Abstract
Booster doses are crucial against severe COVID-19, as rapid virus mutations and variant emergence prolong the pandemic crisis. The virus’s quick evolution, short generation-time, and adaptive changes impact virulence and evolvability, helping predictions about variant of concerns’ (VOCs’) landscapes. Here, in this study, we used a new computational algorithm, to predict the mutational pattern in SARS-CoV-2 ssRNA, proteomics, structural identification, mutation stability, and functional correlation, as well as immune escape mechanisms. Interestingly, the sequence diversity of SARS Coronavirus-2 has demonstrated a predominance of G- > A and C- > U substitutions. The best validation statistics are explored here in seven homologous models of the expected mutant SARS-CoV-2 spike ssRNA and employed for hACE2 and IgG interactions. The interactome profile of SARS-CoV-2 spike with hACE2 and IgG revealed a strong correlation between phylogeny and divergence time. The systematic adaptation of SARS-CoV-2 spike ssRNA influences infectivity and immune escape. Data suggest higher propensity of Adenine rich sequence promotes MHC system avoidance, preferred by A-rich codons. Phylogenetic data revealed the evolution of SARS-CoV-2 lineages’ epidemiology. Our findings may unveil processes governing the genesis of immune-resistant variants, prompting a critical reassessment of the coronavirus mutation rate and exploration of hypotheses beyond mechanical aspects.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coronaviruses (CoVs) were first reported in humans in 1962 [1] and belong to the Order Nidovirales, Family Coronaviridae, Subfamily Orthocoronavirinae, Genus Betacoronavirus, Subgenus Sarbecovirus, Species severe acute respiratory syndrome-related coronavirus, which has a single-stranded RNA (ssRNA) betacoronavirus [2]. This virus could undergo rapid mutations and transmit it within the human population; however, the transmission rate may vary depending upon the vaccination strategy and genetic makeup. At the time of writing, more than 12 strains have already been reported by WHO and CDC [3, 4]. The SARS-CoV-2 surface spike glycoprotein binds with human angiotensin converting enzyme 2 (ACE2) receptors [5]. After anchoring, human TMPRSS2 cleaves and activates spike protein accordingly, to allow SARS-CoV-2 entry, which initiates endocytosis or direct fusion of the viral envelope with the host membrane [6,7,8,9]. The positive-sense single-stranded RNA (ssRNA) genome of SARS-CoV-2 is approximately 30 kb with 14 open reading frames (ORFs), some of them overlapping in nature [10]. The n5′ cap and a 3′-poly(A) tail of SARS-CoV-2 genome serve as an mRNA for translation to produce viral poly-proteins. Highly organized untranslated region; the ssRNA 5′ and 3′ ends contain a (UTR) that is critical in the control of RNA replication and transcription. Seven stem-loop structures in 5′ UTR, while the 3′ UTR contains a stem-loop and a pseudo-knot. The pseudo-knot or stem-loop is thought to have a function in transcriptional control [11]. There are a total of 16 non-structural proteins (NSPs) identified, including ORF1a-encoded proteases nsp5 (chymotrypsin-like protease, 3CLpro or Mpro) and papain-like protease (PLpro) found in subunit nsp3 [10, 12]. The remaining one-third of the genome comprises viral structural protein genes shared by all CoVs (spike, envelope, membrane, and nucleocapsid). Substitution mutation occurs in specified location and codon change from G to A or C to T in the 65% of viral genome. There are ≈100,000 possible single nucleotide substitutions in the SARS-CoV-2 genome [13, 14]. Mutation through natural selection results in evolution. However, most mutations are not beneficial for the organisms with them but may be favorable to the host [15, 16]. It was also reported that interactions between viral and host proteins, including stem loop structures, regulate virus replication and translation. Lowering the number of stem-loop structures in the positive strands of ssRNA viruses enhances RNA expression [17, 18]. The viral replication and transcription are dependent on different factors in the host cell [19]. Nucleotide changes in spike ssRNA that allow evasion of selective pressure reduce antibody and antiviral drug efficacy. SARS CoV 2 spike protein mutations are important for triggering attachment with ACE2 and may result in immunological evasion with IgG [20, 21]. The aim of this study is to predict the structural determination of the SARS-CoV-2 spike protein based on the probabilistic mutation in spike RNA. The mutational impact on the diversification of the stem-looping secondary structure of the viral ssRNA was explored. Further, spike designated ssRNA structural variation has been linked to the pattern of interactions between the viral-spike proteins with ACE2 and human IgG that may reveal the replication and infection mechanisms within the human in the host. We believe that our study will provide a comprehensive understanding of the structural determinants of the SARS-CoV-2 spike protein, offering insights into both the genetic variability of the virus and its infectibility implications. The ultimate goal is to contribute valuable knowledge that can inform public health strategies, vaccine development, and the ongoing battle against the COVID-19 pandemic.
Materials and Methods
Profiling of Mutational Landscape, Nucleotide Statistics, and Substitution Frequency
Furthermore, databases such as GISAID (https://www.gisaid.org), which elucidates its genetic properties, and Nextstrain (https://nextstrain.org), which tracks its mutational profile in the global population, are used. GISAID database represented a total of 7,375,845 SARS-CoV-2 genome sequence entries from patient samples, accessed by the end of January 2022.
We counted the number of nucleotides and their corresponding frequencies of SARS-CoV-2 spike ssRNA sequence. The substitution patterns and rates were estimated using the Tamura 1992 parameter model [22] with MEGA-X [23].
SARS-CoV-2 ssRNA Data Set Curation
Data sets of SARS-CoV-2 spike ssRNA Sequence were primarily obtained from NCBI Genbank Database. Accession number of SARS-CoV-2 (Wuhan): MT079854.1, Alpha: MZ314997, Beta: MZ314998.1, Eta: MZ362451.1, Delta: OK091006.1, Gamma: MZ315141.1, Omicron: OL672836.1.
Prediction and Visualization of the RNA Secondary Structures
The RNA secondary structures of the viral genomic sequences were predicted using the online RNAfold web server at http://rna.tbi.univie.ac.at/cgi-bin/RNAWebSuite/RNAfold.cgi [24]. Predicted RNA secondary structures were then visualized as structural diagram utilizing FORNA server at http://rna.tbi.univie.ac.at/forna [25].
Prediction of Expected Variants of SARS CoV 2
We had significantly characterized predominant mutations occurring in the loop region. Probable mutational hotspots into the different size of the loop regions were identified and edited (G- > A and C- > U substitutions) E.M.1 (9 ± 2), E.M.2 (13 ± 2), E.M.3 (19 ± 3), E.M.4 (27 ± 3), and E.M.5 (39 ± 3) accordingly. The C program was utilized to create different mutant sequences based on the provided probabilistic sequence data sets (Supplementary Fig. 8). Function names like diffchar and sdam were used for this purpose. The scripts were written using pointers. The sample set was specified as a S1 to Sn pointer. nC1 + nC2 + nC3 + …. + nCn sequence sets were generated as expected model (E.M.) to eliminate any cross-platform issues, along with better compatibility all code compiles without all warnings enabled using the C compiler (Fig. 1A). The code has been tested in Windows environment.
ssRNA Structural Stability
Predicted mutant ssRNA sequences were validated based on the thermodynamics and frequency of the MFE structure and diversity [24]. The finding indicated that significant MFE values across the genome, provided the initial evidence for the occurrence of RNA structure formation in SARS-CoV-2 and other coronavirus genomes. MFE calculations reveal the sequence order contribution to RNA folding, with higher values resulting from native sequence folding energies being better than shuffled controls.
Protein Structure Modeling
Suitable mutant sequences of the different spike protein of SARS-COV-2 were selected and the structures were modeled using Swiss-model interactive modeling (https://swissmodel.expasy.org/, accessed February, 2022). The high-quality models were created using the crystal structure of its C-terminal dimerization domain (PDB id: 6VSB). The 3D models were then improved using 3Drefine (http://sysbio.rnet.missouri.edu/3Drefine/) and online protein structure refinement servers in a two-step procedure [25].
The Mutational Impact and Protein–Protein Interaction Analysis of the Modeled Structures
All of the modeled spike protein structures were chosen for docking after being evaluated by Mol Probity Clash, Ramachandran Favoured, QMEANDisCo Global, and QMEAN score. HADDOCK 2.4 was used to perform protein–protein docking of the S (Spike) protein with angiotensin-converting enzyme 2 (PDB id: 6M18) and the heavy and light chains of human IgG (PDB id: 6YOR). To anticipate binding affinities and establish biological interfaces, we accessed the PRODIGY web server (Prodigy Webserver (uu.nl). The structure of molecular interactions between spike proteins and ACE2 and IgG was visualized and analyzed using PyMOL2 [26,27,28].
Surface-Topology Calculation of Proteins
Solvent accessibility factor defines two properties of protein; pocket, where water enters and cavity, where does not. The CASTp: Computed-Atlas-of-Surface-Topography of Protein (http://sts.bioe.uic.edu/castp/index.html?j_5e8c7bec25090) was used to define pocket and cavity.
Evaluation of H-bonding in Docking Structure
Observation of the H-bonding length of amino acid residues between the ACE2 receptor and human IgG of original SARS-CoV-2 isolate Wuhan-Hu-1 (WH1), the D614G mutant, the N501Y mutant, the B.1.617.2 (Delta), and omicron variant were analyzed by the PyMol visualized system to recognize the H-bond length between amino acid residues short or long. By using PyMol software, the result of the free energy network of the stable protein–protein complex was analyzed.
Energetic and Kinetic Data Analysis
The HADDOCK web server was used to generate the protein–protein binding score, cluster-size based RMSD values and other bond energy values, i.e., Van-der-Waals, Electrostatic Desolvation energy were evaluated. Binding affinity [ΔG (kcal mol-1)] Dissociation constant [Kd (M) at 25 °C] explained the strength and the affinity of the interactions.
SARS-CoV-2 Spike Protein Mutational Saturation, Divergence, and Age Estimation
A timetree was constructed by applying the RelTime technique [29, 30] to a user-supplied phylogenetic tree with branch lengths estimated using the Maximum Likelihood (ML) method, utilizing the Tamura-Nei substitution model [30]. Two calibration constraints were used to build the timetree. To define minimum and maximum time constraints for nodes for which calibration densities were supplied. This [31] approach was used to construct confidence intervals. Times were not calculated for out-group nodes because the Reltime technique calculates divergence times using evolutionary rates from the in-group and does not presume that evolutionary rates from the in-group of the clade apply to the out-group. The tree’s estimated log probability value is 6680.52. To describe evolutionary rate variations among sites in 5 categories (+ G, parameter = 0.0500). According to the rate variation hypothesis, some sites [(+ I), 0.00% of sites] were evolutionarily invariable. There were 14 nucleotide sequences in this study. 1st + 2nd + 3rd + noncoding codon locations were included. The total number of places in the final dataset was 4083. MEGA11 [32] was used to execute evolutionary studies.
Results
Phylogenetic Clusters Correlated with Mutational Pattern
We assessed 7,375,845 SARS-CoV-2 genomic sequences uploaded to the GISAID database in January 2022. We also assessed the lineage diversity of SARS-CoV-2 variants through time using phylodynamic time tree analysis. In the SARS-CoV-2 spike gene, phylodynamic time tree analysis revealed mutational hotspots. According to the SARS-CoV-2 global lineage, the VOCs were included in our predicted models (Supplementary Fig. S1A-S1B). Nucleotide frequencies were calculated after retrieving the SARS-CoV-2 spike ssRNA sequence. More A and U (62.9%) and less G and C were shown in spike ssRNA (37.1%). In their spike ssRNA sequence, RSUV analysis revealed more U-rich codon (Supplementary Table S1). We also created a large data collection of 3822 complete coding sequences to build numerous sequence alignments. Furthermore, Tamura model analysis of substitution patterns demonstrated a high probability for all two forms of substitution (G → A and C → U). After examining additional variations, all of the mutations were highlighted in the ssRNA of the Wuhan variant (Supplementary Fig. S2A–S2E). Our findings revealed U-richness in virtually all coding sequences of SARS-CoV-2 spike ssRNA, however this pattern differed at first codon positions [33]. Our research strategy aimed to determine whether the observed patterns were attributable to a biased mutation process or selection.
Expected Mutation Model
To resolve this enigma, we utilized the concept of probable substitution selection (except deleterious mutation), which has been shown to be prevalent in ssRNA viruses [20]. Next, we modeled to construct the proportion of G → A and C → U substitutions in the small loop (9 ± 2 nucleotides) with more mutations, where as in the larger loop (39 ± 3 nucleotides) with fewer expected mutations, thus estimating the differing roles mutation and selection. We expected and designed five sets of ssRNA sequences depending on loop size, where we identified eighty possible mutational spots.
Most of the mutations were observed within the loop region of the spike protein. Prior research indicates that ssRNA loop regions exhibit heightened motion dynamics as a consequence of their minimal covalent connections [34]. We identified most of the mutations in the loop regions. Additionally, we noted that most of the substitution co-occurrence to G- > A and C- > U. The program estimated the probable combination of mutated sequences (Named as E.M.) based on the size of the five types loop structures. Expected mutations were edited into the loops of ssRNA sequence and named as E.M.1 to E.M. 5; small to large accordingly. Using computational program we retrieved a total of 31 sets of Expected Models (E.M.) utilizing nC1 + nC2 + nC3 + …. + nCn equation (Fig. 1B). Utilizing the program generated RNA sequences were then translated into protein sequences.
Free-Energy Landscape and Stability of SARS-CoV-2 Spike ssRNA
MFE (minimum free energy) methods present in the most popular RNAfold web server was used to predict and analyze secondary RNA structures. Free energy principles based on empirical thermodynamic parameters were used in this model. As indicated in the graph (Supplementary Fig. S4) sequence lengths ranging from 1 to 3822 nucleotides. According to ssRNA stability result, MFE average rate was 28.5 ± 0.5 kcal/mol, free energy of the thermodynamic ensemble was −30 ± 0.6 kcal/mol and ensemble diversity was 18 ± 0.5 per 100 nucleotides. Results revealed that eighteen were thermodynamically stable whereas nine were less stable. Preferred twenty-seven number of SARS-CoV-2 spike ssRNA were picked based on the overall minimum free energy associated with the different structural aspects, such as thermodynamic ensemble, MFE and ensemble diversity (Supplementary Fig. S4).
Optimization Toward Spike Protein Structure Modeling and Validation
The best validation statistics were chosen for study from a total of 27 homology models of the mutant SARS-CoV-2 spike ssRNA. Using the Swiss Model workspace, we generated 27 models of glycosylated full-length spike protein by integrating experimental structural data and bioinformatic predictions. The best validation statistics were chosen for this study from a total of seven homologous models of the expected mutant SARS-CoV-2 spike ssRNA. SWISS-MODEL server was used to analyze the stability of homology structures, based on the Z-score, QMEAN, and Ramachandran plot [35]. These computational methods are preferably used to assess the stability of the modeled spike proteins and the hACE2 cellular receptor for molecular docking. The new models resulted a Qmean z-score of −2.00 ± 0.2, a Mol Probity Score of 1.13 ± 0.03, and a Ramachandran Favoured > 91%. For a protein of this size were considered for further analysis, this is within the allowed range. Figure S4 shows a graphic comparing the quality of our model to current x-ray models in terms of Q-mean z-scores (Supplementary Fig. S5). The 3Drefine server was used to improve the models. Five models were generated by the 3D-refine server. The top-ranking models were selected, having favorable properties such as the lowest 3Drefine score, GDT-HA, RMSD, lowest RWPlus score, and MolProbity [36].
Docking Performance and Bonding Pattern
Spike with ACE2 Binding Partner Identification
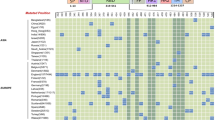
In bound complexes, higher binding affinity was noticed in case of E.M.2 (36), Gamma (32), and Omicron (30) more than 30 ± 3 intercation. Moderate number of bond resulted around the binding region in case of E.M.3 (29), E.M.4 (28), Wuhan (27), Beta (26), E.M.5 (25) and E.M.3.4 (25), Delta (22), and E.M.2.5 (21), whereas lower number of interaction E.M.3.5 (20), Eta (17), and Alpha (14) presented in Fig. 2. Highest number (22 ± 2) of polar amino acids interactome profile found around the binding region in case of omicron, E.M.2, and Beta. Highest number (13 ± 3) of non-polar amino acid interaction noticed in binding complexes, i.e., E.M.2, E.M.3, E.M.4, E.M.5, Gamma, and E.M.3.4. Whereas a lower (7 ± 2) number of non-polar residues interact in the case of E.M.2.5, E.M.3.5, Beta, Delta, Alpha, Eta, and Omicron (Supplementary Fig. S6A-S6N). The binding free energy for viral-spike with ACE2 was calculated. The amino acids contribution to the free energy (−17 ± 1) indicated that the interactions were more favorable for E.M.3, E.M.2, and Omicron.
The hydrogen bonds at the SARS-CoV-2 with H. sapiens ACE2 and SARS-CoV-2 with H. Sapiens IgG interfaces. Percentile stacked bar plot of the residue number of the docked complex revealed after different variants of SARs-CoV-2 spike protein with human ACE2 and IgG heavy with light chain, orange color bar indicating ACE2 with spike; Delta resulted highest number of interaction. Ash color indicates spike with IgG heavy chain and light ash color displays spike with IgG light chain
Spike with IgG Binding Partner Identification
In bound complexes with heavy chain of human IgG revealed that higher number of binding affinity resulted in case of E.M.2 (31) and E.M.3.5 (31). Moderate numbers of bonds were shown in case of E.M.4 (29), E.M.2.5 (28), Omicron (26), E.M.5 (24), Wuhan (24), and E.M.3 (23). Lower number of bond interaction in case of Alpha (20), Eta (17), Gamma (16), and Beta (15) including E.M.3.4 (15) and least number of bonds in case of Delta (7) were identified as primary attractors (Fig. 2). In the cases of E.M.3.4, E.M.4, E.M.2, E.M.2.5, and E.M.3.5, a higher number (18 ± 2) of non-polar amino acid interactome profiles was detected surrounding the binding domain. In the cases of Omicron and E.M.3.5 result reflected higher number (16 ± 1) of polar amino acid bonding patterns.
Whereas a least number of polar and non-polar residues interaction recorded in case of spike of Delta (2) with human IgG heavy chain (Supplementary Fig. S7A-S7N). The binding free energy for the heavy chain of the human IgG antibody with spike was estimated. The contribution of amino acids to the free energy (−18 ± 1) revealed that E.M.3.5, E.M.4, WHCV, and E.M.3 represented a stable interaction. The bound complexes IgG light chain with SARs-CoV-2 spike interestingly showed highest number interaction in case of Omicron (25) and lowest by Delta (4). Other variants including our expected model variants resulted 15 ± 5 number of bonds (Fig. 2). In the cases of E.M.3 (9) highest and Delta, Eta, E.M.5, resulted least number of non-polar amino acid interactome profiles was detected surrounding the binding domain. In the instance of Omicron, the bound complexes of IgG light chain with SARs-CoV-2 spike indicated the maximum number (20) of polar amino acid interactions (Supplementary Fig. S7A-S7N). The interactome profile of ACE2 with the spike complex represented that E.M.2, Gama, Omicron, E.M.3, and E.M.4 showed the largest number of interactions accordingly, in our study. Selected spike protein with IgG-H and IgG-L chain complexes demonstrated that Omicron, E.M.4, and E.M.2.5 showed significant higher number of interaction. Figure 2 revealed that the highest docked surface areas are present in the Delta variants and more of the deleterious nature of the contact areas was documented [37, 38].
The binding free energy of SARS-Cov-2 spike protein with human-ACE2 complex was calculated. Result suggested that the E.M.2, E.M.3, Omicron, and gamma representing stable complex. The binding free energy of the human IgG antibody heavy with light chain with a SARs-CoV-2 spike protein was calculated. Contribution of amino acids to free energy (−12 ± 1) demonstrates a stable bound complex spike of Alpha, Beta E.M.2.5, and E.M.3.4 with human IgG antibody heavy with light chain (Table 1).
Viral membrane glycoproteins, i.e., spikes, are the largest amounts of trans-membrane proteins transcribed from the ssRNA which is in a compact stem-looped structure. So, the fast production of the large number of spikes and the processivity of the RdRp is a mandatory requirement. A large amount of U-A bonding is favorable for maintaining this processivity because lower energy expenditure is required. Additionally, in the case of stability in the protein structure, it was already reported that the trans-membrane proteins encoded with more uracil are more stable and may engage in promiscuous chaperone like activities [39]. In the case of SARS-CoV-2, in our and also in other studies, more U-rich codons are noticed in spike proteins than the viral cytosolic proteins. So spike protein stability by U-rich codon representation may be regarded as one of the functions of the host protein interactivity. A higher ratio of interactions between ACE2: IgG (D + L) represented the more deleterious nature of the variants. In that regard, delta may be judged to be in that category compared to the Wuhan or the Omicron variant. Second position and third position U-rich codons represent Phe, Leu, Ile, Val Cys, Arg, Ser, and Gly amino acids, respectively. The report suggests that the Leu/Ile/Phe-rich domain of some human viruses, i.e., polyomaviruses, including JCV, BKV, and SV40 can form stable dimer/oligomers mediated by a predicted amphipathic α-helix, spanning amino acids. Moreover, deletion of the α-helix renders a replication incompetent virus [40, 41]. Further studies are necessary in relation to the SARS-CoV-2 function. Virus with smaller R group like Val, Gly and Ser and Cys may attribute lower structural hindrance and more liberty to form secondary and tertiary structures which is supportive for more interactive energy ensemble formations. This is supported by Fig. 3. In this regard, transactivation of some Cys-rich proteins in some other RNA viruses like hHIV may be noteworthy, and in addition, Val and Gly are also supportive of its metabolic function and help in anti-termination of the transcriptional regulations [42, 43]. As an individual amino acids thiol containing Cys or hydroxyl Ser has their own capacity for interactivity. Positive charge Arg is helpful for the polar interactions with the contact ligand with an ability to form hydrophobic-hydrophilic combined multi domain structure for higher interactivity. The spike RBD protein and the region of the nucleocapsid protein are demonstrated as nuclear localization signals (NLS). These are enriched with the replacements of positively selected amino acid. These replacements are shown to be linked with apparent epistatic interactions. These are also designated as the blue-prints of major diversification in the SARS-CoV-2 phylogeny [44, 45].
Increased stability and binding affinity interface of spike SARS-CoV-2 with H. sapiens ACE2 protein and IgG. (A) A diagrammatic representation of H. sapiens ACE2 protein and IgG with key amino-acid residues binds more tightly to interact with the spike (S) protein of different SARS-CoV-2 variants, high density color gradients indicates highest number of interactions whereas low density display minimum number of interactions. (B) Multitude of host and different variants of SARS-CoV-2 spike interactions characteristics impacts on disease profiling as per the interaction
Reinfection and recovery from different variants of coronaviruses induce immunological protection. The spike ssRNA mutation rate, which is reflected particularly in the spike glycoprotein, was used to screen for and detect a mutation profile. The assumed E.M.3.4 model may be the variant of concern (VOC). The majority of the mutational impact on expected model variants is in the receptor-binding domain, which increases infectivity by increasing binding affinity with ACE2 [46]. The SARS-CoV-2 Immunity and Reinfection Evaluation (SIREN) project, conducted by Victoria Hall and colleagues in the United Kingdom, revealed that being seropositive to SARS-CoV-2 through natural infection protects against both asymptomatic and symptomatic reinfection [47]. Different study already suggested that antibody through vaccination program enhance protection against a range of SARS-CoV-2 variants. The expected model variants, in particular, exhibit particular immune escape, although most of them have high infectivity (Fig. 3A). However, previous serosurvey and studies suggest that neutral immunity and T-cell responses through vaccine platforms can protect most of the variants, regardless of E.M.3.4 (Fig. 3B).
Time Tree of Empirical Datasets
Phylogenetic evaluation of global viral populations using the GISAID and Nextstrain nomenclature systems revealed multiple clades and lineages (Fig. 4A). A global phylogenetic analysis of the circulating genomes was performed to identify distinct groups and their unique mutational patterns. We used VOC SARS-CoV-2 lineages to create minimal spanning trees (Supplementary Fig. 1A-1B) to visualize genetic links and distances between mutations from various countries. Due to host selection forces, mutations in multiple countries may differ. Some mutation sites in the same nation are constant, and the genome sequence has obvious regional peculiarities [38, 48].
Evolutionary dynamics and time tree of SARS-CoV-2 spike ssRNA. (A)The genome of the SARS-CoV-2 global phylogenetic tree, as extracted from GISAID, as seen in the Nextstrain global phylogenetic tree. Tree-Time was used to create a time-resolved phylogenetic tree with chosen metadata information. (B) Scale lengths represent the divergence time. Phylogeny based on co-evolution divergence times was estimated, and each branch is labeled using the ML method, utilizing the Tamura-Nei substitution model. The sampling interval is similar to the time frame which assesses the long-term rate of evolution with high precision
These findings suggested that, although certain changes are essential in the development of SARS-CoV-2, others may be the consequence of the virus’s adaptability to multiple countries, natural environment, infectivity pattern, and immunoescape mechanisms. The time tree for the SARS-CoV-2 spike was built using ssRNA source trees. By commensalism and amensalism, seven expected variants and five variants of SARS-CoV-2 are highlighted and classified, respectively. The length along the branches is indicated by the bootstrap percentage out of 500 bootstrapping.
The cross-family continuity in reference to time scale phylogeny was estimated using phylogenetic analysis [33]. The SARs-CoV-2 spike ssRNA dataset, which served as a model for SARs-CoV-2 infectivity and escapology in time tree. Due to the former’s higher pace of development, there are also considerable changes in overall tree length. The genomic evolutionary rate of SARs-CoV-2 spike ssRNA was expressed in divergence time, and the values for E.M.5, E.M.4, E.M.3 with E.M.3.5 and E.M.2 with E.M.2.5 & E.M.3.4 displayed, respectively (Fig. 4B). The SARs-CoV-2 expected evolutionary clock was coordinated with future adaptive diversity in humans.
Discussion
We revealed that the spike of SARS-CoV-2 ssRNA studied here had a skewed nucleotide composition in their coding regions, with U and A-rich sequences. The spike ssRNA of SARS-CoV-2 contains 3822 nucleotides, and there are ≈ 11,000 possible single nucleotide substitutions that may occur [13]. One of the experimental goals of this analysis is to explore the fingerprints of the RNA editing ratio in the long-term evolution of the Coronavirus. The existence of a greater rate of Adenine rich sequences, as well as probable selection for amino acids encoded by A-rich codons, promotes MHC system avoidance [49]. We noticed a lower frequency of CG pairs and a higher frequency of U compared to A in the single-stranded RNA (ssRNA). Zinc finger antiviral protein (ZAP), apolipoprotein BmRNA (APOBEC), and adenosine deaminases acting on RNA (ADAR) promote C- > U mutations in the SARS-CoV-2 ssRNA genome [13, 50, 51]. Regardless of the underlying mutational processes, the studies performed in the research revealed an over-representation of C- > U transitions. It was already reported that oxygen tension would alter the induction of bacterial mutations by other mutagens which is regulated by hypoxia-inducible factor-1α [52,53,54]. Predicted ssRNA sequences were chosen and structured to be not only be more thermodynamically stable but also to have a more compact structural ensemble than random sequences. The genomic structural stabilization, at the expense of the lower virulence and pathogenicity that was observed in some recent variants like Omicron resulted in higher spreadability but lower mortality in human host. This suggests a co-surveillance adaptation strategy. Moreover, the selection of more U and A indicates the adaption strategies of simpler nuclear to maintain a lower energy level for structural stabilization. Unlike several other viruses, SARS-CoV-2 has showed purifying selection strategies and has extensively utilized its mutational benefits. Although a significant number of variants might have gone through extirpation steps, a large number of VOCs have survived. Implication in ancestral recombination is a possible game plan of the procurement of the mutational advantages in positive selection procedures. One report reveals that major selective forces are implicated on the ‘501Y lineages’ by repeatedly favored convergent mutations at 35 genome sites of the SARS-CoV-2 [55, 56]. A report revealed that the genomes of SARS-CoV-2 were remarkably structured, with minimum folding energy (expressed as minimum folding energy differences; MFEDs) than previously examined viruses, such as the hepatitis C virus. High MFED values were shared with all coronavirus genomes analyzed, creating several hundred consecutive energetically favored stem-loops throughout the genome. This characteristic provided insight into the selection of the SARS-CoV-2 spike ssRNA for protein modeling. Surprisingly, the findings of the study revealed a strong correlation between phylogeny and divergence time (Fig. 4). The highest number of mutations resulted in the case of E.M.3.4 which is 113 and 87 compared to Omicron and Wuhan reference strain (Supplementary Fig. S8). It also adds to the evidence that systematic adaptation in SARS-CoV-2 spike ssRNA may be a key factor for assuming infectivity and immune-escapology. All the predicted model variants showed moderate or high infective rate but reduced severity due the significant number of IgG light and heavy chain interactions, i.e., E.M.5, E.M.4, E.M.3, E.M.3.5, E.M.2, and E.M.2.5. Only one variant, EM.3.4, appears to cause serious symptoms, like Delta variant. Although a high rate of vaccination results in a neutralization titer against SARS-CoV-2 variants, this may reduce serious symptoms. It was already reported that the decline of the neutralization titer after 250 days represents a considerable loss of protection against SARS-CoV-2 infection [57]. Here, we show that our hypothesized E.M.3.4 provides an evidence-based SARS-CoV-2 variant of concern model that will assist in the design of vaccination strategies to control the pandemic’s future trajectory. Mutational imposition of the fitness cost on the virus is attributed by the stem-loop structural disruption of the ssRNA and lower level of energy ensemble, as observed in the current study. Out of the main three events like infectivity (ACE2-TMPRSS2 role), propagation (replication) and immuno-escape (MHC-IgG recognition), the virus appears to strive for a balance in negotiation. We hypothesize that these balancing features are evolutionary attributed via mutational modifications of the ssRNA of the SARS-CoV-2 following thermodynamic and kinetic properties. Further structural constraints, such as stem-looping, impose genomic reconstruction for molecular epidemiology. As mentioned in this paragraph, not only ACE2 binding or immune interaction, but also RNA-dependent RNA polymerase (RdRp) structural alteration and stability also determine viral fitness. Multiple mutational analyses determined the role of mutations in inducing alterations in RNA secondary structure and RdRp functionality, thereby governing the pathogenicity of the virus. The important finding of more occurrences of neutral variants in asymptomatic or less symptomatic persons versus deleterious variants in the deceased person may suggest viral fitness definition. The killing of the host terminates a large number of viral life-cycle rather co-surveillance with the host increases the propagation and spreadibility of the virus. Therefore, long-term evolutionary fitness is rendered in the second category. The fatal variants that killed large populations did not last long because they had a particular mutation associated with strong ACE2 binding, immune escape, or both. For example, the D614G mutation in the S-glycoprotein, for example, has an impact on immunity and partial vaccination escape [58]. The E484K and N501Y mutations in the receptor-binding domain of spike were the major sources of neutralizing resistance. The 501Y.V2 variants, on the other hand, do not confer improved infectivity in a variety of cell types [59]. With their diverse virulence and epidemiological results, five important mutations (T95I, A222V, G142D, R158G, and K417N) were much more frequent in the Delta Plus variant than in the Delta variant [60]. When comparing the mutational hotspots of the Omicron and Delta variants, it appears that the Omicron variation has a lot of mutations in its spike proteins [61]. However, according to the findings, the probability of severe outcomes after SARS-CoV-2 infection is significantly lower for omicron than for delta [62]. As a result of Omicron infections, booster immunization with mRNA vaccines provides over 70% protection against hospitalization and mortality. The presence of a major cluster of mutations in the S protein has led to the development of more than 12 variants, indicating the beginning of antigenic drift for SARS-CoV-2. The emergence of such variations is particularly concerning since they may evade antibodies and result in increased ACE2 binding. This divergence in time for viral identification might lead to an underestimation of the pandemic’s scale [63, 64]. The predicted S protein, particularly its RBD that interacts with the host cellular receptor to gain access to host cells, is a viable vaccination target for SARS-CoV-2. As illustrated above, E.M.3.4 variant may improve ACE2 affinity but may not necessarily confer antibody resistance. On the other hand, E.M.2, E.M.2.5, E.M.3, E.M.3.5, E.M.4, and E.M.5 resulted in lower ACE2 binding affinity and more IgG interactions. Other potential immunological correlates of protection will be tested and evaluated using our hypothesized models. The WHO has declared the VOCs to be resistant to neutralizing antibodies, making them more infectious and pathogenic [65, 66].
Conclusion
Our hypothesized models for SARS-CoV-2 are resistant to neutralizing antibodies, rendering the virus more infectious and pathogenic. Clade and lineage classifications will be altered due to the virus’s high diversity and high mutability nature. It has already been reported that the molecular evolution pattern of the SARS-CoV-2 amino acid substitution rate is 25.331 per year [67]. Our hypothesized model, E.M.3.4, may evolve in 2024–2025, and E.M.2 is expected to evolve in 2024. On the other hand, our research presents a modeling framework for combine’s imprecise data from previous infected variants with antibody interactome profile, which may provide a strategy for projecting the uncertain future of SARS-CoV-2 immunity. To evaluate the efficacy of control measures and monitor co-evolutionary events in the future, our hypothesized VOC models and evolutionary epidemiology of SARS-CoV-2 lineages may become more significant.
Data Availability
All data generated or analyzed during this study are included in the figures and supporting files.
References
Kendall, E. J., Bynoe, M. L., & Tyrrell, D. A. (1962). Virus isolations from common colds occurring in a residential school. British Medical Journal, 2(5297), 82–86. https://doi.org/10.1136/bmj.2.5297.82
Llanes, A., Restrepo, C. M., Caballero, Z., Rajeev, S., Kennedy, M. A., & Lleonart, R. (2020). Betacoronavirus genomes: How genomic information has been used to deal with past outbreaks and the COVID-19 pandemic. International Journal of Molecular Sciences, 21(12), 4546. https://doi.org/10.3390/ijms21124546
Fehr, A. R., & Perlman, S. (2015). Coronaviruses: An overview of their replication and pathogenesis. Methods in Molecular Biology (Clifton, N.J.), 1282, 1–23. https://doi.org/10.1007/978-1-4939-2438-7_1
Chan, J. F., Kok, K. H., Zhu, Z., Chu, H., To, K. K., Yuan, S., et al. (2020). Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerging Microbes & Infections, 9(1), 221–236. https://doi.org/10.1080/22221751.2020.1719902
Luan, J., Lu, Y., Jin, X., & Zhang, L. (2020). Spike protein recognition of mammalian ACE2 predicts the host range and an optimized ACE2 for SARS-CoV-2 infection. Biochemical and Biophysical Research Communications, 526(1), 165–169. https://doi.org/10.1016/j.bbrc.2020.03.047
Wrapp, D., Wang, N., Corbett, K. S., Goldsmith, J. A., Hsieh, C. L., Abiona, O., et al. (2020). Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science (New York, N.Y.), 367(6483), 1260–1263. https://doi.org/10.1126/science.abb2507
Hoffmann, M., Kleine-Weber, H., Schroeder, S., Krüger, N., Herrler, T., Erichsen, S., et al. (2020). SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell, 181(2), 271–280.e8. https://doi.org/10.1016/j.cell.2020.02.052
Yang, N., & Shen, H. M. (2020). Targeting the endocytic pathway and autophagy process as a novel therapeutic strategy in COVID-19. International Journal of Biological Sciences, 16(10), 1724–1731. https://doi.org/10.7150/ijbs.45498
Xia, S., Zhu, Y., Liu, M., Lan, Q., Xu, W., Wu, Y., et al. (2020). Fusion mechanism of 2019-nCoV and fusion inhibitors targeting HR1 domain in spike protein. Cellular & Molecular Immunology, 17(7), 765–767. https://doi.org/10.1038/s41423-020-0374-2
Snijder, E. J., Bredenbeek, P. J., Dobbe, J. C., Thiel, V., Ziebuhr, J., Poon, L. L., et al. (2003). Unique and conserved features of genome and proteome of SARS-coronavirus, an early split-off from the coronavirus group 2 lineage. Journal of Molecular Biology, 331(5), 991–1004. https://doi.org/10.1016/s0022-2836(03)00865-9
Sola, I., Almazán, F., Zúñiga, S., & Enjuanes, L. (2015). Continuous and discontinuous RNA synthesis in coronaviruses. Annual Review of Virology, 2(1), 265–288. https://doi.org/10.1146/annurev-virology-100114-055218
Angelini, M. M., Akhlaghpour, M., Neuman, B. W., & Buchmeier, M. J. (2013). Severe acute respiratory syndrome coronavirus nonstructural proteins 3, 4, and 6 induce double-membrane vesicles. mBio, 4(4), e00524.13. https://doi.org/10.1128/mBio.00524-13
Sender, R., Bar-On, Y. M., Gleizer, S., Bernshtein, B., Flamholz, A., Phillips, R., et al. (2021). The total number and mass of SARS-CoV-2 virions. Proceedings of the National Academy of Sciences of the United States of America, 118(25), e2024815118. https://doi.org/10.1073/pnas.2024815118
Wang, R., Chen, J., Hozumi, Y., Yin, C., & Wei, G. W. (2020). Decoding asymptomatic COVID-19 infection and transmission. The Journal of Physical Chemistry Letters, 11(23), 10007–10015. https://doi.org/10.1021/acs.jpclett.0c02765
Baer, C. F. (2008). Does mutation rate depend on itself. PLoS Biology, 6(2), e52. https://doi.org/10.1371/journal.pbio.0060052
Loewe, L., & Hill, W. G. (2010). The population genetics of mutations: Good, bad and indifferent. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences, 365(1544), 1153–1167. https://doi.org/10.1098/rstb.2009.0317
Guo, S., & Wong, S. M. (2018). Disruption of a stem-loop structure located upstream of pseudoknot domain in Tobacco mosaic virus enhanced its infectivity and viral RNA accumulation. Virology, 519, 170–179. https://doi.org/10.1016/j.virol.2018.04.009
Liu, Y., Zhang, Y., Wang, M., Cheng, A., Yang, Q., Wu, Y., et al. (2020). Structures and functions of the 3′ untranslated regions of positive-sense single-stranded RNA viruses infecting humans and animals. Frontiers in Cellular and Infection Microbiology, 10, 453. https://doi.org/10.3389/fcimb.2020.00453
Sun, Y., Guo, Y., & Lou, Z. (2012). A versatile building block: The structures and functions of negative-sense single-stranded RNA virus nucleocapsid proteins. Protein & Cell, 3(12), 893–902. https://doi.org/10.1007/s13238-012-2087-5
Harvey, W. T., Carabelli, A. M., Jackson, B., Gupta, R. K., Thomson, E. C., Harrison, E. M., et al. (2021). SARS-CoV-2 variants, spike mutations and immune escape. Nature Reviews. Microbiology, 19(7), 409–424. https://doi.org/10.1038/s41579-021-00573-0
V’kovski, P., Kratzel, A., Steiner, S., Stalder, H., & Thiel, V. (2021). Coronavirus biology and replication: Implications for SARS-CoV-2. Nature Reviews. Microbiology, 19(3), 155–170. https://doi.org/10.1038/s41579-020-00468-6
Tamura, K. (1992). Estimation of the number of nucleotide substitutions when there are strong transition-transversion and G+C-content biases. Molecular Biology and Evolution, 9(4), 678–687. https://doi.org/10.1093/oxfordjournals.molbev.a040752
Kumar, S., Stecher, G., Li, M., Knyaz, C., & Tamura, K. (2018). MEGA X: Molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution, 35(6), 1547–1549. https://doi.org/10.1093/molbev/msy096
Kerpedjiev, P., Hammer, S., & Hofacker, I. L. (2015). Forna (force-directed RNA): Simple and effective online RNA secondary structure diagrams. Bioinformatics (Oxford, England), 31(20), 3377–3379. https://doi.org/10.1093/bioinformatics/btv372
Bhattacharya, D., Nowotny, J., Cao, R., & Cheng, J. (2016). 3Drefine: An interactive web server for efficient protein structure refinement. Nucleic Acids Research, 44(W1), W406–W409. https://doi.org/10.1093/nar/gkw336
Honorato, R. V., Koukos, P. I., Jiménez-García, B., Tsaregorodtsev, A., Verlato, M., Giachetti, A., et al. (2021). Structural biology in the clouds: The WeNMR-EOSC ecosystem. Frontiers in Molecular Biosciences, 8, 729513. https://doi.org/10.3389/fmolb.2021.729513
van Zundert, G. C. P., Rodrigues, J. P. G. L. M., Trellet, M., Schmitz, C., Kastritis, P. L., Karaca, E., et al. (2016). The HADDOCK2.2 Web Server: User-friendly integrative modeling of biomolecular complexes. Journal of Molecular Biology, 428(4), 720–725. https://doi.org/10.1016/j.jmb.2015.09.014
Schrödinger, L., & DeLano, W. (2020). PyMOL. Retrieved from http://www.pymol.org/pymol
Tamura, K., Battistuzzi, F. U., Billing-Ross, P., Murillo, O., Filipski, A., & Kumar, S. (2012). Estimating divergence times in large molecular phylogenies. Proceedings of the National Academy of Sciences of the United States of America, 109(47), 19333–19338. https://doi.org/10.1073/pnas.1213199109
Tamura, K., & Nei, M. (1993). Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Molecular Biology and Evolution, 10(3), 512–526. https://doi.org/10.1093/oxfordjournals.molbev.a040023
Tao, Q., Tamura, K., Mello, B., & Kumar, S. (2020). Reliable confidence intervals for RelTime estimates of evolutionary divergence times. Molecular Biology and Evolution, 37(1), 280–290. https://doi.org/10.1093/molbev/msz236
Tamura, K., Stecher, G., & Kumar, S. (2021). MEGA11: Molecular evolutionary genetics analysis version 11. Molecular Biology and Evolution, 38(7), 3022–3027. https://doi.org/10.1093/molbev/msab120
Tang, X., Wu, C., Li, X., Song, Y., Yao, X., Wu, X., et al. (2020). On the origin and continuing evolution of SARS-CoV-2. National Science Review, 7(6), 1012–1023. https://doi.org/10.1093/nsr/nwaa036
Suleman, M., Luqman, M., Wei, D. Q., Ali, S., Ali, S. S., Khan, A., et al. (2023). Structural insights into the effect of mutations in the spike protein of SARS-CoV-2 on the binding with human furin protein. Heliyon, 9(4), e15083. https://doi.org/10.1016/j.heliyon.2023.e15083
Benkert, P., Biasini, M., & Schwede, T. (2011). Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics (Oxford, England), 27(3), 343–350. https://doi.org/10.1093/bioinformatics/btq662
Williams, C. J., Headd, J. J., Moriarty, N. W., Prisant, M. G., Videau, L. L., Deis, L. N., et al. (2018). MolProbity: More and better reference data for improved all-atom structure validation. Protein Science: A Publication of the Protein Society, 27(1), 293–315. https://doi.org/10.1002/pro.3330
Islam, M. A., Marzan, A. A., Arman, M. S., Shahi, S., Sakif, T. I., Hossain, M., et al. (2023). Some common deleterious mutations are shared in SARS-CoV-2 genomes from deceased COVID-19 patients across continents. Scientific Reports, 13(1), 18644. https://doi.org/10.1038/s41598-023-45517-1
Markov, P. V., Ghafari, M., Beer, M., Lythgoe, K., Simmonds, P., Stilianakis, N. I., et al. (2023). The evolution of SARS-CoV-2. Nature Reviews. Microbiology, 21(6), 361–379. https://doi.org/10.1038/s41579-023-00878-2
Benhalevy, D., Bochkareva, E. S., Biran, I., & Bibi, E. (2015). Model Uracil-Rich RNAs and membrane protein mRNAs interact specifically with cold shock proteins in Escherichia coli. PLoS ONE, 10(7), e0134413. https://doi.org/10.1371/journal.pone.0134413
Sami Saribas, A., Abou-Gharbia, M., Childers, W., Sariyer, I. K., White, M. K., & Safak, M. (2013). Essential roles of Leu/Ile/Phe-rich domain of JC virus agnoprotein in dimer/oligomer formation, protein stability and splicing of viral transcripts. Virology, 443(1), 161–176. https://doi.org/10.1016/j.virol.2013.05.003
Coric, P., Saribas, A. S., Abou-Gharbia, M., Childers, W., White, M. K., Bouaziz, S., et al. (2014). Nuclear magnetic resonance structure revealed that the human polyomavirus JC virus agnoprotein contains an α-helix encompassing the Leu/Ile/Phe-rich domain. Journal of Virology, 88(12), 6556–6575. https://doi.org/10.1128/JVI.00146-14
Kao, S. Y., Calman, A. F., Luciw, P. A., & Peterlin, B. M. (1987). Anti-termination of transcription within the long terminal repeat of HIV-1 by tat gene product. Nature, 330(6147), 489–493. https://doi.org/10.1038/330489a0
Kubota, S., Endo, S., Maki, M., & Hatanaka, M. (1989). Role of the cysteine-rich region of HIV tat protein on its trans-activational ability. Virus Genes, 2(2), 113–118. https://doi.org/10.1007/BF00315255
Holmes, E. C., Goldstein, S. A., Rasmussen, A. L., Robertson, D. L., Crits-Christoph, A., Wertheim, J. O., et al. (2021). The origins of SARS-CoV-2: A critical review. Cell, 184(19), 4848–4856. https://doi.org/10.1016/j.cell.2021.08.017
Kemp, S. A., Collier, D. A., Datir, R. P., Ferreira, I., Gayed, S., Jahun, A., et al. (2021). SARS-CoV-2 evolution during treatment of chronic infection. Nature, 592, 277–282. https://doi.org/10.1038/s41586-021-03291-y
Starr, T. N., Greaney, A. J., Hilton, S. K., Ellis, D., Crawford, K. H. D., Dingens, A. S., et al. (2020). Deep mutational scanning of SARS-CoV-2 receptor binding domain reveals constraints on folding and ACE2 binding. Cell, 182(5), 1295–1310.e20. https://doi.org/10.1016/j.cell.2020.08.012
Hall, V. J., Foulkes, S., Charlett, A., Atti, A., Monk, E. J. M., Simmons, R., et al. (2021). SARS-CoV-2 infection rates of antibody-positive compared with antibody-negative health-care workers in England: A large, multicentre, prospective cohort study (SIREN). Lancet (London, England), 397(10283), 1459–1469. https://doi.org/10.1016/S0140-6736(21)00675-9
Mercatelli, D., & Giorgi, F. M. (2020). Geographic and genomic distribution of SARS-CoV-2 mutations. Frontiers in Microbiology, 11, 1800. https://doi.org/10.3389/fmicb.2020.01800
Kustin, T., & Stern, A. (2021). Biased mutation and selection in RNA viruses. Molecular Biology and Evolution, 38(2), 575–588. https://doi.org/10.1093/molbev/msaa247
Bishop, K. N., Holmes, R. K., Sheehy, A. M., & Malim, M. H. (2004). APOBEC-mediated editing of viral RNA. Science (New York, N.Y.), 305(5684), 645. https://doi.org/10.1126/science.1100658
Samuel, C. E. (2012). ADARs: Viruses and innate immunity. Current Topics in Microbiology and Immunology, 353, 163–195. https://doi.org/10.1007/82_2011_148
Anderson, E. H. (1951). The effect of oxygen on mutation induction by X-rays. Proceedings of the National Academy of Sciences of the United States of America, 37(6), 340–349. https://doi.org/10.1073/pnas.37.6.340
Semenza, G. L., & Wang, G. L. (1992). A nuclear factor induced by hypoxia via de novo protein synthesis binds to the human erythropoietin gene enhancer at a site required for transcriptional activation. Molecular and Cellular Biology, 12(12), 5447–5454. https://doi.org/10.1128/mcb.12.12.5447-5454.1992
Huang, R., Huestis, M., Gan, E. S., Ooi, E. E., & Ohh, M. (2021). Hypoxia and viral infectious diseases. JCI Insight, 6(7), e147190. https://doi.org/10.1172/jci.insight.147190
Martin, D. P., Weaver, S., Tegally, H., San, J. E., Shank, S. D., Wilkinson, E., et al. (2021). The emergence and ongoing convergent evolution of the SARS-CoV-2 N501Y lineages. Cell, 184(20), 5189–5200.e7. https://doi.org/10.1016/j.cell.2021.09.003
Rochman, N. D., Wolf, Y. I., Faure, G., Mutz, P., Zhang, F., & Koonin, E. V. (2021). Ongoing global and regional adaptive evolution of SARS-CoV-2. Proceedings of the National Academy of Sciences of the United States of America, 118(29), e2104241118. https://doi.org/10.1073/pnas.2104241118
Khoury, D. S., Cromer, D., Reynaldi, A., Schlub, T. E., Wheatley, A. K., Juno, J. A., et al. (2021). Neutralizing antibody levels are highly predictive of immune protection from symptomatic SARS-CoV-2 infection. Nature Medicine, 27(7), 1205–1211. https://doi.org/10.1038/s41591-021-01377-8
Bhattacharya, M., Chatterjee, S., Sharma, A. R., Agoramoorthy, G., & Chakraborty, C. (2021). D614G mutation and SARS-CoV-2: Impact on S-protein structure, function, infectivity, and immunity. Applied Microbiology and Biotechnology, 105(24), 9035–9045. https://doi.org/10.1007/s00253-021-11676-2
Li, Q., Nie, J., Wu, J., Zhang, L., Ding, R., Wang, H., et al. (2021). SARS-CoV-2 501Y.V2 variants lack higher infectivity but do have immune escape. Cell, 184(9), 2362–2371.e9. https://doi.org/10.1016/j.cell.2021.02.042
Kannan, S. R., Spratt, A. N., Cohen, A. R., Naqvi, S. H., Chand, H. S., Quinn, T. P., et al. (2021). Evolutionary analysis of the Delta and Delta Plus variants of the SARS-CoV-2 viruses. Journal of Autoimmunity, 124, 102715. https://doi.org/10.1016/j.jaut.2021.102715
Saxena, S. K., Kumar, S., Ansari, S., Paweska, J. T., Maurya, V. K., Tripathi, A. K., et al. (2022). Characterization of the novel SARS-CoV-2 Omicron (B.1.1.529) variant of concern and its global perspective. Journal of Medical Virology, 94(4), 1738–1744. https://doi.org/10.1002/jmv.27524
Nyberg, T., Ferguson, N. M., Nash, S. G., Webster, H. H., Flaxman, S., Andrews, N., et al. (2022). Comparative analysis of the risks of hospitalisation and death associated with SARS-CoV-2 omicron (B.1.1.529) and delta (B.1.617.2) variants in England: A cohort study. Lancet (London, England), 399(10332), 1303–1312. https://doi.org/10.1016/S0140-6736(22)00462-7
Fountain-Jones, N. M., Appaw, R. C., Carver, S., Didelot, X., Volz, E., & Charleston, M. (2020). Emerging phylogenetic structure of the SARS-CoV-2 pandemic. Virus Evolution, 6(2), veaa082. https://doi.org/10.1093/ve/veaa082
Voloch, C. M., da Silva Francisco, R., de Almeida Jr, L. G. P., Brustolini, O. J., Cardoso, C. C., Gerber, A. L., et al. (2021). Intra-host evolution during SARS-CoV-2 prolonged infection. Virus Evolution, 7(2), veab078. https://doi.org/10.1093/ve/veab078
Liu, J., Liu, Y., Xia, H., Zou, J., Weaver, S. C., Swanson, K. A., et al. (2021). BNT162b2-elicited neutralization of B.1.617 and other SARS-CoV-2 variants. Nature, 596(7871), 273–275. https://doi.org/10.1038/s41586-021-03693-y
Liu, C., Ginn, H. M., Dejnirattisai, W., Supasa, P., Wang, B., Tuekprakhon, A., et al. (2021b). Reduced neutralization of SARS-CoV-2 B.1.617 by vaccine and convalescent serum. Cell, 184(16), 4220–4236.e13. https://doi.org/10.1016/j.cell.2021.06.020
Rambaut, A. (2020). Phylogenetic analysis of nCoV-2019 genomes. Virological. https://virological.org/t/phylodynamic-analysis-176-genomes-6-mar-2020/356.
Acknowledgements
The authors thank the Oriental Institute of Science and Technology for supporting our research work.
Funding
There was no funding for this work from grants or other sources to aid in the preparation of this article.
Author information
Authors and Affiliations
Contributions
AS, Data curation, Formal analysis, Software, Visualization; TAG, Development of expected model sequences by C program; BB, Writing-review and editing; SM, Conceptualization, Formal analysis, Writing – original draft, Writing-review and editing; ASP, Conceptualization, Data curation, Methodology, Resources, Software, Supervision, Writing – original draft, Writing – review and editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Consent for Publication
Not applicable.
Ethical approval
No ethical approval was required for this work. This article does not contain studies with human or animal subjects performed by any of the authors.
Research Involving Human and Animal Rights
This study was performed at in silico level.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sarkar, A., Ghosh, T.A., Bandyopadhyay, B. et al. Prediction of Prospective Mutational Landscape of SARS-CoV-2 Spike ssRNA and Evolutionary Basis of Its Host Interaction. Mol Biotechnol (2024). https://doi.org/10.1007/s12033-024-01146-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12033-024-01146-1