Abstract
Cellular transcriptomes are frequently adorned by a variety of chemical modification marks, which in turn have a profound influence on its functioning. Of these modifications, the one which has invited a lot of attention in the recent years is m6A RNA methylation, leading to the development of RNA epigenetics or epitranscriptomics as a frontier research area. m6A RNA methylation is one of the most abundant reversible internal modification seen in cellular RNAs. Studies in the last few years have not only shed light on the molecular machinery involved in m6A RNA methylation but also on the impact of this modification in regulating gene expression and hence biological processes. In this review, we will emphasize the biological impact of this modification in normal organismal development and diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The flow of genetic information in biological systems from DNA to RNA to protein is explained by the central dogma of molecular biology, which forms the backbone of modern biology [1]. This flow of genetic information is with checks and balances and is often subjected to a wide variety of regulatory controls operational at multiple levels. Adding to this regulatory complexity is the reversible chemical modification of DNA and nucleosomal histones, which are collectively termed as epigenetic marks or signatures. One such well-studied epigenetic modification is DNA methylation, wherein the enzyme DNA methyl transferase adds the methyl group to cytosine in the CpG islands of DNA sequence, leading to transcriptional silencing [2]. The loss of DNA methylation, i.e., hypomethylation has been reported to promote tumorigenicity in various cancer types [3, 4]. Aside from DNA, the tails of nucleosomal histone proteins, which are essential for chromatin formation, are often subjected to numerous chemical modifications like acetylation, deacetylation, methylation, phosphorylation and ubiquitination [5]. These modifications are catalyzed by specific enzymes, for example, histone acetyl transferase and deacetylase, histone methyl transferase and demethylase facilitate acetylation, deacetylation, methylation and demethylation of histones respectively. Acetylation of histones 3 and 4 at specific amino acid residues, activate transcription by loosening up the chromatin, whereas methylation of histones 3 and 4 at specific positions can activate transcription in certain instances and repress in others [6].
Cellular RNAs, coding as well as non-coding, since their birth by transcription, have been known to be subjected to > 100 different chemical modifications, most irreversible or static in nature, such as 5′ cap, pseudouridine, 5-hydroxymethylcytosine, 5-methylcytosine to name a few, which often impacts their structure as well as function [7]. Intriguingly about two-thirds of these chemical modifications involve addition of methyl group and the structure and topology of the most common methylated nucleosides reported in eukaryotic messenger RNAs (mRNA) are illustrated in Fig. 1. The list includes N7-methylguanosine (m7G), which aids in translation, splicing, nuclear export and prevents degradation; 2′-O-methylation (Nm), which modulates translation efficiency; 5-methyl cytosine (m5C), which regulates translation and nuclear export; N1-methyladenosine (m1A), which impacts translation; N6-methyladenosine (m6A), which is critical for various aspects of mRNA metabolism; and 3-methylcytidine (m3C), whose function is yet to be uncovered [8]. Of these methylation events, the one that has attracted immense attention and is the focus of this review is the chemical modification termed as ‘N6-methyladenosine’ or ‘m6A’, as it adds a methyl group to the nitrogenous base at the sixth position of the adenine residue of the RNA. m6A RNA methylation has emerged as one of the most abundant internal modification of RNAs and was first discovered in eukaryotic messenger RNAs in Novikoff hepatoma cells [9] and mouse L cells [10]. However, the precise function of m6A mark remained elusive and it was just regarded as any other post transcriptional modification, presumed to have a role in RNA processing. The dynamic and reversible nature of this RNA modification came to the fore, when a demethylating enzyme, FTO (fat mass and obesity-associated protein) which removes the methyl group from RNA was reported [11]. The reversibility of m6A mark, unlike other previously known modifications, which were irreversible in nature, gives it an added flexibility desired for regulating gene expression. Not only in messenger RNAs, m6A has been reported to be present in all classes of cellular RNAs, be it ribosomal RNAs, transfer RNAs and various non-coding RNAs like microRNAs, long non-coding RNAs, circular RNAs [12,13,14,15]. m6A RNA methylation has emerged as a key regulator of various post transcriptional gene regulatory processes like RNA splicing, stability, export, degradation and also as a signal for translational initiation machinery to act upon it to start protein synthesis [16]. Various reports also have shown the importance of this reversible RNA methylation in development and as well diseases like cancer, diabetes [17,18,19,20]. m6A modification is evolutionarily conserved, but only adenosine residues present in the consensus motif, RRACH (where R = G/A, H = A/C/U, underlined letter stands for adenosine being modified to m6A) are methylated [21]. Topologically, this modification is predominantly clustered in regions near 3′-UTR, stop codons [22, 23], long internal exons and 5′-UTR [24]. It was also observed in introns [25] and are added to nascent pre-mRNA associated with chromatin for their cytoplasmic turnover rate [26].
Spectrum of methylated nucleosides in eukaryotic messenger RNA. Top panel depicts the chemical structure of the various methylated nucleosides. Bottom panel highlights the distribution of the methylated nucleosides to the mRNA cap structure, 5′ or 3′ untranslated region (UTR) or the coding region of mRNA. m7G (7-methylguanosine); Nm (2′-O-methylation); m5C (5-methylcytosine); m1A (N1-methyladenosine); m6A (N6-methyladenosine); m3C (3-methylcytidine)
The in-depth study of m6A pathway was revolutionized in 2012 by two research groups in parallel, wherein they developed technique for mapping and detecting m6A methylome, referred to as m6A-seq or MeRIP-seq, i.e., immunoprecipitation of RNAs with m6A mark using an anti-m6A antibody, followed by next-generation sequencing [22, 27, 28]. This was further facilitated by development of methods such as PA-m6A-SEQ (photo-crosslinking-assisted m6A-sequencing) [29], SCARLET (site-specific cleavage and radioactive-labeling followed by ligation-assisted extraction and thin-layer chromatography) [30] and LC–MS/MS (liquid chromatography linked to tandem mass spectrometry) [31]. In addition, bioinformatics tools such as SRAMP [32], TargetM6A [33], RNA-methylPred [34], iRNA-methyl [35] and pRNAm-PC [36] have been helpful in predicting m6A sites. Taken together these technological advancements for detection and prediction of m6A sites have made a huge impact on the nascent field of RNA epigenetics or epitranscriptomics [37,38,39].
A multi-subunit methyl transferase complex, i.e., m6A ‘writers’, adds the methyl group to adenosine; demethylases, i.e., m6A ‘erasers’ aids in the removal of methyl group and specific RNA binding proteins, i.e., m6A ‘readers’ recognize the m6A deposits and bind to the m6A methylated RNA and modulate specific downstream functions. In this review, we attempt to summarize the key players involved in m6A RNA methylation pathway and their implications in human health.
m6A ‘Writers’
m6A ‘writers’ are complex of methyl transferases, which install the m6A mark onto specific sites on RNA (Fig. 2). Using a combination of labeling and chromatographic techniques, the presence of internal methylated adenosines in eukaryotic messenger RNAs was first demonstrated [9, 24]. Following this, employing in vitro methylation together with mutational studies, the consensus motif associated with m6A deposition was identified to be ‘RRACH’ (where R = G/A, H = A/C/U, underlined letter stands for adenosine being modified to m6A) motif and the frequency of deposition was ~ 1 to 3 methyl group per mRNA molecule, primarily in the 3′ end of mRNA [21, 40,41,42]. The first indication for the existence of a RNA methyl transferase in nucleus was provided by in vitro methylation of prolactin mRNA upon incubation with HeLa nuclear extract in the presence of S-adenosyl methionine (SAM) [41]. However, the signals which regulate the deposition of m6A were not fully understood till 2018, when Bertero et al., for the first time showed that TGFβ can modulate m6A dynamics in human stem cells [43]. In another work published in 2017, R2 hydroxyglutarate (R2HG), an oncometabolite having anti-tumor activity in certain leukemia and gliomas, inhibits the demethylase, FTO, leading to increased m6A levels and decrease in stability of CEBPA and MYC transcripts [44]. Taken together, these reports, shed some light on the external and internal cues regulating the dynamics and specificity of m6A methylation.
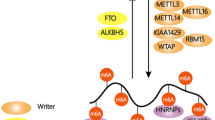
Schematic representation of reversible m6A RNA methylation and m6A-mediated regulation of gene expression. m6A ‘writer’ complex or the methyl transferase complex is composed of METTL3/METTL14/WTAP along with other auxiliary proteins such as VIRMA, ZC3H13 and RBM15; facilitates the deposition of m6A methyl mark on cellular RNAs in the nucleus; the writer METTL16 is a specific methyl transferase for U6 small nuclear RNA. m6A ‘readers’ or m6A binding proteins consist primarily of proteins such as YTHDF1-3, YTHDC2 and eIF3, which are cytoplasmic readers, and HNRNPA2B1, HNRNPC, HNRNPG and YTHDC1, which are nuclear readers. m6A readers binds to the target RNAs in a m6A-dependent manner and determine the downstream fate of the transcripts by regulating various steps of RNA metabolism such as splicing, 3′-end processing, primary microRNA (pri-miRNA) processing, nuclear export, degradation, stability and translation. m6A readers act in conjunction with specific proteins to elicit their function, for example SRSF3 facilitates the nuclear export by serving as an adaptor between the export protein NXF1 and reader YTHDC1; SRFSF3/10 facilitate m6A-dependent splicing functions of the reader YTHDC1. m6A ‘erasers’ or demethylase consist of FTO and ALKBH5, both of which oxidatively remove the m6A signatures from the transcripts in the nucleus, thus making the process reversible
Methyl Transferase like 3 (METTL3)
Bokar et al. uncovered different components of the SAM-dependent methyl transferase complex (MTase complex), using classical biochemical techniques and named them as MT-A1 (30 kDa), MT-A2 (200 kDa) and MT-B (875 kDa) [42]. The 200 kDa fragment of the complex, now termed as methyl transferase like 3 (METTL3) has a SAM-binding site and is essential for m6A methylation activity [42, 45]. METTL3 functions as heterodimer with methyl transferase like 14 (METT14), and exists in a multi-subunit complex with other auxiliary factors like Wilm’s tumor associated protein (WTAP). METTL3 via its impact on m6A deposition and hence RNA homeostasis, has been reported to influence normal organismal development and several human diseases like cancer (Table 1).
m6A deposition often serves as molecular switches, referred to as ‘m6A switch’, by facilitating the binding of specific RNA binding proteins and such interactions are important for cellular RNA transactions [46, 47]. During development, METTL3 and METTL14 were implicated in regulating cell fate transition in embryonic stem cells [48, 49] and their knockdown led to embryonic lethality in mice [50]. METTL3 also played role in regulation of osteogenic differentiation [51,52,53] and may be a potential therapeutic target in reversing osteoporosis condition [54]. Interestingly, it was also reported to enhance long term memory in mice [55] and mediate T-cell homeostasis [56]. METTL3 was also shown to have role in autophagy of cardiomyocytes, wherein the enzymes with opposing activities such as methyl transferase, METTL3 and demethylase, ALKBH5 were demonstrated to regulate autophagy via TFEB, which in turn regulated the expression of both METTL3 and ALKBH5 [57]. Inhibition of m6A methylation either by knockdown of METTL3 or depletion of consensus sequences had an impact on the expression level of circadian clock genes [58, 59]. Furthermore, METTL3 also promotes processing of primary microRNA in a m6A-dependent manner and its depletion shows global reduction in mature microRNAs [60]. METTL3 also facilitates translation of mRNA in a m6A-dependent manner and promotes a variety of cancer types [61,62,63,64,65,66]. Not only linear RNAs, METTL3-dependent m6A marks were found extensively in a subset of circular RNAs and were reported to promote translation initiation of the circular RNAs [14, 15].
Methyl Transferase like 14 (METTL14)
Results on structure-based computational studies from two different groups revealed the existence of methyl transferase like 14 (METTL14), an additional m6A methyl transferase bearing significant sequence and structural homology with METTL3; however, the functions of METTL14 remained elusive [67, 68]. Study on m6A RNA methylation employing mouse embryonic stem cells demonstrated that m6A writers function as a multi-subunit complex with METTL14 and METTL3 acting in tandem as heterodimers to deposit m6A on target genes and knocking down both, led to derailment of m6A methylation in mRNA transcripts and loss of self-renewal ability [49]. The interaction between METTL3 and METTL14 was further reinforced by crystal structure-based analysis, whereby METTL14 was shown to be required for binding of METTL3 to its RNA substrate [69, 70]. Further to this, Schöller et al., in 2018, provided insight into the overall architecture of this heterodimer complex and their interactions and requirement of splicing regulator WTAP (Wilm’s tumor-1 associated protein) for their nuclear localization. This work also shed some light on function of METTL14, particularly the role of carboxyl terminus RGG domain [71]. In higher eukaryotes, the existence of homologs of METTL14 and other proteins involved in mammalian m6A RNA methylation showcased their evolutionary and functional significance [72].
METTL14 impacts developmental processes and a plethora of human diseases (Table 1). The deletion of METTL14 in mouse highlighted its role in early embryonic development [73] and spermatogenesis [74]. Neuron-specific deletion of METTL14 in adult mice brain had a profound effect on its learning ability, which gave further validation to the importance of the m6A mRNA modification during embryonic and postnatal development [75]. METTL14 deletion in neuron stem cell (NSC) in a mouse model displayed severe proliferation and differentiation changes affecting normal neuronal development [76]. Aberrant regulation of METTL14 was implicated in tumor metastasis in hepatocellular carcinoma [77] and malignant hematopoiesis [78]. Strikingly, a cross-talk between histone modification and m6A RNA methylation was revealed in 2019 by Huang et al., where it was reported that histone H3 trimethylation at lysine 36, a mark for transcriptional elongation, directly binds to METTL14 and guides the methyl transferase complex to nascent transcripts for m6A deposition [79].
Wilm’s Tumor-1 Associated Protein (WTAP) and Other Writers
Wilm’s tumor-1 associated protein (WTAP) first came into picture as an interaction partner of Wilm’s tumor-1 (WT1) protein [80]. WTAP was first identified as a regulatory subunit of m6A methyl transferase complex and its knockdown inhibited methyl transferase activity of METTL3-METTL14 complex demonstrating the importance of WTAP in m6A installation [81].
Extensive work done to probe the protein–protein interactions, led to the discovery of other protein partners involved in this dynamic modification, like KIAA1429 or VIRMA (Vir like m6A methyl transferase associated), which is the human homolog of Drosophila’s Virilizer protein) [82, 83], RBM15 (RNA binding protein 15) [84] which interact with methyl transferase complex via WTAP. Furthermore, RBM15, interacts with BAF155 subunit of chromatin remodeling complex BAF and recruits m6A methyl transferase complex to BAF155 to modulate its stability and regulate cortical development [85]. RBM15 is also critical for deposition of m6A by METTL3 on X-inactive specific transcript (XIST), which promotes XIST-mediated transcriptional repression [84]. Another component of the complex KIAA0853 or ZC3H13, is a zinc finger protein, which was identified in mouse embryonic stem cell and functions as a recruiter protein facilitating localization of methyl transferase complex to nucleus and its ablation led to loss of self-renewing ability of the mouse embryonic stem cells due to failure of the methyl transferase complex to reach the nucleus [86]. WTAP and other m6A writers, too, have a role in various human pathologies (Table 1).
Methyl Transferase like 16 (METTL16)
Aside from METTL3-METTL14-WTAP m6A writer complex, mammalian cell bear additional methyl transferase—METTL16, which is involved in m6A deposition on a limited subset of RNAs and U6 small nuclear RNA [87]. Interestingly, METTL16, modulates SAM homeostasis by depositing m6A in 3′-UTR of SAM synthetase (MAT2A) transcript and regulates its expression through altered splicing and stability [88]. Using crosslinking studies, m6A methylation targets of METTL16 like long non-coding RNA, U6 snRNA and mRNAs were deciphered [89]. In contrast to METTL3-METTL14 heterodimer complex, METTL16 works as a homodimer [90] and mouse embryo lacking METTL16 showed severe deregulation in transcriptomes and embryonic lethality [91].
m6A ‘Readers’
Gene regulatory impact of m6A deposition are mediated by specific proteins termed as m6A ‘readers’, which read the m6A signature in the RNA transcript and elicit the downstream response and fate of the transcript either in nucleus or cytoplasm (Fig. 1). Dominissini et al. presented the first evidence for the existence of RNA binding proteins that recognize and bind to the m6A mark in RNAs [27]. A majority of the mammalian m6A ‘readers’ bear the YTH (YT521-B homology) domain, which has an aromatic cage for specifically accommodating the methyl mark. Aside from this, numerous other proteins, which lack the YTH domain, can also function as m6A ‘readers’ like IGF2BP, HNRNP, eIF3 to name a few. Facilitated by structural studies, YTH family of proteins were suggested to bind to m6A signature [92] with a preferential binding to the consensus sequence, GG(m6A)C [93]. Like m6A writers, most of the reader proteins show an aberrant expression pattern in a variety of human diseases and have a role in disease progression and facilitation (Table 1).
YTH Domain-Containing Family Protein 1 (YTHDF1)
The cytoplasmic m6A ‘reader’ protein, YTHDF1, binds to m6A marks on transcripts and promotes protein synthesis through its interaction with the translation initiation machinery, particularly the translation initiation factor, eIF3 [94]. YTHDF1 was reported to regulate translation of several m6A marked neuronal transcripts like ROBO3.1, which helps in axon guidance during neuronal development [95]. Furthermore, it also enabled the translation of certain target transcripts in response to neuronal excitation in mouse brain’s hippocampal region and thus influencing learning and memory processes [96]. Notably, pluripotency of induced pluripotent stem cells (iPSCs) was regulated in a m6A-dependent manner by YTHDF1, via binding and promoting translation of JAK2 transcript, to maintain the stemness [97].
YTH Domain-Containing Family Protein 2 (YTHDF2)
Chuan He’s lab first shed light on the functional relevance of the cytoplasmic reader protein, YTHDF2, highlighting its role as a regulator of mRNA stability by promoting degradation of m6A marked transcripts and reported its co-localization with decay proteins like DCP1A, GW182 and DDX6 in P-bodies [98]. Furthermore, YTHDF2 interacts with CNOT1 leading to the recruitment of CCR4-NOT complex and subsequent deadenylation and degradation of m6A marked transcripts [99]. Structural studies have further illuminated the binding of this reader protein to methylated adenosine on its target RNAs [100, 101]. YTHDF2 has functions in development and disease and this has been emphasized by multiple studies. Interestingly, in response to heat shock stress, YTHDF2 moves to the nucleus and protects the m6A marks in 5′-UTR of stress-induced transcripts from demethylation, which subsequently leads to their enhanced translation [102]. YTHDF2 functions in adult stem cell maintenance and in ex vivo expansion of hematopoietic stem cells (HSC), by regulating the stability of many mRNAs crucial for HSC self-renewal [103]. Its deletion causes embryonic lethality in mice and in heterozygous condition exhibited severe neural deregulation affecting embryonic brain development [104]. In zebrafish and mice model, YTHDF2 deficiency has been reported to cause dysregulations in maternal to zygotic transition process [105] and female specific infertility respectively [106]. Like other m6A players, YTHDF2 has also been reported to have aberrant expression pattern in a plethora of pathological conditions such as hepatocellular carcinoma [107], prostate cancer [108] and in liver cancer [65] (Table 1).
YTH Domain-Containing Family Protein 3 (YTHDF3)
The cytoplasmic m6A reader, YTHDF3, works in close co-operation with the other cytoplasmic readers, YTHDF1 and YTHDF2, in deciding the fate of m6A bearing transcripts and often there is an overlap between targets of YTHDF3 with targets of YTHDF1 and YTHDF2 [109, 110]. It acts in co-operation with YTHDF1 and promotes translation efficiency of the targets, and by mediating decay of target transcripts, its synergy with YTHDF2 is displayed [109].
YTH Domain-Containing Protein 1 (YTHDC1)
YTHDC1 is a nuclear m6A reader, unlike other members of this protein family, which are cytoplasmic readers. It modulates alternative splicing of target transcripts via exon inclusion in a m6A-dependent manner, by binding and recruiting the splicing factor, SRSF3 to pre-mRNAs and in turn blocking SRSF10 from binding [111]. Apart from pre-mRNA splicing regulation, YTHDC1 also functions in nuclear mRNA export of methylated target mRNAs, by facilitating their binding to nuclear export factor 1 (NXF1) in a SRSF3-dependent manner [112]. Interestingly, YTHDC1 plays a pivotal role in maintaining intracellular levels of SAM, via degradation of m6A marked methionine adenosyltransferase 2A (MAT2A) transcripts, wherein the m6A deposition is catalyzed by the writer, METLL16, in a SAM-dependent manner [113]. Furthermore, this reader protein is essential for mice embryo development and fertility, particularly oocyte growth and maturation, via its impact on alternative splicing and polyadenylation, in a m6A-dependent manner [114]. In addition to this, YTHDC1 mediates XIST-mediated transcriptional silencing by binding to methylated XIST long non-coding RNA [84].
YTH Domain-Containing Protein 2 (YTHDC2)
The cytoplasmic reader protein, YTHDC2, is characterized by the presence of 3′–5′ RNA helicase activity, in addition to the YTH domain and plays an essential role in male and female fertility in mice via regulation of methylated transcripts to modulate meiosis in germ cells [115, 116]. YTHDC2 exerts its regulatory impact by enhancing translation efficiency of its targets by interacting with the ribosomal small subunit as well as by destabilization and degradation of the target transcripts via interacting with and recruiting 5′–3′ exoribonuclease, XRN1 to the transcripts [117]. The expression of YTHDC2 is elevated in several cancer cell lines and its role in colon tumor metastasis was reported [118] (Table 1).
Insulin-like Growth Factor 2 Binding Protein (IGF2BP)
As opposed to the YTH domain-containing m6A readers, the insulin-like growth factor 2 mRNA-binding proteins (IGF2BPs; IGF2BP1/2/3) represent a distinct family of m6A readers which target a large number of transcripts, by selectively binding to the m6A mark via their K homology domain [119]. Mechanistically, unlike the mRNA destabilization function of the reader, YTHDF2, IGF2BPs promotes mRNA stability and translation of its targets [119]. IGF2BPs play critical oncogenic roles in cancer cells, by stabilizing m6A methylated mRNAs of oncogenic targets like MYC [119].
Heterogeneous Nuclear Ribonucleoproteins (hnRNPs)
HnRNPs are a large family of RNA binding proteins having roles in multiple facets of RNA metabolism including readers of m6A methylation [120]. HNRNPA2B1 acts in concert with METTL3 and influences alternative splicing outcome of a subset of transcripts, as well as promotes primary microRNA processing of a subset of primary microRNA transcripts [121]. However, the precise binding specificity of hnRNPA2B1 to m6A methylation is a matter of debate and a combination of structural, biochemical and bioinformatics studies suggest that instead of direct binding to m6A mark, the mark possibly facilitates accessibility of hnRNPA2B1 to certain binding sites on the RNA in the vicinity of the m6A mark, the so called concept of ‘m6A switches’ [122]. Further support to this concept was provided by studies on the reader protein, hnRNPG, wherein m6A mark increases the accessibility of the surrounding RNA sequence to bind hnRNPG, instead of a direct binding to the m6A signature [123].
Eukaryotic Initiation Factor 3 (eIF3)
eIF3 serves as a m6A reader by binding specifically to m6A signatures in 5′-UTR of target transcripts and promotes their cap-independent translation [124]. Such a regulatory mechanism is frequently used during cellular stresses whereby 5′-UTR m6A increases leading to increase in cap-independent translation of mRNAs like HSP70 mRNA [124].
Proline Rich Coiled-Coil 2A (PRRC2A)
PRRC2A is a novel m6A reader, bearing proline rich coiled-coil domain, and was reported to modulate oligodendrocyte specification and myelination, primarily by binding and stabilizing its target OLIG2 mRNA in a m6A-dependent manner [125].
m6A ‘Erasers’
Erasers are RNA demethylases which remove the m6A mark installed by the writer complex and thus makes the m6A methylation reversible and dynamic in nature (Fig. 2). Even though the m6A writer, METTL3 was discovered decades ago, but the first m6A eraser, fat mass and obesity-associated protein (FTO) was reported only in 2011, and was shown to exhibit m6A demethylase activity [11]. Soon after this, the second m6A eraser, AlkB homology 5 (ALKBH5) was reported [126]. Interestingly, the substrate specificity of the m6A erasers is defined by a m6A-mediated substrate discrimination mechanism [127].
Fat Body Mass and Obesity-Associated Protein (FTO)
Pioneering work from Chuan He’s lab was instrumental in uncovering the m6A erasers involved in removing the m6A mark from methylated transcripts. In 2008, employing in vitro experiments, recombinant FTO was demonstrated to catalyze oxidative demethylation of 3-methylthymine in single-stranded DNA and 3-methyluracil in single-stranded RNA substrates, with a greater preference being for the RNA [128]. FTO is an iron(II) and α-ketoglutarate dependent dioxygenase, homologous to the DNA repair protein, AlkB, and has been reported to be associated with obesity and increased body mass in humans [129, 130], as well as other developmental processes like cilia formation [131]. In 2011, Jia et al., showed the demethylase activity of FTO in vivo and its depletion led to increase in m6A levels of polyadenylated RNAs [11]. Furthermore, FTO demethylates its substrate through an oxidative process via formation of two intermediate products, N6-hydroxylmethyladenosine and N6-formyladenosine, whose function is a matter of further examination [132]. FTO shuttles between nucleus and cytoplasm mediated by interaction with exportin 2 (XPO2) and is suggestive of its different substrates in different cellular compartments [133]. Strikingly, a report suggested that FTO can also demethylate m6Am, present at the first encoded nucleotide after the 7-methylguanosine cap in a subset of mRNAs and destabilizes them [134].
Multiple research studies have established the role of FTO in several facets of RNA metabolism such as pre-mRNA splicing and degradation [135,136,137,138,139] and its impact on sculpting the transcriptome [140, 141]. FTO modulates adipogenesis in a m6A-dependent manner, by regulating proliferation and differentiation of preadipocytes and impacts triglyceride metabolism [142,143,144,145]. Aside from adipogenesis, FTO impacts other developmental processes in a m6A-dependent manner, like bone development [146], postnatal brain development in mice where it is essential for adult neural stem cells proliferation [147, 148] and local mRNA translation in axons [149]. Increased expression of FTO was reported to reverse ischemic induced heart failure in mouse models [150] and skeletal muscle specific FTO deletion both under in vitro and in vivo scenarios, demonstrated its requirement for mitochondrial biogenesis [151] and for modulating lipid accumulation in skeletal muscle [152]. Interestingly, glycogen synthase kinase 3 (GSK3) was shown to regulate pluripotency in mouse embryonic stem cells through modulating m6A level by phosphorylating FTO, leading to its polyubiquitination and degradation [153]. FTO was suggested to contribute to pancreas islet β cells impairment and inhibition of its activity might be a potential target for the treatment of diabetes [154]. In accordance with and supporting this finding, elevated expression of FTO was often observed in diabetic patients [17]. Circadian rhythm is closely linked to obesity and FTO was shown to regulate the circadian rhythms by forming complexes with cryptochromes [155, 156]. Interestingly, last year a rather surprising function of FTO was uncovered. In a m6A-dependent manner, it was shown to modulate biogenesis of small nuclear RNAs and may influence RNA splicing since small nuclear RNAs are integral part of spliceosomes, which mediate pre-mRNA splicing [134]. FTO is aberrantly expressed in multiple cancer types and has been shown to influence various aspects of cancer development, progression and therapeutic resistance in a m6A-dependent manner (Table 1).
AlkB Homology 5 (ALKBH5)
ALKBH5, the second m6A eraser, identified by Chuan He’s group in 2013, like FTO, belongs to the AlkB family of proteins and oxidatively reverses the m6A modification [126]. Initial eraser function of ALKBH5 was probed by in vitro experiments as in case of FTO; however, eventual in vivo studies using ALKBH5 knockout mice revealed its role in spermatogenesis and fertility in males [126, 157]. Interestingly, expression of ALKBH5 is stimulated under oxygen limiting hypoxic conditions, in a hypoxia-inducible factor (HIF)-dependent fashion and this mediates enrichment of breast cancer stem cells in hypoxic tumor microenvironment [158,159,160]. Furthermore, a role for ALKBH5 in recurrent miscarriages was reported, wherein it controls trophoblast invasion at the maternal–fetal interface by modulating the stability of CRY61 transcripts, in a m6A-dependent manner [161]. Like, FTO, ALKBH5 has diverse functional and regulatory role in human diseases like cancer (Table 1).
Conclusions and Future Perspectives
The discovery of reversible m6A RNA methylation a few years back and subsequent functional characterization of the writers, readers and erasers of the m6A mark, has taken the field of gene expression by storm and has emerged as one of the key regulators of gene expression via its impact on virtually every aspect of RNA processing. Not only does it influence normal developmental processes, it also impinges upon a plethora of human diseases. Despite this, several pertinent questions are yet to be addressed to get a holistic view of this regulatory process. Some of the key questions are: What are the regulators of m6A methylation? What are the extracellular and intracellular signaling cues which regulate the m6A writers, readers and erasers? How is the substrate specificity of m6A writing, reading and erasing regulated? What is the cross-talk between m6A methylation and other reversible modifications of RNA? What are the fine and interwoven regulatory networks operating between m6A methylation and other RNA processing events? Deciphering the answers to these questions will provide a deeper insight into the role of m6A RNA methylation and in particular propel designing of novel therapies targeting this regulatory pathway.
References
Crick, F. (1970). Central dogma of molecular biology. Nature, 227, 561–563.
Jones, P. L., Veenstra, G. J., Wade, P. A., Vermaak, D., Kass, S. U., Landsberger, N., et al. (1998). Methylated DNA and MeCP2 recruit histone deacetylase to repress transcription. Nature Genetics, 19, 187–191.
Feinberg, A. P., & Vogelstein, B. (1983). Hypomethylation distinguishes genes of some human cancers from their normal counterparts. Nature, 301, 89–92.
Cheah, M. S., Wallace, C. D., & Hoffman, R. M. (1984). Hypomethylation of DNA in human cancer cells: a site-specific change in the c-myc oncogene. Journal of the National Cancer Institute, 73, 1057–1065.
Dong, X., & Weng, Z. (2013). The correlation between histone modifications and gene expression. Epigenomics, 5, 113–116.
Javaid, N., & Choi, S. (2017). Acetylation- and methylation-related epigenetic proteins in the context of their targets. Genes (Basel). https://doi.org/10.3390/genes8080196.
Dezi, V., Ivanov, C., Haussmann, I. U., & Soller, M. (2016). Nucleotide modifications in messenger RNA and their role in development and disease. Biochemical Society Transactions, 44, 1385–1393.
Roundtree, I. A., Evans, M. E., Pan, T., & He, C. (2017). Dynamic RNA modifications in gene expression regulation. Cell, 169, 1187–1200.
Desrosiers, R., Friderici, K., & Rottman, F. (1974). Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proc Natl Acad Sci U S A, 71, 3971–3975.
Perry, R. P., & Kelley, D. E. (1974). Existence of methylated messenger RNA in mouse L cells. Cell, 1, 37–42.
Jia, G., Fu, Y., Zhao, X., Dai, Q., Zheng, G., Yang, Y., et al. (2011). N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nature Chemical Biology, 7, 885–887.
Chen, T., Hao, Y. J., Zhang, Y., Li, M. M., Wang, M., Han, W., et al. (2015). m(6)A RNA methylation is regulated by microRNAs and promotes reprogramming to pluripotency. Cell Stem Cell, 16, 289–301.
Linder, B., Grozhik, A. V., Olarerin-George, A. O., Meydan, C., Mason, C. E., & Jaffrey, S. R. (2015). Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nature Methods, 12, 767–772.
Yang, Y., Fan, X., Mao, M., Song, X., Wu, P., Zhang, Y., et al. (2017). Extensive translation of circular RNAs driven by N(6)-methyladenosine. Cell Research, 27, 626–641.
Zhou, C., Molinie, B., Daneshvar, K., Pondick, J. V., Wang, J., Van Wittenberghe, N., et al. (2017). Genome-wide maps of m6A circRNAs identify widespread and cell-type-specific methylation patterns that are distinct from mRNAs. Cell Rep, 20, 2262–2276.
Zhao, B. S., Roundtree, I. A., & He, C. (2017). Post-transcriptional gene regulation by mRNA modifications. Nature Reviews Molecular Cell Biology, 18, 31–42.
Shen, F., Huang, W., Huang, J. T., Xiong, J., Yang, Y., Wu, K., et al. (2015). Decreased N(6)-methyladenosine in peripheral blood RNA from diabetic patients is associated with FTO expression rather than ALKBH5. Journal of Clinical Endocrinology and Metabolism, 100, E148–154.
Ding, C., Zou, Q., Ding, J., Ling, M., Wang, W., Li, H., et al. (2018). Increased N6-methyladenosine causes infertility is associated with FTO expression. Journal of Cellular Physiology, 233, 7055–7066.
Chandola, U., Das, R., & Panda, B. (2015). Role of the N6-methyladenosine RNA mark in gene regulation and its implications on development and disease. Brief Funct Genomics, 14, 169–179.
Dai, D., Wang, H., Zhu, L., Jin, H., & Wang, X. (2018). N6-methyladenosine links RNA metabolism to cancer progression. Cell Death Dis, 9, 124.
Csepany, T., Lin, A., Baldick, C. J., Jr., & Beemon, K. (1990). Sequence specificity of mRNA N6-adenosine methyltransferase. Journal of Biological Chemistry, 265, 20117–20122.
Meyer, K. D., Saletore, Y., Zumbo, P., Elemento, O., Mason, C. E., & Jaffrey, S. R. (2012). Comprehensive analysis of mRNA methylation reveals enrichment in 3' UTRs and near stop codons. Cell, 149, 1635–1646.
Ke, S., Alemu, E. A., Mertens, C., Gantman, E. C., Fak, J. J., Mele, A., et al. (2015). A majority of m6A residues are in the last exons, allowing the potential for 3' UTR regulation. Genes & Development, 29, 2037–2053.
Wei, C. M., Gershowitz, A., & Moss, B. (1976). 5'-Terminal and internal methylated nucleotide sequences in HeLa cell mRNA. Biochemistry, 15, 397–401.
Carroll, S. M., Narayan, P., & Rottman, F. M. (1990). N6-methyladenosine residues in an intron-specific region of prolactin pre-mRNA. Molecular and Cellular Biology, 10, 4456–4465.
Ke, S., Pandya-Jones, A., Saito, Y., Fak, J. J., Vagbo, C. B., Geula, S., et al. (2017). m(6)A mRNA modifications are deposited in nascent pre-mRNA and are not required for splicing but do specify cytoplasmic turnover. Genes & Development, 31, 990–1006.
Dominissini, D., Moshitch-Moshkovitz, S., Schwartz, S., Salmon-Divon, M., Ungar, L., Osenberg, S., et al. (2012). Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature, 485, 201–206.
Dominissini, D., Moshitch-Moshkovitz, S., Salmon-Divon, M., Amariglio, N., & Rechavi, G. (2013). Transcriptome-wide mapping of N(6)-methyladenosine by m(6)A-seq based on immunocapturing and massively parallel sequencing. Nature Protocols, 8, 176–189.
Chen, K., Lu, Z., Wang, X., Fu, Y., Luo, G. Z., Liu, N., et al. (2015). High-resolution N(6) -methyladenosine (m(6) A) map using photo-crosslinking-assisted m(6) A sequencing. Angewandte Chemie (International ed. in English), 54, 1587–1590.
Liu, N., Parisien, M., Dai, Q., Zheng, G., He, C., & Pan, T. (2013). Probing N6-methyladenosine RNA modification status at single nucleotide resolution in mRNA and long noncoding RNA. RNA, 19, 1848–1856.
Thuring, K., Schmid, K., Keller, P., & Helm, M. (2016). Analysis of RNA modifications by liquid chromatography-tandem mass spectrometry. Methods, 107, 48–56.
Zhou, Y., Zeng, P., Li, Y. H., Zhang, Z., & Cui, Q. (2016). SRAMP: prediction of mammalian N6-methyladenosine (m6A) sites based on sequence-derived features. Nucleic Acids Research, 44, e91.
Li, G. Q., Liu, Z., Shen, H. B., & Yu, D. J. (2016). Target M6A: identifying N(6)-methyladenosine sites from RNA sequences via position-specific nucleotide propensities and a support vector machine. IEEE Transactions of Nanobioscience, 15, 674–682.
Jia, C. Z., Zhang, J. J., & Gu, W. Z. (2016). RNA-MethylPred: A high-accuracy predictor to identify N6-methyladenosine in RNA. Analytical Biochemistry, 510, 72–75.
Chen, W., Feng, P., Ding, H., Lin, H., & Chou, K. C. (2015). iRNA-Methyl: Identifying N(6)-methyladenosine sites using pseudo nucleotide composition. Analytical Biochemistry, 490, 26–33.
Liu, Z., Xiao, X., Yu, D. J., Jia, J., Qiu, W. R., & Chou, K. C. (2016). pRNAm-PC: Predicting N(6)-methyladenosine sites in RNA sequences via physical-chemical properties. Analytical Biochemistry, 497, 60–67.
Liu, N., & Pan, T. (2015). Probing RNA modification status at single-nucleotide resolution in total RNA. Methods in Enzymology, 560, 149–159.
Zhu, Y., Zhou, G., Yu, X., Xu, Q., Wang, K., Xie, D., et al. (2017). LC-MS-MS quantitative analysis reveals the association between FTO and DNA methylation. PLoS ONE, 12, e0175849.
Zhao, X., Zhang, Y., Ning, Q., Zhang, H., Ji, J., & Yin, M. (2019). Identifying N(6)-methyladenosine sites using extreme gradient boosting system optimized by particle swarm optimizer. Journal of Theoretical Biology, 467, 39–47.
Narayan, P., Ludwiczak, R. L., Goodwin, E. C., & Rottman, F. M. (1994). Context effects on N6-adenosine methylation sites in prolactin mRNA. Nucleic Acids Research, 22, 419–426.
Narayan, P., & Rottman, F. M. (1988). An in vitro system for accurate methylation of internal adenosine residues in messenger RNA. Science, 242, 1159–1162.
Bokar, J. A., Rath-Shambaugh, M. E., Ludwiczak, R., Narayan, P., & Rottman, F. (1994). Characterization and partial purification of mRNA N6-adenosine methyltransferase from HeLa cell nuclei. Internal mRNA methylation requires a multisubunit complex. Journal of Biological Chemistry, 269, 17697–17704.
Bertero, A., Brown, S., Madrigal, P., Osnato, A., Ortmann, D., Yiangou, L., et al. (2018). The SMAD2/3 interactome reveals that TGFbeta controls m(6)A mRNA methylation in pluripotency. Nature, 555, 256–259.
Su, R., Dong, L., Li, C., Nachtergaele, S., Wunderlich, M., Qing, Y., et al. (2018). R-2HG exhibits anti-tumor activity by targeting FTO/m(6)A/MYC/CEBPA signaling. Cell, 172(90–105), e123.
Bokar, J. A., Shambaugh, M. E., Polayes, D., Matera, A. G., & Rottman, F. M. (1997). Purification and cDNA cloning of the AdoMet-binding subunit of the human mRNA (N6-adenosine)-methyltransferase. RNA, 3, 1233–1247.
Liu, N., Dai, Q., Zheng, G., He, C., Parisien, M., & Pan, T. (2015). N(6)-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature, 518, 560–564.
Zhou, K. I., Parisien, M., Dai, Q., Liu, N., Diatchenko, L., Sachleben, J. R., et al. (2016). N(6)-methyladenosine modification in a long noncoding RNA hairpin predisposes its conformation to protein binding. Journal of Molecular Biology, 428, 822–833.
Lin, S., & Gregory, R. I. (2014). Methyltransferases modulate RNA stability in embryonic stem cells. Nature Cell Biology, 16, 129–131.
Wang, Y., Li, Y., Toth, J. I., Petroski, M. D., Zhang, Z., & Zhao, J. C. (2014). N6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells. Nature Cell Biology, 16, 191–198.
Geula, S., Moshitch-Moshkovitz, S., Dominissini, D., Mansour, A. A., Kol, N., Salmon-Divon, M., et al. (2015). m6A mRNA methylation facilitates resolution of naive pluripotency toward differentiation. Science, 347, 1002–1006.
Tian, C., Huang, Y., Li, Q., Feng, Z., & Xu, Q. (2019). Mettl3 regulates osteogenic differentiation and alternative splicing of Vegfa in bone marrow mesenchymal stem cells. International Journal of Molecular Sciences. https://doi.org/10.3390/ijms20030551.
Yu, J., Shen, L., Liu, Y., Ming, H., Zhu, X., Chu, M., et al. (2020). The m6A methyltransferase METTL3 cooperates with demethylase ALKBH5 to regulate osteogenic differentiation through NF-kappaB signaling. Molecular and Cellular Biochemistry, 463, 203–210.
Zhang, Y., Gu, X., Li, D., Cai, L., & Xu, Q. (2019). METTL3 regulates osteoblast differentiation and inflammatory response via Smad signaling and MAPK signaling. International Journal of Molecular Sciences. https://doi.org/10.3390/ijms21010199.
Yan, G., Yuan, Y., He, M., Gong, R., Lei, H., Zhou, H., et al. (2019). m(6)A methylation of precursor-miR-320/RUNX2 controls osteogenic potential of bone marrow-derived mesenchymal stem cells. Mol Ther Nucleic Acids, 19, 421–436.
Zhang, Z., Wang, M., Xie, D., Huang, Z., Zhang, L., Yang, Y., et al. (2018). METTL3-mediated N(6)-methyladenosine mRNA modification enhances long-term memory consolidation. Cell Research, 28, 1050–1061.
Li, H. B., Tong, J., Zhu, S., Batista, P. J., Duffy, E. E., Zhao, J., et al. (2017). m(6)A mRNA methylation controls T cell homeostasis by targeting the IL-7/STAT5/SOCS pathways. Nature, 548, 338–342.
Song, H., Feng, X., Zhang, H., Luo, Y., Huang, J., Lin, M., et al. (2019). METTL3 and ALKBH5 oppositely regulate m(6)A modification of TFEB mRNA, which dictates the fate of hypoxia/reoxygenation-treated cardiomyocytes. Autophagy, 15, 1419–1437.
Fustin, J. M., Doi, M., Yamaguchi, Y., Hida, H., Nishimura, S., Yoshida, M., et al. (2013). RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell, 155, 793–806.
Fustin, J. M., Kojima, R., Itoh, K., Chang, H. Y., Ye, S., Zhuang, B., et al. (2018). Two Ck1delta transcripts regulated by m6A methylation code for two antagonistic kinases in the control of the circadian clock. Proc Natl Acad Sci U S A, 115, 5980–5985.
Alarcon, C. R., Lee, H., Goodarzi, H., Halberg, N., & Tavazoie, S. F. (2015). N6-methyladenosine marks primary microRNAs for processing. Nature, 519, 482–485.
Vu, L. P., Pickering, B. F., Cheng, Y., Zaccara, S., Nguyen, D., Minuesa, G., et al. (2017). The N(6)-methyladenosine (m(6)A)-forming enzyme METTL3 controls myeloid differentiation of normal hematopoietic and leukemia cells. Nature Medicine, 23, 1369–1376.
Choe, J., Lin, S., Zhang, W., Liu, Q., Wang, L., Ramirez-Moya, J., et al. (2018). mRNA circularization by METTL3-eIF3h enhances translation and promotes oncogenesis. Nature, 561, 556–560.
Lin, S., Choe, J., Du, P., Triboulet, R., & Gregory, R. I. (2016). The m(6)A methyltransferase METTL3 promotes translation in human cancer cells. Molecular Cell, 62, 335–345.
Barbieri, I., Tzelepis, K., Pandolfini, L., Shi, J., Millan-Zambrano, G., Robson, S. C., et al. (2017). Promoter-bound METTL3 maintains myeloid leukaemia by m(6)A-dependent translation control. Nature, 552, 126–131.
Chen, M., Wei, L., Law, C. T., Tsang, F. H., Shen, J., Cheng, C. L., et al. (2018). RNA N6-methyladenosine methyltransferase-like 3 promotes liver cancer progression through YTHDF2-dependent posttranscriptional silencing of SOCS2. Hepatology, 67, 2254–2270.
Wu, Y., Xie, L., Wang, M., Xiong, Q., Guo, Y., Liang, Y., et al. (2018). Mettl3-mediated m(6)A RNA methylation regulates the fate of bone marrow mesenchymal stem cells and osteoporosis. Nature Communications, 9, 4772.
Bujnicki, J. M., Feder, M., Radlinska, M., & Blumenthal, R. M. (2002). Structure prediction and phylogenetic analysis of a functionally diverse family of proteins homologous to the MT-A70 subunit of the human mRNA:m(6)A methyltransferase. Journal of Molecular Evolution, 55, 431–444.
Petrossian, T. C., & Clarke, S. G. (2011). Uncovering the human methyltransferasome. Molecular and Cellular Proteomics, 10(M110), 000976.
Wang, X., Feng, J., Xue, Y., Guan, Z., Zhang, D., Liu, Z., et al. (2016). Structural basis of N(6)-adenosine methylation by the METTL3-METTL14 complex. Nature, 534, 575–578.
Wang, P., Doxtader, K. A., & Nam, Y. (2016). Structural basis for cooperative function of Mettl3 and Mettl14 methyltransferases. Molecular Cell, 63, 306–317.
Scholler, E., Weichmann, F., Treiber, T., Ringle, S., Treiber, N., Flatley, A., et al. (2018). Interactions, localization, and phosphorylation of the m(6)A generating METTL3-METTL14-WTAP complex. RNA, 24, 499–512.
Ruzicka, K., Zhang, M., Campilho, A., Bodi, Z., Kashif, M., Saleh, M., et al. (2017). Identification of factors required for m(6) A mRNA methylation in Arabidopsis reveals a role for the conserved E3 ubiquitin ligase HAKAI. New Phytologist, 215, 157–172.
Meng, T. G., Lu, X., Guo, L., Hou, G. M., Ma, X. S., Li, Q. N., et al. (2019). Mettl14 is required for mouse postimplantation development by facilitating epiblast maturation. The FASEB Journal, 33, 1179–1187.
Lin, Z., Hsu, P. J., Xing, X., Fang, J., Lu, Z., Zou, Q., et al. (2017). Mettl3-/Mettl14-mediated mRNA N(6)-methyladenosine modulates murine spermatogenesis. Cell Research, 27, 1216–1230.
Koranda, J. L., Dore, L., Shi, H., Patel, M. J., Vaasjo, L. O., Rao, M. N., et al. (2018). Mettl14 is essential for epitranscriptomic regulation of striatal function and learning. Neuron, 99(283–292), e285.
Wang, Y., Li, Y., Yue, M., Wang, J., Kumar, S., Wechsler-Reya, R. J., et al. (2018). N(6)-methyladenosine RNA modification regulates embryonic neural stem cell self-renewal through histone modifications. Nature Neuroscience, 21, 195–206.
Ma, J. Z., Yang, F., Zhou, C. C., Liu, F., Yuan, J. H., Wang, F., et al. (2017). METTL14 suppresses the metastatic potential of hepatocellular carcinoma by modulating N(6)-methyladenosine-dependent primary MicroRNA processing. Hepatology, 65, 529–543.
Weng, H., Huang, H., Wu, H., Qin, X., Zhao, B. S., Dong, L., et al. (2018). METTL14 inhibits hematopoietic stem/progenitor differentiation and promotes leukemogenesis via mRNA m(6)A modification. Cell Stem Cell, 22(191–205), e199.
Huang, H., Weng, H., Zhou, K., Wu, T., Zhao, B. S., Sun, M., et al. (2019). Histone H3 trimethylation at lysine 36 guides m(6)A RNA modification co-transcriptionally. Nature, 567, 414–419.
Little, N. A., Hastie, N. D., & Davies, R. C. (2000). Identification of WTAP, a novel Wilms' tumour 1-associating protein. Human Molecular Genetics, 9, 2231–2239.
Ping, X. L., Sun, B. F., Wang, L., Xiao, W., Yang, X., Wang, W. J., et al. (2014). Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Research, 24, 177–189.
Schwartz, S., Mumbach, M. R., Jovanovic, M., Wang, T., Maciag, K., Bushkin, G. G., et al. (2014). Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5' sites. Cell Rep, 8, 284–296.
Yue, Y., Liu, J., Cui, X., Cao, J., Luo, G., Zhang, Z., et al. (2018). VIRMA mediates preferential m(6)A mRNA methylation in 3'UTR and near stop codon and associates with alternative polyadenylation. Cell Discov, 4, 10.
Patil, D. P., Chen, C. K., Pickering, B. F., Chow, A., Jackson, C., Guttman, M., et al. (2016). m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature, 537, 369–373.
Xie, Y., Castro-Hernandez, R., Sokpor, G., Pham, L., Narayanan, R., Rosenbusch, J., et al. (2019). RBM15 modulates the function of chromatin remodeling factor BAF155 through RNA methylation in developing cortex. Molecular Neurobiology, 56, 7305–7320.
Wen, J., Lv, R., Ma, H., Shen, H., He, C., Wang, J., et al. (2018). Zc3h13 regulates nuclear RNA m(6)A methylation and mouse embryonic stem cell self-renewal. Molecular Cell, 69(1028–1038), e1026.
Shimba, S., Bokar, J. A., Rottman, F., & Reddy, R. (1995). Accurate and efficient N-6-adenosine methylation in spliceosomal U6 small nuclear RNA by HeLa cell extract in vitro. Nucleic Acids Research, 23, 2421–2426.
Pendleton, K. E., Chen, B., Liu, K., Hunter, O. V., Xie, Y., Tu, B. P., et al. (2017). The U6 snRNA m(6)A methyltransferase METTL16 regulates SAM synthetase intron retention. Cell, 169(824–835), e814.
Warda, A. S., Kretschmer, J., Hackert, P., Lenz, C., Urlaub, H., Hobartner, C., et al. (2017). Human METTL16 is a N(6)-methyladenosine (m(6)A) methyltransferase that targets pre-mRNAs and various non-coding RNAs. EMBO Reports, 18, 2004–2014.
Ruszkowska, A., Ruszkowski, M., Dauter, Z., & Brown, J. A. (2018). Structural insights into the RNA methyltransferase domain of METTL16. Sci Rep, 8, 5311.
Mendel, M., Chen, K. M., Homolka, D., Gos, P., Pandey, R. R., McCarthy, A. A., et al. (2018). Methylation of structured RNA by the m(6)A writer METTL16 Is essential for mouse embryonic development. Molecular Cell, 71(986–1000), e1011.
Theler, D., Dominguez, C., Blatter, M., Boudet, J., & Allain, F. H. (2014). Solution structure of the YTH domain in complex with N6-methyladenosine RNA: a reader of methylated RNA. Nucleic Acids Research, 42, 13911–13919.
Xu, C., Liu, K., Ahmed, H., Loppnau, P., Schapira, M., & Min, J. (2015). Structural Basis For The Discriminative Recognition of N6-methyladenosine RNA by the human YT521-B homology domain family of proteins. Journal of Biological Chemistry, 290, 24902–24913.
Wang, X., Zhao, B. S., Roundtree, I. A., Lu, Z., Han, D., Ma, H., et al. (2015). N(6)-methyladenosine modulates messenger RNA translation efficiency. Cell, 161, 1388–1399.
Zhuang, M., Li, X., Zhu, J., Zhang, J., Niu, F., Liang, F., et al. (2019). The m6A reader YTHDF1 regulates axon guidance through translational control of Robo3.1 expression. Nucleic Acids Research, 47, 4765–4777.
Shi, H., Zhang, X., Weng, Y. L., Lu, Z., Liu, Y., Lu, Z., et al. (2018). m(6)A facilitates hippocampus-dependent learning and memory through YTHDF1. Nature, 563, 249–253.
Wu, R., Liu, Y., Zhao, Y., Bi, Z., Yao, Y., Liu, Q., et al. (2019). m(6)A methylation controls pluripotency of porcine induced pluripotent stem cells by targeting SOCS3/JAK2/STAT3 pathway in a YTHDF1/YTHDF2-orchestrated manner. Cell Death & Disease, 10, 171.
Wang, X., Lu, Z., Gomez, A., Hon, G. C., Yue, Y., Han, D., et al. (2014). N6-methyladenosine-dependent regulation of messenger RNA stability. Nature, 505, 117–120.
Du, H., Zhao, Y., He, J., Zhang, Y., Xi, H., Liu, M., et al. (2016). YTHDF2 destabilizes m(6)A-containing RNA through direct recruitment of the CCR4-NOT deadenylase complex. Nature Communications, 7, 12626.
Li, F., Zhao, D., Wu, J., & Shi, Y. (2014). Structure of the YTH domain of human YTHDF2 in complex with an m(6)A mononucleotide reveals an aromatic cage for m(6)A recognition. Cell Research, 24, 1490–1492.
Zhu, T., Roundtree, I. A., Wang, P., Wang, X., Wang, L., Sun, C., et al. (2014). Crystal structure of the YTH domain of YTHDF2 reveals mechanism for recognition of N6-methyladenosine. Cell Research, 24, 1493–1496.
Zhou, J., Wan, J., Gao, X., Zhang, X., Jaffrey, S. R., & Qian, S. B. (2015). Dynamic m(6)A mRNA methylation directs translational control of heat shock response. Nature, 526, 591–594.
Li, Z., Qian, P., Shao, W., Shi, H., He, X. C., Gogol, M., et al. (2018). Suppression of m(6)A reader Ythdf2 promotes hematopoietic stem cell expansion. Cell Research, 28, 904–917.
Li, M., Zhao, X., Wang, W., Shi, H., Pan, Q., Lu, Z., et al. (2018). Ythdf2-mediated m(6)A mRNA clearance modulates neural development in mice. Genome Biology, 19, 69.
Zhao, B. S., Wang, X., Beadell, A. V., Lu, Z., Shi, H., Kuuspalu, A., et al. (2017). m(6)A-dependent maternal mRNA clearance facilitates zebrafish maternal-to-zygotic transition. Nature, 542, 475–478.
Ivanova, I., Much, C., Di Giacomo, M., Azzi, C., Morgan, M., Moreira, P. N., et al. (2017). The RNA m(6)A reader YTHDF2 is essential for the post-transcriptional regulation of the maternal transcriptome and oocyte competence. Molecular Cell, 67(1059–1067), e1054.
Yang, Z., Li, J., Feng, G., Gao, S., Wang, Y., Zhang, S., et al. (2017). MicroRNA-145 modulates N(6)-methyladenosine levels by targeting the 3'-untranslated mRNA region of the N(6)-Methyladenosine Binding YTH domain family 2 protein. Journal of Biological Chemistry, 292, 3614–3623.
Li, J., Meng, S., Xu, M., Wang, S., He, L., Xu, X., et al. (2018). Downregulation of N(6)-methyladenosine binding YTHDF2 protein mediated by miR-493-3p suppresses prostate cancer by elevating N(6)-methyladenosine levels. Oncotarget, 9, 3752–3764.
Shi, H., Wang, X., Lu, Z., Zhao, B. S., Ma, H., Hsu, P. J., et al. (2017). YTHDF3 facilitates translation and decay of N(6)-methyladenosine-modified RNA. Cell Research, 27, 315–328.
Li, A., Chen, Y. S., Ping, X. L., Yang, X., Xiao, W., Yang, Y., et al. (2017). Cytoplasmic m(6)A reader YTHDF3 promotes mRNA translation. Cell Research, 27, 444–447.
Xiao, W., Adhikari, S., Dahal, U., Chen, Y. S., Hao, Y. J., Sun, B. F., et al. (2016). Nuclear m(6)A reader YTHDC1 regulates mRNA splicing. Molecular Cell, 61, 507–519.
Roundtree, I. A., Luo, G. Z., Zhang, Z., Wang, X., Zhou, T., Cui, Y., et al. (2017). YTHDC1 mediates nuclear export of N(6)-methyladenosine methylated mRNAs. Elife. https://doi.org/10.7554/eLife.31311.
Shima, H., Matsumoto, M., Ishigami, Y., Ebina, M., Muto, A., Sato, Y., et al. (2017). S-adenosylmethionine synthesis is regulated by selective N(6)-adenosine methylation and mRNA degradation involving METTL16 and YTHDC1. Cell Reports, 21, 3354–3363.
Kasowitz, S. D., Ma, J., Anderson, S. J., Leu, N. A., Xu, Y., Gregory, B. D., et al. (2018). Nuclear m6A reader YTHDC1 regulates alternative polyadenylation and splicing during mouse oocyte development. PLoS Genetics, 14, e1007412.
Wojtas, M. N., Pandey, R. R., Mendel, M., Homolka, D., Sachidanandam, R., & Pillai, R. S. (2017). Regulation of m(6)A transcripts by the 3'–>5' RNA helicase YTHDC2 is essential for a successful meiotic program in the mammalian germline. Molecular Cell, 68(374–387), e312.
Hsu, P. J., Zhu, Y., Ma, H., Guo, Y., Shi, X., Liu, Y., et al. (2017). Ythdc2 is an N(6)-methyladenosine binding protein that regulates mammalian spermatogenesis. Cell Research, 27, 1115–1127.
Kretschmer, J., Rao, H., Hackert, P., Sloan, K. E., Hobartner, C., & Bohnsack, M. T. (2018). The m(6)A reader protein YTHDC2 interacts with the small ribosomal subunit and the 5'-3' exoribonuclease XRN1. RNA, 24, 1339–1350.
Tanabe, A., Tanikawa, K., Tsunetomi, M., Takai, K., Ikeda, H., Konno, J., et al. (2016). RNA helicase YTHDC2 promotes cancer metastasis via the enhancement of the efficiency by which HIF-1alpha mRNA is translated. Cancer Letters, 376, 34–42.
Huang, H., Weng, H., Sun, W., Qin, X., Shi, H., Wu, H., et al. (2018). Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nature Cell Biology, 20, 285–295.
Konig, J., Zarnack, K., Rot, G., Curk, T., Kayikci, M., Zupan, B., et al. (2010). iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nature Structural & Molecular Biology, 17, 909–915.
Alarcon, C. R., Goodarzi, H., Lee, H., Liu, X., Tavazoie, S., & Tavazoie, S. F. (2015). HNRNPA2B1 is a mediator of m(6)A-Dependent Nuclear RNA processing events. Cell, 162, 1299–1308.
Wu, B., Su, S., Patil, D. P., Liu, H., Gan, J., Jaffrey, S. R., et al. (2018). Molecular basis for the specific and multivariant recognitions of RNA substrates by human hnRNP A2/B1. Nature Communications, 9, 420.
Liu, N., Zhou, K. I., Parisien, M., Dai, Q., Diatchenko, L., & Pan, T. (2017). N6-methyladenosine alters RNA structure to regulate binding of a low-complexity protein. Nucleic Acids Research, 45, 6051–6063.
Meyer, K. D., Patil, D. P., Zhou, J., Zinoviev, A., Skabkin, M. A., Elemento, O., et al. (2015). 5' UTR m(6)A promotes cap-independent translation. Cell, 163, 999–1010.
Wu, R., Li, A., Sun, B., Sun, J. G., Zhang, J., Zhang, T., et al. (2019). A novel m(6)A reader Prrc2a controls oligodendroglial specification and myelination. Cell Research, 29, 23–41.
Zheng, G., Dahl, J. A., Niu, Y., Fedorcsak, P., Huang, C. M., Li, C. J., et al. (2013). ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Molecular Cell, 49, 18–29.
Zou, S., Toh, J. D., Wong, K. H., Gao, Y. G., Hong, W., & Woon, E. C. (2016). N(6)-Methyladenosine: a conformational marker that regulates the substrate specificity of human demethylases FTO and ALKBH5. Science Reports, 6, 25677.
Jia, G., Yang, C. G., Yang, S., Jian, X., Yi, C., Zhou, Z., et al. (2008). Oxidative demethylation of 3-methylthymine and 3-methyluracil in single-stranded DNA and RNA by mouse and human FTO. FEBS Letters, 582, 3313–3319.
Frayling, T. M., Timpson, N. J., Weedon, M. N., Zeggini, E., Freathy, R. M., Lindgren, C. M., et al. (2007). A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science, 316, 889–894.
Freathy, R. M., Timpson, N. J., Lawlor, D. A., Pouta, A., Ben-Shlomo, Y., Ruokonen, A., et al. (2008). Common variation in the FTO gene alters diabetes-related metabolic traits to the extent expected given its effect on BMI. Diabetes, 57, 1419–1426.
Osborn, D. P., Roccasecca, R. M., McMurray, F., Hernandez-Hernandez, V., Mukherjee, S., Barroso, I., et al. (2014). Loss of FTO antagonises Wnt signaling and leads to developmental defects associated with ciliopathies. PLoS ONE, 9, e87662.
Fu, Y., Jia, G., Pang, X., Wang, R. N., Wang, X., Li, C. J., et al. (2013). FTO-mediated formation of N6-hydroxymethyladenosine and N6-formyladenosine in mammalian RNA. Nat Commun, 4, 1798.
Gulati, P., Avezov, E., Ma, M., Antrobus, R., Lehner, P., O'Rahilly, S., et al. (2014). Fat mass and obesity-related (FTO) shuttles between the nucleus and cytoplasm. Bioscience Reports. https://doi.org/10.1042/BSR20140111.
Mauer, J., Sindelar, M., Despic, V., Guez, T., Hawley, B. R., Vasseur, J. J., et al. (2019). FTO controls reversible m(6)Am RNA methylation during snRNA biogenesis. Nature Chemical Biology, 15, 340–347.
Widagdo, J., Zhao, Q. Y., Kempen, M. J., Tan, M. C., Ratnu, V. S., Wei, W., et al. (2016). Experience-dependent accumulation of N6-methyladenosine in the prefrontal cortex is associated with memory processes in mice. Journal of Neuroscience, 36, 6771–6777.
Walters, B. J., Mercaldo, V., Gillon, C. J., Yip, M., Neve, R. L., Boyce, F. M., et al. (2017). The role of The RNA demethylase FTO (fat mass and obesity-associated) and mRNA methylation in hippocampal memory formation. Neuropsychopharmacology, 42, 1502–1510.
Zhao, X., Yang, Y., Sun, B. F., Shi, Y., Yang, X., Xiao, W., et al. (2014). FTO-dependent demethylation of N6-methyladenosine regulates mRNA splicing and is required for adipogenesis. Cell Research, 24, 1403–1419.
Sun, L., Ma, L., Zhang, H., Cao, Y., Wang, C., Hou, N., et al. (2019). Fto deficiency reduces anxiety- and depression-like behaviors in mice via alterations in gut microbiota. Theranostics, 9, 721–733.
Spychala, A., & Ruther, U. (2019). FTO affects hippocampal function by regulation of BDNF processing. PLoS ONE, 14, e0211937.
Berulava, T., Ziehe, M., Klein-Hitpass, L., Mladenov, E., Thomale, J., Ruther, U., et al. (2013). FTO levels affect RNA modification and the transcriptome. European Journal of Human Genetics, 21, 317–323.
Merkestein, M., McTaggart, J. S., Lee, S., Kramer, H. B., McMurray, F., Lafond, M., et al. (2014). Changes in gene expression associated with FTO overexpression in mice. PLoS ONE, 9, e97162.
Jiao, Y., Zhang, J., Lu, L., Xu, J., & Qin, L. (2016). The Fto gene regulates the proliferation and differentiation of pre-adipocytes in vitro. Nutrients, 8, 102.
Ben-Haim, M. S., Moshitch-Moshkovitz, S., & Rechavi, G. (2015). FTO: linking m6A demethylation to adipogenesis. Cell Research, 25, 3–4.
Zhang, M., Zhang, Y., Ma, J., Guo, F., Cao, Q., Zhang, Y., et al. (2015). The demethylase activity of FTO (fat mass and obesity associated protein) is required for preadipocyte differentiation. PLoS ONE, 10, e0133788.
Wang, C. Y., Shie, S. S., Wen, M. S., Hung, K. C., Hsieh, I. C., Yeh, T. S., et al. (2015). Loss of FTO in adipose tissue decreases Angptl4 translation and alters triglyceride metabolism. Science Signal. https://doi.org/10.1126/scisignal.aab3357.
Sachse, G., Church, C., Stewart, M., Cater, H., Teboul, L., Cox, R. D., et al. (2018). FTO demethylase activity is essential for normal bone growth and bone mineralization in mice. Biochimica et Biophysica Acta (BBA) - Molecular Basis of Diseases, 1864, 843–850.
Gao, X., Shin, Y. H., Li, M., Wang, F., Tong, Q., & Zhang, P. (2010). The fat mass and obesity associated gene FTO functions in the brain to regulate postnatal growth in mice. PLoS ONE, 5, e14005.
Li, L., Zang, L., Zhang, F., Chen, J., Shen, H., Shu, L., et al. (2017). Fat mass and obesity-associated (FTO) protein regulates adult neurogenesis. Human Molecular Genetics, 26, 2398–2411.
Yu, J., Chen, M., Huang, H., Zhu, J., Song, H., Zhu, J., et al. (2018). Dynamic m6A modification regulates local translation of mRNA in axons. Nucleic Acids Research, 46, 1412–1423.
Mathiyalagan, P., Adamiak, M., Mayourian, J., Sassi, Y., Liang, Y., Agarwal, N., et al. (2019). FTO-dependent N(6)-methyladenosine regulates cardiac function during remodeling and repair. Circulation, 139, 518–532.
Wang, X., Huang, N., Yang, M., Wei, D., Tai, H., Han, X., et al. (2017). FTO is required for myogenesis by positively regulating mTOR-PGC-1alpha pathway-mediated mitochondria biogenesis. Cell Death Diseases, 8, e2702.
Wu, W., Feng, J., Jiang, D., Zhou, X., Jiang, Q., Cai, M., et al. (2017). AMPK regulates lipid accumulation in skeletal muscle cells through FTO-dependent demethylation of N(6)-methyladenosine. Sci Rep, 7, 41606.
Faulds, K. J., Egelston, J. N., Sedivy, L. J., Mitchell, M. K., Garimella, S., Kozlowski, H., et al. (2018). Glycogen synthase kinase-3 (GSK-3) activity regulates mRNA methylation in mouse embryonic stem cells. Journal of Biological Chemistry, 293, 10731–10743.
Fan, H. Q., He, W., Xu, K. F., Wang, Z. X., Xu, X. Y., & Chen, H. (2015). FTO inhibits insulin secretion and promotes NF-kappaB activation through positively regulating ROS production in pancreatic beta cells. PLoS ONE, 10, e0127705.
Froy, O. (2012). Circadian rhythms and obesity in mammals. ISRN Obes, 2012, 437198.
Wang, C. Y., Shie, S., Hsieh, I. C., Tsai, M. L., & Wen, M. S. (2015). FTO modulates circadian rhythms and inhibits the CLOCK-BMAL1-induced transcription. Biochemical and Biophysical Research Communications, 464(3), 826–832. https://doi.org/10.1016/j.bbrc.2015.07.046.
Tang, C., Klukovich, R., Peng, H., Wang, Z., Yu, T., Zhang, Y., et al. (2018). ALKBH5-dependent m6A demethylation controls splicing and stability of long 3'-UTR mRNAs in male germ cells. Proc Natl Acad Sci U S A, 115, E325–E333.
Thalhammer, A., Bencokova, Z., Poole, R., Loenarz, C., Adam, J., O'Flaherty, L., et al. (2011). Human AlkB homologue 5 is a nuclear 2-oxoglutarate dependent oxygenase and a direct target of hypoxia-inducible factor 1alpha (HIF-1alpha). PLoS ONE, 6, e16210.
Zhang, C., Samanta, D., Lu, H., Bullen, J. W., Zhang, H., Chen, I., et al. (2016). Hypoxia induces the breast cancer stem cell phenotype by HIF-dependent and ALKBH5-mediated m(6)A-demethylation of NANOG mRNA. Proceedings of the National Academy of Sciences of the United States of America, 113, E2047–2056.
Zhang, C., Zhi, W. I., Lu, H., Samanta, D., Chen, I., Gabrielson, E., et al. (2016). Hypoxia-inducible factors regulate pluripotency factor expression by ZNF217- and ALKBH5-mediated modulation of RNA methylation in breast cancer cells. Oncotarget, 7, 64527–64542.
Li, X. C., Jin, F., Wang, B. Y., Yin, X. J., Hong, W., & Tian, F. J. (2019). The m6A demethylase ALKBH5 controls trophoblast invasion at the maternal-fetal interface by regulating the stability of CYR61 mRNA. Theranostics, 9, 3853–3865.
Lin, S., Liu, J., Jiang, W., Wang, P., Sun, C., Wang, X., et al. (2019). METTL3 promotes the proliferation and mobility of gastric cancer cells. Open Med (Wars), 14, 25–31.
Cheng, M., Sheng, L., Gao, Q., Xiong, Q., Zhang, H., Wu, M., et al. (2019). The m(6)A methyltransferase METTL3 promotes bladder cancer progression via AFF4/NF-kappaB/MYC signaling network. Oncogene, 38, 3667–3680.
Han, J., Wang, J. Z., Yang, X., Yu, H., Zhou, R., Lu, H. C., et al. (2019). METTL3 promote tumor proliferation of bladder cancer by accelerating pri-miR221/222 maturation in m6A-dependent manner. Molecular Cancer, 18, 110.
Dahal, U., Le, K., & Gupta, M. (2019). RNA m6A methyltransferase METTL3 regulates invasiveness of melanoma cells by matrix metallopeptidase 2. Melanoma Research, 29, 382–389.
Visvanathan, A., Patil, V., Abdulla, S., Hoheisel, J. D., & Somasundaram, K. (2019). N(6)-methyladenosine landscape of glioma stem-like cells: METTL3 is essential for the expression of actively transcribed genes and sustenance of the oncogenic signaling. Genes (Basel). https://doi.org/10.1038/onc.2017.351.
Visvanathan, A., Patil, V., Arora, A., Hegde, A. S., Arivazhagan, A., Santosh, V., et al. (2018). Essential role of METTL3-mediated m(6)A modification in glioma stem-like cells maintenance and radioresistance. Oncogene, 37, 522–533.
Hua, W., Zhao, Y., Jin, X., Yu, D., He, J., Xie, D., et al. (2018). METTL3 promotes ovarian carcinoma growth and invasion through the regulation of AXL translation and epithelial to mesenchymal transition. Gynecologic Oncology, 151, 356–365.
Liu, J., Eckert, M. A., Harada, B. T., Liu, S. M., Lu, Z., Yu, K., et al. (2018). m(6)A mRNA methylation regulates AKT activity to promote the proliferation and tumorigenicity of endometrial cancer. Nature Cell Biology, 20, 1074–1083.
Taketo, K., Konno, M., Asai, A., Koseki, J., Toratani, M., Satoh, T., et al. (2018). The epitranscriptome m6A writer METTL3 promotes chemo- and radioresistance in pancreatic cancer cells. International Journal of Oncology, 52, 621–629.
Li, X., Tang, J., Huang, W., Wang, F., Li, P., Qin, C., et al. (2017). The M6A methyltransferase METTL3: acting as a tumor suppressor in renal cell carcinoma. Oncotarget, 8, 96103–96116.
Yue, B., Song, C., Yang, L., Cui, R., Cheng, X., Zhang, Z., et al. (2019). METTL3-mediated N6-methyladenosine modification is critical for epithelial-mesenchymal transition and metastasis of gastric cancer. Molecular Cancer, 18, 142.
Liu, L., Wang, J., Sun, G., Wu, Q., Ma, J., Zhang, X., et al. (2019). m(6)A mRNA methylation regulates CTNNB1 to promote the proliferation of hepatoblastoma. Molecular Cancer, 18, 188.
Cai, J., Yang, F., Zhan, H., Situ, J., Li, W., Mao, Y., et al. (2019). RNA m(6)A methyltransferase METTL3 promotes the growth of prostate cancer by regulating Hedgehog pathway. Onco Targets Therapy, 12, 9143–9152.
Wang, J., Yan, S., Lu, H., Wang, S., & Xu, D. (2019). METTL3 attenuates LPS-induced inflammatory response in macrophages via NF-kappab signaling pathway. Mediators of Inflammation, 2019, 3120391.
Xie, W., Ma, L. L., Xu, Y. Q., Wang, B. H., & Li, S. M. (2019). METTL3 inhibits hepatic insulin sensitivity via N6-methyladenosine modification of Fasn mRNA and promoting fatty acid metabolism. Biochemical and Biophysical Research Communications, 518, 120–126.
Miao, W., Chen, J., Jia, L., Ma, J., & Song, D. (2019). The m6A methyltransferase METTL3 promotes osteosarcoma progression by regulating the m6A level of LEF1. Biochemical and Biophysical Research Communications, 516, 719–725.
Jin, D., Guo, J., Wu, Y., Du, J., Yang, L., Wang, X., et al. (2019). m(6)A mRNA methylation initiated by METTL3 directly promotes YAP translation and increases YAP activity by regulating the MALAT1-miR-1914-3p-YAP axis to induce NSCLC drug resistance and metastasis. Journal of Hematology & Oncology, 12, 135.
Zhao, W., Cui, Y., Liu, L., Ma, X., Qi, X., Wang, Y., et al. (2020). METTL3 facilitates oral squamous cell carcinoma tumorigenesis by enhancing c-Myc stability via YTHDF1-mediated m(6)A modification. Molecular Therapy - Nucleic Acids, 20, 1–12.
Xie, H., Li, J., Ying, Y., Yan, H., Jin, K., Ma, X., et al. (2020). METTL3/YTHDF2 m(6) A axis promotes tumorigenesis by degrading SETD7 and KLF4 mRNAs in bladder cancer. Journal of Cellular and Molecular Medicine. https://doi.org/10.2139/ssrn.3429876.
Deng, R., Cheng, Y., Ye, S., Zhang, J., Huang, R., Li, P., et al. (2019). m(6)A methyltransferase METTL3 suppresses colorectal cancer proliferation and migration through p38/ERK pathways. Onco Targets Ther, 12, 4391–4402.
Zhu, W., Si, Y., Xu, J., Lin, Y., Wang, J. Z., Cao, M., et al. (2020). Methyltransferase like 3 promotes colorectal cancer proliferation by stabilizing CCNE1 mRNA in an m6A-dependent manner. Journal of Cellular and Molecular Medicine, 24, 3521–3533.
Li, T., Hu, P. S., Zuo, Z., Lin, J. F., Li, X., Wu, Q. N., et al. (2019). METTL3 facilitates tumor progression via an m(6)A-IGF2BP2-dependent mechanism in colorectal carcinoma. Molecular Cancer, 18, 112.
Hao, H., Hao, S., Chen, H., Chen, Z., Zhang, Y., Wang, J., et al. (2019). N6-methyladenosine modification and METTL3 modulate enterovirus 71 replication. Nucleic Acids Research, 47, 362–374.
Wei, W., Huo, B., & Shi, X. (2019). miR-600 inhibits lung cancer via downregulating the expression of METTL3. Cancer Management and Research, 11, 1177–1187.
Hesser, C. R., Karijolich, J., Dominissini, D., He, C., & Glaunsinger, B. A. (2018). N6-methyladenosine modification and the YTHDF2 reader protein play cell type specific roles in lytic viral gene expression during Kaposi's sarcoma-associated herpesvirus infection. PLoS Pathogens, 14, e1006995.
Lichinchi, G., Gao, S., Saletore, Y., Gonzalez, G. M., Bansal, V., Wang, Y., et al. (2016). Dynamics of the human and viral m(6)A RNA methylomes during HIV-1 infection of T cells. Nat Microbiol, 1, 16011.
Courtney, D. G., Kennedy, E. M., Dumm, R. E., Bogerd, H. P., Tsai, K., Heaton, N. S., et al. (2017). Epitranscriptomic enhancement of influenza A virus gene expression and replication. Cell Host & Microbe, 22(377–386), e375.
Imam, H., Khan, M., Gokhale, N. S., McIntyre, A. B. R., Kim, G. W., Jang, J. Y., et al. (2018). N6-methyladenosine modification of hepatitis B virus RNA differentially regulates the viral life cycle. Proceedings of the National Academy of Sciences USA, 115, 8829–8834.
Lichinchi, G., Zhao, B. S., Wu, Y., Lu, Z., Qin, Y., He, C., et al. (2016). Dynamics of human and viral RNA methylation during Zika virus infection. Cell Host & Microbe, 20, 666–673.
Chen, X., Xu, M., Xu, X., Zeng, K., Liu, X., Sun, L., et al. (2020). METTL14 suppresses CRC progression via regulating N6-methyladenosine-dependent primary miR-375 processing. Molecular Therapy, 28, 599–612.
Jo, H. J., Shim, H. E., Han, M. E., Kim, H. J., Kim, K. S., Baek, S., et al. (2013). WTAP regulates migration and invasion of cholangiocarcinoma cells. Journal of Gastroenterology, 48, 1271–1282.
Xi, Z., Xue, Y., Zheng, J., Liu, X., Ma, J., & Liu, Y. (2016). WTAP expression predicts poor prognosis in malignant glioma patients. Journal of Molecular Neuroscience, 60, 131–136.
Huang, H., Weng, H., Sun, W., Qin, X., Shi, H., Wu, H., et al. (2018). Author correction: recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nature Cell Biology, 20, 1098.
Muller, S., Glass, M., Singh, A. K., Haase, J., Bley, N., Fuchs, T., et al. (2019). IGF2BP1 promotes SRF-dependent transcription in cancer in a m6A- and miRNA-dependent manner. Nucleic Acids Research, 47, 375–390.
Zhao, X., Chen, Y., Mao, Q., Jiang, X., Jiang, W., Chen, J., et al. (2018). Overexpression of YTHDF1 is associated with poor prognosis in patients with hepatocellular carcinoma. Cancer Biomarkers, 21, 859–868.
Nishizawa, Y., Konno, M., Asai, A., Koseki, J., Kawamoto, K., Miyoshi, N., et al. (2018). Oncogene c-Myc promotes epitranscriptome m(6)A reader YTHDF1 expression in colorectal cancer. Oncotarget, 9, 7476–7486.
Li, Z., Weng, H., Su, R., Weng, X., Zuo, Z., Li, C., et al. (2017). FTO plays an oncogenic role in acute myeloid leukemia as a N(6)-methyladenosine RNA demethylase. Cancer Cell, 31, 127–141.
Liu, J., Ren, D., Du, Z., Wang, H., Zhang, H., & Jin, Y. (2018). m(6)A demethylase FTO facilitates tumor progression in lung squamous cell carcinoma by regulating MZF1 expression. Biochemical and Biophysical Research Communications, 502, 456–464.
Zhou, S., Bai, Z. L., Xia, D., Zhao, Z. J., Zhao, R., Wang, Y. Y., et al. (2018). TO regulates the chemo-radiotherapy resistance of cervical squamous cell carcinoma (CSCC) by targeting beta-catenin through mRNA demethylation. Molecular Carcinogenesis, 57, 590–597.
Tang, X., Liu, S., Chen, D., Zhao, Z., & Zhou, J. (2019). The role of the fat mass and obesity-associated protein in the proliferation of pancreatic cancer cells. Oncology Letters, 17, 2473–2478.
Zhuang, C., Zhuang, C., Luo, X., Huang, X., Yao, L., Li, J., et al. (2019). N6-methyladenosine demethylase FTO suppresses clear cell renal cell carcinoma through a novel FTO-PGC-1alpha signalling axis. Journal of Cellular and Molecular Medicine, 23, 2163–2173.
Niu, Y., Lin, Z., Wan, A., Chen, H., Liang, H., Sun, L., et al. (2019). RNA N6-methyladenosine demethylase FTO promotes breast tumor progression through inhibiting BNIP3. Molecular Cancer, 18, 46.
Zhang, S., Zhao, B. S., Zhou, A., Lin, K., Zheng, S., Lu, Z., et al. (2017). m(6)A demethylase ALKBH5 maintains tumorigenicity of glioblastoma stem-like cells by sustaining FOXM1s. Cancer Cell, 31(591–606), e596.
Acknowledgements
This work was supported by an extramural grant from DST-SERB, Government of India [Grant Number CRG/2018/000492 (to P.K.)]. P.K. acknowledges infrastructural and financial support from BITS Pilani-Hyderabad Campus, India. K.R. acknowledges fellowship from BITS Pilani-Hyderabad Campus, India. We apologize to our colleagues whose work we could not mention because of space limitations.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Karthiya, R., Khandelia, P. m6A RNA Methylation: Ramifications for Gene Expression and Human Health. Mol Biotechnol 62, 467–484 (2020). https://doi.org/10.1007/s12033-020-00269-5
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-020-00269-5