Abstract
Breast cancer is the second most commonly diagnosed cancer, worldwide. Human epidermal growth factor receptor 2 (HER2)-overexpressing breast cancer is correlated with poor prognosis. HER2-targeting monoclonal antibodies resulted in longer survival of HER2+ breast cancer. Single-chain variable fragment (scFv) demonstrates improved penetrability into tumors. Due to the presence of two disulfide bond, scFv expression in reducing bacterial cytoplasm may cause formation of inclusion bodies. Disulfide bond can be formed properly in cytoplasm of SHuffle® strain as it is trxB−, gor−, and overexpresses cytoplasmic DsbC chaperone. In this study, the anti-HER2 scFv was successfully expressed and purified in BL21 (DE3) and SHuffle® cells. Here, significant higher soluble anti-HER2 scFv was produced in SHuffle® than in BL21 strain. The specific binding of anti-HER2 scFv to HER2 was shown by flow cytometry analysis and ELISA. Moreover, it was demonstrated that the anti-HER2 scFv produced in SHuffle® binds to HER2 at higher level as compared to that expressed in BL21 cells. Furthermore, competitive ELISA-based study suggested that anti-HER2 scFv recognizes the same epitope of HER2 receptor as the trastuzumab antibody. Our findings indicated that correct disulfide bond formation in SHuffle® strain can result in enhanced solubility and higher biological activity level of anti-HER2 scFv.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is one of the most commonly diagnosed cancers, worldwide [1] and also among Iranian women [2]. 25–30% of breast cancers are associated with human epidermal growth factor receptor 2 (HER2) overexpression, a signaling tyrosine kinase receptor causing epithelial cell growth and differentiation, tumor invasiveness, accelerated angiogenesis, and decreased apoptosis [3,4,5,6]. HER2-overexpressing breast cancers are correlated with poor prognosis and decreased overall survival [5, 7]. The humanized monoclonal antibody against HER2 (trastuzumab, Herceptin®) binds to the extracellular domain of HER2 and has been reported to improve the overall survival in metastatic breast cancer when administered in the adjuvant setting [8, 9]. Variable regions of heavy (VH) and light (VL) chains of immunoglobulins are linked via a flexible linker encoding a short peptide to form single-chain variable fragments (scFv) [10]. Due to lack of Fc domain in scFvs and consequently their small size, they demonstrated better tumor penetration and faster pharmacokinetics as compared to full-length antibody [11]. The targeting abilities of scFvs have been used in several therapeutic and diagnostic applications. Many studies have been reported the application of antibody fragment targeting HER2 for construction of immunotoxins [12, 13] and anti-HER2 immunoliposomes [14,15,16] as targeted cancer therapeutics.
Escherichia coli (E. coli) is the most common platform for expression of recombinant proteins and has many advantages such as relatively low cost, simplicity, and high yields compared to eukaryotic biological systems [17]. Due to lack of glycosylation in scFvs, they can be expressed in E. coli [18]. Two specialized systems including thioredoxin (thioredoxins and thioredoxin reductase) as well as glutaredoxin (glutaredoxins, glutathione, and glutathione oxidoreductase) pathways result in holding free thiol groups in a reduced state in E. coli cytoplasm [10, 19]. Therefore, formation of disulfide bond is rare in the reducing cytoplasmic environment of E. coli. Consequently, inactive insoluble aggregates called inclusion bodies are formed [20, 21]. To enhance the solubility of disulfide bond containing protein, a complicated and detailed oxidative folding process is required [22]. In addition, the refolding methods may not completely be able to reform the native protein fold and may decrease the functionality of the recombinant protein [23, 24]. Despite all the limitation mentioned above, functional expression of disulfide bond containing proteins in the cytoplasm is possible by using engineered E. coli strains with oxidative cytoplasmic environment [25]. Recently, the genetically engineered expression strain, E. coli SHuffle® T7 Express (SHuffle®) (BL21 (DE3) background New England BioLab, USA), has become available especially for expression of proteins requiring disulfide bonds for their activity. This strain has trxB−, gor− mutations as the commercial Origami (Novagen), and also overexpresses cytoplasmic disulfide bond isomerase (DsbC) [26, 27]. DsbC acts as both protein disulfide isomerase and chaperone and subsequently its overexpression lead to lesser protein aggregates into inactive inclusion bodies [28, 29].
Although the expression of scFvs against HER2 in E. coli strains have been studied by several research groups [11, 30], the expression of scFv in the oxidative cytoplasmic environment of E. coli SHuffle® has not been reported. Herein, the scFv version of trastuzumab (anti-HER2 scFv) as a disulfide bonds containing protein was expressed in E. coli SHuffle® and BL21 (DE3) strains. Furthermore, the soluble expression of anti-HER2 scFv in E. coli cytoplasm was compared in this two strains. In the present study, we provided the first report, to our knowledge, for comparing the biological activity of a disulfide bond containing protein expressed in E. coli SHuffle® and BL21 (DE3) strains. Herein, different methods including HER2− and cell-based ELISA as well as flowcytometry were used to compare the biological activity of anti-HER2 scFv expressed in E. coli SHuffle® and BL21 (DE3) strains.
Methods and Materials
Bacterial Strains, Plasmid, and Reagents
Escherichia coli strains BL21 (DE3) and SHuffle® (NEB) and also the protein expression vector pET-22b (+) were kindly gifted from Dr. Nematollahi and Dr. Behdani, respectively (Pasteur Institute of Iran). E. coli strain was grown in Luria–Bertani (LB) medium (Floka). The growth medium was supplemented with the ampicillin (100 µg/mL) and spectinomycin (Fisiopharma S.r.I.-Salerno-Italy) at final concentration of 50 µg/mL, when required. All chemicals and reagents used were provided from standard commercial sources such as Merck, unless otherwise stated.
The human breast carcinoma cell line SK-BR-3, and HER2-low-expressing cells MDA-MB-231 were obtained from National Cell Bank of Iran (NCBI) (Pasture Institute, Iran, Tehran) and grown in RPMI 1640 medium (Biosera) supplemented with 10% (v/v) fetal bovine serum (FBS) (Gibco, Scotland) and 1% penicillin–streptomycin (100 IU/mL penicillin and 100 μg/mL streptomycin) (Gibco, Scotland) at 37 °C and in a humidified atmosphere containing 5% CO2.
Construction of Recombinant Anti-HER2 scFv Expressing Plasmid
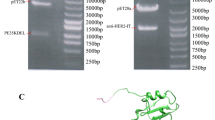
The variable heavy (VH) and light (VL) chains of trastuzumab (drug bank number DB00072) were linked using (G4S)3 linker and then codon optimized for high expression in E. coli (Generay biotech, Shanghai, China). The NcoI/XhoI-digested fragment containing the anti-HER2 scFv gene (756 bp) was gel-extracted (MEGA quick gel extraction kit, iNtRON biotechnology) and then cloned into pET-22b (+) expression vector, in frame with His-tag to detect and also purify the expressed protein (Fig. 1). Additionally, the anti-HER2 scFv gene was cloned in the same sites of pET-28b (+) (without pelB). Eventually, the recombinant purified pET-22 and pET-28 plasmids containing anti-HER2 scFv gene were sequenced (Bioneer, Korea) to confirm the correct expression frame.
Expression of Anti-HER2 scFv Protein
The recombinant pET-22 plasmid containing anti-HER2 scFv gene was transformed into competent SHuffle® and BL21 (DE3) strain. Furthermore, pET-28 (anti-HER2 scFv) was transformed into competent SHuffle® strain. Recombinant SHuffle® clones containing pET-22 (anti-HER2 scFv) were selected and grown on LB agar plates containing 100 μg/mL ampicillin and 50 μg/mL spectinomycin. The selection of recombinant SHuffle® clones containing pET-28 (anti-HER2 scFv) was carried on the plates containing 25 μg/mL kanamycin and 50 μg/mL spectinomycin. A positive colony of each host was used to inoculate pre-culture LB broth medium containing the proper antibiotic and incubated with shaking overnight. The cultures were shaken at 30 °C and 200 rpm for SHuffle® cells and at 37 °C and 180 rpm for BL21 (DE3). Then, LB medium containing the appropriate antibiotic was inoculated with the overnight culture to a final OD600 of 0.1. When OD600 reached 0.6 to 0.7, the expression was induced by adding 1 mM isopropyl ß-d-1 thiogalactopyranoside (IPTG) and the induced culture was shaken for another 2 h under the previously stated conditions for both strains. After the induction step, centrifugation (4 °C, 10,000×g for 5 min) was applied to collect the bacterial biomass and then, the bacterial pellet was resuspended in sample buffer 2X (4% SDS, 20% glycerol, 2% 2-mercaptoethanol (2-ME), 0.01% bromophenol blue, 500 mM Tris–HCl, pH 6.5). To analyze protein expression, the samples were subjected to 12.5% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) gel. Then, total Lab TL 120 software was used to determine the percentage of anti-HER2 scFv band in stained SDS-PAGE gels. BCA assay (Parstous Company) using the bovine serum albumin (BSA, Atocell) as the protein standard was conducted to determine the total protein concentration of each sample. To define the concentration of anti-HER2 scFv protein, the band percentage was multiplied by total protein concentration of the sample.
Determining the Effect of Different Induction Conditions on Anti-HER2 scFv Expression
The role of three factors including induction temperature, concentration of inducer (IPTG), and harvesting time on the expression of anti-HER2 scFv in BL21 (DE3) and SHuffle® strains was analyzed. To identify the effect of induction temperature on anti-HER2 scFv expression, the bacteria were grown at 30 °C for SHuffle® and at 37 °C for BL21 (DE3) cells to reach the optimal density (OD600 of 0.6–0.7). Afterward, anti-HER2 scFv expression was induced using 1 mM IPTG and the growth culture was carried out at different temperatures (15, 25, 30, or 37 °C) in both SHuffle® and BL21 (DE3) strains. Then, the samples were collected 2, 4, 6, and 24 h after induction. To determine the influence of IPTG concentration, after reaching the desired OD600 of 0.6–0.7, different concentrations of inducer (0.01, 0.05, 0.1, 0.25, 0.5, and 1 mM for SHuffle® and 0.25, 0.5, 1, and 2 mM for BL21 (DE3) cells) were added. Then, the bacteria cultures were carried out at the optimum induction temperature of the examined for each strain.
Recombinant Anti-HER2 scFv Protein Purification
The induced bacterial cells were thawed on ice and resuspended in the lysis buffer (NaH2PO4 50 mM, NaCl 300 mM, Imidazole 10 mM; pH 8). Then, 1 mg/mL of lysozyme, DNase (100 μg/mL), and MgSO4 (100 mM) were added and incubation on ice was carried out for 30 min. Afterward, the suspension was disrupted using sonication on ice for 30 min [300 W, 7 s working and 8 s resting, (Topsonics, Iran)] and subsequently the cell debris was removed by centrifugation at 10,000×g at 4 °C for 25 min. In the next step, the presence of anti-HER2 scFv in supernatant (the soluble fraction) was verified by SDS-PAGE analysis and then was subjected to the affinity chromatography column packed with high-capacity Ni–NTA agarose beads under native condition as described by the manufacturer (Qiagen, Netherlands). To wash the column and then elute the anti-HER2 scFv from the Ni–NTA column, buffers containing 20 and 250 mM imidazole were used, respectively. Next, the eluted fractions were dialyzed against phosphate-buffered solution (PBS, pH 7.4) using 14 kDa molecular weight cut-off (MWCO) dialysis tubing (Sigma-Aldrich) at 4 °C and then syringe-filtered for further experiments.
Western Blotting
The separated bands of total proteins from recombinant BL21 (DE3) and SHuffle® strains were electrophoretically transferred to a polyvinylidene difluoride (PVDF) membrane (Roche) (100 V for 90 min). Then, blocking of the nonspecific binding on blotted membrane was performed using 2.5% BSA in tris buffered saline with 0.1% tween 20 (TBS-T) overnight at 4 °C. After performing three times washing procedure using wash buffer (TBS-T), mouse monoclonal anti-polyhistidine antibody (Sigma-Aldrich) (diluted 1:10,000 in blocking buffer) was added to the membrane and incubated at room temperature for 90 min. After another washing procedure, goat peroxidase-conjugated anti-mouse IgG antibody (Sigma-Aldrich) (diluted 1:5000 in blocking buffer) was added as the secondary antibody and the membrane was incubated at room temperature for 60 min. Finally, membrane was visualized by adding 0.6 mg/mL 3,3′-diaminobenzidine (DAB, Sigma) in 0.12% H2O2 and 1 M Tris–HCl.
Flow Cytometry Analysis
The cultured SK-BR-3 and MDA-MB-231 cells were washed and resuspended in PBS/BSA 1% and approximately 106 cells were plated in 96-well U-type bottom plate (SPL, South Korea). Then, cells were incubated with 100 ng/µL of anti-HER2 scFv expressed in BL21 (D3) and SHuffle® (diluted in PBS/BSA 1%) for 30 min at 4 °C. The treated cells were centrifuged (4 °C, 500×g for 5 min) and then resuspended in PBS/BSA 1%. After performing two times washing procedures with PBS/BSA 1%, the cells were incubated with 100 µL mouse anti-polyhistidine antibody (diluted in PBS/BSA 1%, 1:1500) for 30 min at 4 °C in dark place. Then, after two times washing with PBS/BSA 1%, the cells were incubated with 100 μL of FITC-conjugated anti-mouse IgG (FC specific) (diluted in PBS/BSA 1%, 1:200) in the same conditions as stated in the previous step. After two times washing, cells were resuspended in PBS. Flow cytometry data were acquired on a FACSCalibur flow cytometer (BD, USA), then analyzed and presented using Flow Jo software (Treestar, USA). Cell incubation with anti-polyhistidine antibody and FITC anti-mouse IgG without any treatment of scFv was performed to determine the background fluorescence.
Enzyme‑Linked Immuno Sorbent Assays (ELISA)
Recombinant HER2 antigen (500 pg/μL, Invitrogen, USA) in coating buffer (0.2 M Na2CO3/NaHCO3, pH 9.6; 100 μL per well) was adsorbed to 96-well flat-bottom ELISA plate (SPL, South Korea) by overnight incubation at 4 °C. Then, the plate was washed 3 times using phosphate-buffered saline with 0.05% tween 20 (PBS-T) to remove unbound antigen. Blocking was accomplished by incubating the plate with 100 μL/well of 5% skim milk (Merck) in PBS for 120 min at room temperature on shaker and subsequently the wells were washed. Then, 100 μL of purified anti-HER2 scFv per each well (10 and 100 ng/μL) was bound to the HER2 antigen by incubating at 37 °C for 60 min. After five times washing, the plate was incubated by monoclonal anti-polyhistidine antibody produced in mouse (diluted in blocking buffer, 1:1500) for 120 min at room temperature on shaker. After another rinsing procedure, incubation was done in the presence of anti-mouse IgG (FC specific) peroxidase antibody in goat (diluted in blocking buffer, 1:5000) and then the wells were washed in the same conditions. Finally, 100 μL of TMB solution (BD bioscience) containing 1.5% H2O2 was added to each well for 20 min. After adding stop solution (1 M H2SO4), the absorbance was measured at 450 nm (ELISA plate reader, Synergy Biotech, USA).
Competitive Binding Assay to HER2
To evaluate whether anti-HER2 scFv recognizes the same epitope of HER2 receptor as the trastuzumab, competitive binding of trastuzumab (AryoTrustTM, Aryogen pharmed, Iran) to HER2 in the presence of anti-HER2 scFv was assessed. Accordingly, as described earlier recombinant HER2 antigen was coated to 96-well ELISA plate and then, incubated by blocking buffer. After washing procedure, purified anti-HER2 scFv (100 ng/μL) was added and incubated for 60 min at 37 °C. The plate was washed five times with PBS-T and then trastuzumab (1 and 10 ng/μL) was added to compete with anti-HER2 scFv for 60 min at 37 °C. Then, after rinsing the wells, they were incubated with anti-Human IgG (FC specific) peroxidase antibody in goat (diluted in blocking buffer, 1:5000) for 120 min at room temperature. Subsequently, the absorbance was measured as previously stated.
Cell-Based ELISA to Analyze HER2 Binding
The binding ability of anti-HER2 scFv to HER2-overexpressing SK-BR-3 and HER2-low-expressing MDA-MB-231 cells was performed by cell-based ELISA assay. For this purpose, SK-BR-3 and MDA-MB-231 cells were seeded (103–105 cells/mL) into 96-well plates for attachment overnight. The cells were washed with PBS and then fixed by 10% neutral formaldehyde (Merck) (0.1 mol/L PBS, 10% formaldehyde, pH 7.4) at room temperature for 1 h. Then, after washing the fixed cells were blocked with PBS containing 5% skim milk (Merck) for 2 h. After another washing procedure, the cells were incubated with 100 ng/μL of purified anti-HER2 scFv at 37 °C for 1 h. Subsequently, the cells were washed five times with PBS-T (PBS with 0.5% tween 20) and incubated with monoclonal anti-polyhistidine antibody produced in mouse (diluted in blocking buffer 1:1500) at room temperature for 2 h and washed. Further, the cells were incubated in the presence of mouse anti-polyhistidine antibody and then anti-mouse IgG (FC specific) peroxidase antibody as previously explained. Finally, the cells were washed 5 times with PBS-T and the color was developed as mentioned earlier.
Statistical Analysis
To perform statistical analysis, GraphPad Prism 6.0 for Windows (GraphPad Prism, San Diego, California, USA) was used. One-way ANOVA with Tukey’s post hoc test was applied to analyze the data of ELISA experiments. The data were presented as mean ± standard deviation (SD) of two independent experiments in duplicate. The p value less than 0.05 (p < 0.05) was considered statistically significant.
Results
Expression, Purification, and Identification of the Recombinant Anti-HER2 scFv
Recombinant pET-22 plasmid containing anti-HER2 scFv gene pET-22 (anti-HER2 scFv) (Fig. 1) was transformed into E. coli strains, BL21 (DE3) and SHuffle®. pET-28 (anti-HER2 scFv) was also transformed into SHuffle®. As indicated in Fig. 2, both BL21 (DE3) and SHuffle® reached their log phase (OD600 ~ 0.6) approximately 2 h after bacterial inoculation. However, the growth curve demonstrated slower growth rate of SHuffle® than that of BL21 (DE3), after induction. Then, the recombinant bacteria were induced by 1 mM IPTG at 37 °C in BL21 (DE3) and 30 °C in SHuffle®. The results showed that a recombinant protein with the size of 28 kDa was induced successfully (Fig. 3a). The expressed scFv protein was also analyzed by Western blot, with an antibody directed against the His-tag. The blot revealed a single band of approximately 28 kDa (Fig. 3b). Anti-HER2 scFv was purified using Ni–NTA affinity chromatography under native condition. SDS-PAGE results displayed the presence of highly purified protein by a single band of approximately 28 kDa (Fig. 3a). As demonstrated in Fig. 4, there was no significant difference between the amount of anti-HER2 scFv expressed by SHuffle® containing pET-22 (anti-HER2 scFv) and pET-28 (anti-HER2 scFv). Even, the amount of anti-HER2 scFv produced by SHuffle® containing pET-22 (anti-HER2 scFv) was higher than that expressed by SHuffle® containing pET-28 (anti-HER2 scFv), 24 h after induction. Therefore, as it is more suitable to decrease the variables for comparing BL21 (DE3) and SHuffle®, SHuffle® containing pET-22 (anti-HER2 scFv) was selected for comparison between these two strains.
Anti-HER2 scFv expression, purification, and Western blot analysis. a SDS-PAGE analysis of anti-HER2 scFv expressed in BL21 (DE3) and SHuffle® strains. Total protein from E. coli SHuffle® containing pET-22 (anti-HER2 scFv) plasmid after induction with 1 mM IPTG for 24 h at 30 °C (lane 1) and before induction (lane 2), protein marker (Fermentas, MW, lane 3), total protein from E. coli BL21 (DE3) containing pET-22 (anti-HER2 scFv) plasmid before induction (lane 4) and after induction with 1 mM IPTG for 24 h at 37 °C (lane 5), purified anti-HER2 scFv expressed from E. coli BL21 (DE3) (lane 6). b Western blot analysis of the expressed anti-HER2 scFv using anti-His-tag antibody. Prestained protein marker (sinaclon) (lane 1), Total protein from E. coli BL21 (DE3) containing pET-22 (anti-HER2 scFv) plasmid before induction (lane 2) and after induction with 1 mM IPTG for 24 h at 37 °C (lane 3), total protein from SHuffle® strain containing pET-22 (anti-HER2 scFv) plasmid after 24-h induction at 30 °C (lane 4)
Expression of anti-HER2 scFv by SHuffle® strains containing pET-22 (anti-HER2 scFv) and pET-28 (anti-HER2 scFv). The expression of anti-HER2 scFv protein was analyzed at various harvesting times following IPTG induction (1 mM) at 30 °C in SHuffle® strains containing pET-22 (anti-HER2 scFv) and pET-28 (anti-HER2 scFv)
The Expression of Anti-HER2 scFv Under Different Induction Conditions in BL21 (DE3) and SHuffle®E. coli Strains
In our study, different factors including the duration and temperature of induction and also the concentration of inducer (IPTG) were investigated on anti-HER2 scFv expression level. To optimize the induction duration, the cell pellet was collected by centrifugation 2, 4, 6, and 24 h after induction. In both strains, significant higher amount of anti-HER2 scFv was expressed 24 h after IPTG induction in comparison with other times (Fig. 5a).
Expression of anti-HER2 scFv under different induction conditions. a The expression of anti-HER2 scFv protein at various harvesting times following IPTG induction (1 mM) at 37 °C in BL21 and at 30 °C in SHuffle®, p < 0.01 (filled rhombus) was considered as significant difference between 24- and 4-h post-induction times in BL21 and p < 0.0001 (filled circle) demonstrated significant difference between 24- and 6-h post-induction times in SHuffle®. b Anti-HER2 scFv expression at different temperatures induced by IPTG (1 mM) after 24 h in BL21 and SHuffle® strains. c Induction of recombinant anti-HER2 scFv by different concentrations of IPTG at 37 °C for 24 h in BL21 and at 30 °C for 24 h in SHuffle® strain. Data represented as mean ± SD of two independent experiments. p < 0.01 (**) and p < 0.001 (***) were considered as significant difference
The effect of temperature on anti-HER2 scFv expression in both hosts was also examined. In BL21 (DE3), the expression level of anti-HER2 scFv at 37 °C was significantly higher than those at 25 and 30 °C (anti-HER2 scFv expressed in BL21 (DE3) cells at 37 °C vs. 25 and 30 °C, p < 0.001) (Fig. 5b). However, the highest anti-HER2 scFv was expressed at 30 °C in SHuffle® strain, (anti-HER2 scFv expressed in SHuffle® at 30 °C vs. 37 °C, p < 0.01) (Fig. 5b).
In the next step, we sought to determine the optimal IPTG inducing concentration. As shown in Fig. 5c, induction of BL21 (DE3) strain with 0.25 mM IPTG resulted in significant higher amount of anti-HER2 scFv as compared to induction with 1 and 2 mM IPTG (anti-HER2 scFv induced by 0.25 mM vs. 1 mM IPTG, p < 0.001). Also, we found that the concentrations of IPTG had no obvious effect on anti-HER2 scFv expression in SHuffle® (Fig. 5c). (The SDS-PAGE images of different expression conditions are shown in supplementary 1–6). The final production yields of the anti-HER2 scFv at optimal condition for BL21 (DE3) (24 h after induction with 0.25 mM IPTG at 37 °C) and SHuffle® (24 h after induction with 0.05 mM IPTG at 30 °C) were 214 and 147 mg/L, respectively.
Comparison of BL21 (DE3) and SHuffle®E. coli Strains for Soluble Anti-HER2 scFv Expression
In order to investigate the effect of temperature on anti-HER2 scFv soluble expression in both E. coli BL21 (DE3) and SHuffle® strains, the transformed cells were grown to OD600 0.6 and then induced using 0.25 mM IPTG for BL21 (DE3) and 0.05 mM IPTG for SHuffle® at different temperatures (25, 30, and 37 °C). The solubility of anti-HER2 scFv protein expressed in BL21 (DE3) strain decreased as the induction temperature increased, such that the solubility of anti-HER2 scFv was significantly higher when induced at 25 °C as compared to 37 °C. However, the solubility of anti-HER2 scFv in SHuffle® was not affected by temperature (Fig. 6, supplementary 7 and 8).
The solubility of anti-HER2 scFv expressed in E. coli BL21 (DE3) and SHuffle®. E. coli BL21 (DE3) and SHuffle® containing pET-22 (anti-HER2 scFv) were induced at different temperatures (25, 30, and 37 °C) using 0.25 and 0.05 mM IPTG, respectively. Data represented as mean ± SD of two independent experiments. (*) and (**) represent for p < 0.05 and p < 0.01, respectively, and considered as significant difference
As demonstrated in Fig. 6, the soluble/insoluble ratio for anti-HER2 scFv expressed in SHuffle® cells at 30 °C was significantly higher than those expressed in BL21 (DE3) at 30 and also 37 °C (soluble/insoluble ratio of anti-HER2 scFv expressed in SHuffle® at 30 °C vs. anti-HER2 scFv expressed in BL21 (DE3) strain at 30 and 37 °C, p < 0.01).
Flow Cytometry Analysis of Anti-HER2 scFv Binding to HER2-Positive Cells
The binding ability and specificity of anti-HER2 scFv expressed in BL21 (DE3) and SHuffle® strains were examined by flowcytometry analysis using SK-BR-3 cells and MDA-MB-231 cell line as HER2-positive and HER2-negative cell lines, respectively. Anti-HER2 scFv expressed in BL21 (DE3) (Fig. 7a) and SHuffle® strains (Fig. 7b) binds selectively to SK-BR-3 cells as shown by the shift in fluorescence intensity value compared with background staining (untreated cells). However, no obvious shift in fluorescence value was observed in the flow cytometry of purified anti-HER2 scFv on MDA-MB-231 cells as compared to background staining (Fig. 7a, b). Moreover, anti-HER2 scFv expressed in SHuffle® demonstrated higher binding to SK-BR-3 cells as compared with anti-HER2 scFv expressed in BL21 (DE3) (Fig. 7).
Cell binding assay of anti-HER2 scFv by flowcytometry analysis. The binding of purified anti-HER2 scFv expressed in a BL21 (DE3) and b SHuffle® on the cell surface of (1) SK-BR-3 and (2) MDA-MB-231 cells is shown. Cell staining in the presence of PBS and absence of anti-HER2 scFv was demonstrated as untreated control
Binding Assay of Anti-HER2 scFv to HER2
To analyze the biological activity of purified anti-HER2 scFv, the binding activity of scFv proteins to HER2 was assessed by HER2- and cell-based ELISA. The HER2-specificity was confirmed by significant higher signal of anti-HER2 scFv bound to HER2 antigen comparing to negative control (wells without recombinant HER2 antigen) (Fig. 8a). Furthermore, the anti-HER2 scFv expressed in SHuffle® (100 ng/µL) bound to HER2 antigen at significant higher level than the one expressed in BL21 (DE3) cells (p < 0.001). In addition, anti-HER2 scFv expressed in SHuffle® demonstrated significant higher binding to HER2 when applied at 100 ng/µL concentration as compared to 10 ng/µL (p < 0.05). However, this significant difference between two different concentrations of anti-HER2 scFv expressed in BL21 (DE3) (100 ng/µL and 10 ng/µL) was not seen. Also, as presented in Fig. 8b, purified anti-HER2 scFv expressed in both BL21 and SHuffle® shows significant higher binding ability to SK-BR-3 compared to MDA-MB-231 cells. Therefore, it can be concluded that anti-HER2 scFv is able to bind specifically to HER2 antigen on SK-BR-3 cells. The anti-HER2 scFv expressed in SHuffle® could bind to SK-BR-3 cells at significant higher level than anti-HER2 scFv expressed in BL21 (DE3) (anti-HER2 scFv expressed in SHuffle® vs. BL21 (DE3) (10 and 100 ng/μL) p < 0.05). However, no significant difference between two concentrations of anti-HER2 scFv (expressed in both BL21 and SHuffle® strains) in binding ability to HER2-overexpressing cells was demonstrated.
Binding assay of the purified anti-HER2 scFv expressed from BL21 (DE3) and Shuffle® to HER2. The HER2-binding activity was evaluated by ELISA using both a purified HER2 antigen and b HER2-overexpressing cell line. Data are represented as mean ± SD of two independent experiments in duplicate. (*), (**), and (***) represent for p < 0.05, p < 0.01, and p < 0.001, respectively
In conclusion, scFv expressed from SHuffle® demonstrated more HER2-binding activity. The result of this study suggested that anti-HER2 scFv is more correctly folded in SHuffle® as compared to BL21 (DE3).
Competition of Trastuzumab and Anti-HER2 scFv for Binding to HER2 Antigen
Competition study showed that pre-incubation with the recombinant anti-HER2 scFv expressed from BL21 and SHuffle® causes significant reduction in HER2-binding ability of trastuzumab (10 ng/μL) (Fig. 9). At lower concentration of trastuzumab (1 ng/μL), pre-incubation with the anti-HER2 scFv expressed in SHuffle® caused significant decrease in HER2-binding ability of trastuzumab (p < 0.05). However, pre-incubation with anti-HER2 scFv expressed in BL21 did not cause significant reduction in HER2-binding ability of trastuzumab (1 ng/μL). Totally, this result suggested that anti-HER2 scFv recognizes the same epitope of HER2 receptor as the trastuzumab antibody.
Competitive binding assay of Trastuzumab and anti-HER2 scFv to HER2. The binding of trastuzumab (AryoTrustTM) to purified HER2 in the presence of anti-HER2 scFv was determined. Mean ± SD of two independent experiments in duplicate are shown. p < 0.05, p < 0.001, and p < 0.0001 are demonstrated with (*), (***), and (****), respectively
Discussion
Due to the presence of two disulfide bonds in scFv, its expression in reducing environment of E. coli cytoplasm is aggregation-prone and generally results in misfolded and biologically inactive inclusion bodies [31]. Refolding of inclusion bodies elongates the time of recombinant protein production. The efficiency of refolding may be low and refolded proteins might be unstable. Therefore, it is often desirable to express high degrees of soluble protein [32].
In the current study, an attempt was made to express anti-HER2 scFv in two different strains of E. coli and compare their ability for soluble and functional expression. E. coli BL21 (DE3) is deficient in two main proteases OmpT and Lon. It has become as the gold standard among expression hosts since it has been commercialized [33]. In SHuffle®, not only glutaredoxin reductase and thioredoxin reductase are deleted (Δgor ΔtrxB), but also the periplasmic disulfide bond isomerase DsbC without its signal sequence is expressed in cytoplasm. The correct disulfide bonds are formed by reduction and isomerization of the wrongly formed mis-oxidized disulfide bonds via DsbC in SHuffle® [34]. It has been claimed that the SHuffle® is able to express recombinant proteins with several disulfide bonds and also to refine the mis-oxidized bonds and progress proper folding [27].
In this study, the effects of duration and temperature of induction and also concentration of inducer (IPTG) on the total yield of anti-HER2 scFv were investigated to obtain optimal expression condition for each E. coli strains. The most anti-HER2 scFv was expressed in BL21 (DE3) 24 h after induction with 0.25 mM IPTG at 37 °C. The optimal condition for expression in SHuffle® strain was obtained after 24 h of induction with 0.05 mM IPTG at 30 °C (Fig. 5). Consistent with the result of our study, several studies reported 37 and 30 °C as the optimal induction temperature for maximum protein production in BL21 (DE3) [35, 36] SHuffle® [29, 37], respectively. Our results are also consistent with the previous studies in which the quantity of newly expressed protein was decreased by lowering the induction temperature in BL21 (DE3) [11, 38]. IPTG has the ability to inhibit cell growth and also has impact on the level of protein expression [38]. The expression rate can be lowered by decreasing IPTG concentration and consequently intracellular folding efficiency can be enhanced [39]. Therefore, the optimization of IPTG concentration is highly required. In this study, IPTG concentration demonstrated significant effect on total anti-HER2 scFv expression in BL21 (DE3) strain (Fig. 5c). Our results in BL21 (DE3) strain are consistent with the study of Heo et al. in which reducing the IPTG concentration from 1 to 0.05 mM caused approximately 1.6-fold increase in the productivity of functional anti-c Met scFv expressed in Origami (DE3) [39]. The anti-HER2 scFv expression in SHuffle® did not significantly affect IPTG concentration. Napathorn and his colleagues also demonstrated that the total protein expression in Rosetta-gami-B did not alter by varying IPTG concentrations. However, they showed that lower transcription rate using lesser concentration of IPTG than standard (1 mM) results in proper folding and more soluble fragments [38]. Furthermore, in agreement with our study, Peciak et al. demonstrated that varying IPTG concentration had little effect on SUMO-Interferon consensus fusion protein expression in SHuffle®E. coli: Totally, it can be concluded that the effect of IPTG concentration on protein expression varies by different E. coli host [40]. To optimize the culture conditions for protein expression, it is essential to evaluate the effect of involved variables together and simultaneously, which can be obtained using statistical design of experiments [41, 42]. The results of the current study can be further used to apply experimental design methods for achieving the optimized expression conditions for anti-HER2 scFv protein with a minimum number of experiments [41, 42].
The solubility of a protein is an important indicator of its correct folding as determined by functional binding [43]. It was found that the decreases of induction temperature caused improvement of protein solubility [44]. In this study, the solubility of anti-HER2 scFv expressed in BL21 (DE3) and induced at 25 °C was significantly higher than that induced at 37 °C (Fig. 6). In agreement with our data, the most soluble GST-PTEN protein was expressed in BL21 at the low induction temperature of 20 °C in the study of Hu et al. [45]. Temperature reduction may eliminate heat shock proteases that are induced under overexpression conditions. Moreover, the activity and expression of many E. coli chaperones are increased at lower temperature and thus the corrected folding of the target protein is facilitated [46]. Totally, despite lowering the induction temperature, soluble fraction was approximately half (0.43–0.52) of insoluble fraction in recombinant BL21 (DE3) strain (Fig. 6).
Herein, due to lack of difference between the amount of anti-HER2 scFv expressed by pET-22 (with pelB) and pET-28 (without pelB) (Fig. 4), the experiment for comparison of SHuffle® and BL21 (DE3) strains was conducted using E. coli strains containing pET-22 (anti-HER2 scFv). Ritthisan et al. also used pET-22 plasmid for expression of an antibody-enzyme fusion gene in E. coli SHuffle® [47]. Our results demonstrated that the solubility of anti-HER2 scFv expressed in SHuffle® at 30 °C was higher than that expressed in BL21 (DE3) at 37 and 30 °C (Fig. 6). In this study, as it is anticipated oxidative environment in SHuffle® provided appropriate folding in recombinant protein [22]. Our results are consistent with the recent study in which fully soluble and active recombinant EhCP1 enzyme was expressed using E. coli SHuffle® cells. Furthermore, an antibody-enzyme fusion protein containing antigen-binding fragment (Fab) fused to E. coli alkaline phosphatase (AP) was successfully expressed in soluble form having both antigen-binding and AP activity in E. coli SHuffle® [47]. Safarpour et al. reported that TNF-α expression in E. coli SHuffle® results in about 1.5 times higher disulfide band formation compared to expression in BL21 (DE3) [29]. Lauber et al. demonstrated that the human glycosyltransferase expressed in SHuffle® host cells produces the same CD spectra as the commercially available recombinant glycosyltransferase from NS0 murine myeloma indicating similar folding state for both glycosyltransferases [48].
To our knowledge, we describe the first study comparing the biological activity of disulfide bond containing protein expressed in BL21 (DE3) and SHuffle®. Herein, the solubility of anti-HER2 scFv protein expressed in SHuffle® at 30 °C was significantly higher than anti-HER2 scFv protein expressed in BL21 (DE3) strain at both 30 and 37 °C. Furthermore, the solubility of anti-HER2 scFv protein expressed in BL21 (DE3) strain at 30 °C was not significantly higher than the one expressed at 37 °C (Fig. 6). Therefore, we compared anti-HER2 scFv biological activity in the optimal induction temperature related to each host (37 °C for BL21 (DE3) and 30 °C for SHuffle®). Subsequently, our results can be extrapolated to many studies that induced protein expression at 37 °C in BL21 (DE3) and 30 °C in SHuffle®.
Binding to a cell-embedded tumor-associated antigen is the first key step in the mechanism of antitumor immune agents [49]. Thus, further to study the binding properties of anti-HER2 scFv using HER2 antigen, we applied cell-based ELISA and also flow cytometry analysis to evaluate HER2 binding ability of anti-HER2 scFv. In some studies, flow cytometry methods appeared to be more sensitive and specific than ELISA methods, due to losing the least number of specificities [50, 51]. Flow cytometry data (Fig. 7) and the results of cell-based ELISA (Fig. 8B) showed that the purified anti-HER2 scFv from BL21 (DE3) and SHuffle® bind specifically to HER2-positive SK-BR-3 cells and not to HER2-negative MDA-MB-231. A similar observation was previously reported for anti-HER2 scFv (from pertuzumab) produced in BL21 (DE3) with a high binding affinity to HER2-positive BT-474 breast cancer cells but no binding affinity to MDA-MB-231 cells [11]. In addition, the results of HER2-based ELISA indicated that the expressed anti-HER2 scFv in both strains can specifically recognize HER2 antigen (Fig. 8a). In the competitive ELISA, inhibition of HER2-binding of trastuzumab by the expressed anti-HER2 scFv showed that anti-HER2 scFv recognizes the same epitope of HER2 receptor as the trastuzumab antibody [49]. Competition assay was also used in the study of Agha miri et al. to demonstrate recognizing the same epitope of CD22 receptor by the expressed fusion protein and RFB4 antibody (anti-CD22 Mab). They demonstrated that an anti-CD22 scFv-apoptin fusion protein expressed in BL21 (DE3) decreased MFI value (measured by flow cytometry) of RFB4 to 60% [52].
In this study, the scFv expressed in SHuffle® demonstrated higher HER2-binding ability than the one expressed in BL21 (DE3) using flow cytometry analysis as well as both recombinant HER2- and cell-based ELISA. The significant difference between two different doses of anti-HER2 scFv (10 and 100 ng/µL) was only observed in the one expressed in SHuffle® due to the presence of DsbC in oxidative environment of SHuffle®. Anti-HER2 scFv expressed in BL21 (DE3) at higher concentration (100 ng/µL) may form more miss-folded anti-HER2 scFv protein and subsequently did not cause significant higher HER2 binding as compared to its lower concentration (10 ng/µL). The significant difference between two different doses of anti-HER2 scFv was only demonstrated in HER2-based ELISA and not cell-based ELISA which may be due to the difference in purity of the capture antigen [53]. Furthermore, significant decrease in HER2-binding ability of low concentration of trastuzumab (1 ng/µL) was only observed with pre-incubation of anti-HER2 scFv expressed in SHuffle® strain and the one expressed in BL21 (DE3) strain did not reduce the HER2-binding ability of low concentration of trastuzumab, significantly. To have more quantitative comparison between anti-HER2 scFv expressed in BL21 (DE3) and SHuffle®, the constants that characterize the strength and kinetics of HER2 and anti-HER2 scFv reaction need to be further studied. The affinity values can be obtained by ELISA and also an optical method surface plasmon resonance (SPR). According to the results of Heinrich et al., it is highly required to use both ELISA-based and SPR methods to obtain the affinity values of a biological reaction between HER2 and anti-HER2 scFv [54].
Totally, it can be concluded that expression in SHuffle® at 30 °C resulted in enhanced solubility and higher level of HER2-binding ability as compared to its expression in BL21 (DE3) at 37 °C that may be due to correct disulfide bond formation and proper folding [25]. It is highly suggested to provide further support for proper folding of the recombinant protein produced in SHuffle® host cells by measuring CD-spectroscopy in future study.
References
Ferlay, J., Soerjomataram, I., Dikshit, R., Eser, S., Mathers, C., Rebelo, M., et al. (2015). Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. International Journal of Cancer,136, E359–E386.
Taslimi, Y., Zahedifard, F., Habibzadeh, S., Taheri, T., Abbaspour, H., Sadeghipour, A., et al. (2016). Antitumor effect of IP-10 by using two different approaches: Live delivery system and gene therapy. Journal of Breast Cancer,19, 34–44.
Alirezapour, B., Jalilian, A. R., Bolourinovin, F., & Moradkhani, S. (2013). Production and quality control of [67 Ga]-DOTA-trastuzumab for Radioimmunoscintigraphy. Iranian Journal of Pharmaceutical Research,12, 355–366.
Hajighasemlou, S., Alebouyeh, M., Rastegar, H., Manzari, M. T., Mirmoghtadaei, M., Moayedi, B., et al. (2015). Preparation of immunotoxin herceptin-botulinum and killing effects on two breast cancer cell lines. Asian Pacific Journal of Cancer Prevention,16, 5977–5981.
Moghimi, S. M., Rahbarizadeh, F., Ahmadvand, D., & Parhamifar, L. (2013). Heavy chain only antibodies: A new paradigm in personalized HER2+ breast cancer therapy. BioImpacts,3, 1–4.
Spigel, D. R., & Burstein, H. J. (2002). HER2 overexpressing metastatic breast cancer. Current Treatment Options in Oncology,3, 163–174.
Borg, Å., Tandon, A. K., Sigurdsson, H., Clark, G. M., Fernö, M., Fuqua, S. A., et al. (1990). HER-2/neu amplification predicts poor survival in node-positive breast cancer. Cancer Research,50, 4332–4337.
Dressman, M. A., Baras, A., Malinowski, R., Alvis, L. B., Kwon, I., Walz, T. M., et al. (2003). Gene expression profiling detects gene amplification and differentiates tumor types in breast cancer. Cancer Research,63, 2194–2199.
Gajria, D., & Chandarlapaty, S. (2011). HER2-amplified breast cancer: Mechanisms of trastuzumab resistance and novel targeted therapies. Expert Review of Anticancer Therapy,11, 263–275.
Jurado, P., Ritz, D., Beckwith, J., de Lorenzo, V., & Fernandez, L. A. (2002). Production of functional single-chain Fv antibodies in the cytoplasm of Escherichia coli. Journal of Molecular Biology,320, 1–10.
Akbari, V., Sadeghi, H. M. M., Jafrian-Dehkordi, A., Abedi, D., & Chou, C. P. (2014). Functional expression of a single-chain antibody fragment against human epidermal growth factor receptor 2 (HER2) in Escherichia coli. Journal of Industrial Microbiology and Biotechnology,41, 947–956.
Cao, Y., Marks, J. D., Huang, Q., Rudnick, S. I., Xiong, C., Hittelman, W. N., et al. (2012). Single-chain antibody-based immunotoxins targeting Her2/neu: Design optimization and impact of affinity on antitumor efficacy and off-target toxicity. Molecular Cancer Therapeutics,11, 143–153.
Cao, Y., Marks, J. D., Marks, J. W., Cheung, L. H., Kim, S., & Rosenblum, M. G. (2009). Construction and characterization of novel, recombinant immunotoxins targeting the Her2/neu oncogene product: in vitro and in vivo studies. Cancer Research,69, 8987–8995.
Nikkhoi, S. K., Rahbarizadeh, F., Ranjbar, S., Khaleghi, S., & Farasat, A. (2018). Liposomal nanoparticle armed with bivalent bispecific single-domain antibodies, novel weapon in HER2 positive cancerous cell lines targeting. Molecular Immunology,96, 98–109.
Park, J., Kirpotin, D., Hong, K., Shalaby, R., Shao, Y., Nielsen, U., et al. (2001). Tumor targeting using anti-her2 immunoliposomes. Journal of Controlled Release,74, 95–113.
Park, J. W., Hong, K., Kirpotin, D. B., Colbern, G., Shalaby, R., Baselga, J., et al. (2002). Anti-HER2 immunoliposomes: Enhanced efficacy attributable to targeted delivery. Clinical Cancer Research,8, 1172–1181.
Rosano, G. L., & Ceccarelli, E. A. (2014). Recombinant protein expression in Escherichia coli: Advances and challenges. Frontiers in Microbiology,5, 172.
Guglielmi, L., & Martineau, P. (2009). Expression of single-chain Fv fragments in E. coli cytoplasm. Antibody Phage Display Springer,2, 215–224.
Ritz, D., & Beckwith, J. (2001). Roles of thiol-redox pathways in bacteria. Annual Reviews in Microbiology,55, 21–48.
Stewart, E. J., Åslundm, F., & Beckwith, J. (1998). Disulfide bond formation in the Escherichia coli cytoplasm: An in vivo role reversal for the thioredoxins. The EMBO Journal,17, 5543–5550.
Villaverde, A., & Carrió, M. M. (2003). Protein aggregation in recombinant bacteria: Biological role of inclusion bodies. Biotechnology Letters,25, 1385–1395.
Jalomo-Khayrova, E., Mares, R. E., Muñoz, P. L., Meléndez-López, S. G., Rivero, I. A., & Ramos, M. A. (2018). Soluble expression of an amebic cysteine protease in the cytoplasm of Escherichia coli SHuffle Express cells and purification of active enzyme. BMC Biotechnology,18, 20.
Fathi-Roudsari, M., Akhavian-Tehrani, A., & Maghsoudi, N. (2016). Comparison of three Escherichia coli strains in recombinant production of reteplase. Avicenna Journal of Medical Biotechnology,8, 16–22.
Rabhi-Essafi, I., Sadok, A., Khalaf, N., & Fathallah, D. M. (2007). A strategy for high-level expression of soluble and functional human interferon α as a GST-fusion protein in E. coli. Protein Engineering, Design & Selection,20, 201–209.
Derman, A. I., Prinz, W. A., Belin, D., & Beckwith, J. (1993). Mutations that allow disulfide bond formation in the cytoplasm of Escherichia coli. Science,262, 1744–1747.
De Marco, A. (2009). Strategies for successful recombinant expression of disulfide bond-dependent proteins in Escherichia coli. Microbial Cell Factories,8, 26.
Lobstein, J., Emrich, C. A., Jeans, C., Faulkner, M., Riggs, P., & Berkmen, M. (2012). SHuffle, a novel Escherichia coli protein expression strain capable of correctly folding disulfide bonded proteins in its cytoplasm. Microbial Cell Factories,11, 56.
McCarthy, A. A., Haebel, P. W., Törrönen, A., Rybin, V., Baker, E. N., & Metcalf, P. (2000). Crystal structure of the protein disulfide bond isomerase, DsbC, from Escherichia coli. Nature Structural & Molecular Biology,7, 196–199.
Safarpour, H., Banadkoki, S. B., Keshavarzi, Z., Morowvat, M. H., Soleimanpour, M., Pourmolaei, S., et al. (2017). Expression analysis and ATR-FTIR characterization of the secondary structure of recombinant human TNF-α from Escherichia coli SHuffle® T7 Express and BL21 (DE3) cells. International Journal of Biological Macromolecules,99, 173–178.
Balandin, T. G., Edelweiss, E., Andronova, N. V., Treshalina, E. M., Sapozhnikov, A. M., & Deyev, S. M. (2011). Antitumor activity and toxicity of anti-HER2 immunoRNase scFv 4D5-dibarnase in mice bearing human breast cancer xenografts. Investigational New Drugs,29, 22–32.
Sørensen, H. P., & Mortensen, K. K. (2005). Soluble expression of recombinant proteins in the cytoplasm of Escherichia coli. Microbial Cell Factories,4, 1.
Misawa, S., & Kumagai, I. (1999). Refolding of therapeutic proteins produced in Escherichia coli as inclusion bodies. Peptide Science,51, 297–307.
Terpe, K. (2006). Overview of bacterial expression systems for heterologous protein production: From molecular and biochemical fundamentals to commercial systems. Applied Microbiology and Biotechnology,72, 211–222.
Ren, G., Ke, N., & Berkmen, M. (2016). Use of the SHuffle strains in production of proteins. Current Protocols in Protein Science,85(1), 5–26.
Jaliani, H. Z., Farajnia, S., Safdari, Y., Mohammadi, S. A., Barzegar, A., & Talebi, S. (2014). Optimized condition for enhanced soluble-expression of recombinant mutant anabaena variabilis phenylalanine ammonia lyase. Advanced Pharmaceutical Bulletin,4, 261–266.
Naderi, S., Alikhani, M. Y., Karimi, J., Shabab, N., Mohamadi, N., Jaliani, H. Z., et al. (2015). Cytoplasmic expression, optimization and catalytic activity evaluation of recombinant mature lysostaphin as an anti-staphylococcal therapeutic in Escherichia coli. Acta Medica International,2, 72–77.
Ke, N., & Berkmen, M. (2014). Production of disulfide-bonded proteins in Escherichia coli. Current Protocols in Molecular Biology. https://doi.org/10.1002/0471142727.mb1601bs108.
Napathorn, S. C., Kuroki, M., & Kuroki, M. (2014). High expression of fusion proteins consisting of a single-chain variable fragment antibody against a tumor-associated antigen and interleukin-2 in Escherichia coli. Anticancer Research,34, 3937–3946.
Heo, M. A., Kim, S. H., Kim, S. Y., Kim, Y. J., Chung, J., Oh, M. Km., et al. (2006). Functional expression of single-chain variable fragment antibody against c-Met in the cytoplasm of Escherichia coli. Protein Expression and Purification,47, 203–209.
Peciak, K., Tommasi, R., Choi, J-w, Brocchini, S., & Laurine, E. (2014). Expression of soluble and active interferon consensus in SUMO fusion expression system in E. coli. Protein Expression and Purification,99, 18–26.
Akbari, V., Mir MohammadSadeghi, H., Jafarian-Dehkordi, A., Perry Chou, C., & Abedi, D. (2015). Optimization of a single-chain antibody fragment overexpression in Escherichia coli using response surface methodology. Research in Pharmaceutical Sciences,10, 75–83.
Agha Amiri, S., Zarei, N., Enayati, S., Azizi, M., Khalaj, V., & Shahhosseini, S. (2018). Expression optimization of anti-CD22 scFv-apoptin fusion protein using experimental design methodology. Iranian Biomedical Journal,22, 66–69.
Drees, J. J., Augustin, L. B., Mertensotto, M. J., Schottel, J. L., Leonard, A. S., & Saltzman, D. A. (2014). Soluble production of a biologically active single-chain antibody against murine PD-L1 in Escherichia coli. Protein Expression and Purification,94, 60–66.
Schein, C. H., & Noteborn, M. H. (1988). Formation of soluble recombinant proteins in Escherichia coli is favored by lower growth temperature. Bio/technology,6, 291–294.
Hu, Y., An, Y., Fang, N., Li, Y., Jin, H., Nazarali, A., et al. (2015). The optimization of soluble PTEN expression in Escherichia coli. The Open Biochemistry Journal,9, 42–48.
Zhu, Y. Q., Tong, W. Y., Wei, D. Z., Zhou, F., & Zhao, J. B. (2007). Environmental stimuli on the soluble expression of anti-human ovarian carcinoma × anti-human CD3 single-chain bispecific antibody in recombinant Escherichia coli. Biochemical Engineering Journal,37, 184–19144.
Ritthisan, P., Ojima-Kato, T., Damnjanović, J., Kojima, T., & Nakano, H. (2018). SKIK-zipbody-alkaline phosphatase, a novel antibody fusion protein expressed in Escherichia coli cytoplasm. Journal of Bioscience and Bioengineering,126, 705–709.
Lauber, J., Handrick, R., Leptihn, S., Dürre, P., & Gaisser, S. (2015). Expression of the functional recombinant human glycosyltransferase GalNAcT2 in Escherichia coli. Microbial Cell Factories,14, 3.
Troise, F., Cafaro, V., Giancola, C., D’Alessio, G., & De Lorenzo, C. (2008). Differential binding of human immunoagents and Herceptin to the ErbB2 receptor. The FEBS Journal,275, 4967–4979.
Dasso, J., Lee, J., Bach, H., & Mage, R. G. (2002). A comparison of ELISA and flow microsphere-based assays for quantification of immunoglobulins. Journal of Immunological Methods,263, 23–33.
Worthington, J., Robson, A., Sheldon, S., Langton, A., & Martin, S. (2001). A comparison of enzyme-linked immunoabsorbent assays and flow cytometry techniques for the detection of HLA specific antibodies. Human Immunology,62, 1178–1184.
Amiri, S. A., Shahhosseini, S., Zarei, N., Khorasanizadeh, D., Aminollahi, E., Rezaie, F., et al. (2017). A novel anti-CD22 scFv–apoptin fusion protein induces apoptosis in malignant B-cells. AMB Express,7, 112.
Jamieson, D., Cresti, N., Verrill, M. W., & Boddy, A. V. (2009). Development and validation of cell-based ELISA for the quantification of trastuzumab in human plasma. Journal of Immunological Methods,345, 106–111.
Heinrich, L., Tissot, N., Hartmann, D. J., & Cohen, R. (2010). Comparison of the results obtained by ELISA and surface plasmon resonance for the determination of antibody affinity. Journal of Immunological Methods,352, 13–22.
Acknowledgements
This work was supported by the grant from the research deputy of Shahid Beheshti University of Medical Sciences (SBMU).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.

12033_2019_221_MOESM1_ESM.jpg
Supplementary material 1 Anti-HER2 scFv expression in BL21 (DE3). Analysis of anti-HER2 scFv expressed in BL21 (DE3). Protein marker (Fermentas) (MW), total protein from BL21 (DE3) containing pET-22 (anti-HER2 scFv) plasmid before induction (lane 1), 2 (lane 2), 4 (lane 3), 6 (lane 4) and 24 (lane 5) h after induction with 1 mM IPTG at 37 °C. Total protein from BL21 (DE3) containing pET-22 (without insert) before induction (lane 6), 2 (lane 7), 4 (lane 8), 6 (lane 9) and 24 (lane 10) h after induction with 1 mM IPTG at 37 °C. (JPEG 851 kb)

12033_2019_221_MOESM2_ESM.jpg
Supplementary material 2 Anti-HER2 scFv expression in SHuffle®. Analysis of anti-HER2 scFv expressed in SHuffle®. Protein marker (Fermentas) (MW), total protein from SHuffle® containing pET-22 (anti-HER2 scFv) plasmid before induction (lane 1), 2 (lane 2), 4 (lane 3), 6 (lane 4) and 24 h (lane 5) after induction with 1 mM IPTG at 30 °C. Total protein from SHuffle® containing pET-22 (without insert) before induction (lane 6), 2 (lane 7), 4 (lane 8) and 6 (lane 9) h after induction with 1 mM IPTG at 30 °C. (JPEG 575 kb)

12033_2019_221_MOESM3_ESM.jpg
Supplementary material 3 Effect of temperature on anti-HER2 scFv expression in BL21 (DE3). BL21 (DE3) containing pET-22 (anti-HER2 scFv) was induced by IPTG (1 mM), incubated at different induction temperatures. Protein marker (Fermentas) (MW), total protein from recombinant BL21 (DE3) before induction (lane 1), after induction with 1 mM IPTG for 6 (lane 2) and 24 h (lane 3) at 37 °C; 6 (lane 4) and 24 h (lane 5) at 30 °C, 6 (lane 6) and 24 h (lane 7) at 25 °C. (JPEG 332 kb)

12033_2019_221_MOESM4_ESM.jpg
Supplementary material 4 Induction of anti-HER2 scFv expression in SHuffle® at different temperatures. SHuffle® containing pET-22 (anti-HER2 scFv) was induced by IPTG (1 mM), incubated at different induction temperatures. Protein marker (Fermentas) (MW), total protein from recombinant SHuffle® after induction with 1 mM IPTG for 24 h at 37 (lane 1), 25 (lane 2), 30 (lane 3) and 15 °C (lane 4). Total protein from recombinant BL21 (DE3) before induction (lane 5). (JPEG 1792 kb)

12033_2019_221_MOESM5_ESM.jpg
Supplementary material 5 Induction of recombinant anti-HER2 scFv expression in BL21 (DE3) by different concentrations of IPTG. BL21 (DE3) containing pET-22 (anti-HER2 scFv) was induced by different concentrations of IPTG at 37 °C for 24 h. Protein marker (Fermentas) (MW), total protein from BL21 (DE3) containing pET-22 (anti HER2-scFv) before induction (lane 1), 24 h after induction with 0.25 (lane 2), 0.5 (lane 3), 1 (lane 4), and 2 (lane 5) mM IPTG at 37 °C. (JPEG 556 kb)

12033_2019_221_MOESM6_ESM.jpg
Supplementary material 6 Induction of recombinant anti-HER2 scFv expression in SHuffle® by different concentrations of IPTG. SHuffle® containing pET-22 (anti-HER2 scFv) was induced by different concentrations of IPTG at 30 °C for 24 h. Protein marker (Fermentas) (MW), total protein from BL21 (DE3) containing pET-22 (anti HER2-scFv) before induction (lane 1), 24 h after induction with 0.01 (lane 2), 0.05 (lane 3), 0.1 (lane 4), 0.5 (lane 5) and 1 (lane 6) mM IPTG at 30 °C. (JPEG 1500 kb)

12033_2019_221_MOESM7_ESM.jpg
Supplementary material 7 The effect of induction temperature on the solubility of anti-HER2 scFv expressed in BL21 (DE3). SDS-PAGE analysis of solubility of anti-HER2 scFv expressed in BL21 (DE3). Protein marker (Fermentas, MW, lane 1), total protein from E. coli BL21 (DE3) containing pET-22 (anti-HER2 scFv) plasmid before induction (lane 2) and after induction with 1 mM IPTG for 24 h at 37 °C (lane 3), the soluble (lane 4) and insoluble (lane 5) fractions of BL21 (DE3) lysates after induction with 0. 25 mM IPTG at 25 °C, the soluble (lane 6) and insoluble (lane 7) fractions of BL21 (DE3) lysates after induction with 0. 25 mM IPTG at 30 °C, the soluble (lane 8) and insoluble (lane 9) fractions of BL21 (DE3) lysates after induction with 0. 25 mM IPTG at 37 °C. (JPEG 237 kb)

12033_2019_221_MOESM8_ESM.jpg
Supplementary material 8 The effect of induction temperature on the solubility of anti-HER2 scFv expressed in SHuffle®. SDS-PAGE analysis of solubility of anti-HER2 scFv expressed in SHuffle®. Protein marker (Fermentas, MW), the soluble (lane 1) and insoluble (lane 2) fractions of SHuffle® lysates after induction with 0.05 mM IPTG at 25 °C, the soluble (lane 3) and insoluble (lane 4) fractions of SHuffle® lysates after induction with 0.05 mM IPTG at 30 °C, the soluble (lane 5) and insoluble (lane 6) fractions of SHuffle® lysates after induction with 0.05 mM IPTG at 37 °C. (JPEG 403 kb)
Rights and permissions
About this article
Cite this article
Ahmadzadeh, M., Farshdari, F., Nematollahi, L. et al. Anti-HER2 scFv Expression in Escherichia coli SHuffle®T7 Express Cells: Effects on Solubility and Biological Activity. Mol Biotechnol 62, 18–30 (2020). https://doi.org/10.1007/s12033-019-00221-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-019-00221-2