Abstract
Nucleic acid immunization has recently exhibited a great promise for immunotherapy of various diseases. However, it is now clear that powerful strategies are imminently needed to improve their efficiency. In this regard, whole bacteriophage particles have been described as efficient DNA vaccine delivery vehicles, capable of circumventing the limitations of naked DNA immunization. Moreover, phage particles could be engineered to display specific peptides on their surfaces. Given these inherent characteristics of phages, we have designed a novel hybrid phage-DNA immunization vector using both M13 and pAAV plasmid elements. Following the construction and in vitro confirmation of the designed vectors, they were used for comparative mice immunization, carrying the same DNA sequence. The results indicated the efficacy of the designed hybrid phage particles, to elicit higher humoral immunity, in comparison to conventional DNA-immunization vectors (pCI). In light of these findings, it could be concluded that using adeno-associated virus (AAV) expression cassette along with displaying TAT peptide on the surface of the phage particle could be deemed as an appealing strategy to enhance the DNA-immunization and vaccination efficacy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nucleic acid immunization has recently exhibited a great promise in immunotherapy for various diseases. This immunization method is basically the expression of protective antigens within the host cells by inoculation of a target animal with specific gene of interest under the control of an exogenous eukaryotic promoter [1, 2]. Comparing with conventional immunization methods like attenuated, killed, or subunit vaccines, nucleic acid immunization potentially represents several superiorities, such as good safety, specificity to induce immune response for the antigen of interest, induction of both B- and T-cell responses, relatively low cost of production, and ease of manufacturing. However, inherent barriers of the body, poor stability, and inappropriate in vivo distribution are among the possible reasons to hinder the efficiency of nucleic acid immunizations [3]. Given these limitations, it is now clear that powerful strategies are imminently needed to improve the responses in DNA-immunization experiments. However, many of improvement methods are associated with increased costs or cumbersome nature in addition to regulatory hurdles. It is obvious that a sustainable strategy is required to enhance the efficiency of DNA-immunization components particularly via a more robust delivery system and antigen design approach like in silico studies [4,5,6,7,8,9,10,11].
Based on prior investigations, whole bacteriophage particles as immunogene carriers have been described as DNA vaccine delivery vehicles which are capable of circumventing the limitations of the naked DNA-immunization method [12, 13]. Due to their low cost of production, stability, and high valency of foreign antigen display on their surfaces, phages have garnered a lot of attention as vaccine carriers [14, 15]. Filamentous bacteriophages are among the non-pathogenic, non-lytic viruses capable of infecting and replicating in Escherichia coli cells carrying an F′ episome. These phages could be easily manipulated for peptide display on their surface proteins and insertion of DNA-immunization cassettes [16]. However, Hajitou et al. have previously introduced AAVP vectors as phage-based hybrids between adeno-associated virus (AAV) and M13 phage [17]. These hybrids of two single-stranded DNA viruses could overcome the limitation of phage particles. Compared to conventional phages, the AAVP vectors are associated with more promising gene delivery potentials and long-term in vivo gene expression following their systemic delivery [17]. Given these circumstance, AAVP-based design of vectors could improve the phage-based immunization results. Dickkopf-1 (DKK1) is a secretory glycoprotein, acting as an antagonist of the canonical Wnt signaling pathway. Expression rate discrepancies reported in various tumor cells make DKK1 an ideal target for cancer detection and prevention (reviewed in [18]). Given the potential of this antigen to be a cancer therapy or detection target, we have selected DKK1 as the target antigen to build our immunogene. The antibodies against this antigen could be employed for both cancer detection and prevention purposes.
In the present study, we have constructed a novel phage-DNA immunization vehicle to evaluate its effectivity eliciting humoral immune responses against the target antigens. The constructed vectors were assessed both in vivo and in vitro to evaluate their functionality and immunogenicity. Our results indicated that the employed approach is capable of successful immunization against the challenging target antigen. Moreover, the obtained results revealed that the engineered phage-DNA immunization construct triggers a better humoral immunity in comparison to conventional DNA vaccination constructs. To the best of our knowledge, it is the first study to design an engineered phage-based DNA immunization vehicle which is a combination of M13 bacteriophage with surface-located TAT peptide and AAV eukaryotic expression cassette.
Materials and Methods
Ethics Statement
To alleviate the suffering, mice were housed in spacious plexiglas breeding cages (two mice in each cage) with woodchip bedding and free access to food and water. They were monitored on daily basis (the bedding was changed regularly) and all of the animals were kept at 20–25 °C on a 12-h light/dark cycle (with lights on at 7 a.m.). All painful procedures (like blood sample collection) were performed under general anesthesia. All procedures were performed according to the ethical guidelines of Faculty of Medical Sciences, Tarbiat Modares University, based on the United States NIH Guide for the Care and Use of Laboratory Animals (publication no. 85-23).
In Silico Antigen Design
The tools for linear and conformational B cell epitope prediction, available at IEDB server (http://www.iedb.org/), were used to determine the immunologically important regions of the DKK1 protein. The 3D structure of Dickkopf 1 (DKK1) tumor antigen (required to predict the conformational epitopes) along with the data about the regions with functional significance was adapted from the study we previously performed in our lab [5]. The most immunologically potent regions were linked by a linker sequence and fused to an immunization agent adapted from a study performed by Chambers et al. [19]. The corresponding gene sequence was optimized for eukaryotic protein expression using JCat server at http://www.jcat.de/.
Construction of pAK8–GVO–AAV–DKK1–TAT Phagemid Vector
In order to obtain phage-DNA vaccination constructs, three consecutive sub-clonings were performed. The gene sequence corresponding to the designed DKK1 gene was chemically synthesized by Generay Biotech Co, Ltd. The synthesized gene was sub-cloned into the pAAV–MCS vector (Agilent Technologies Company) between the XbaI and BglII restriction sites to construct the pAAV–MCS–DKK1 vector. The whole eukaryotic expression cassette of the pAAV–MCS–DKK1 plasmid was amplified with PCR method using specific forward (AGATAGACGCTCACATGTCCTGG) and reverse (TAATAAACATGTAATACCGCATCAGG) primers. The amplified sequence encompasses the region between the left and right ITRs of the plasmid. The PCR product was sub-cloned into the pAK8–GVO plasmid (a king’s gift from Dr. Klimka, University of Cologne, Cologne, Germany) using the PciI restriction site to construct the pAK8–GVO–AAV–DKK1 phagemid. Thereafter, a gene sequence was designed for trans-activating transcriptional activator (TAT) to be displayed at N terminus of the phage PVIII protein. The sequence of the gene was optimized for expression in E. coli gene. Ultimately, the gene sequence for TAT was chemically synthesized by Generay Biotech Co, Ltd. and sub-cloned into the pAK8–GVO–AAV–DKK1 phagemid between the SfiI and NotI restriction sites to construct the pAK8–GVO–AAV–DKK1–TAT phagemid. All of the restriction enzymes, T4 ligase and high-fidelity DNA polymerase were purchased from Thermo Fisher Scientific Company, while the DNA extraction kit was purchased from GeneAll Company. The enzymatic reactions and plasmid extractions were done according to the manufacturer’s instructions. All of the plasmids were transformed into the E. coli TG1 competent cells. The transformations were performed by the conventional CaCl2-heat shock method. The accuracy of the sub-cloning was confirmed by the sequencing of the final plasmids. The specific DKK1 primers (forward: CGGGGATCCTCTAGACTCGC and reverse: GATCACTAAAGATCTACGGGTGGC) were used to confirm the sequences of pAAV–MCS–DKK1 and pAK8–GVO–AAV–DKK1 constructs, while specific TAT primers (forward: CGGCCCACCACCACCACCACCACG and reverse: GCGGAGCCGCCGCCGCCCTG) were used to confirm the pAK8–GVO–AAV–DKK1–TAT sub-cloning.
Construction of pCI–DKK1 Vector
The same immunogene which was used to construct the pAK8–GVO–AAV–DKK1–TAT vector, was sub-cloned into the pCI vector (Promega Corporation product E1721) between the XhoI and SalI restriction sites. The same procedures of “Materials and Methods” were employed to perform and confirm the sub-cloning. The DNA sequencing of the final plasmid was done by specific DKK1 primers (forward: CGGGGATCCTCTAGACTCGC and reverse: GATCACTAAAGATCTACGGGTGGC).
Phage Rescue, Amplification, and Purification
The routinely used helper phage rescue and PEG precipitation method was adapted from the study performed by Wu et al. [20] to prepare and purify the phage particles. Following phage amplification and purification, phages were tittered using colony forming unit (CFU) assay as previously described [21].
Evaluation of Displayed TAT Peptide with ELISA
An ELISA test was designed to assess the correct peptide display on the surface of phage particles. In this regard, microtiter (Maxisorb) plates were coated with 1012 phage particles in 100 µL of PBS buffer followed by an overnight incubation at 4 °C. After three times of washing with PBS, the wells were blocked with PBS-5% skimmed milk and incubated for an hour at 37 °C. Then wells were washed five times with PBS and PBST (PBS containing 0.05% Tween-20) and 100 µL of (1:10,000) HRP-conjugated anti-His tag antibody (Pharmacia) was added to each well and incubated at 37 °C for another hour. Then 100 µL of TMB substrate was added and after 15 min of dark incubation (color development). 50 µL of 2 N HCl was used to stop the reaction. Finally, the OD was determined at 450 nm. A phage particle displaying an unrelated peptide was used as control.
In Vitro Assessment of Constructed Plasmids
The sequence for a green florescence protein (GFP) was sub-cloned downstream of the DKK1 sequence to assess the ability of the construct for eukaryotic expression and overall correctness of the constructed plasmid. Three consecutive sub-cloning has been performed to construct the GFP-containing constructs. The DKK1 gene was first sub-cloned into the pEGFP–N1 plasmid (CLONTECH Laboratories, Inc.) between the SalI and XhoI restriction sites (upstream of the GFP gene) to build the pEGFP–N1–DKK1 plasmid. As the second sub-cloning, the sequences of the DKK1 and GFP genes were amplified together using specific forward (CCTGCTCGAGGCCACCATGGATG) and reverse (CCGCAGATCTTTACTTGTACAGCTCGTCC) primers. Then the amplified fragment was sub-cloned into pAK8–GVO–AAV–DKK1–TAT (by replacing the DKK1 gene). The sub-cloning was performed by XbaI and BglII restriction sites to construct pAK8–GVO–AAV–DKK1–GFP–TAT phagemid. On the other hands, the construction of pCI–DKK1–GFP vector was performed by the same strategy using XhoI and NotI restriction enzymes. The accuracy of the sub-cloning was confirmed by sequencing the final plasmids using the previously designed primers. It should be noted that the EGFP gene was under the control of CMV promoter of the pEGFP-N1 plasmid which is located upstream of the MCS. However, after the sub-cloning process, both DKK1 and EGFP sequences are under the control of the CMV promoter from final plasmids, upstream of the DKK1 sequence. This means that the translation process would start with DKK1 and go through GFP protein. Thus, any GFP expression would guarantee the DKK1 expression. Thereafter, the embryonic kidney 293T (HEK293T) cells (293T; ATCC CCL-11268) were selected for in vitro assessment of eukaryotic expression. The cells were maintained in minimal essential medium (DMEM; Life Technologies, Inc., Grand Island, NY, USA) supplemented with 10% fetal bovine serum (FBS; HyClone Laboratories, Logan, UT, USA) and 1% penicillin–streptomycin solution (Life Technologies, Inc.). They were grown at 37 °C in air enriched with 5% CO2. Upon reaching the appropriate number, the cells were transfected by calcium phosphate method [22] with the pAK8–GVO–AAV–DKK1–GFP–TAT and pCI–DKK1–GFP plasmids. The GFP-expressing cells were excited at 491 nm, and fluorescence emission was detected at 515 nm. Fluorescence emission of expressed GFP was observed by a fluorescence microscope.
Mice Immunization
Four groups (seven mice per each group) of BALB/c female mice (Razi Institute, Karaj, Iran) were obtained (4–6 weeks old). Two groups were immunized with constructed immunogenes, while two other groups were immunized with the same immunogenes without the DKK1 gene as control groups. Six mice from each group were immunized intra-muscularly by the immunogenes and one of them was used as the control (injected by PBS). Approximately 1012 PFU phages (in 100 µL of PBS) harboring pAK8–GVO–AAV–DKK1–GFP–TAT construct (displaying TAT peptide on their surfaces) were injected to one group, while 100 µg of pCI–DKK1 plasmid (equal to 1013 copies of the plasmid) was injected to the other group. The immunizations were boosted three times with 10 day intervals and the same regimen. The test and control mice were bled 3 days after the last injection. The control immunogenes which lack the DKK1 gene were used with the same regimen.
Assessing the Antibody Response by ELISA Method
The wells of microtiter ELISA plate were coated with 3 µg/mL of whole eukaryotic DKK1 (purchased from Abcam Company) protein and incubated at 4 °C overnight. After the blocking and washing steps, mice serum (immunized with the pAK8–GVO–AAV–DKK1–GFP–TAT phages, pCI–DKK1 plasmid, pAK8–GVO–AAV–GFP–TAT phages, and pCI plasmid) were added into the wells at dilutions of 1:500, 1:1000, 1:1500, and 1:2000 and incubated for 1 h at 37 °C. Then wells were washed five times with PBS and PBST. Thereafter, 100 µL of 1/10,000 diluted HRP-conjugated anti-mouse antibody was added to the wells and incubated at 37 °C for 1 h. The reaction was stopped by 50 µL of 2 N HCl. Finally, the OD was determined at 450 nm using the ELISA reader.
Statistical Analysis
Statistical analyses were performed with the Mann–Whitney U test to evaluate the statistical comparison among all experiments. Values of p < 0.05 were considered significant.
Results
In Silico Antigen Design
Two segments of DKK1 protein, spanning 11–80 and 186–235 amino acids, were selected as the regions harboring the best immunogenicity properties (Fig. 1). Moreover, to achieve better immune responses, a fusion protein was added to the antigen sequence. The included fusion consisted of PII and P30 epitopes (to induce the CD4+ T cells which are necessary to have strong humoral responses [23]) along with the COMP domain (which is responsible for solubility of the final protein and it helps to form pentamers and may enhance the antigen uptake by antigen-presenting cells (APCs) or alleviate the B cell activation with a T-helper independent response [24,25,26]). To have a gene with high translational capability, the sequence of the final gene was optimized according to the Eukaryotic codon usage (the sequences are presented in Supplementary Fig. 1).
Immuno-informatics analysis. The results of predicting linear B cell epitopes (a) and conformational B cell epitopes (b) are presented. The regions in yellow are the B cell epitope regions. The regions with highest epitope score are presented. The regions in C terminus and N terminus of the DKK1 protein showed the highest epitope density
Construction of Phage-DNA Immunization Plasmid
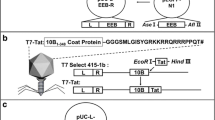
The designed TAT protein (Supplementary Fig. 2) was successfully sub-cloned and the sequencing results indicated the accuracy of the performed sub-cloning processes. Figure 2 schematically illustrates the employed strategy to construct the required plasmids for in vitro and in vivo experiments. The sub-cloning procedure to construct pAK8–GVO–AAV–DKK1–TAT and pCI–DKK1 along with the schematic drawing of these plasmids is presented in Fig. 2.
Sub-cloning strategies and constructed plasmids. The sub-cloning strategies to construct the pCI–DKK1, pCI–DKK1–GFP, pAK8–GVO–AAV–DKK1–TAT, and pAK8–GVO–AAV–DKK1–GFP–TAT plasmids are presented within the figure. The sub-cloning processes are started with sub-cloning of the chemically synthesized DKK1 and TAT genes into the target plasmids. The figure includes all intermediate plasmids employed to construct the final plasmids. Each important element within the sequence of the plasmids is depicted in a distinguished color with an accompanying description
Phage Preparation and In Vitro Assessments
The ELISA test result (Fig. 3) indicated that the TAT peptide is displayed on the surface of the phage particle. On the other hands, the emitted green florescence light from the HEK293T cells suggests the accuracy of transfection process. It indicates the ability of the constructed plasmids for eukaryotic gene expression. Figure 2 shows the schematic drawings of the GFP-containing plasmids and their sub-cloning process. Figure 4 shows the results of GFP expression within the HEK293T cells.
In vitro confirmation of the constructs. On the left is the GFP expression result for pAK8–GVO–AAV–DKK1–GFP–TAT, while GFP expression results for the pCI–DKK1–GFP is in the middle and the cells without transfection are in the right (negative control). The cells were excited at 491 nm and fluorescence emission was detected at 515 nm using a fluorescence microscope
Mice Immunization Results
The ELISA results (Fig. 5) for the immunized mice revealed that the immunization regimen have successfully elicited humoral immunity against the designed DKK1 antigen. Since the coated antigen is the eukaryotic active DKK1 protein, the antibodies are interacting solely against the DKK1 protein and the accompanying fusion protein is not included to achieve the reported ODs. Moreover, these results indicated that the produced antibody is capable of interacting with the eukaryotic whole DKK1 protein. This property implies the immunological similarity between the eukaryotic whole DKK1 protein and the designed antigen. This could be in part due to the possibility of getting similar 3D structure, disulfide bond formation, and glycosylation pattern. The higher ODs obtained from phage-immunization reveals the higher efficiency of constructed immunization vehicle. The obtained ODs for control groups reveal the specificity of the raised antibodies against the designed antigen.
Mice immunization results. Total IgG assay between mice groups was measured at 1:500 dilution of mice sera by ELISA method. Data are shown as mean values of eight samples ± SD per groups and asterisk indicates the significant differences (*p < 0.05) according to Mann–Whitney U analysis. The differences between pAK8–GVO–AAV–TAT, pCI vector, PBS, and the blank control were not significant
Discussion
Due to their particulate nature, phages are taken up by APCs, cleared from the circulation and targeted to the spleen and liver Kupffer cells which make them ideal immunogene delivery systems. Given the appealing features of the phage-DNA-immunization systems, we have devised a novel immunogene delivery construct with improved characteristics. Our immunization results indicated that using an improved phage delivery system would lead to more efficient immunity against the target antigen in comparison to conventional DNA-immunization approaches. Although the AAVP vector is introduced by Hajitou et al. [17], the studies using this sort of constructs are mostly focused on their gene delivery and cell-targeting applications. Using an AAVP-based design, we have constructed a novel hybrid phage particle (the combination of M13 phage displaying CPP–TAT on the surface and AAV–ITR eukaryotic cassette have never been reported) which aside from its common applications could be used as a successful immunogene to elicit humoral immune responses. Given our construct as the immunization agent, the researchers could circumvent all the arduous procedures associated with the protein expression and purification (as the ongoing method to produce immunogenes for poly- and monoclonal antibodies). Moreover, a significant reduction in the time needed to prepare the immunogene would be the direct consequence of developing these constructs. The issue of time is more outstanding regarding the vaccine development efforts.
Despite the various advantages offered by phages, they have not reached their full potential as immunization vehicles [27]. The rate-limiting step of phage-immunization might be mechanistically related to the post-targeting fate of the single-stranded DNA of the phage genome [28]. In an attempt to improve phage-immunization results, Hajitou et al. hypothesized that post-targeting transgene expression of the phage could be improved by genetic incorporation of compatible cis-elements [such as inverted terminal repeats (ITRs)] from a mammalian single-stranded DNA virus such as recombinant AAV [29]. A hybrid between AAV and phage genome would improve the post-transfection fate of the delivered transgene via maintenance of the entire mammalian transgene cassette, better persistence of episomal DNA, and formation of concatamers of the transgene cassette [28]. These compelling features have convinced us to design a hybrid between the elements of AAV plasmid and phage genome.
Cell-penetrating peptides (CPPs) like TAT peptide, derived from HIV-1 transactivator protein, could penetrate into living mammalian cells [30, 31]. Owing to their positive charges, these peptides use negatively charged glycosaminoglycans (GAGs) such as heparan sulfate (HS) and chondroitin sulfate (CS) as cell surface receptors for their cellular uptake [32]. The ability of cationic CPPs like TAT for Cell type-independent internalization is rooted in the presence of HS and CS on the surface of virtually all mammalian cells [30]. Although there is an agreement regarding the sulfated proteoglycans to be the cell surface receptor, the mechanism of TAT peptide endocytosis is still a place for argument [31]. Several mechanisms including clathrin-mediated endocytosis [33], caveolae-mediated endocytosis [34], or macropinocytosis [32] have been proposed for their action mechanism. TAT peptide, displayed on the surface of bacteriophage, facilitates cellular internalization of the recombinant phage into mammalian cells. It has been shown that TAT peptide on the surface of large bacteriophages such as filamentous M13 (approximately 900 nm in length and 9 nm in width) facilitated the internalization of the phage particles into various tumor cells [35, 36]. This property was the rationale behind the inclusion of this peptide in our final construct. The displayed TAT peptide on the surface of our hybrid phage could be one the contributing factors to the obtained good immune response.
It has been shown that the efficiency of DNA immunization is depending on the route of administration. Successful transfection of the cells at the administration site plays a pivotal role in developing high immune responses. The transfection rate is even more important than the amount of administered immunization agent [37]. In this regard, our results apparently corroborate the results of prior investigations. Using two immunization constructs, we have underscored the importance of the transfection rate. The employed constructs mainly differ in the parameters that ultimately matter in the transfection rate. In an idealized model of DNA and phage-DNA immunization mechanism, the local cells of administration site (the myocyte cells and APC) should present the target antigen in the context of major histocompatibility complex I and II. Direct transfection of the cells or cross presentation of the target antigen is the main cellular event that enforce antigen presentation. Afterwards, macrophages could travel into draining lymph nodes and consequently activate both humoral and cellular branches of the immune response [38]. It seems that all of the engineered features of our final phage construct helps for better deployment of immunization mechanism which ends with better results. However, we have selected a challenging antigen for our study to assess the ability of the designed construct to raise antibodies against poor immunogenes. The DKK1 protein is a secretory antigen with high identity to the DKK1 sequence of Mus musculus (the selected N terminus region for the immunogene harbors 88% and the selected C terminus region harbors 96% sequence identity to the equivalent sequences of mouse DKK1 protein). The secretory proteins are freely exposed to the host immune system and are diagnosed as self-antigens, therefore getting strong immune responses against these antigens is more challenging. Moreover, the immunizations were performed without any aid from common adjuvants or post-administration manipulations like electroporation. Despite the potential effects of aforementioned issues, the achieved immunity is stronger than the responses obtained from similar studies using other phage-immunization agents [39, 40]. Our better immunization results are more discernible in comparison to the study performed by Zhang et al. [41]. They have designed a novel DKK1 multi-epitope DNA vaccine and measured its high level expression. Although they have used electroporation after each injection, they have obtained poor humoral responses in comparison to our results. This could be construed as low immunogenicity of DKK1 molecule as well as higher efficacy of our immunization construct.
Conclusion
It could be concluded that using AAV expression cassette along with displaying the TAT peptide on the surface of the phage particle could be deemed as an effective strategy to enhance the DNA-immunization results. These kind of hybrid delivery vehicles could be used for better immunization of various hosts.
References
Wolff, J. A., Malone, R. W., Williams, P., Chong, W., Acsadi, G., Jani, A., & Felgner, P. L. (1990). Direct gene transfer into mouse muscle in vivo. Science, 247(4949 Pt 1), 1465–1468.
Tang, D. C., DeVit, M., & Johnston, S. A. (1992). Genetic immunization is a simple method for eliciting an immune response. Nature, 356(6365), 152–154.
Yang, J., Li, Y., Jin, S., Xu, J., Wang, P. C., Liang, X. J., & Zhang, X. (2015). Engineered biomaterials for development of nucleic acid vaccines. Biomaterials Research, 19, 5.
Khalili, S., Mohammadpour, H., Barough, M. S., & Kokhaei, P. (2016). ILP-2 modeling and virtual screening of an FDA-approved library: A possible anticancer therapy. Turkish Journal of Medical Sciences, 46(4), 1135–1143.
Khalili, S., Rasaee, M., & Bamdad, T. (2017). 3D structure of DKK1 indicates its involvement in both canonical and non-canonical Wnt pathways. Molecular Biology, 51(1), 155–166.
Khalili, S., Zakeri, A., Hashemi, Z. S., Masoumikarimi, M., Manesh, M. R. R., Shariatifar, N., & Sani, M. J. (2017). Structural analyses of the interactions between the thyme active ingredients and human serum albumin. Turkish Journal of Biochemistry, 42(4), 459–467.
Khalili, S., Rasaee, M. J., Mousavi, S. L., Amani, J., Jahangiri, A., & Borna, H. (2017). In silico prediction and in vitro verification of a novel multi-epitope antigen for HBV detection. Molecular Genetics, Microbiology and Virology, 32(4), 230–240.
Mard-Soltani, M., Rasaee, M. J., Khalili, S., Sheikhi, A., Hedayati, M., Ghaderi-Zefrehi, H., & Alasvand, M. (2018). The effect of differentially designed fusion proteins to elicit efficient anti-human thyroid stimulating hormone immune responses. Iranian Journal of Allergy, Asthma and Immunology, 17(2), 158–170.
Jahangiri, A., Amani, J., & Halabian, R. (2017). In silico analyses of Staphylococcal enterotoxin B as a DNA vaccine for cancer therapy. International Journal of Peptide Research and Therapeutics, 24, 1–12.
Jahangiri, A., Rasooli, I., Owlia, P., Fooladi, A. A. I., & Salimian, J. (2018). An integrative in silico approach to the structure of Omp33-36 in Acinetobacter baumannii. Computational Biology And Chemistry, 72, 77–86.
Jahangiri, A., Rasooli, I., Owlia, P., Fooladi, A. A. I., & Salimian, J. (2018). Highly conserved exposed immunogenic peptides of Omp34 against Acinetobacter baumannii: An innovative approach. Journal of Microbiological Methods, 144, 79–85.
Clark, J. R., & March, J. B. (2004). Bacteriophage-mediated nucleic acid immunisation. FEMS Immunology And Medical Microbiology, 40(1), 21–26.
March, J. B., Clark, J. R., & Jepson, C. D. (2004). Genetic immunisation against hepatitis B using whole bacteriophage lambda particles. Vaccine, 22(13–14), 1666–1671.
Jafari, N., & Abediankenari, S. (2015). Phage particles as vaccine delivery vehicles: Concepts, applications and prospects. Asian Pacific Journal of Cancer Prevention, 16(18), 8019–8029.
Clark, J. R., & March, J. B. (2006). Bacteriophages and biotechnology: Vaccines, gene therapy and antibacterials. Trends in Biotechnology, 24(5), 212–218.
Sartorius, R., Russo, D., D’Apice, L., & De Berardinis, P. (2012) Filamentous bacteriophages: An antigen and gene delivery system. In S. Baschieri (ed.), Innovation in vaccinology: From design, through to delivery and testing (pp. 123–134).Dordrecht: Springer.
Hajitou, A., Trepel, M., Lilley, C. E., Soghomonyan, S., Alauddin, M. M., Marini, F. C., Restel, B. H., Ozawa, M. G., Moya, C. A., & Rangel, R. (2006). A hybrid vector for ligand-directed tumor targeting and molecular imaging. Cell, 125(2), 385–398.
Kagey, M. H., & He, X. (2017). Rationale for targeting the Wnt signaling modulator DKK1 for oncology. British Journal of Pharmacology, 174(24), 4637–4650
Chambers, R. S., & Johnston, S. A. (2003). High-level generation of polyclonal antibodies by genetic immunization. Nature Biotechnology, 21(9), 1088–1092.
Wu, Y., Wan, Y., Bian, J., Zhao, J., Jia, Z., Zhou, L., Zhou, W., & Tan, Y. (2002). Phage display particles expressing tumor-specific antigens induce preventive and therapeutic anti-tumor immunity in murine p815 model. International Journal Of Cancer, 98(5), 748–753.
Coia, G., Hudson, P. J., & Irving, R. A. (2001). Protein affinity maturation in vivo using E. coli mutator cells. Journal of Immunological Methods, 1(251), 187–193.
Nag, A., Datta, A., Yoo, K., Bhattacharyya, D., Chakrabortty, A., Wang, X., Slagle, B. L., Costa, R. H., & Raychaudhuri, P. (2001). DDB2 induces nuclear accumulation of the hepatitis B virus X protein independently of binding to DDB1. Journal of Virology, 75(21), 10383–10392.
Panina-Bordignon, P., Tan, A., Termijtelen, A., Demotz, S., Corradin, G., & Lanzavecchia, A. (1989). Universally immunogenic T cell epitopes: Promiscuous binding to human MHC class II and promiscuous recognition by T cells. European Journal Of Immunology, 19(12), 2237–2242.
Terskikh, A. V., Le Doussal, J.-M., Crameri, R., Fisch, I., Mach, J.-P., & Kajava, A. V. (1997). “Peptabody”: A new type of high avidity binding protein. Proceedings of the National Academy of Sciences, 94(5), 1663–1668.
Clair, N. S., Shenoy, B., Jacob, L. D., & Margolin, A. L. (1999). Cross-linked protein crystals for vaccine delivery. Proceedings of the National Academy of Sciences, 96(17), 9469–9474.
Barbacid, M., Breitman, M., Lauver, A., Long, L., Vogt, P., Beemon, K., & Wang, L. (1982). Synthesis and assembly of hepatitis B virus surface antigen particles in yeast. Nature, 298, 347.
Hashemi, H., Bamdad, T., Jamali, A., Pouyanfard, S., & Mohammadi, M. G. (2010). Evaluation of humoral and cellular immune responses against HSV-1 using genetic immunization by filamentous phage particles: A comparative approach to conventional DNA vaccine. Journal of Virological Methods, 163(2), 440–444.
Hajitou, A., Trepel, M., Lilley, C. E., Soghomonyan, S., Alauddin, M. M., Marini, F. C. 3rd, Restel, B. H., Ozawa, M. G., Moya, C. A., Rangel, R., Sun, Y., Zaoui, K., Schmidt, M., von Kalle, C., Weitzman, M. D., Gelovani, J. G., Pasqualini, R., & Arap, W. (2006). A hybrid vector for ligand-directed tumor targeting and molecular imaging. Cell, 125(2), 385–398.
Hajitou, A., Rangel, R., Trepel, M., Soghomonyan, S., Gelovani, J. G., Alauddin, M. M., Pasqualini, R., & Arap, W. (2007). Design and construction of targeted AAVP vectors for mammalian cell transduction. Nature Protocols, 2(3), 523–531.
Foged, C., & Nielsen, H. M. (2008). Cell-penetrating peptides for drug delivery across membrane barriers. Expert Opinion on Drug Delivery, 5(1), 105–117.
Marschall, A. L., Frenzel, A., Schirrmann, T., Schungel, M., & Dubel, S. (2011). Targeting antibodies to the cytoplasm. mAbs, 3(1), 3–16.
Gump, J. M., & Dowdy, S. F. (2007). TAT transduction: The molecular mechanism and therapeutic prospects. Trends in Molecular Medicine, 13(10), 443–448.
Richard, J. P., Melikov, K., Brooks, H., Prevot, P., Lebleu, B., & Chernomordik, L. V. (2005). Cellular uptake of unconjugated TAT peptide involves clathrin-dependent endocytosis and heparan sulfate receptors. The Journal of Biological Chemistry, 280(15), 15300–15306.
Ferrari, A., Pellegrini, V., Arcangeli, C., Fittipaldi, A., Giacca, M., & Beltram, F. (2003). Caveolae-mediated internalization of extracellular HIV-1 tat fusion proteins visualized in real time. Molecular Therapy: The Journal of the American Society of Gene Therapy, 8(2), 284–294.
Eguchi, A., Akuta, T., Okuyama, H., Senda, T., Yokoi, H., Inokuchi, H., Fujita, S., Hayakawa, T., Takeda, K., Hasegawa, M., & Nakanishi, M. (2001). Protein transduction domain of HIV-1 Tat protein promotes efficient delivery of DNA into mammalian cells. The Journal of Biological Chemistry, 276(28), 26204–26210.
Mukai, Y., Sugita, T., Yamato, T., Yamanada, N., Shibata, H., Imai, S., Abe, Y., Nagano, K., Nomura, T., Tsutsumi, Y., Kamada, H., Nakagawa, S., & Tsunoda, S. (2006). Creation of novel protein transduction domain (PTD) mutants by a phage display-based high-throughput screening system. Biological & Pharmaceutical Bulletin, 29(8), 1570–1574.
Wang, S., Zhang, C., Zhang, L., Li, J., Huang, Z., & Lu, S. (2008). The relative immunogenicity of DNA vaccines delivered by the intramuscular needle injection, electroporation and gene gun methods. Vaccine, 26(17), 2100–2110.
Kutzler, M. A., & Weiner, D. B. (2008). DNA vaccines: Ready for prime time? Nature Reviews Genetics, 9(10), 776–788.
Ou, C., Tian, D., Ling, Y., Pan, Q., He, Q., Eko, F. O., & He, C. (2013). Evaluation of an ompA-based phage-mediated DNA vaccine against Chlamydia abortus in piglets. International Immunopharmacology, 16(4), 505–510.
Ling, Y., Liu, W., Clark, J. R., March, J. B., Yang, J., & He, C. (2011). Protection of mice against Chlamydophila abortus infection with a bacteriophage-mediated DNA vaccine expressing the major outer membrane protein. Veterinary Immunology and Immunopathology, 144(3–4), 389–395.
Zhang, X., Liu, S., Li, S., Du, Y., Dou, Y., Li, Z., Yuan, H., & Zhao, W. (2015). Designation of a novel DKK1 multiepitope DNA vaccine and inhibition of bone loss in collagen-induced arthritic mice. BioMed Research International. https://doi.org/10.1155/2015/765490.
Acknowledgements
The authors wish to thank Tarbiat Modares University for supporting the conduct of this research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Informed Consent
Informed consent was obtained from all individual participants included in the study.
Research Involving Human Participants and/or Animals
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Khalili, S., Rasaee, M.J., Bamdad, T. et al. A Novel Molecular Design for a Hybrid Phage-DNA Construct Against DKK1. Mol Biotechnol 60, 833–842 (2018). https://doi.org/10.1007/s12033-018-0115-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-018-0115-2