Abstract
Atherosclerosis (AS) is a chronic and progressive inflammatory disease. Ubiquitin-conjugating enzyme E2G 2 (UBE2G2) has been reported to be differentially expressed in subjects with abnormal coronary endothelial function. We intended to further explore the effect of UBE2G2 in AS using apolipoprotein E-deficient (ApoE−/−) mice. Relative UBE2G2 expression in aortic sinus tissues was examined by Real-time reverse transcriptase-polymerase chain reaction and immunohistochemical staining. Atherosclerotic plaque formation was observed through hematoxylin–eosin staining. The protein levels of adhesion biomarkers and inflammatory cytokines was analyzed by western blotting. The direct interaction between UBE2G2 and miR-204-5p was predicted by bioinformatic analysis, and the correlation was analyzed by Pearson's correlation test, and verified by luciferase reporter assay. Human vascular smooth muscle cells (VSMCs) development was detected by 5-ethynyl-2′-deoxyuridine labeling assay and wound healing assays. UBE2G2 was highly expressed in the aortic sinus tissues of high-fat diet-fed ApoE−/− mice. The atherosclerotic plaque formation was increased in ApoE−/− mice, while UBE2G2 knockdown reduced it. Silencing of UBE2G2 also inhibited the expression and protein levels of adhesion biomarkers and inflammatory cytokines in ApoE−/− mice. MiR-204-5p was the upstream effector of UBE2G2 and miR-204-5p overexpression was found to inhibit the proliferation and migration of human VSMCs through regulating UBE2G2 expression. UBE2G2 inhibition attenuated AS in ApoE−/− mice and UBE2G2 expression was negatively regulated by miR-204-5p in human VSMCs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Atherosclerosis (AS) is characterized by lipids accumulation, inflammatory factor activation and thrombosis in the arterial vessel wall, which is a major cause of death and disability worldwide [1,2,3]. The alterations of proliferation, migration, and secretion function in vascular smooth muscle cell (VSMCs) also contribute to the AS development [4]. Moreover, apolipoprotein E-deficient (ApoE−/−) is well characterized to develop atherosclerotic lesions similar to those in humans [5]. Therefore, we chose ApoE−/− mice as the model of in vivo experiment, and VSMCs were used for in vitro study to explore novel research strategy for AS from a molecular level.
Ubiquitin-conjugating enzyme E2G 2 (UBE2G2) has the typical secondary structure of members of the ubiquitin carrier proteins family, which has been transcribed in most tissues but at particularly high levels in skeletal muscle and heart [6,7,8]. In addition, UBE2G2 involvement has been reported in several diseases, like hypoglycemia [9], type 2 diabetes [10], and liver diseases [11]. A recent study orders the genes that are differentially expressed in subjects with normal (n = 6) or abnormal (n = 13) coronary endothelial function [12]. UBE2G2 expression shows the most significant difference with the P value of 2.9 × 10−6, indicating its potential role in early coronary AS. Interestingly, Robichaud et al. proposed that the ubiquitination factor UBE2G2 regulates cholesterol efflux in macrophage foam cells, suggesting its potential role in atherogenesis [13]. However, the specific role and mechanism of UBE2G2 in AS mice model and human vascular smooth muscle cells (VSMCs) are unclear.
MicroRNAs (miRNAs) are a subset of non-coding RNAs of 18–22 nt. The major function of miRNAs is the regulation of mRNA expression by pairing with 3'UTR target sequence of mRNA [14, 15]. Numerous miRNAs have been reported to be involved in the development of VSMCs [16]. MiR-204-5p is predicted to be the upstream miRNA of UBE2G2 by the starBase [17]. Moreover, prior studies indicate that miR-204-5p is involved in the growth and migration of VSMCs via regulating Matrix metalloproteinase-9 (MMP-9) expression [18] and is closely related to AS [19]. In addition, the anti-inflammatory role of miR-204-5p has been revealed in steoarthritis [20]. Notably, Wang et al. reported that Cyanidin-3–O–glucoside (C3G) decreases blood lipid levels and suppresses pro-inflammatory cytokine production in in rabbits using high-fatty diet (HFD), which was associated with the down-regulation of miR-204-5p [21]. Nevertheless, no study discusses the UBE2G2/mir-204-5p axis in the development of AS.
In this study, we constructed an animal model of AS by feeding ApoE−/− mice with HFD to investigate the function of UBE2G2 in AS. We hypothesized that UBE2G2 may regulate the AS development in HFD-fed ApoE−/− mice and VSMCs.
Materials and Methods
Animal Models
Thirty male normal C57BL/6 J and ApoE−/− C57BL/6 J mice at 8 weeks of age were obtained from the Jackson Laboratory (Beijing, China). All animals were kept in constant temperature (22–25 °C) and with a 12/12 h light–dark cycle and divided into two groups: a normal chow diet (control group, n = 10) or a high-fat diet (n = 20). HFD-fed mice were then randomly divided into HFD group (n = 10) and HFD + adeno-associated virus (AAV)-sh-UBE2G2 group (n = 10). Mice from HFD + AAV-sh-UBE2G2 group were intraperitoneally injected with AAV vector-expressing UBE2G2 knockdown (AAV-sh-UBE2G2) at a dose of 4 mg·kg−1⋅d−1, while mice in the high-fat diet group were injected with an equal volume of saline which contains 0.1% dimethyl sulfoxide (DMSO), and mice in the normal chow diet group were only injected with equal volume of saline. The total duration of the HFD feeding paradigm was 20 weeks (9 weeks before AAV vector administration and 11 weeks after that). Mice from different groups were sacrificed by cervical dislocation after the ether anesthesia. Finally, the aortic sinus tissues were collected. All animal experimental procedures were approved by the Animal Care and Use Committee of the Affiliated Zhangjiagang Hospital of Soochow University (Jiangsu, China) according to institutional guidelines for animal ethics.
Cell Culture and Transfection
Human VSMCs were purchased from ATCC (Manassas, VA, USA) and cultured in DMEM (Thermo Fisher Scientific, HyClone, Logan, UT, USA) which contains 10% fetal bovine serum (Thermo Fisher Scientific). VSMCs were grown at 37 °C in 5% CO2. The medium was changed every 48 h. MiR-204-5p mimics, pcDNA3.1-UBE2G2, and corresponding negative controls were all purchased from GenePharma (Shanghai, China) and transfected into VSMCs using lipofectamine 3000 (Invitrogen, Carlsbad, CA, USA) for 48 h.
Reverse Transcription Quantitative Polymerase Chain Reaction (RT-qPCR)
Total RNA of VSMCs and aortic sinus tissues was extracted using TRIzol reagent (ThermoFisher Scientific) and reversely transcribed to cDNA using a SuperScript II kit (Invitrogen). Then, the expression of miR-204-5p and UBE2G2 was detected using QuantiTect SYBR Green PCR Master Mix (Qiagen, Valencia, CA). The 2−ΔΔCT method was used to calculate the relative RNA levels with U6 as a loading control. The primer sequences are shown in Table 1.
Immunohistochemical Staining
Aortic sinus tissues were first dehydrated in alcohol and xylene, and then embedded in paraffin. Each tissue was cut into five Sections (8 μm-thick), which were then incubated with H2O2 (3%) in methanol for 10 min. Subsequently, the sections were probed with primary antibodies (50 μL) against UBE2G2 (ab235790; 1:200) at 4 °C overnight. After washing with PBS, the sections were incubated with secondary antibody for 1 h at 37 °C. The sections were then developed with diaminobenzidine (DAB) substrate for visualization. After being dehydrated and sealed with the neutral gum, the sections were subjected to the fluorescence microscopy (IX-51, Olympus).
Hematoxylin and Eosin (H&E) Staining
After 24 h of fixation in 4% paraformaldehyde, the tissues were embedded in paraffin. Sections (5 μm-thick) were prepared and mounted on glass slides. Subsequently, paraffin-embedded aortic sinus tissues were dewaxed and rehydrated. Slides were then stained with hematoxylin and eosin for 5 min respectively. Images were captured using an optical microscope (40×; DP73; Olympus, Tokyo, Japan).
Western Blot
Total protein was extracted from VSMCs and aortic sinus tissues using RIPA buffer (Roche, Shanghai, China) and subjected to sodium dodecylsulfate polyacrylamide gel electrophoresis. Subsequently, the proteins were transferred to preactivated polyvinylidene difluoride membrane (Millipore, Bedford, MA, USA). After being blocked with Tris-buffered saline-tween (TBST) containing 5% skimmed milk for 1 h at room temperature, the membrane was then incubated with the primary antibody [anti-intercellular adhesion molecule 1 (ICAM-1), ab222736, 1:1000; anti-The vascular cellular adhesion molecule-1 (VCAM-1), ab134047, 1:2000; anti-Tumour Necrosis Factor alpha (TNF alpha); ab183218, 1:1000; anti- interleukin-1beta (IL-1β), ab254360, 1:1000; anti-IL-6, ab259341, 1:1000; anti-C-X-C motif chemokine-2 (CXCL-2), ab271209, 1:1000; anti-UBE2G2, ab174296, 1:1000; anti-GAPDH, ab8245, 1:500] at 4 °C overnight. After rinsing with TBST, the membrane was incubated with the secondary antibody for 1 h at room temperature. ECL (Biosci, Wuhan, China) was used to visualize the protein bands on the membrane. Total protein levels were normalized to GAPDH and each sample was replicated triple.
Bioinformatic Analysis
The potential upstream targets of UBE2G2 are predicted by the starBase (https://starbase.sysu.edu.cn/) [22]. By reviewing the published literature to understand the functions of all predicted target genes, we selected miR-204-5p that might be associated with UBE2G2 for validation. Subsequently, the binding sites between miR-204-5p and UBE2G2 were provided by the PITA database [23].
Dual-Luciferase Reporter Gene Assay
The wild (wt) and mutated (mut) target fragments of UBE2G2 3′UTR were synthesized and cloned into pGL3 vector (Promega, Madison, WI, USA) to construct UBE2G2 3′UTR-wt and UBE2G2 3′UTR-mut reporter gene vectors, respectively. UBE2G2 3′UTR-wt and UBE2G2 3′UTR-mut reporter was co-transfected into VSMCs with miR-204-5p mimics (100 nM) or NC mimics (100 nM) using lipofectamine 3000 (Invitrogen). Forty-eight hours after transfection, dual-luciferase reporter assay system kit (Promega)was prepared to measure the luciferase activity.
5-Ethynyl-2′-Deoxyuridine (EdU) Assay
VSMCs were cultured in 96-well plate for 48 h and treated with 100 μL of medium containing 50 μM EdU for 2 h. The cells were then fixed with paraformaldehyde (4%; Beyotime) for 30 min and incubated with Triton-X-100 (0.5%) in PBS for 20 min. The nuclei were stained with DAPI. The proliferation rate was calculated according to the manufacturer’s instructions. Images were captured using a fluorescence microscope (80i; Nikon, Inc., Tokyo, Japan). Afterwards EdU-positive cells and total cells were counted within each field.
Wound Healing Assay
VSMCs were cultured in six-well plates at a density of 4 × 105 cells/well for a whole day. The wounds were then made by a sterile pipette tip (200 μL), and immediately washed three times with PBS. Cell migration to the wounded gap was then monitored by microscopy after 24 and 48 h. Image J software was used to visualize the distance on both sides of the scratch and take photos at different time points.
Statistical Analysis
All the corresponding experiments were independently repeated more than three times. The independent sample t-test was used for the comparison between 2 groups. One-way ANOVA followed by Tukey’s post hoc analysis was used for multiple groups. The correlation of UBE2G2 and miR-506-3p was analyzed using Spearman’s rank correlation analysis. Values of P < 0.05 were considered significant. We used SPSS 22.0 software to process the data.
Results
UBE2G2 Expression in Aortic Sinus Tissues
According to the results of RT-qPCR, UBE2G2 expression was obviously increased in aortic sinus tissues of ApoE−/− mice in comparison with that in the control group (P < 0.001). UBE2G2 was silenced in ApoE−/− mice using an adenovirus-associated virus (AAV)-mediated system, and the expression of UBE2G2 was sharply suppressed (P < 0.001), compared with that in ApoE−/− group (Fig. 1A). Similarly, immunohistochemistry staining of the aortic sinus section for UBE2G2 showed strong expression in ApoE−/− mice relative to that in control group. Inversely, UBE2G2 knockdown downregulated the UBE2G2 expression (Fig. 1B). Thus, The UBE2G2 expression was elevated in aortic sinus tissues of ApoE−/− mice with AS.
UBE2G2 expression in aortic sinus tissues. A Real-time reverse transcriptase-polymerase chain reaction (RT-qPCR) analysis of UBE2G2 expression in aortic sinus tissues of normal mice, high-fat diet (HFD)-fed ApoE−/− mice, and HFD-fed ApoE−/− mice with UBE2G2 knockdown. B Immunohistochemistry staining of aortic sinus tissues from the normal mice, HFD-fed ApoE−/− mice and HFD-fed ApoE−/− mice with UBE2G2 knockdown *P < 0.05, **P < 0.01, ***P < 0.001
Silencing of UBE2G2 Reduces Atherosclerotic Plaque Formation in ApoE−/− Mice
H&E staining revealed a significantly smaller atherosclerotic plaques characteristic features of AS in ApoE−/− + AAV-sh-UBE2G2 group, compared with that in ApoE−/− mice (Fig. 2A). Mice in the ApoE−/−-treated groups or in ApoE−/− + AAV-sh-UBE2G2 group had no significant changes in body weight compared with mice in the control group (Table 2). In addition, higher concentrations of TG, TC, HDL, and LDL was shown by the hematological analysis in the ApoE−/− group. However, silencing of UBE2G2 had no signifcant efects on blood lipid concentrations of AS mice (Table 3). Thus, UBE2G2 knockdown suppressed atherosclerotic plaque formation in ApoE−/− mice.
UBE2G2 Knockdown Inhibits the Inflammatory Pathway
Vascular inflammation is the early marker of AS, which is accompanied by elevated levels of the adhesion molecule and biomarkers of inflammation [24]. The mRNA expression and proteins levels of adhesion biomarkers and inflammatory cytokines (ICAM-1, VCAM-1, TNF-α, IL1β, IL-6, and CXCL-2) were promoted in ApoE−/− mice relative to those in control group. However, the silencing of UBE2G2 suppressed the expression of adhesion biomarkers and inflammatory cytokines, compared with that in ApoE−/− group (Fig. 3A, B). Collectively, UBE2G2 inhibition suppressed the inflammatory pathway in ApoE−/− mice.
MiR-204-5p Targets UBE2G2 3′UTR
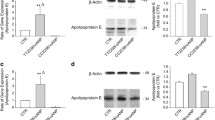
The starBase website predicted miR-204-5p as an upstream target gene of UBE2G2 [17]. We detected miR-204-5p expression in aortic sinus tissues of ApoE−/− mice with AS and found a significantly downregulated expression level in aortic sinus tissues of ApoE−/− mice (P < 0.01) (Fig. 4A). Spearman’s rank correlation analysis subsequently revealed that miR-204-5p expression was negatively correlated with UBE2G2 expression in aortic sinus tissues of ApoE−/− mice (Fig. 4B). Moreover, miR-204-5p expression was significantly elevated in VSMCs overexpressing miR-204-5p (Fig. 4C). While UBE2G2 mRNA expression and protein levels were downregulated in VSMCs overexpressing miR-204-5p (Fig. 4D, E). PITA predicted the binding sites and luciferase reporter assay confirmed it. The luciferase activity of UBE2G2 3′UTR-wt was obviously decreased in the cells overexpressing miR-204-5p relative to that in control group. While the luciferase activity of UBE2G2 3′UTR-mut remained unchanged (Fig. 4F, G). These results all indicated that miR-204-5p targeted UBE2G2 3′UTR.
MiR-204-5p targets UBE2G2 3′UTR. A RT-qPCR of miR-204-5p in aortic sinus tissues from HFD-fed ApoE−/− mice. B Spearman’s rank correlation analysis revealed the correlation between miR-204-5p expression and UBE2G2 expression in aortic sinus tissues of HFD-fed ApoE−/− mice. C RT-qPCR of miR-204-5p expression in VSMCs overexpressing miR-204-5p. D and E RT-qPCR and western blotting of UBE2G2 mRNA expression and protein levels in VSMCs overexpressing miR-204-5p. F and G PITA predicted the binding site between miR-204-5p and UBE2G2, and which is verified by luciferase reporter assay. **P < 0.01, ***P < 0.001
MiR-204-5p Inhibits the Proliferation and Migration of Human VSMCs by UBE2G2
The proliferation of VSMCs was inhibited by miR-204-5p overexpression. However, UBE2G2 overexpression partially reversed the inhibitory effect of miR-204-5p overexpression and promoted the VSMCs proliferation, as shown by EdU assay (Fig. 5A). Consistently, the migratory ability of VSMCs was inhibited by miR-204-5p overexpression, while UBE2G2 overexpression promoted the decreased wound heealing percentage induced by miR-204-5p overexpression in VSMCs (Fig. 5B). Collectively, miR-204-5p inhibited the proliferation and migration of human VSMCs by regulating UBE2G2 expression.
Discussion
Accumulating studies have shown that various inflammatory cells and a large number of inflammatory mediators are always involved from lipid streaks to fibrous plaques and atherosclerotic plaques during the occurrence and development of AS lesions [25]. As a chronic inflammatory disease, AS involves many different cells and molecules.
Recently, ubiquitin proteasome system is proved to affect numerous cardiovascular processes [26]. As part of the ubiquitin proteasome system, enzymes of three different types (E1, E2, and E3) have the key role of directing the addition of ubiquitin to specific target proteins [26]. Ubiquitin-conjugating enzyme E2 relevant proteins have been found to be associated with AS development like UBE2Z [27], UBE2J2 [28], and UBE2G2 [13], which has been predicted as a novel biomarker for treating AS [12]. In our study, ApoE−/− mice with AS were established to explore the concrete role and mechanism of UBE2G2. Interestingly, UBE2G2 expression was obviously increased in aortic sinus tissues of ApoE−/− mice, and UBE2G2 knockdown suppressed atherosclerotic plaque formation, as well as the expression of adhesion biomarkers and inflammatory cytokines in ApoE−/− mice with AS. Thus, we demonstrated for the first time the protective effect of UBE2G2 knockdown on atherosclerotic plaque formation and the inflammatory response in HFD-fed ApoE−/− mice.
Mechanically, miRNA exerts its role in the occurrence and development of diseases by cleaving mRNA and inhibiting its translation [29, 30]. Importantly, increasing miRNAs are reported to be crucial regulators for the abnormal proliferation and migration of VSMCs, such as miR-377-3p [31], miR-128-3p [32], and miR-126-5p [33]. In our study, miR-204-5p has been predicted as an upstream target gene of UBE2G2. Previous study has reported that miR-204-5p exerted its effect on the growth and migration of VSMCs by targeting MMP-9 [18]. Another research also confirmed the the downregulation of miR-204-5p in endothelial cells of a rabbit model was associated with AS [21]. In addition, miR-204-5p is also involved in the regulatory mechanism of X-inactive specific transcript (XIST)/toll-like receptor 4 (TLR4) and plays a protective role in ox-LDL-induced human umbilical vein endothelial cells injury, thus inhibiting the development of AS [34]. Notably, in our study, miR-204-5p expression was decreased in aortic sinus tissues of ApoE−/− mice with AS and was negatively correlated with UBE2G2 expression. Interestingly, miR-204-5p overexpression inhibited the proliferation and migration of human VSMCs. However, UBE2G2 overexpression significantly reversed the effect of miR-204-5p mimics.
Our research revealed the potential role of UBE2G2 in AS. However, some limitations still exist in this study. First, the effect of its upstream target miR-204-5p on AS development needs to be confirmed through establishing AS mice model in the subsequent experiments. Secondly, many signaling pathways including p38, ERK1/2, and JNK pathway have been found to participate in the AS development and mediate the inflammation process in AS [35]. We should further find out the relevant signaling pathways that affect the role of UBE2G2.
Conclusion
In conclusion, inhibition of UBE2G2 protected ApoE−/− mice from AS. We also found that miR-204-5p targets UBE2G2, and miR-204-5p overexpression regulated proliferation and migration of human VSMCs via UBE2G2 axis. Thus, UBE2G2 may be a potential biomarker for the prevention of AS.
Data Availability
The datasets generated and/or analyzed during the current study are not publicly available due research design, but are available from the corresponding author on reasonable request.
Change history
03 October 2022
A Correction to this paper has been published: https://doi.org/10.1007/s12033-022-00579-w
References
Libby, P. (2002). Inflammation in atherosclerosis. Nature, 420(6917), 868–874. https://doi.org/10.1038/nature01323
Glass, C. K., & Witztum, J. L. (2001). Atherosclerosis. The road ahead. Cell, 104(4), 503–516. https://doi.org/10.1016/s0092-8674(01)00238-0
Ross, R. (1999). Atherosclerosis–an inflammatory disease. The New England Journal of Medicine, 340(2), 115–126. https://doi.org/10.1056/nejm199901143400207
Wan, Q., Liu, Z., & Yang, Y. (2018). Puerarin inhibits vascular smooth muscle cells proliferation induced by fine particulate matter via suppressing of the p38 MAPK signaling pathway. BMC Complementary and Alternative Medicine, 18(1), 146. https://doi.org/10.1186/s12906-018-2206-9
Rask-Madsen, C., Li, Q., Freund, B., Feather, D., Abramov, R., Wu, I. H., Chen, K., Yamamoto-Hiraoka, J., Goldenbogen, J., Sotiropoulos, K. B., Clermont, A., Geraldes, P., Dall’Osso, C., Wagers, A. J., Huang, P. L., Rekhter, M., Scalia, R., Kahn, C. R., & King, G. L. (2010). Loss of insulin signaling in vascular endothelial cells accelerates atherosclerosis in apolipoprotein E null mice. Cell Metabolism, 11(5), 379–389. https://doi.org/10.1016/j.cmet.2010.03.013
Rose, S. A., Leek, J. P., Moynihan, T. P., Ardley, H. C., Markham, A. F., & Robinson, P. A. (1998). Assignment1 of the ubiquitin conjugating enzyme gene, UBE2G2, to human chromosome band 21q22.3 by in situ hybridization. Cytogenetics and Cell Genetics, 83(1–2), 98–99. https://doi.org/10.1159/000015141
Katsanis, N., & Fisher, E. M. (1998). Identification, expression, and chromosomal localization of ubiquitin conjugating enzyme 7 (UBE2G2), a human homologue of the Saccharomyces cerevisiae ubc7 gene. Genomics, 51(1), 128–131. https://doi.org/10.1006/geno.1998.5263
Reyes, L. F., Sommer, C. A., Beltramini, L. M., & Henrique-Silva, F. (2006). Expression, purification, and structural analysis of (HIS)UBE2G2 (human ubiquitin-conjugating enzyme). Protein Expression and Purification, 45(2), 324–328. https://doi.org/10.1016/j.pep.2005.08.018
Atkin, A. S., Moin, A. S. M., Nandakumar, M., Al-Qaissi, A., Sathyapalan, T., Atkin, S. L., & Butler, A. E. (2021). Impact of severe hypoglycemia on the heat shock and related protein response. Scientific Reports, 11(1), 17057. https://doi.org/10.1038/s41598-021-96642-8
Atkin, A. S., Moin, A. S. M., Al-Qaissi, A., Sathyapalan, T., Atkin, S. L., & Butler, A. E. (2021). Plasma heat shock protein response to euglycemia in type 2 diabetes. BMJ Open Diabetes Research & Care. https://doi.org/10.1136/bmjdrc-2020-002057
Filali-Mouncef, Y., Hunter, C., Roccio, F., Zagkou, S., Dupont, N., Primard, C., Proikas-Cezanne, T., & Reggiori, F. (2021). The ménage à trois of autophagy, lipid droplets and liver disease. Autophagy. https://doi.org/10.1080/15548627.2021.1895658
Hebbel, R. P., Wei, P., Milbauer, L., Corban, M. T., Solovey, A., Kiley, J., Pattee, J., Lerman, L. O., Pan, W., & Lerman, A. (2020). Abnormal endothelial gene expression associated with early coronary atherosclerosis. Journal of the American Heart Association, 9(14), e016134. https://doi.org/10.1161/jaha.120.016134
Robichaud, S., Fairman, G., Vijithakumar, V., Mak, E., Cook, D. P., Pelletier, A. R., Huard, S., Vanderhyden, B. C., Figeys, D., Lavallée-Adam, M., Baetz, K., & Ouimet, M. (2021). Identification of novel lipid droplet factors that regulate lipophagy and cholesterol efflux in macrophage foam cells. Autophagy. https://doi.org/10.1080/15548627.2021.1886839
Mohr, A. M., & Mott, J. L. (2015). Overview of microRNA biology. Seminars in Liver Disease, 35(1), 3–11. https://doi.org/10.1055/s-0034-1397344
Pasquinelli, A. E. (2012). MicroRNAs and their targets: Recognition, regulation and an emerging reciprocal relationship. Nature Reviews Genetics, 13(4), 271–282. https://doi.org/10.1038/nrg3162
Tian, S., Yuan, Y., Li, Z., Gao, M., Lu, Y., & Gao, H. (2018). LncRNA UCA1 sponges miR-26a to regulate the migration and proliferation of vascular smooth muscle cells. Gene, 673, 159–166. https://doi.org/10.1016/j.gene.2018.06.031
Witkos, T. M., Koscianska, E., & Krzyzosiak, W. J. (2011). Practical aspects of microRNA target prediction. Current Molecular Medicine, 11(2), 93–109. https://doi.org/10.2174/156652411794859250
Wang, N., Yuan, Y., Sun, S., & Liu, G. (2020). microRNA-204-5p Participates in Atherosclerosis Via Targeting MMP-9. Open Medicine, 15, 231–239. https://doi.org/10.1515/med-2020-0034
Zhang, F., Zhang, R., Zhang, X., Wu, Y., Li, X., Zhang, S., Hou, W., Ding, Y., Tian, J., Sun, L., & Kong, X. (2018). Comprehensive analysis of circRNA expression pattern and circRNA-miRNA-mRNA network in the pathogenesis of atherosclerosis in rabbits. Aging, 10(9), 2266–2283. https://doi.org/10.18632/aging.101541
He, X., & Deng, L. (2021). miR-204-5p inhibits inflammation of synovial fibroblasts in osteoarthritis by suppressing FOXC1. Journal of Orthopaedic Science. https://doi.org/10.1016/j.jos.2021.03.014
Wang, Z., Zhang, M., Wang, Z., Guo, Z., Wang, Z., & Chen, Q. (2020). Cyanidin-3-O-glucoside attenuates endothelial cell dysfunction by modulating miR-204-5p/SIRT1-mediated inflammation and apoptosis. BioFactors, 46(5), 803–812. https://doi.org/10.1002/biof.1660
Li, J. H., Liu, S., Zhou, H., Qu, L. H., & Yang, J. H. (2014). starBase v2.0: Decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Research, 42, D92-97. https://doi.org/10.1093/nar/gkt1248
Kertesz, M., Iovino, N., Unnerstall, U., Gaul, U., & Segal, E. (2007). The role of site accessibility in microRNA target recognition. Nature Genetics, 39(10), 1278–1284. https://doi.org/10.1038/ng2135
Libby, P. (2012). Inflammation in atherosclerosis. Arteriosclerosis, Thrombosis, and Vascular Biology, 32(9), 2045–2051. https://doi.org/10.1161/atvbaha.108.179705
Libby, P., Ridker, P. M., & Hansson, G. K. (2009). Inflammation in atherosclerosis: From pathophysiology to practice. Journal of the American College of Cardiology, 54(23), 2129–2138. https://doi.org/10.1016/j.jacc.2009.09.009
Willis, M. S., Bevilacqua, A., Pulinilkunnil, T., Kienesberger, P., Tannu, M., & Patterson, C. (2014). The role of ubiquitin ligases in cardiac disease. Journal of Molecular and Cellular Cardiology, 71, 43–53. https://doi.org/10.1016/j.yjmcc.2013.11.008
Rodríguez-Pérez, J. M., Posadas-Sánchez, R., Vargas-Alarcón, G., Blachman-Braun, R., García-Flores, E., Cazarín-Santos, B. G., Castillo-Avila, R. G., Borgonio-Cuadra, V. M., Tovilla-Zárate, C. A., González-Castro, T. B., López-Bautista, F., & Pérez-Hernández, N. (2020). The rs46522 polymorphism of the ubiquitin-conjugating enzyme E2Z gene is associated with abnormal metabolic parameters in patients with myocardial infarction: The genetics of atherosclerosis disease mexican study. DNA and Cell Biology, 39(7), 1155–1161. https://doi.org/10.1089/dna.2020.5477
Tan, J. M. E., Cook, E. C. L., van den Berg, M., Scheij, S., Zelcer, N., & Loregger, A. (2019). Differential use of E2 ubiquitin conjugating enzymes for regulated degradation of the rate-limiting enzymes HMGCR and SQLE in cholesterol biosynthesis. Atherosclerosis, 281, 137–142. https://doi.org/10.1016/j.atherosclerosis.2018.12.008
Chen, K., & Rajewsky, N. (2007). The evolution of gene regulation by transcription factors and microRNAs. Nature Reviews Genetics, 8(2), 93–103. https://doi.org/10.1038/nrg1990
Lewis, B. P., Burge, C. B., & Bartel, D. P. (2005). Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell, 120(1), 15–20. https://doi.org/10.1016/j.cell.2004.12.035
Wang, H., Wei, Z., Li, H., Guan, Y., Han, Z., Wang, H., & Liu, B. (2020). MiR-377-3p inhibits atherosclerosis-associated vascular smooth muscle cell proliferation and migration via targeting neuropilin2. Bioscience Reports. https://doi.org/10.1042/bsr20193425
Farina, F. M., Hall, I. F., Serio, S., Zani, S., Climent, M., Salvarani, N., Carullo, P., Civilini, E., Condorelli, G., Elia, L., & Quintavalle, M. (2020). miR-128-3p is a novel regulator of vascular smooth muscle cell phenotypic switch and vascular diseases. Circulation Research, 126(12), e120–e135. https://doi.org/10.1161/circresaha.120.316489
Chen, Z., Pan, X., Sheng, Z., Yan, G., Chen, L., & Ma, G. (2019). Baicalin suppresses the proliferation and migration of Ox-LDL-VSMCs in atherosclerosis through upregulating miR-126-5p. Biological & Pharmaceutical Bulletin, 42(9), 1517–1523. https://doi.org/10.1248/bpb.b19-00196
Lu, G., Tian, P., Zhu, Y., Zuo, X., & Li, X. (2020). LncRNA XIST knockdown ameliorates oxidative low-density lipoprotein-induced endothelial cells injury by targeting miR-204–5p/TLR4. Journal of Biosciences. https://doi.org/10.1007/s12038-020-0022-0
Yang, Y., Pei, K., Zhang, Q., Wang, D., Feng, H., Du, Z., Zhang, C., Gao, Z., Yang, W., & Wu, J. (1865). Li Y (2020) Salvianolic acid B ameliorates atherosclerosis via inhibiting YAP/TAZ/JNK signaling pathway in endothelial cells and pericytes. Biochimica et Biophysica Acta Molecular and Cell Biology of Lipids, 10, 158779. https://doi.org/10.1016/j.bbalip.2020.158779
Acknowledgements
Not applicable.
Funding
No funding was received.
Author information
Authors and Affiliations
Contributions
All authors were the main designers of this study. YL, ZL, ZW, KL, LL, CW, and YX performed the experiments. YL, ZL, and ZW analyzed the data. YL drafted the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical Approval
All animal experimental procedures were approved by the Animal Care and Use Committee of the Affiliated Zhangjiagang Hospital of Soochow University (Jiangsu, China) according to institutional guidelines for animal ethics.
Informed Consent
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, Y., Luo, Z., Wu, Z. et al. The Protective Effect of UBE2G2 Knockdown Against Atherosclerosis in Apolipoprotein E-Deficient Mice and Its Association with miR-204-5p. Mol Biotechnol 64, 1045–1054 (2022). https://doi.org/10.1007/s12033-022-00482-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-022-00482-4