Abstract
Metabolism of non-glucose carbon sources is often highly regulated at the transcriptional and post-translational levels. This level of regulation is lacking in Saccharomyces cerevisiae strains engineered to metabolize xylose. To better control transcription in S. cerevisiae, the xylose-dependent, DNA-binding repressor (XylR) from Caulobacter crescentus was used to block transcription from synthetic promoters based on the constitutive Ashbya gossypii TEF promoter. The new hybrid promoters were repressed in the absence of xylose and showed up to a 25-fold increase in the presence of xylose. Activation of the promoter was highly sensitive to xylose with activity seen at concentrations below 2 μM xylose. These new xylose-inducible promoters allow improved control of gene expression for engineered strains of Saccharomyces yeasts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Economical production of bio-renewable fuels and chemicals will require the utilization of all of the sugars available in lignocellulosic materials. While glucose is the most abundant and is readily metabolized by industrial organisms like Saccharomyces cerevisiae, xylose can make up a significant percentage of the available sugar [1]. Natural xylose utilizing organisms coordinately regulate the expression of genes required for xylose metabolism. The XYL1 and XYL2 genes, encoding xylose reductase and xylitol dehydrogenase, from five xylose utilizing yeasts are highly induced by xylose, indicating the presence of a xylose-specific transcription factor in yeasts like Scheffersomyces stipitis [2]. These transcription factors are lacking in S. cerevisiae as expression of the S. stipitis XYL1 and XYL2 genes from their native promoters in S. cerevisiae resulted in low, constitutive expression of the genes [3, 4].
There is increasing interest in generating a variety of promoters for enhanced control of gene expression in S. cerevisiae. Much of this effort has focused on constitutive expression, and there are now a wide variety of options for constitutive expression from low to high levels [5, 6]. Inducible promoters are available for several conditions including the presence of galactose, copper, oxygen depletion, and glucose concentration [6, 7]. Xylose-dependent gene regulation has been lacking for S. cerevisiae until recently, where XylR repressors from six different bacteria were investigated for their ability to repress transcription from promoters modified to contain the repressor binding sites [8, 9].
Strains of S. cerevisiae engineered for xylose fermentation have been described with up to eight genes over expressed [10]. Constitutive, high level, expression of all the required genes is not necessarily beneficial to the cell. Even when one gene is over expressed, it can be a waste of resources to express the gene when it is not needed. In certain cases, constitutive expression has been shown to be detrimental. Constitutive expression of XYL1, XYL2, and TAL1, while beneficial for xylose metabolism, decreased specific growth rate and ethanol production from glucose [11]. Additionally, other genes such as ZWF1 (glucose-6-P dehydrogenase) and PGI1 (phosphoglucose isomerase) are induced in xylose utilizing yeasts, but have decreased levels in S. cerevisiae grown on xylose medium [12–14]. The availability of a xylose-inducible expression system for S. cerevisiae would allow fine-tuning the expression of genes required for xylose metabolism, while minimizing the impact of expressing numerous genes.
One possible approach to developing a xylose-specific promoter for S. cerevisiae would be to utilize xylose-inducible promoters from fungi. Transcriptional activation of xylanases in Aspergillus niger is mediated by the transcription factor XlnR, a zinc binuclear cluster-type DNA-binding protein similar to Gal4p in S. cerevisiae [15]. Similarly, Xyr1 from Trichoderma reesei, a homolog of XlnR, has been shown to bind to xylanase promoters [16]. XlnR is highly phosphorylated when it is activated by the presence of xylose [17]. However, the identity of the kinase responsible for the xylose-induced phosphorylation is not known. Without knowledge of the protein/signal responsible for activation, the expression system cannot be reconstituted in S. cerevisiae.
Transcriptional regulation has also been well-studied in bacteria, and two different types of xylose-regulated transcription have been found. These include promoters based on either transcriptional activation or repression, the latter due to DNA-binding protein interference [18–20]. Activation-type systems potentially have the same problem as using a transcription factor from fungi, which is, not having the proper activation signal in the heterologous organism. Repressor-based transcriptional control can be readily transferred to another organism and typically requires only the modified promoter sequence and expression of the repressor in the target organism [21]. In this study, we expressed the sequence-specific DNA-binding repressor protein XylR from Caulobacter crescentus in S. cerevisiae. Strains expressing the XylR protein were then used to identify synthetic hybrid promoters containing the XylR binding site sequence that also showed xylose-dependent regulation.
Materials and Methods
Strains, Media, and General Methods
Escherichia coli strain NEB10β (NEB; Beverly, MA, USA) was used for routine maintenance and preparation of plasmids, and cells were grown in LB medium [22]. Yeast strains and plasmids used in this study are listed in Tables 1 and 2, respectively. Plasmid DNA was transformed into yeast cells using a standard lithium acetate method [23]. Synthetic complete medium (SC) consisted of 6.7 g/l Difco yeast nitrogen base (YNB) (United States Biological; Marblehead, MA, USA) and was supplemented with amino acids [24]. For maintenance of plasmids, media was made without tryptophan or leucine as necessary. Synthetic medium was filter sterilized, and sterile carbon sources were added separately. For integration of the repressor and promoter::lacZ fusion cassettes into the genome, NotI-linearized pRH561 (P TEF ) and NotI-linearized pRH564 (P TEF-xylO2-1) plasmids were transformed into S. cerevisiae strain CEN.PK2-1C. After a 4-h recovery in YPD, cells that had integrated the plasmid were selected on YPD plates containing G418 (200 mg/L).
Transcriptional Activity Assays
The Beta-Glo Assay system (Promega; Madison, WI, USA) was used to determine the level of transcriptional activity from promoter::lacZ fusions, essentially as reported in [25]. Cells from cultures in log-phase (0.2–0.5 OD660) grown in SC medium were diluted in fresh medium to a final OD660 = 0.004. Unless specified, carbon sources used were either 2% sucrose or 2% each sucrose and xylose. Each assay was initiated using the same amount of cell mass to minimize variation due to differing cell concentrations. Assays were started by adding 50 µL of diluted cells to 50 µL of Beta-Glo reagent, mixed thoroughly, and incubated at room temperature. Using these conditions, activity measurements were stable from 60 to 120 min. All assays were performed in 96-well, opaque (white), flat-bottomed microtiter plates. During 60–90 min, the samples were read using the luminescence mode of a SpectraMax M5 microplate reader (Molecular Devices; Sunnyvale, CA, USA). Promoter activity (i.e., β-galactosidase activities) reported in figures is based on the Relative Light Units (RLU in millions), normalized to starting cell mass. Unless specified, promoter activity assays were performed using centromere-based plasmids to express both the repressor and lacZ marker. Assays were performed at least in triplicate.
Construction of XylR Repressor Variants and Wild-Type and Xylose-Regulated Promoters
DNA fragments for cloning were amplified using Phusion High-fidelity DNA polymerase (NEB). The codon-optimized XylR gene from Caulobacter crescentus (accession # AAK25027) was synthesized (DNA2.0; Menlo Park, CA, USA) and cloned into vectors for expression in S. cerevisiae using restriction endonuclease sites added to the ends of the gene. To facilitate entry into the nucleus, the SV40 large T-antigen nuclear localization signal (PKKKRKV) was added to the amino terminus of the XylR gene by PCR amplification of the XylR gene using primers SpeI-NLS-XylR-F (5′-GGACTAGTGCATGCCCAAGAAGAAAAGGAAAGTTAATCAACCAGTAGAAAGACAGCGTAGG-3′) and SalI-XylR-R (5′-GCGTCGACTTATTAGACTCCAGCAGGGCCTGAGG-3′) which contained the NLS sequence. The SpeI-SalI fragment-containing NLS-XylR was cloned into SpeI-SalI digested expression vector pRH164 to create pRH467. Fusion of the NLS-XylR repressor to the S. cerevisiae SSN6 gene was done by overlap PCR using primers SpeI-NLS-XylR-F (5′-GGACTAGTGCATGCCCAAGAAGAAAAGG-3′), XylR-linker-R (5′- GAACCTCCACCTCCGGAACCGACTCCAGCAGGGCCTGAG-3′), linker-SSN6-F (5′- GGTTCCGGAGGTGGAGGTTCTATGAATCCGGGCGGTGAAC-3′), and SSN6-SalI-R (5′-GGTCGACTTAGTCGTCGTAGTTTTCATCTTCTTCCACTTG-3′). A short flexible linker peptide (GSGGGGS) was used to separate the XylR and SSN6 coding regions. The fusion product from the overlap PCR was ligated to pCR2.1, and the plasmid (pRH481) was sequenced to confirm that no mutations occurred during PCR amplification of the DNA fragments. The SpeI-SalI fragment from pRH481, containing NLS-XylR-SSN6, was then sub-cloned into SpeI-SalI digested pRH164 to create pRH483, which was used as the main repressor throughout this study.
The Ashbya gossypii TEF promoter (Accession # X73978) and derivatives of the promoter containing varying numbers of the operator sequence (xylO) for binding the XylR repressor were synthesized (GenScript; Piscataway, NJ, USA). Promoters were sub-cloned into vectors for expression in S. cerevisiae using SacI/SpeI restriction endonuclease sites added to the ends of the synthesized DNA fragments. To create expression vectors pRH511 − pRH514 and pRH546 − pRH549 containing the various promoter::lacZ fusions, a DNA fragment containing a his3-lacZ fusion from pRH498 was sub-cloned into plasmids containing the various promoters using SpeI and SalI restriction sites. Plasmid pRH498 was created by replacing the HindIII-SalI fragment of HIS3 with a HindIII-SalI lacZ fragment that was PCR amplified from the lacZ fusion vector YEp353 using primers HindIII-lacZ-F (5′-CGACCTGCAGCCAAGCTTGCGATCC-3′) and SalI-lacZ-R (5′-GCGTCGACCCTGCCCGGTTATTATTATTTTTGACAC-3′). This latter cloning step generated an in-frame fusion of the lacZ gene to a short N-terminal fragment of the HIS3 gene and served as the lacZ marker for transcription activity assays. Nucleotide sequences for all promoter and repressor variants are available in supplemental Table S1.
Statistical Analyses
For experiments with three or greater biological replicates, probability analyses were performed using the Student’s t test with a two-tailed distribution and compared to the appropriate control strain. Values with P < 0.05 were considered significant for this study. Statistical analysis was performed using Microsoft Excel.
Results and Discussion
Comparison of Xylose-Dependent Regulation from Hybrid Promoters
The Ashbya gossypii TEF promoter is a constitutive promoter that works well in S. cerevisiae [26]. The A. gossypii TEF promoter has two TATA sites with transcription most likely initiating from the TATA at position -102, relative to the A. gossypii TEF ATG codon [27]. In this study, the second TATA sequence present at position -63 was mutated to reduce the possibility of transcription initiation at this location under conditions where the main site is inhibited. This type of regulation is observed with the CYC1 promoter, where inhibition of the main TATA sites leads to increased use of downstream sites [28]. Activity of the TEF promoter containing the mutated TATA site was comparable to constitutive S. cerevisiae promoters, indicating that mutation of the TATA at position -63 had minimal effect on promoter activity (supplemental Fig. S1).
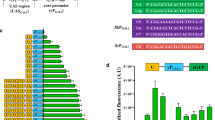
Our initial efforts using this promoter focused on inhibiting assembly of the RNA polymerase complex by placing xylO operator sites for binding XylR around the TATA element (Fig. 1). A twenty-base pair xylO sequence, 5′-ACATGTTAGCGCTACCAAGT-3′, encompassing the XylR binding site from the promoter region of the Caulobacter crescentus xylX gene was incorporated at various positions in the TEF promoter. In the absence of xylose, the XylR repressor protein from C. crescentus binds to a 14 bp DNA sequence in the xylO operator. When xylose is present, XylR was shown to dissociate from DNA containing this sequence [20], indicating that this operator sequence could be used to conditionally inhibit transcription from a synthetic promoter (Fig. 1). A similar strategy was used with tetO 2 operator sites that bind to the TetR repressor protein in the absence of tetracycline to create a Tet de-repressible promoter based on the GAL1 promoter from S . cerevisiae [29]. This approach was also used more recently to create xylose-regulated gene expression in S. cerevisiae [8, 9].
Diagram showing the synthetic promoters. a Cartoon depiction showing XylR binding the sequence-specific xylO operator in the absence of xylose. b In the presence of xylose, XylR dissociates from the DNA allowing transcription to occur. c Using the Ashbya gossypii TEF promoter, hybrid promoters were generated using xylO sequence to replace DNA around the TATA element. Positions shown are relative to the lacZ start codon
The repressor used to assay promoter activity from hybrid promoters containing the C. crescentus xylO sites was modified to contain an amino-terminal nuclear localization signal (NLS) and a C-terminal yeast chromatin remodeling protein, Ssn6p. The nuclear localization tag was added to the N-terminus of the C. crescentus XylR repressor (37.4 kDa) to facilitate transport through the nuclear pore, which has a 20-40 kDa diffusion limit, and into the yeast nucleus [30, 31]. Fusion of XylR to the S. cerevisiae protein Ssn6p (i.e., NLS-XylR-Ssn6p) was done to enhance repression when bound to the hybrid promoters. Ssn6p interacts with Tup1p and the Ssn6p/Tup1p complex is a general repressor of transcription which is thought to modify the chromatin structure around the TATA element and transcription initiation site [32]. Fusing Ssn6p to the DNA-binding protein lexA and targeting it to a hybrid promoter-containing lexA binding sites has been shown to repress transcription [33].
It was possible that replacing DNA sequence around the TATA element with xylO sequence could result in a non-functional promoter. To determine if incorporating xylO sites into the TEF promoter had a negative effect on promoter activity, strains with and without expression of the repressor were analyzed (Fig. 2). In cells without the repressor, addition of 1 xylO site immediately downstream of the TATA element (i.e., promoter P TEF-xylO1) resulted in decreased promoter activity. When assayed in cells expressing repressor, activity from promoter P TEF-xylO1 was significantly repressed relative to cells not expressing the repressor. Adding xylose to the medium reversed the repression. Promoter P TEF-xylO2-2 contained 2 xylO sites downstream of the TATA element. This promoter showed even less activity compared to the parent promoter but was still regulated by xylose. The best performing promoter, P TEF-xylO2-1, contained 2 xylO sites flanking the TATA element. This promoter had activity comparable to the strong constitutive S. cerevisiae promoters (Fig. 2 and supplemental Fig. S1). It also showed the most repression of transcription in the absence of xylose. An 8.4-fold induction (or de-repression) was observed when xylose was added to the medium (Fig. 2, strain 1061, + repressor). Induction with this promoter/repressor system was slightly higher than that obtained with other xylose-regulated expression systems which led to 1.8- to 8-fold induction [8, 9].
Promoter activity of hybrid TEF promoters containing varying numbers of xylO sites. Promoter activity was measured as beta-galactosidase activity from expression of the lacZ marker gene fused to each promoter. Both the repressor and promoter::lacZ fusions were maintained on low-copy plasmids. Activity measurements are in relative light units (RLU, millions) and are normalized to cell mass. Activity from each promoter was measured at least three times. Error bars represent the standard deviation
Disrupting the interaction of enhancers at UAS sites has also been an effective approach for regulating promoters. The A. gossypii TEF promoter has two UASrpg sites, and removal of these sites results in a non-functional promoter [27]. The yeast protein Rap1p has been shown to bind UASrpg sites, and these sites are able to activate transcription when added to minimal promoters [34]. To interfere with Rap1p binding and further increase repression, xylO sequence was used to replace sequence at various positions around the UASrpg activation sites. With the exception of P TEF-xylO3, increased repression and/or xylose-dependent gene expression were not observed, and this approach was not pursued further (supplemental Fig. S2).
Integration of the P TEF and P TEF-xylO2-1 Promoters into the Genome
Plasmid stability in cells is rarely near 100% in S. cerevisiae. Selective pressure was maintained for plasmids used in this study to determine promoter activity; however, even under these conditions, 20–30% of the cells may have lost the plasmid [35]. Cells without the repressor plasmid would be able to activate the hybrid promoter, leading to elevated promoter activity under repressive conditions. To ensure that every cell expressed the repressor, both the repressor and promoter::lacZ fusion were integrated into the genome. When grown on medium lacking xylose, transcription was repressed in strains with the hybrid promoter compared to strains with the WT promoter (Fig. 3). As seen previously with plasmid-based promoters, activity increased when xylose was present in the medium (Fig. 3, strain 1211). Compared to promoter activity with plasmids (Fig. 2, strain 1061), transcriptional repression was not significantly increased when the repressor was integrated into genome, and some promoter activity was still seen in the absence of xylose. To determine if the low level of promoter activity seen in strain 1211 was due to the repressor not fully occupying the xylO sites, additional repressor was expressed from the low-copy plasmid pRH483 (strains 1216 and 1217). A larger increase in repression on was observed in cells with extra repressor, resulting in a 25-fold induction in the presence of xylose (Fig. 3, strain YRH1217). This latter result suggested that repressor expression may be limiting and improvement in repression can be made by increasing the amount of repressor available in the cell.
Promoter activity of the chromosomally integrated xylose-regulated promoter P TEF-xylO2-1. Promoter activity was measured as beta-galactosidase activity from expression of the lacZ marker gene fused to each promoter. In all the strains, repressor was also integrated into the genome. For strains with additional repressor, extra repressor was plasmid-based (i.e., episomal repressor). Activity measurements are in relative light units (RLU, millions) and are normalized to cell mass. Activity from each promoter was measured at least three times. Error bars represent the standard deviation
Effect of XylR Modifications on Repression of the P TEF-xylO2-1 Promoter
As mentioned previously, the repressor used in these previous experiments was modified to contain an NLS and a C-terminal fusion to the yeast chromatin remodeling protein Ssn6p. To determine the impact of the added elements, we tested different combinations lacking the modifications (e.g., NLS-XylR and XylR). When xylose was added to the medium, activity from the P TEF-xylO2-1 promoter was similar for all repressor variants (Fig. 4). In the absence of xylose, the highest level of repression was observed with the NLS-XylR-Ssn6p repressor (Fig. 4, strain YRH1061). Removing the Ssn6p fusion decreased repression slightly, but the difference was not statistically significant (Fig. 4, strain 1227). Strain 1276, expressing just the native XylR, lacking both the NLS and Ssn6 fusion, further decreased the repression. These data suggested that while the NLS was beneficial, it was not absolutely required. The C. crescentus XylR (37.4 kDa) is at the top range of the passive diffusion limit through nuclear pore complexes and would not be expected to enter the nucleus efficiently in the absence of an NLS. Further inspection of the XylR sequence showed a highly charged amino terminus with a run of basic amino acids (e.g., RQRRR). Many NLS sequences contain a short cluster of basic amino acids which have been shown to contribute to NLS function [36]. It is possible that the N-terminal region of the C. crescentus XylR protein (MNQPVERQRRR) provides some function as a NLS in S. cerevisiae.
Promoter activity of strains expressing varying XylR proteins. The effect of modifying the XylR repressor was determined by measuring promoter activity from strains expressing the fully modified repressor (NLS-XylR-Ssn6p) in comparison to variants of the repressor lacking modification (e.g., NLS-XylR and XylR). Promoter activity was measured as beta-galactosidase activity from expression of the lacZ marker gene fused to the P TEF-xylO2-1 promoter. Both the repressor and P TEF-xylO2-1::lacZ fusion were maintained on low-copy plasmids. Activity measurements are in relative light units (RLU, millions) and are normalized to cell mass. Activity from each promoter was measured at least three times. Error bars represent the standard deviation
Promoter Sensitivity
In order for de-repression of transcription to occur, xylose has to be present in the cell at a high enough concentration to bind to XylR. The C. crescentus XylR protein has an estimated dissociation constant of 20–50 μM xylose [20]. However, xylose enters S. cerevisiae through hexose transporters, and their affinity for xylose is very poor (K m ~ 190 mM to 1.5 M) [37]. Additionally, glucose is a competitive inhibitor of xylose and is preferentially transported. To determine how sensitive the hybrid promoter was to xylose, and if glucose was inhibitory to de-repression, activity of the P TEF-xylO2-1 promoter was measured at low xylose concentration in medium that contained either sucrose or glucose as a carbon source. Sucrose enters the cell through a sucrose-permease, and this disaccharide would not be expected to compete with xylose for uptake. In the absence of xylose, as expected, cells lacking repressor showed high promoter activity compared to cells expressing the repressor (Fig. 5). De-repression was observed at very low levels of xylose. When sucrose was used as the carbon source, 0.2 mg/L xylose (1.33 μM) resulted in a 3.3-fold increase in promoter activity. A sixfold increase in promoter activity was seen with 2 mg/L xylose (13.3 μM). When cells were grown in medium containing glucose, xylose also increased promoter activity at low concentrations. However, after 2-h exposure to xylose, promoter activity was approximately 1.8-fold less than what was observed with sucrose. After 4-h exposure to 2 mg/L xylose, the difference in promoter activity between cells grown in glucose vs. sucrose was much less, suggesting that any inhibitory effect due to glucose occurs only at very low xylose concentration and early time points after induction.
Promoter activity for strains expressing P TEF-xylO2-1 in the presence of low xylose concentration. Promoter activity was measured as beta-galactosidase activity from expression of the lacZ marker gene fused to the P TEF-xylO2-1 promoter. Both the repressor and P TEF-xylO2-1::lacZ fusion were maintained on low-copy plasmids. Activity measurements are in relative light units (RLU, millions) and are normalized to cell mass. Activity from each promoter was measured at least three times. Error bars represent the standard deviation
Our observation of induction with 2 mg/L xylose suggests that this hybrid promoter is as sensitive to induction as the Tet- inducible promoter is in yeast with doxycycline [38]. It is also considerably more sensitive than the endogenous galactose-inducible promoter which has been shown to require between 1 and 3 g/L galactose for optimum activation [39]. Additionally, the P TEF-xylO2-1 promoter is rapidly induced, compared again to galactose induction which can take more than 20 h to reach maximal expression [39]. The P TEF-xylO2-1 promoter is also functional in the presence of glucose, making this promoter an excellent candidate for protein expression in situations where glucose is the carbon source of choice. The P TEF-xylO2-1 promoter can also be used with strains not engineered for xylose consumption. In this scenario, xylose would be an inexpensive, stable inducer that would not be significantly degraded or consumed. When used at such low concentrations, the effect of xylose on cellular metabolism would also be minimal.
Conclusions
Metabolic engineering of S. cerevisiae to convert biomass-derived xylose to fuels and chemicals will require the expression of numerous genes. Expressing genes at high levels can be detrimental to the cell, resulting in decreased cell growth and productivity. Thus, for optimum yield and productivity, it is important to control the level of expression. Correct timing of expression is also important. For example, use of a constitutive promoter for expression of the 2-PS gene for production of triacetic acid lactone resulted in 4.5-fold less product compared to using the late phase ADH2 promoter [40]. In the case of xylose metabolism, it would not be beneficial to express all of the required genes constitutively at elevated levels. To address this issue, hybrid, xylose-regulated synthetic promoters were developed. When xylose was present, promoter P TEF-xylO2-1 had activity comparable to S. cerevisiae promoters such as P PGK1 , P ADH1 , or P HXT7 and showed up to a 25-fold difference in activity compared to when xylose was not available. These new promoters provide improved control of gene expression for engineered Saccharomyces strains and are a starting point for generating new promoters with additional regulation (e.g., glucose repression), increased XylR repression, and more tunable expression levels.
References
Saha, B. C. (2003). Hemicellulose bioconversion. Journal of Industrial Microbiology and Biotechnology, 30, 279–291.
Wohlbach, D. J., Kuo, A., Sato, T. K., Potts, K. M., Salamov, A. A., Labutti, K. M., et al. (2011). Comparative genomics of xylose-fermenting fungi for enhanced biofuel production. Proceedings of the National Academy of Sciences, 108, 13212–13217.
Amore, R., Kötter, P., Kuster, C., Ciriacy, M., & Hollenberg, C. P. (1991). Cloning and expression in Saccharomyces cerevisiae of the NAD(P)H-dependent xylose reductase-encoding gene (XYL1) from the xylose-assimilating yeast Pichia stipitis. Gene, 109, 89–97.
Kötter, P., Amore, R., Hollenberg, C. P., & Ciriacy, M. (1990). Isolation and characterization of the Pichia stipitis xylitol dehydrogenase gene, XYL2, and construction of a xylose-utilizing Saccharomyces cerevisiae transformant. Current Genetics, 18, 493–500.
Nevoigt, E., Kohnke, J., Fischer, C. R., Alper, H., Stahl, U., & Stephanopoulos, G. (2006). Engineering of promoter replacement cassettes for fine-tuning of gene expression in Saccharomyces cerevisiae. Applied and Environmental Microbiology, 72, 5266–5273.
Blazeck, J., Garg, R., Reed, B., & Alper, H. S. (2012). Controlling promoter strength and regulation in Saccharomyces cerevisiae using synthetic hybrid promoters. Biotechnology and Bioengineering, 109, 2884–2895.
Da Silva, N. A., & Srikrishnan, S. (2012). Introduction and expression of genes for metabolic engineering applications in Saccharomyces cerevisiae. FEMS Yeast Research, 12, 197–214.
Teo, W. S., & Chang, M. W. (2015). Bacterial XylRs and synthetic promoters function as genetically encoded xylose biosensors in Saccharomyces cerevisiae. Biotechnology Journal, 10, 315–322.
Wang, M., Li, S., & Zhao, H. (2016). Design and engineering of intracellular-metabolite-sensing/regulation gene circuits in Saccharomyces cerevisiae. Biotechnology and Bioengineering, 113, 206–215.
Bera, A. K., Ho, N. W., Khan, A., & Sedlak, M. (2011). A genetic overhaul of Saccharomyces cerevisiae 424A(LNH-ST) to improve xylose fermentation. Journal of Industrial Microbiology and Biotechnology, 38, 617–626.
Meinander, N. Q., Boels, I., & Hahn-Hägerdal, B. (1999). Fermentation of xylose/glucose mixtures by metabolically engineered Saccharomyces cerevisiae strains expressing XYL1 and XYL2 from Pichia stipitis with and without overexpression of TAL1. Bioresource technology, 68, 79–87.
Hector, R. E., Mertens, J. A., Bowman, M. J., Nichols, N. N., Cotta, M. A., & Hughes, S. R. (2011). Saccharomyces cerevisiae engineered for xylose metabolism requires gluconeogenesis and the oxidative branch of the pentose phosphate pathway for aerobic xylose assimilation. Yeast, 28, 645–660.
Jeffries, T. W., & Van Vleet, J. R. (2009). Pichia stipitis genomics, transcriptomics, and gene clusters. FEMS Yeast Research, 9, 793–807.
Yuan, T., Ren, Y., Meng, K., Feng, Y., Yang, P., Wang, S., et al. (2011). RNA-Seq of the xylose-fermenting yeast Scheffersomyces stipitis cultivated in glucose or xylose. Applied Microbiology and Biotechnology, 92, 1237–1249.
van Peij, N. N., Visser, J., & de Graaff, L. H. (1998). Isolation and analysis of xlnR, encoding a transcriptional activator co-ordinating xylanolytic expression in Aspergillus niger. Molecular Microbiology, 27, 131–142.
Rauscher, R., Wurleitner, E., Wacenovsky, C., Aro, N., Stricker, A. R., Zeilinger, S., et al. (2006). Transcriptional regulation of xyn1, encoding xylanase I, in Hypocrea jecorina. Eukaryotic Cell, 5, 447–456.
Noguchi, Y., Tanaka, H., Kanamaru, K., Kato, M., & Kobayashi, T. (2011). Xylose triggers reversible phosphorylation of XlnR, the fungal transcriptional activator of xylanolytic and cellulolytic genes in Aspergillus oryzae. Bioscience, Biotechnology, and Biochemistry, 75, 953–959.
Gartner, D., Degenkolb, J., Ripperger, J. A., Allmansberger, R., & Hillen, W. (1992). Regulation of the Bacillus subtilis W23 xylose utilization operon: interaction of the Xyl repressor with the xyl operator and the inducer xylose. Molecular and General Genetics, 232, 415–422.
Song, S., & Park, C. (1997). Organization and regulation of the D-xylose operons in Escherichia coli K-12: XylR acts as a transcriptional activator. Journal of Bacteriology, 179, 7025–7032.
Stephens, C., Christen, B., Watanabe, K., Fuchs, T., & Jenal, U. (2007). Regulation of D-xylose metabolism in Caulobacter crescentus by a LacI-type repressor. Journal of Bacteriology, 189, 8828–8834.
Gari, E., Piedrafita, L., Aldea, M., & Herrero, E. (1997). A set of vectors with a tetracycline-regulatable promoter system for modulated gene expression in Saccharomyces cerevisiae. Yeast, 13, 837–848.
Sambrook, J., & Russell, D. W. (2001). Molecular cloning: A laboratory manual. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press.
Gietz, D., & Woods, R. A. (2002). Transformation of yeasts by the lithium acetate/single-stranded carrier/polyethylene glycol method. Methods in Enzymology, 350, 87–96.
Amberg, B. C., Burke, D. J., & Strathern, J. N. (2005). Methods in yeast genetics: A cold spring harbor laboratory course manual. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press.
Hector, R. E., Bowman, M. J., Skory, C. D., & Cotta, M. A. (2009). The Saccharomyces cerevisiae YMR315 W gene encodes an NADP(H)-specific oxidoreductase regulated by the transcription factor Stb5p in response to NADPH limitation. New Biotechnology, 26, 171–180.
Wach, A., Brachat, A., Pohlmann, R., & Philippsen, P. (1994). New heterologous modules for classical or PCR-based gene disruptions in Saccharomyces cerevisiae. Yeast, 10, 1793–1808.
Steiner, S., & Philippsen, P. (1994). Sequence and promoter analysis of the highly expressed TEF gene of the filamentous fungus Ashbya gossypii. Molecular and General Genetics, 242, 263–271.
Li, W. Z., & Sherman, F. (1991). Two types of TATA elements for the CYC1 gene of the yeast Saccharomyces cerevisiae. Molecular and Cellular Biology, 11, 666–676.
Murphy, K. F., Balázsi, G., & Collins, J. J. (2007). Combinatorial promoter design for engineering noisy gene expression. Proceedings of the National Academy of Sciences, 104, 12726–12731.
Kalderon, D., Roberts, B. L., Richardson, W. D., & Smith, A. E. (1984). A short amino acid sequence able to specify nuclear location. Cell, 39, 499–509.
Paine, P. L., Moore, L. C., & Horowitz, S. B. (1975). Nuclear envelope permeability. Nature, 254, 109–114.
Smith, R. L., & Johnson, A. D. (2000). Turning genes off by Ssn6-Tup1: A conserved system of transcriptional repression in eukaryotes. Trends in Biochemical Sciences, 25, 325–330.
Keleher, C. A., Redd, M. J., Schultz, J., Carlson, M., & Johnson, A. D. (1992). Ssn6-Tup1 is a general repressor of transcription in yeast. Cell, 68, 709–719.
Idrissi, F.-Z., Fernández-Larrea, J. B., & Piñ, B. (1998). Structural and functional heterogeneity of rap1p complexes with telomeric and UASrpg-like DNA sequences. Journal of Molecular Biology, 284, 925–935.
Sikorski, R. S., & Hieter, P. (1989). A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics, 122, 19–27.
Makkerh, J. P. S., Dingwall, C., & Laskey, R. A. (1996). Comparative mutagenesis of nuclear localization signals reveals the importance of neutral and acidic amino acids. Current Biology, 6, 1025–1027.
Kotter, P., & Ciriacy, M. (1993). Xylose fermentation by Saccharomyces cerevisiae. Applied Microbiology and Biotechnology, 38, 776–783.
Bellí, G., Garí, E., Piedrafita, L., Aldea, M., & Herrero, E. (1998). An activator/repressor dual system allows tight tetracycline-regulated gene expression in budding yeast. Nucleic Acids Research, 26, 942–947.
Li, J., Wang, S., VanDusen, W. J., Schultz, L. D., George, H. A., Herber, W. K., et al. (2000). Green fluorescent protein in Saccharomyces cerevisiae: Real-time studies of the GAL1 promoter. Biotechnology and Bioengineering, 70, 187–196.
Cardenas, J., & Da Silva, N. A. (2014). Metabolic engineering of Saccharomyces cerevisiae for the production of triacetic acid lactone. Metabolic Engineering, 25, 194–203.
Christianson, T. W., Sikorski, R. S., Dante, M., Shero, J. H., & Hieter, P. (1992). Multifunctional yeast high-copy-number shuttle vectors. Gene, 110, 119–122.
Myers, A. M., Tzagoloff, A., Kinney, D. M., & Lusty, C. J. (1986). Yeast shuttle and integrative vectors with multiple cloning sites suitable for construction of lacZ fusions. Gene, 45, 299–310.
Hector, R. E., Dien, B. S., Cotta, M. A., & Qureshi, N. (2011). Engineering industrial Saccharomyces cerevisiae strains for xylose fermentation and comparison for switchgrass conversion. Journal of Industrial Microbiology and Biotechnology, 38, 1193–1202.
Voth, W. P., Richards, J. D., Shaw, J. M., & Stillman, D. J. (2001). Yeast vectors for integration at the HO locus. Nucleic Acids Research, 29, E59.
Hauf, J., Zimmermann, F. K., & Muller, S. (2000). Simultaneous genomic overexpression of seven glycolytic enzymes in the yeast Saccharomyces cerevisiae. Enzyme and Microbial Technology, 26, 688–698.
Acknowledgements
We thank Katherine Card for her technical assistance throughout this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Ronald Hector is an inventor on U.S. Patent No. 9,506,097 B1, which pertains to the xylose-regulated promoters used in this study. Jeffrey Mertens declares that he has no competing interests.
Additional information
Disclaimer
Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hector, R.E., Mertens, J.A. A Synthetic Hybrid Promoter for Xylose-Regulated Control of Gene Expression in Saccharomyces Yeasts. Mol Biotechnol 59, 24–33 (2017). https://doi.org/10.1007/s12033-016-9991-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-016-9991-5