In this study, significance was determined based on Student’s t-test for two groups and one-way ANOVA with Tukey’s tests for multiple groups at a significance level of P < 0.05. Expression association was analyzed using Pearson correlation analysis. The results were analyzed using GraphPad Prism7 and exhibited as the mean ± standard deviation (SD).
Results
Curcumin Treatment Repressed NSCLC Malignant Behaviors In vitro
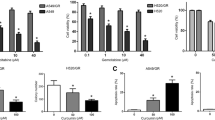
First of all, to check the biological function of curcumin on NSCLC, cell viability was evaluated under curcumin treatment (10 μM, 20 μM, and 40 μM) for 24 h. As shown in Fig. 1A, curcumin exposure did not change the viability of 16HBE cells, but it decreased the viability of NSCLC cells (LK-2 and H1650) in a concentration-dependent way. However, there were no significant changes observed in cell viability inhibition between the groups of LK-2 and H1650 cells treated with 20 μM or 40 μM curcumin. Thus, 20 μM was selected and used for the following study. After that, EdU assay displayed that curcumin addition might apparently hinder NSCLC cell proliferation in vitro relative to the control group (Fig. 1B). Beyond that, flow cytometry analysis exhibited that NSCLC cell apoptosis rate was clearly induced after curcumin exposure compared with the control group (Fig. 1C). Apart from that, the tube formation ability of HUVEC cells was also reduced by the treatment with the preconditioned medium of the LK-2 and H1650 cells exposed to curcumin (Fig. 1D). Lipid peroxidation, GSH depletion, and the accumulation of iron have been reported as the key events in ferroptosis. In order to explore whether curcumin might trigger ferroptosis in vitro, we first assessed the levels of the general oxidative stress indicators (SOD, MDA, GSH, and LDH) levels. As displayed in Fig. 1E–H, curcumin treatment elicited an obvious decline in SOD and GSH levels and a substantial enhancement in MDA and LDH levels. Simultaneously, our data also found that applying curcumin might greatly improve the Fe+ level of LK-2 and H1650 (Fig. 1I). Then, the biomarkers of ferroptosis were determined by western blot assay. Results presented that the protein levels of SLC7A11 and GPX4 were remarkably downregulated, and ACSL4 and TFR1 protein levels were evidently enhanced after curcumin treatment (Fig. 1J and k). Overall, these data suggested that curcumin treatment restrained NSCLC cell proliferation, apoptosis, and ferroptosis in vitro.
Effects of curcumin on NSCLC cell proliferation, apoptosis, angiopoiesis, and ferroptosis. A CCK-8 assay was used to analyze cell viability in 16HBE, LK-2, and H1650 cells treated with Curcumin (10 μM, 20 μM, and 40 μM) or 0 μM (control) for 24 h. (B-K) LK-2 and H1650 cells were treated with 0 μM Curcumin (control) or 20 μM Curcumin for 24 h. B Cell proliferation was detected using EdU assay in treated LK-2 and H1650 cells. C Apoptosis rate was analyzed in treated LK-2 and H1650 cells using flow cytometry assay. D The tube formation of HUVECs was detected by treatment with the preconditioned medium of LK-2 and H1650 cells treated with control or Curcumin. E–H Special assay kits identified products of SOD, MDA, GSH, and LDH. I Fe+ level in LK-2 and H1650 cells was detected using Iron assay kit. J and K Western blot assay was performed to determine the protein levels of SLC7A11, GPX4, ACSL4, and TFR1 in treated LK-2 and H1650 cells. *P < 0.05
Upregulation of DMRT3 Partly Abolished the Repression of Curcumin Exposure on the Malignancy of NSCLC Cells
A recent study has discovered that DMRT3 expression was aberrantly upregulated in NSCLC tissues and cells [23]. In order to identify the differential expression of DMRT3 in NSCLC, its expression was first detected in 33 paired NSCLC tissues and adjacent normal tissues. As shown in Fig. 2A and, b, DMRT3 expression was clearly increased in NSCLC samples. Moreover, we further validated that DMRT3 was highly expressed in NSCLC cell lines in comparison with 16HBE cells (Fig. 2C and) d). First of all, the overexpression efficiency of DMRT3 in LK-2 and H1650 cells was detected and shown in Figure S1A. Simultaneously, DMRT3 expression was apparently increased in pcDNA-DMRT3-transfected LK-2 and H1650 cells, which were partially overturned by curcumin treatment (Figure S1A). Subsequently, loss-of-function analyses were implemented to investigate the effects of DMRT3 and curcumin in NSCLC. At first, our data noticed that the mRNA level and protein level of DMRT3 were significantly suppressed by curcumin treatment, which was partially overturned after pcDNA-DMRT3 introduction (Fig. 2E and) F). Functionally, reduced cell viability and proliferation caused by curcumin exposure were effectively abrogated by DMRT3 upregulation (Fig. 2G and H). Moreover, flow cytometry assay presented that the forced expression of DMRT3 could strikingly mitigate the positive effect of curcumin on apoptosis rate in LK-2 and H1650 cells (Fig. 2I). In addition, culture with curcumin CM apparently inhibited HUVEC tube formation capacity, while these impacts were reversed by DMRT3 upregulation (Fig. 2J). In terms of ferroptosis, curcumin treatment-mediated SOD and GSH inhibition, and MDA and LDH increase in LK-2 and H1650 cells were significantly abolished after DMRT3 overexpression (Fig. 3A–D). In parallel, curcumin treatment-induced Fe+ level enhancement was effectively reversed after pcDNA-DMRT3 transfection (Fig. 3E). Meanwhile, the expression level of ferroptosis-markers such as ACSL4, SLC7A11, GPX4 and TFR1 was determined using western blot. Results showed that the upregulation of DMRT3 might drastically abrogated curcumin-triggered decrease in SLC7A11 and GPX4 protein levels and an increase in ACSL4 and TFR1 protein levels in LK-2 and H1650 cells (Fig. 3F and G). Collectively, these data illuminated that curcumin treatment might block NSCLC cell proliferation, apoptosis, and ferroptosis via regulating DMRT3.
Effects of DMRT3 on proliferation, apoptosis, and angiopoiesis in curcumin-treated NSCLC cells. A and B The mRNA level and protein level of DMRT3 were detected in 33 NSCLC tissues and 33 normal tissues using RT-qPCR and western blot. C and D RT-qPCR and western blot analysis of DMRT3 content in 16HBE, LK-2, and H1650 cells. E–J LK-2 and H1650 cells were treated with control, Curcumin, Curcumin + pcDNA, or Curcumin + DMRT3. E and F DMRT3 content was analyzed in treated LK-2 and H1650 cells using RT-qPCR and western blot. G CCK-8 analysis of cell viability in treated LK-2 and H1650. H EdU analysis of cell proliferation using EdU assay in treated LK-2 and H1650 cells. I Flow cytometry analysis of cell apoptosis rate treated LK-2 and H1650 cells. J We collected pre-conditioned medium from treated LK-2 and H1650 cells, cultured HUVECs in these two conditioned medium (CM) samples, and measured for the tube formation capacity. *P < 0.05
Effects of DMRT3 on ferroptosis in curcumin-treated NSCLC cells. LK-2 and H1650 cells were treated with control, Curcumin, Curcumin + pcDNA, or Curcumin + DMRT3. A–D SOD, MDA, GSH, and LDH levels were examined using Corresponding kits. E Fe+ level in LK-2 and H1650 cells was assessed using Iron assay kit. F and G SLC7A11, GPX4, ACSL4, and TFR1 protein levels were measured using Western blot. *P < 0.05
SLC7A11 Expression was Upregulated in Curcumin-Treated NSCLC Cells
Interestingly, our data identified that the mRNA level and protein level of SLC7A11, a key suppressor of ferroptosis, were significantly upregulated in NSCLC tissues relative to normal tissues (Fig. 4A and B). Furthermore, we found that the SLC7A11 level had a positive correlation with DMRT3 expression in NSCLC tissues (Fig. 4C). Beyond that, RT-qPCR and western blot also confirmed that SLC7A11 content was prominently higher in NSCLC cell lines (LK-2 and H1650) than in 16HBE cells (Fig. 4D and E). Besides, the significant downregulation of SLC7A11 was observed in curcumin-treated LK-2 and H1650 cells versus the control groups (Fig. 4F and G). In total, these data indicated the involvement of DMRT3 and SLC7A11 in the NSCLC process.
Effects of SLC7A11 on curcumin-treated NSCLC cells. A and B RT-qPCR and western blot analysis of SLC7A11 content in 33 NSCLC tissues and 33 normal tissues. C Pearson correlation analysis was applied to evaluate the expression association between DMRT3 and SLC7A11 in NSCLC tissues. D and E SLC7A11 mRNA level and protein level were determined using RT-qPCR and western blot assay in 16HBE, LK-2, and H1650 cells. F and G RT-qPCR and western blot analysis of SLC7A11 content in LK-2 and H1650 cells treated with control or Curcumin. *P < 0.05
DMRT3 Directly Bound the Promoter of SLC7A11
To better map the interaction between DMRT3 and SLC7A11, JASPAR database was applied, and a DMRT3 motif was observed in the − 409 to − 399 promoter region of SLC7A11 (Fig. 5A). Subsequently, to further validate whether there exists a targeted regulation between DMRT3 and SLC7A11 in LK-2 and H1650 cells, ChIP experiment was carried out. Data exhibited that DMRT3 bound to the region sites of the SLC7A11 promoter (Fig. 5B). Furthermore, a dual-luciferase reporter assay presented that DMRT3 overexpression might obviously enhance the luciferase activity of SLC7A11 reporter systems in LK-2 and H1650 cells, rather than the mutant group (Fig. 5C and D), implying the promoting effect of DMRT3 for SLC7A11 promoter might be dependent on the DMRT3 motif. In addition, western blot analysis exhibited that the SLC7A11 protein level in LK-2 and H1650 cells was clearly constrained by DMRT3 depletion, and distinctly improved by DMRT3 overexpression (Fig. 5E). Together, these data illuminated that transcription factor DMRT3 might directly bind to the SLC7A11 promoter and promote its expression.
DMRT3 might activate the transcription of SLC7A11. A The JASPAR website predicted DMRT3 to possess binding sites with the SLC7A11 promoter sequence. B ChIP assay was performed to confirm the binding ability between DMRT3 and SLC7A11 promoter. C and D Their interaction was verified using a dual-luciferase reporter assay in LK-2 and H1650 cells. E SLC7A11 protein level was determined in LK-2 and H1650 cells transfected with si-NC, si-DMRT3, pcDNA, or DMRT3. *P < 0.05
Curcumin Treatment might Suppress the Malignancy of NSCLC Cells via Regulating SLC7A11
Subsequently, to clarify whether the inhibitory impact of curcumin treatment was mediated by the DMRT3/SLC7A11 axis, the influences of SLC7A11 on curcumin-mediated NSCLC progression. At first, the overexpression efficiency of SLC7A11 in LK-2 and H1650 cells was measured and presented in Figure S1B. Meanwhile, SLC7A11 mRNA level was obviously upregulated in pcDNA-SLC7A11-transfected LK-2 and H1650 cells, which were partly abolished by curcumin treatment (Figure S1B). Then, western blot analysis displayed that curcumin treatment might impede SLC7A11 protein level in LK-2 and H1650 cells, whereas these effects were partially overturned after pcDNA-SLC7A11 transfection (Fig. 6A). Functional analysis showed that the repression of curcumin treatment on NSCLC cell viability and proliferation was effectively ameliorated by SLC7A11 upregulation (Fig. 6B and C). Furthermore, the curcumin-induced apoptosis rate in LK-2 and H1650 cells was relieved by SLC7A11 overexpression (Fig. 6D). Furthermore, the suppressive effects of HUVEC tube formation after curcumin exposure were significantly reversed by SLC7A11 overexpression (Fig. 6E). In addition, curcumin treatment-mediated ferroptosis inhibition in LK-2 and H1650 cells was greatly ameliorated through SLC7A11 upregulation, accompanied by enhanced SOD and GSH levels (Fig. 6F and H), reduced MDA and LDH levels (Fig. 6G and I), and decreased Fe+ level (Fig. 6J). Consistently, western blot analysis showed that curcumin-mediated SLC7A11 and GPX4 protein expression inhibition, and ACSL4 and TFR1 protein expression promotion in LK-2 and H1650 cells were partly overturned by SLC7A11 overexpression (Fig. 6K and L). All of these results indicated that the upregulation of SLC7A11 might attenuate the repression of curcumin on NSCLC cell development.
Effects of curcumin and SLC7A11 on NSCLC cell development. LK-2 and H1650 cells were treated with 0 μM Curcumin (control), Curcumin, Curcumin + pcDNA, or Curcumin + SLC7A11. A Western blot analysis of SLC7A11 protein level in treated LK-2 and H1650 cells. B and C CCK-8 and EdU assays were performed to examine cell proliferative ability in treated LK-2 and H1650 cells. D Flow cytometry assay was conducted to assess LK-2 and H1650 cell apoptosis rate. E HUVECs were cultured in treated LK-2 and H1650 cell CM and assayed for tube formation capacity. F–I SOD, MDA, GSH, and LDH levels were determined using special assay kits. J Fe+ level in treated LK-2 and H1650 cells was monitored using Iron assay kit. K and L The protein levels of SLC7A11, GPX4, ACSL4, and TFR1 in treated LK-2 and H1650 cells were analyzed using western blot assay. *P < 0.05
DMRT3 Overexpression Mitigated the Repression of Curcumin on Xenograft Tumor Growth
Based on the above findings, a xenograft tumor mouse model was established to further validate that applying curcumin might regulate the malignancy of NSCLC cells through DMRT3. As shown in Fig. 7A and B, curcumin treatment significantly repressed tumor volume and weight, whereas pcDNA-DMRT3 might abolish the repression of curcumin on tumor growth. Moreover, western blot assay showed that DMRT3 and SLC7A11 protein levels were strikingly lower in tumor tissues from the curcumin + pcDNA group than the curcumin + DMRT3 group (Fig. 7C). Additionally, immunohistochemical staining uncovered that the curcumin + pcDNA group repressed DMRT3, SLC7A11, and Bcl2 positive expression, and promoted Bax positive expression relative to the curcumin + DMRT3 group (Fig. 7D). Accordingly, it is concluded that curcumin treatment inhibited xenograft tumor growth by regulating DMRT3.
Curcumin might dampen the tumorigenicity of LK-2 cells in a xenograft model by regulating DMRT3. LK-2 cells transfected with pcDNA or DMRT3 were inoculated subcutaneously into the nude mice. Then, these mice with tumor formation were given 50 mg/kg curcumin. A Growth curve of xenografted tumors was displayed. B Weight of resected tumor masses was presented. C Protein levels of DMRT3 and SLC7A11 in the xenografts were assessed using western blot. F DMRT3, SLC7A11, Bax, and Bcl2 expression was gauged in xenografts using Immunohistochemical staining. *P < 0.05
Discussion
In recent years, the benefits of curcumin in different organ systems have been widely described in several human cancers [28]. In fact, it has become evident that curcumin might hinder lung cancer growth by repressing cell proliferation, inducing apoptosis, and displaying less adverse effects and toxicity [29, 30]. Consistent with these studies, our data identified the anti-proliferation, pro-apoptosis, and anti-angiopoiesis effects of curcumin on NSCLC cell lines. Of note, many types of research have reported that cell death plays an important part in preventing and treating hyper-proliferative cancers [31, 32]. In terms of morphology, biochemistry, and genetics, ferroptosis is a new form of programmed cell death, different from necrosis, apoptosis, and autophagy [33]. In general, intracellular iron accumulation, ROS, GSH, and lipid peroxidation are known as the indispensable hallmarks of ferroptosis [34]. Emerging evidence has implicated that induced ferroptosis in NSCLC is responsible for the inhibition of tumor progression both in vitro and in vivo [35]. Interestingly, recent literature has exhibited that curcumin in NSCLC cells might induce ferroptosis by activating autophagy [17]. In agreement with these reports, applying curcumin might boost ferroptosis, which is useful in treating NSCLC.
Notably, some studies have indicated that curcumin and its analogs might mediate the anti-cancer effect in NSCLC through regulating various mechanisms [18, 36, 37], containing transcription factors [38]. Furthermore, it has been reported that DMRT3, a transcription factor, is an abnormal expression in pan-cancer and might facilitate tumor occurrence and development [39, 40], containing NSCLC [23]. Moreover, DMRT3 might partake in the TP63/SOX2 circuit for modulating squamous cell differentiation or survival, and these three factors might co-regulate gene functioning in augmenting NSCLC development and formation [24]. Consistent with these former works, our data demonstrated that DMRT3 content was clearly upregulated in NSCLC samples and cell lines. Of interest, our results found a significant downregulation of DMRT3 in curcumin-treated NSCLC cells for the first time. Apart from that, functional analysis verified that DMRT3 overexpression might partially abolish the effects of curcumin on NSCLC cell growth and ferroptosis in vitro. Consistently, our data validated that curcumin treatment might block tumor growth in vivo via regulating DMRT3 expression. These outcomes implied that the repression role of curcumin on NSCLC progression might be partly mediated by downregulating DMRT3.
It has been widely reported that transcription factors might modulate the transcription of target genes via binding to the specific sequences at the promoter region [41]. Herein, the current work identified the positive association between DMRT3 and SLC7A11. Moreover, our data presented that DMRT3 might improve the transcription of SLC7A11 via binding to its promoter region, indicating that SLC7A11 might be a novel DMRT3-target gene. Furthermore, SLC7A11 (also known as xCT or CCBR1), a component of the cysteine-glutamate transporter, might take part in different stages of human cancer development via regulating cell growth, cell death, and cell metabolism [42]. Beyond that, the upregulation of SLC7A11 might expedite the malignant behaviors and hinder ferroptosis of NSCLC cells [43, 44]. Consistent with the former study [17], our data verified that SLC7A11 content was clearly reduced in curcumin-treated NSCLC cell lines. In contrast to these previous studies, our results provide the first evidence that SLC7A11 upregulation might significantly mitigate the effects of curcumin on NSCLC cell growth and ferroptosis repression. The above findings further supported that the suppressive role of curcumin on NSCLC progression might be partly mediated by the DMRT3/SLC7A11 axis. Targeting the DMRT3/SLC7A11 axis might be an important focus in exploring effective anti-lung cancer therapies. The combination of curcumin with the DMRT3/SLC7A11 axis may further enhance its anti-lung cancer efficacy compared to curcumin alone.
Conclusion
In summary, these results elucidated compelling evidence that curcumin might repress cell growth and induce ferroptosis in NSCLC cells by targeting the DMRT3/SLC7A11 axis, providing a novel avenue of therapy for NSCLC treatment.
References
Siegel, R. L., Miller, K. D., Wagle, N. S., & Jemal, A. (2023). Cancer statistics, 2023. CA: A Cancer Journal for Clinicians, 73(1), 17–48. https://doi.org/10.3322/caac.21763
Leiter, A., Veluswamy, R. R., & Wisnivesky, J. P. (2023). The global burden of lung cancer: Current status and future trends. Nature Reviews. Clinical Oncology, 20(9), 624–639. https://doi.org/10.1038/s41571-023-00798-3
Sung, H., Ferlay, J., Siegel, R. L., Laversanne, M., Soerjomataram, I., Jemal, A., & Bray, F. (2021). Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA: A Cancer Journal for Clinicians, 71(3), 209–249. https://doi.org/10.3322/caac.21660
Rodak, O., Peris-Díaz, M. D., Olbromski, M., Podhorska-Okołów, M., & Dzięgiel, P. (2021). Current landscape of non-small cell lung cancer: epidemiology, histological classification, targeted therapies, and immunotherapy. Cancers (Basel). https://doi.org/10.3390/cancers13184705
Duma, N., Santana-Davila, R., & Molina, J. R. (2019). Non-small cell lung cancer: epidemiology, screening, diagnosis, and treatment. Mayo Clinic Proceedings, 94(8), 1623–1640. https://doi.org/10.1016/j.mayocp.2019.01.013
Reck, M., Heigener, D. F., Mok, T., Soria, J. C., & Rabe, K. F. (2013). Management of non-small-cell lung cancer: Recent developments. Lancet, 382(9893), 709–719. https://doi.org/10.1016/s0140-6736(13)61502-0
Imran, M., Ullah, A., Saeed, F., Nadeem, M., Arshad, M. U., & Suleria, H. A. R. (2018). Cucurmin, anticancer, & antitumor perspectives: A comprehensive review. Critical Reviews in Food Science and Nutrition, 58(8), 1271–1293. https://doi.org/10.1080/10408398.2016.1252711
Kotha, R. R., & Luthria, D. L. (2019). Curcumin: Biological, Pharmaceutical, Nutraceutical, and Analytical Aspects. Molecules. https://doi.org/10.3390/molecules24162930
Kocaadam, B., & Şanlier, N. (2017). Curcumin, an active component of turmeric (Curcuma longa), and its effects on health. Critical Reviews in Food Science and Nutrition, 57(13), 2889–2895. https://doi.org/10.1080/10408398.2015.1077195
Górnicka, J., Mika, M., Wróblewska, O., Siudem, P., & Paradowska, K. (2023). Methods to improve the solubility of curcumin from turmeric. Life (Basel). https://doi.org/10.3390/life13010207
Luis, P. B., Kunihiro, A. G., Funk, J. L., & Schneider, C. (2020). Incomplete hydrolysis of curcumin conjugates by β-glucuronidase: Detection of complex conjugates in plasma. Molecular Nutrition & Food Research, 64(6), e1901037. https://doi.org/10.1002/mnfr.201901037
Wang, R., Han, J., Jiang, A., Huang, R., Fu, T., Wang, L., Zheng, Q., Li, W., & Li, J. (2019). Involvement of metabolism-permeability in enhancing the oral bioavailability of curcumin in excipient-free solid dispersions co-formed with piperine. International Journal of Pharmaceutics. https://doi.org/10.1016/j.ijpharm.2019.02.027
Soleimani, V., Sahebkar, A., & Hosseinzadeh, H. (2018). Turmeric (Curcuma longa) and its major constituent (curcumin) as nontoxic and safe substances: Review. Phytotherapy Research, 32(6), 985–995. https://doi.org/10.1002/ptr.6054
Ming, T., Tao, Q., Tang, S., Zhao, H., Yang, H., Liu, M., Ren, S., & Xu, H. (2022). Curcumin: An epigenetic regulator and its application in cancer. Biomedicine & Pharmacotherapy. https://doi.org/10.1016/j.biopha.2022.113956
Giordano, A., & Tommonaro, G. (2019). Curcumin and cancer. Nutrients. https://doi.org/10.3390/nu11102376
Mehta, H. J., Patel, V., & Sadikot, R. T. (2014). Curcumin and lung cancer—a review. Targeted Oncology, 9(4), 295–310. https://doi.org/10.1007/s11523-014-0321-1
Tang, X., Ding, H., Liang, M., Chen, X., Yan, Y., Wan, N., Chen, Q., Zhang, J., & Cao, J. (2021). Curcumin induces ferroptosis in non-small-cell lung cancer via activating autophagy. Thoracic Cancer, 12(8), 1219–1230. https://doi.org/10.1111/1759-7714.13904
Tang, C., Liu, J., Yang, C., Ma, J., Chen, X., Liu, D., Zhou, Y., Zhou, W., Lin, Y., & Yuan, X. (2022). Curcumin and its analogs in non-small cell lung cancer treatment: Challenges and expectations. Biomolecules. https://doi.org/10.3390/biom12111636
Zhang, T., & Zarkower, D. (2017). DMRT proteins and coordination of mammalian spermatogenesis. Stem Cell Research. https://doi.org/10.1016/j.scr.2017.07.026
Li, H. L., Zou, Z. C., Fang, C., Zheng, Y. P., Guo, X. M., & Yang, W. H. (2023). Mammalian DMRTs: Structure, function and relationship with cancer. Sheng Li Xue Bao, 75(2), 269–278.
Bellefroid, E. J., Leclère, L., Saulnier, A., Keruzore, M., Sirakov, M., Vervoort, M., & De Clercq, S. (2013). Expanding roles for the evolutionarily conserved Dmrt sex transcriptional regulators during embryogenesis. Cellular and Molecular Life Sciences, 70(20), 3829–3845. https://doi.org/10.1007/s00018-013-1288-2
Kopp, A. (2012). Dmrt genes in the development and evolution of sexual dimorphism. Trends in Genetics, 28(4), 175–184. https://doi.org/10.1016/j.tig.2012.02.002
Yang, D., Liu, M., Jiang, J., Luo, Y., Wang, Y., Chen, H., Li, D., Wang, D., Yang, Z., & Chen, H. (2022). Comprehensive analysis of DMRT3 as a potential biomarker associated with the immune infiltration in a pan-cancer analysis and validation in lung adenocarcinoma. Cancers (Basel). https://doi.org/10.3390/cancers14246220
Zhang, S., Li, M., Ji, H., & Fang, Z. (2018). Landscape of transcriptional deregulation in lung cancer. BMC Genomics, 19(1), 435. https://doi.org/10.1186/s12864-018-4828-1
Ma, L., Chen, T., Zhang, X., Miao, Y., Tian, X., Yu, K., Xu, X., Niu, Y., Guo, S., Zhang, C., Qiu, S., Qiao, Y., Fang, W., Du, L., Yu, Y., & Wang, J. (2021). The m(6)A reader YTHDC2 inhibits lung adenocarcinoma tumorigenesis by suppressing SLC7A11-dependent antioxidant function. Redox Biology. https://doi.org/10.1016/j.redox.2020.101801
He, F., Zhang, P., Liu, J., Wang, R., Kaufman, R. J., Yaden, B. C., & Karin, M. (2023). ATF4 suppresses hepatocarcinogenesis by inducing SLC7A11 (xCT) to block stress-related ferroptosis. Journal of Hepatology, 79(2), 362–377. https://doi.org/10.1016/j.jhep.2023.03.016
Zhang, W., Sun, Y., Bai, L., Zhi, L., Yang, Y., Zhao, Q., Chen, C., Qi, Y., Gao, W., He, W., Wang, L., Chen, D., Fan, S., Chen, H., Piao, H. L., Qiao, Q., Xu, Z., Zhang, J., Zhao, J., … Wang, Y. (2021). RBMS1 regulates lung cancer ferroptosis through translational control of SLC7A11. Journal of Clinical Investigation. https://doi.org/10.1172/jci152067
Shah, M., Murad, W., Mubin, S., Ullah, O., Rehman, N. U., & Rahman, M. H. (2022). Multiple health benefits of curcumin and its therapeutic potential. Environmental Science and Pollution Research International, 29(29), 43732–43744. https://doi.org/10.1007/s11356-022-20137-w
Ashrafizadeh, M., Najafi, M., Makvandi, P., Zarrabi, A., Farkhondeh, T., & Samarghandian, S. (2020). Versatile role of curcumin and its derivatives in lung cancer therapy. Journal of Cellular Physiology, 235(12), 9241–9268. https://doi.org/10.1002/jcp.29819
Salehi, M., Movahedpour, A., Tayarani, A., Shabaninejad, Z., Pourhanifeh, M. H., Mortezapour, E., Nickdasti, A., Mottaghi, R., Davoodabadi, A., Khan, H., Savardashtaki, A., & Mirzaei, H. (2020). Therapeutic potentials of curcumin in the treatment of non-small-cell lung carcinoma. Phytotherapy Research, 34(10), 2557–2576. https://doi.org/10.1002/ptr.6704
Tong, X., Tang, R., Xiao, M., Xu, J., Wang, W., Zhang, B., Liu, J., Yu, X., & Shi, S. (2022). Targeting cell death pathways for cancer therapy: Recent developments in necroptosis, pyroptosis, ferroptosis, and cuproptosis research. Journal of Hematology & Oncology, 15(1), 174. https://doi.org/10.1186/s13045-022-01392-3
Strasser, A., & Vaux, D. L. (2020). Cell death in the origin and treatment of cancer. Molecular Cell, 78(6), 1045–1054. https://doi.org/10.1016/j.molcel.2020.05.014
Yan, H. F., Zou, T., Tuo, Q. Z., Xu, S., Li, H., Belaidi, A. A., & Lei, P. (2021). Ferroptosis: Mechanisms and links with diseases. Signal Transduction and Targeted Therapy, 6(1), 49. https://doi.org/10.1038/s41392-020-00428-9
Rochette, L., Dogon, G., Rigal, E., Zeller, M., Cottin, Y., & Vergely, C. (2022). Lipid peroxidation and iron metabolism: Two corner stones in the homeostasis control of ferroptosis. International Journal of Molecular Sciences. https://doi.org/10.3390/ijms24010449
Zou, J., Wang, L., Tang, H., Liu, X., Peng, F., & Peng, C. (2021). Ferroptosis in non-small cell lung cancer: Progression and therapeutic potential on it. International Journal of Molecular Sciences. https://doi.org/10.3390/ijms222413335
Xu, X., Zhang, X., Zhang, Y., & Wang, Z. (2021). Curcumin suppresses the malignancy of non-small cell lung cancer by modulating the circ-PRKCA/miR-384/ITGB1 pathway. Biomedicine & Pharmacotherapy. https://doi.org/10.1016/j.biopha.2021.111439
Wang, J. Y., Wang, X., Wang, X. J., Zheng, B. Z., Wang, Y., Wang, X., & Liang, B. (2018). Curcumin inhibits the growth via Wnt/β-catenin pathway in non-small-cell lung cancer cells. European Review for Medical and Pharmacological Sciences, 22(21), 7492–7499. https://doi.org/10.26355/eurrev_201811_16290
Xu, X., & Zhu, Y. (2017). Curcumin inhibits human non-small cell lung cancer xenografts by targeting STAT3 pathway. American Journal of Translational Research, 9(8), 3633–3641.
Katsuta, E., Huyser, M., Yan, L., & Takabe, K. (2021). A prognostic score based on long-term survivor unique transcriptomic signatures predicts patient survival in pancreatic ductal adenocarcinoma. American Journal of Cancer Research, 11(9), 4294–4307.
Meng, Y., Li, S., Gu, D., Xu, K., Du, M., Zhu, L., Chu, H., Zhang, Z., Wu, Y., Fu, Z., & Wang, M. (2020). Genetic variants in m6A modification genes are associated with colorectal cancer risk. Carcinogenesis, 41(1), 8–17. https://doi.org/10.1093/carcin/bgz165
Lambert, S. A., Jolma, A., Campitelli, L. F., Das, P. K., Yin, Y., Albu, M., Chen, X., Taipale, J., Hughes, T. R., & Weirauch, M. T. (2018). The Human Transcription Factors. Cell, 172(4), 650–665. https://doi.org/10.1016/j.cell.2018.01.029
Li, S., Lu, Z., Sun, R., Guo, S., Gao, F., Cao, B., & Aa, J. (2022). The role of SLC7A11 in Cancer: Friend or Foe? Cancers (Basel). https://doi.org/10.3390/cancers14133059
Lu, X., Kang, N., Ling, X., Pan, M., Du, W., & Gao, S. (2021). MiR-27a-3p promotes non-small cell lung cancer through SLC7A11-mediated-ferroptosis. Frontiers in Oncology. https://doi.org/10.3389/fonc.2021.759346
Xu, Y., Lv, D., Yan, C., Su, H., Zhang, X., Shi, Y., & Ying, K. (2022). METTL3 promotes lung adenocarcinoma tumor growth and inhibits ferroptosis by stabilizing SLC7A11 m(6)A modification. Cancer Cell International, 22(1), 11. https://doi.org/10.1186/s12935-021-02433-6
Acknowledgements
None.
Funding
None.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12033_2024_1166_MOESM1_ESM.tif
Supplementary file1 (TIF 220 KB)—Figure S1 DMRT3 and SLC7A11 expression were detected in NSCLC cells. (A) RT-qPCR assay was used to measure DMRT3 mRNA level in LK-2 and H1650 cells treated with pcDNA, DMRT3, Curcumin + pcDNA, or Curcumin + DMRT3. (B) SLC7A11 mRNA level was assessed in LK-2 and H1650 cells treated with pcDNA, SLC7A11, Curcumin + pcDNA, or Curcumin + SLC7A11 using RT-qPCR assay. *P < 0.05
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, B., Zhou, L. & Zhang, Q. Curcumin Inhibits the Progression of Non-small Cell Lung Cancer by Regulating DMRT3/SLC7A11 Axis. Mol Biotechnol (2024). https://doi.org/10.1007/s12033-024-01166-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12033-024-01166-x