Abstract
Bioinformatics methods were used to analyze the role of PCAT6 in a variety of tumors and verify its role in oesophageal squamous cell carcinoma (ESCC) EC109 cells. The pan-cancer dataset was downloaded from the University of California Santa Cruz (UCSC) database to analyze the expression of PCAT6 in pan-cancer and its relationship with prognosis, clinical features, and immune infiltration. The expression and prognosis of PCAT6 in ESCC were verified by Gene Expression Omnibus (GEO) and Kaplan–Meier database. CCK8, colony formation, wound healing, Transwell cell invasion (CI), and cell migration (CM) assays were used to detect the effect of PCAT6 knockdown on the ability of ESCC cell proliferation (CP), CI and CM. Gene Set Enrichment Analysis was used to analyze the signaling pathways involved in the regulation of PCAT6. Quantitative real-time PCR and western blotting were used to examine the expression of cancer stem cell-related markers and the activation of JAK/STAT pathway in ESCC after PCAT6 knockdown. PCAT6 is significantly up-regulated in a variety of tumor tissues, and its expression is closely related to prognosis, clinical features and immune infiltration. High expression of PCAT6 leads to poor prognosis in ESCC patients. In ESCC EC109 cells, PCAT6 knockdown inhibited the ability of CP, CI, CM, and stemness, and inhibited the activation of JAK/STAT signaling pathway. PCAT6 expression is elevated in a variety of tumors. PCAT6 plays an oncogene role in ESCC by activating the JAK/STAT signaling pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Oesophageal cancer (ESCA) is a widespread gastrointestinal cancer, with an incidence rate ranking seventh worldwide (604,000 new cases) and a total mortality rate ranking sixth in the world (544,000 new cases) [1]. Oesophageal squamous cell carcinoma (ESCC) and oesophageal adenocarcinoma (EA) are two of the most prevalent histological subtypes of ESCA. More than 90% of ESCA cases in China are ESCC, which is the leading cause of death in patients with ESCA [2]. Surgical and medical treatments are evolving, but many patients with ESCC are still in the advanced stages when diagnosed [3]. The 5-year overall survival (OS) rate of patients with ESCC is less than 13%, owing to recurrence, extensive cell invasion (CI), and metastasis [4]. Therefore, exploring new markers has important clinical significance for the early detection and timely therapeutic intervention of ESCC.
Long noncoding RNAs (lncRNAs) are over 200 nucleotides long and do not code for proteins [5]. Increasing evidence has shown that lncRNAs play essential function in tumour cell proliferation (CP), cell differentiation (CD), CI, and cell migration (CM) [6]. LncRNAs can not only be used as markers for the early diagnosis of tumours but also as therapeutic targets for tumours [7]. Therefore, an in-depth study of the role of LncRNAs in tumours has important clinical significance. Prostate cancer-associated transcript 6 (PCAT6) is a newly discovered lncRNA, 764 bp in length, located on chromosome 1q32.1 [8]. PCAT6 was first discovered in keratinocytes. Studies have shown that prostate cancer cell metastasis and keratinocyte proliferation in prostate cells are significantly associated with PCAT6 [9, 10]. A number of studies have reported that PCAT6 is carcinogenic, including prostate cancer [11], bladder cancer [12], ovarian cancer [13], and colon cancer [14].However, there is no report on PCAT6 in pan-cancer, and no study on the biological function and potential mechanism of PCAT6 in ESCC.
In this study, bioinformatics was used to analyze the expression of PCAT6 in pan-cancer and its relationship with prognosis, clinical features and immune infiltration of various tumors. In addition, we demonstrated that PCAT6 was significantly upregulated in ESCC tissues, and its high expression was related to worse patient prognosis. To the best of our knowledge, these results are the first to demonstrate that PCAT6 accelerates ESCC migration and stemness via the JAK/STAT pathway.
Materials and methods
Experimental material
ESCC EC109 cells were obtained from Nanjing Kebai Biotechnology Co. Ltd. The cell culture reagents were RPMI 1640 medium (Gibco, Australia), foetal bovine serum (FBS; Gibco, Australia), and penicillin–streptomycin. The materials used for cell transfection were Opti-MEM™ I reduced serum medium (ORSM, Gibco, USA), Lipofectamine 3000 reagent (Invitrogen, USA), and small interfering RNA (siRNA). siRNA was purchased from RiboBio (Guangzhou, China), and the sequences are shown in Supplementary Table 1. RNA extraction and quantitative real-time PCR (qRT-PCR) were performed using TRIzol (Sangon Biotech, China), lnRcute lncRNA First-Strand cDNA Kit (TIANGEN, Code NO. KR202), and lnRcute lncRNA qPCR Kit (SYBR Green) (TIANGen, Code NO. FP402). Primers for PCAT6, JAK2, STAT3, cyclin D1 (CCND1), SOX2, SOX9, OCT4, and NANOG were purchased from GENERAL BIOL (Anhui, China), and the sequences are shown in Supplementary Table 2. The reagents for protein isolation and western blotting were RIPA Lysis Buffer (Beyotime Biotech, China), PMSF (Biosharp, China), phosphatase inhibitor 100× (Biosharp, China), Modified BCA Protein Assay Kit (Sangon Biotech, China), nitrocellulose membrane (Beyotime Biotech, China), HRP-labelled Goat Anti-Mouse IgG (H + L) (Beyotime Biotech, China), HRP-labelled Goat Anti-Rabbit IgG (H + L) (Beyotime Biotech, China), ECL chemiluminescence kit (Beyotime Biotech, China), CCND1 (Cat# 60,186–1-Ig, 1:5000, mouse monoclonal antibody, Proteintech Group), JAK2 (Cat#A19629, 1:2000, rabbit monoclonal antibody, ABclonal), STAT3 (Cat# 60,199–1-Ig, 1:2000, mouse monoclonal antibody, Proteintech Group), phosphor (p)-STAT3 (Cat#AP0705, 1:1000, rabbit monoclonal antibody, ABclonal), phosphor (p)-JAK2 (Cat#AP0531, 1:1000, rabbit monoclonal antibody, ABclonal), GAPDH (Cat# 60,004–1-Ig, 1:20,000, mouse monoclonal antibody, Proteintech Group), and Cell Counting Kit-8 (Beyotime Biotech, China).
Cell culture
ESCC EC109 cells were grown in complete medium (90% RPMI 1640 medium, 10% FBS, and 1% penicillin–streptomycin). The cells were then incubated at 37 °C and 5% CO2 until 2–3 days after replacing the medium when the cells grew to about 80% confluency; cells in the logarithmic growth phase were employed for subsequent experiments.
Transfection
EC109 cells were seeded in six-well plates and transfected with Lipofectamine 3000 reagent when cell confluence reached 70–90%. To increase transfection efficiency, only basic medium and ORSM were introduced to the wells. Three wells were used as an example: configuration A solution (ORSM 375 μL + 22.5 μL Lipofectamine 3000 reagent) and B solution (ORSM 375 μL + 11.25 μL siRNA). The mixture of solution A and solution B was kept at 25 ± 5 °C for 10–15 min, and then 250 μL of the mixture was introduced to each well. The cells were collected for further analysis after 24–72 h of incorporation.
RNA isolation and qRT-PCR
TRIzol reagent was used to isolate total RNA from cells, following the manufacturer’s instructions. The lnRcute lncRNA First-Strand cDNA Kit was used to remove genomic DNA and reverse synthesise the first cDNA strand. Finally, qRT-PCR was performed using the lnRcute lncRNA qPCR kit under the following conditions: 95 °C for 3 min, 40 cycles of 95 °C for 5 s, and 60 °C for 15 s. Relative lncRNA PCAT6 levels in EC109 cells were quantified using the Ct method following completion of the reaction.
Western blotting
Total proteins were extracted using the RIPA Lysis Buffer (Strong), PMSF, and phosphatase inhibitor at a ratio of 100:1:1. The absorbance of the protein samples was measured using an enzyme-linked immunosorbent assay (ELISA; Biotek, USA), and the protein concentration was calculated based on the standard curve. The target protein was separated using 10% SDS-PAGE, followed by transfer to a nitrocellulose membrane. CCND1 and STAT3 were blocked in 5% skim milk at 25 ± 5 °C for 2 h, JAK2 and p-STAT3 were blocked in 3% skim milk at 25 ± 5 °C for 2 h, and p-JAK2 was blocked in 3% bovine albumin at 25 ± 5 °C for 2 h. Blots were incubated with the primary antibody at 4 °C for 18–24 h, followed by three 1 × TBST-based rinses of 7 min each. Then, blots were incubated with the secondary antibody at 25 ± 5 °C for 2 h, followed by three 1× TBST rinses of 7 min each. Next, chemiluminescence solution was added to the blots before exposure. The grey value of protein expression was analysed using the ImageJ software.
Cell proliferation (CP) and colony formation (CF) assay
The Cell Counting Kit-8 was employed to determine the proliferative ability of ESCC EC109 cells. Cells in the logarithmic growth phase were seeded in 12-well plates at a density of 2 × 105 cells/well, followed by incubation for 0, 24, 48, 72, and 96 h. Before the measurement, cells were incubated with the reagent for 1 h. Then, the liquid was added to the 96-well plate at 100 μL/well, followed by absorbance measurement via ELISA. The CF ability of EC109 cells was determined using a CF assay. Subsequently, 0.5 × 103 cells/well were seeded in six-well plates, grown for 15 days, fixed with paraformaldehyde for 30 min, and subsequently stained with 1% crystal violet for 30 min.
Wound healing assay
A wound healing assay was performed to determine the migratory ability of ESCC EC109 cells. A marker was used to mark the line under the back of the 6-well plate to ensure that each photograph was in the same position. When the cells were evenly distributed, and the cell fusion rate was 90–95%, a 200 μL gun was used for scratching. Scratches were straight and uniform in width. After washing 3–6 times with PBS, the first photograph was taken under a microscope. RPMI 1640 medium containing 1% FBS was added, and the cells were photographed after 24 and 48 h of culture in a cell incubator. Finally, potential changes in cell migratory ability were determined according to the width of wound healing.
Transwell assay
Transwell invasion and migration assays were performed to determine the CI and CM abilities of the ESCC EC109 cells. Transwell CI assay: A dilution gel (Matrigel gel: RPMI 1640 medium = 1:8) was added to the upper chamber and incubated in a cell culture box for 1 h. 20% FBS RPMI 1640 medium was added to the hole as an inducer to induce the cells in the upper chamber through the hole on the membrane to reach the lower chamber. Then, 200 μL of cell suspension without FBS (1 × 105 cells/well) was added to the upper chamber. After 48 h of culture, the cells were fixed with paraformaldehyde for 30 min and stained with 1% crystal violet for 20 min after air drying. Finally, the cells in the upper chamber were wiped clean, and the cells at the bottom of the upper chamber were observed under a microscope. The dilution gel was not added for the Transwell CM assay, and the cells were cultured in a cell incubator for 24 h. The rest of the steps were as reported above.
Bioinformatics analysis
The RNA-seq data and clinical data of 33 tumors in the TCGA database were downloaded from the University of California Santa Cruz (UCSC, https://xena.ucsc.edu) database, and the differential expression level of PCAT6 in Pan-Cancer and its relationship with the survival and clinical characteristics of tumor patients were analyzed. Due to the small number of normal samples in the TCGA database, we used the UCSC database to download the TCGA TARGET GTEx dataset (which integrates the data of normal tissues in the GTEx database and the data of TCGA tumor tissues) to further analyze the differential expression level of PCAT6 in Pan-Cancer (Tumor name abbreviations see Supplementary Table 3). The score data of six kinds of immune infiltrating cells in 33 kinds of tumors were downloaded from the Tumor lMmune Estimation Resource (TIMER, https://cistrome.shinyapps.io/timer/) database, and the correlation between PCAT6 expression and immune cell score was analyzed. The R language estimate package was used to analyze the ImmuneScore and StromalScore of each tumor sample, and the correlation between PCAT6 expression and immune score was observed in 33 tumors. The gene expression data for ESCA and normal oesophageal epithelial tissues were downloaded from Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/) (GSE100942, GPL570), and the expression data of PCAT6 were extracted for differential expression analysis. The online website of the Kaplan–Meier plotter (http://kmplot.com/analysis/) was used to analyse the survival of PCAT6 patients with ESCA. Gene Set Enrichment Analysis (GSEA) was used to analyze the signaling pathways involved in the regulation of PCAT6.
Statistical analysis
All data analyses were performed using the R software v4.0 and GraphPad Prism 8. Kruskal-Walis test was used to analyze the expression level of PCAT6 in different normal tissues and different tumor tissues. Kaplan–Meier (K-M) survival analysis was used to compare the relationship between PCAT6 expression and tumor prognosis. Univariate Cox regression analysis was used to calculate the correlation between PCAT6 expression in tumor and patient survival. When p < 0.05 and |R|> 0.20, the correlation is considered significant. The experimental data are expressed as the mean ± SD, and the t-test was used to compare the two groups. p < 0.05 was considered statistically significant (*p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001).
Results
Expression of PCAT6 in various tumor tissues
The results of the TCGA database pan-cancer analysis showed that PCAT6 was expressed in a variety of tumors. Compared with normal tissues, PCAT6 mRNA expression was significantly up-regulated in GBMLGG, LGG, LUAD, COAD, COADREAD, BRCA, ESCA, STES, KIRP, KIPAN, PRAD, UCEC, HNSC, LUSC, LIHC, THCA, READ, BLCA, and CHOL, and significantly down-regulated in KICH (Fig. 1a). The results of TCGA database combined with GTEx database showed that the mRNA expression of PCAT6 in GBM, STAD, WT, OV, PAAD, UCS, ALL, LAML was also significantly up-regulated, and the mRNA expression in SKCM and TGCT was also significantly down-regulated (Fig. 1b).
The effect of PCAT6 expression on the prognosis of tumor patients
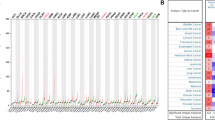
The relationship between PCAT6 expression and OS showed that PCAT6 had prognostic significance only in ACC, COAD, KICH, KIRC, LGG, LIHC, SARC, SKCM, THCA and UVM (Fig. 2a). The results of K-M survival curve showed that the high expression of PCAT6 in ACC, COAD, KICH, KIRC, LGG, LIHC, SARC, SKCM and THCA was associated with poor prognosis, and the low expression of PCAT6 in UVM was associated with poor prognosis (Fig. 3a). Considering that there may be factors that cause death not caused by tumors during follow-up, we analyzed the relationship between PCAT6 expression and prognosis Disease Specific Survival (DSS). The results showed that PCAT6 had prognostic significance in ACC, KICH, KIRC, LGG, SKCM, STAD, THCA, THYM and UVM (Fig. 2b). The results of K-M survival curve showed that the high expression of PCAT6 in ACC, KICH, KIRC, LGG, SKCM, STAD, THCA and THYM was associated with poor prognosis, and the low expression of PCAT6 in UVM was associated with poor prognosis (Fig. 3b). Subsequently, we further analyzed the relationship between PCAT6 expression and prognosis Disease Free Survival (DFS). The results showed that PCAT6 had prognostic significance in ACC, BRCA, COAD and ESCA (Fig. 2c). The results of K-M survival curve showed that high expression of PCAT6 in ACC, COAD and ESCA was associated with poor prognosis (Fig. 3c). Finally, we analyzed the relationship between PCAT6 expression and progression free survival (PFS). The results showed that PCAT6 had prognostic significance in ACC, COAD, KICH, KIRC, LICH, PCPG, STAD and UVM (Fig. 2d). The results of K-M survival curve showed that high expression of PCAT6 in ACC, COAD, KICH, KIRC, LICH, PCPG and STAD was associated with poor prognosis and the low expression of PCAT6 in UVM was associated with poor prognosis (Fig. 3d).
Relationship between PCAT6 expression and prognosis of tumor patients. (a) Univariate COx regression analysis of the relationship between PCAT6 and OS in 33 tumors (red represents p < 0.05). (b) Univariate COx regression analysis of the relationship between PCAT6 and DSS in 33 tumors (red represents p < 0.05). (c) Univariate COx regression analysis of the relationship between PCAT6 and DFS in 33 tumors (red represents p < 0.05). (d) Univariate COx regression analysis of the relationship between PCAT6 and PFS in 33 tumors (red represents p < 0.05)
Relationship between the expression of PCAT6 and the clinical characteristics of patients with various tumors
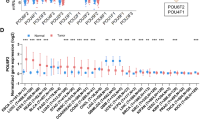
The correlation between the expression of PCAT6 and the clinical characteristics of patients with multiple cancers was analyzed after excluding the cancers with less than 3 samples in a single cancer. The results showed that the expression of PCAT6 was closely related to the tumor T stage and N stage of ESCA, STES, KIPAN, PRAD, HNSC, KIRC, LIHC, THCA (Fig. 4a, b; Supplementary Tables 4, 5) and the tumor M stage of CESC, COADREAD, KIRC, PAAD (Fig. 4c; Supplementary Table 6). In addition, the expression of PCAT6 was closely related to the tumor stage of COAD, COADREAD, KIPAN, KIRC, LIHC, THCA (Fig. 4d; Supplementary Table 7) and the tumor grade of STES, KIPAN, STAD, HNSC, KIRC, PAAD (Fig. 4e, Supplementary Table 8). Finally, the expression of PCAT6 was closely related to the gender of STES, KIRC, LUSC patients (Fig. 4f, Supplementary Table 9) and the age of ESCA, GBMLGG, LUAD, STES patients (Fig. 4g, Supplementary Table 10).
Relationship between the expression of PCAT6 and the clinical characteristics of patients with various tumors. (a–c) The relationship between PCAT6 expression and tumor TNM stage. (d) The relationship between PCAT6 expression and tumor stage. (e) The relationship between PCAT6 expression and tumor grade. (f) The relationship between the expression level of PCAT6 and the gender of tumor patients. (g) The relationship between the expression level of PCAT6 and the age of tumor patients. (*p < 0.05, **p < 0.01, ***p < 0.001)
Relationship of PCAT6 expression with tumor immune invasion and tumor microenvironment
The correlation between PCAT6 expression and the data of six immune infiltrating cells in 33 tumors showed that the expression level of PCAT6 was significantly correlated with the level of immune infiltration in LGG, HNSC, SKCM, PRAD, BRCA, GBMLGG, LUAD, STAD. The expression level of PCAT6 was negatively correlated with the levels of six tumor-infiltrating immune cells in LGG, HNSC, SKCM, PRAD, BRCA, GBMLGG, LUAD, STAD, namely B cells, T cells CD4, T cells CD8, dendritic cells, macrophages and neutrophils (P < 0.05, Supplementary Fig. 1a). The relationship between PCAT6 expression and ImmuneScore, StromalScore and ImmuneScore of ESTIMATE in 33 tumors showed that PCAT6 expression was negatively correlated with ImmuneScore of LGG, BRCA and LUAD, negatively correlated with StromalScore of BLCA, HNSC and STAD, and negatively correlated with ImmuneScore of ESTIMATE of LUAD, LUSC and HNSC (Supplementary Fig. 1b; Table 11–13).
PCAT6 is upregulated in ESCC and correlates with the prognosis of patients with ESCC
The expression level of PCAT6 in ESCC was further verified by GEO chip. The results showed that PCAT6 was highly expressed in ESCC (p < 0.05, Supplementary Fig. 2a). The Kaplan–Meier survival plot showed that elevated PCAT6 expression correlated with shorter OS in ESCC patients (HR = 2.26 (1.03–6.63), p = 0.036) (Supplementary Fig. 2b). These results indicate that the PCAT6 was upregulated in ESCC and correlated with the poor prognosis of patients with oesophageal carcinoma.
PCAT6 acts as an oncogene in ESCC
The experiment was divided into si-LncRNA PCAT6-1, si-LncRNA PCAT6-2, si-LncRNA PCAT6-3, and si-NC. The si-LncRNA PCAT6-1, with the highest transfection efficiency, was selected via qRT-PCR for further experiments (Fig. 5a). CP assay showed that PCAT6 knockdown inhibited ESCC EC109 CP (p < 0.05, Fig. 5b). The CF assay showed that the CF ability of ESCC EC109 cells was significantly inhibited after PCAT6 knockdown (p < 0.05, Fig. 5c). The wound-healing assay revealed that the CM distance of the si-LncRNA PCAT6 group was shorter relative to that of the si-NC group (p < 0.05, Fig. 5d). Transwell CI and CM assays showed that the number of invasive and migratory cells in the si-LncRNA PCAT6 group was markedly reduced compared to those in the si-NC group (p < 0.05, Fig. 5e). These results suggest that PCAT6 acts as an oncogene in ESCC.
PCAT6 through the JAK/STAT pathway to accelerate proliferation, invasion, migration, and stemness of ESCC. (a) The efficiency of PCAT6 knockdown in ESCC EC109 cells was assessed via qRT-PCR. (b) The outcome of PCAT6 knockdown on proliferation of ESCC EC109 cells by CCK8 assay. (c) The outcome of PCAT6 knockdown on the colony formation of ESCC EC109 cells assessed via colony formation assay. (d) The outcome of PCAT6 knockdown on wound healing of ESCC EC109 cells via wound healing assay. (e) The outcome of PCAT6 knockdown on migration and invasion of ESCC EC109 cells via Tanswell migration and invasion assay. (f–g) Outcome of PCAT6 knockdown on the SOX2, SOX9, OCT4, and SOX2-related gene expression in ESCC EC109 cells. (h–j) Outcome of PCAT6 knockdown on the expression of JAK/STAT network-related genes and proteins, namely, JAK2, STAT3, CCND1, P-JAK2, and P-STAT3 in ESCC EC109 cells. (*p < 0.05, **p < 0.01, ***p < 0.001)
PCAT6 contributes to the maintenance of ESCC Cancer Stem cell (CSC) characteristics
The qRT-PCR results showed that after PCAT6 knockdown, the mRNA expression of SOX2 (p = 0.0098), SOX9 (p = 0.002), OCT4 (p = 0.003), and NANOG (p = 0.014) reduced dramatically (Fig. 5f). Western blot analysis revealed that PCAT6 knockdown decreased the protein levels of SOX2 (p = 0.022), SOX9 (p = 0.027), OCT4 (p = 0.045), and NANOG (p = 0.016) (Fig. 5g). Therefore, PCAT6 regulates the CSC characteristics of ESCC EC109 cells by modulating the expression of SOX2, SOX9, OCT4, and NANOG.
PCAT6 accelerates the ESCC EC109 CP via the JAK/STAT pathway
We performed KEGG enrichment analysis to determine the possible signalling pathways involved in PCAT6 function in ESCC. The screening enrichment results showed the top 10 signalling pathways (Table 1). The Janus kinase/signal transducer of activators of transcription (JAK/STAT) axis is a commonly utilised network for cytokine signal transduction affecting CP, CD, cell apoptosis (CA), and inflammation (NES = 2.02, NOM p-value = 0.000, FDR q-val = 0.008). qRT-PCR data indicates that PCAT6 knockdown reduced the transcript levels of JAK2 (p = 0.016) and CCND1 (p = 0.001) but did not affect STAT3 transcript levels (p = 0.404) (Fig. 5h). Western blot analysis revealed that after PCAT6 knockdown, the protein levels of JAK2 (p = 0.025), p-JAK2 (p = 0.045), p-STAT3 (p = 0.027), and CCND1 (p = 0.023) decreased, while the difference in STAT3 protein expression level was not statistically significant (p = 0.969) (Fig. 5i). These results suggest that PCAT6 regulates the progression of oesophageal squamous cell carcinoma EC109 cells through the JAK/STAT pathway.
Discussion
Increasingly, studies have shown that lncRNA is widely involved in regulating gene expression and plays a vital role in CP, CD, CA, and tumourigenesis[15]. PCAT6 is a newly discovered lncRNA that promotes the progression of a variety of tumors [8, 10]. It has been reported that the expression of PCAT6 is up-regulated in most tumors such as STAD [16], Ovarian Cancer [17] and Cervical Cancer [18] compared with normal tissues. This study integrated the samples of TCGA and GTEx databases, and observed that PCAT6 was highly expressed in various tumors in the integrated data. Then according to the expression level of PCAT6, the samples were divided into high and low expression groups, and the prognosis of PCAT6 in pan-cancer was further analyzed. The results showed that the expression of PCAT6 affected the prognosis of patients with ACC, BRCA, COAD, ESCA, KICH, KIRC, LGG, LIHC, SARC, SKCM, THCA, UVM, STAD, THYM and PCPG. We also analyzed the relationship between PCAT6 expression and clinical characteristics of patients. The results showed that the expression of PCAT6 was closely related to TNM stage, stage, grade, gender and age of different types of cancer. In addition, we also studied the relationship between PCAT6 expression and immune infiltration level and immune score in different types of cancer. The results showed that PCAT6 was involved in tumor immune invasion, and the expression level of PCAT6 was correlated with immune score. These results suggest that PCAT6 is expected to be a prognostic indicator in the above tumors, but further research is needed.
ESCC is a malignant tumour derived from the oesophageal epithelial cells. Since most ESCC patients are at an advanced stage at the time of diagnosis, the 5-year OS rate for ESCC patients is less than 13% [4]. Therefore, there is an urgent need for new and specific molecular markers that enhance the efficiency of early diagnosis of ESCC to detect and treat diseases early and prolong the 5-year OS rate of patients with ESCC. In recent years, the study of lncRNA in ESCC has attracted the attention of researchers. For example, lncRNAs FER1L4 and NLIPMT are downregulated in ESCC and inhibit ESCC growth and invasion [19, 20]. LINC01296, lncRNA CASC9, and lncRNA GHET1 are highly expressed in ESCC, and their high expression promotes the development of ESCC and adversely affects patient prognosis [21,22,23]. Therefore, we verified the relationship between PCAT6 and ESCC from biological function and molecular mechanism. Firstly, we used GEO database and Kaplan–Meier database to analyze the expression and prognosis of PCAT6 in ESCC. The results showed that PCAT6 was highly expressed in ESCC and associated with poor prognosis. Then we knocked down PCAT6 and found that PCAT6 knockdown significantly inhibited the ability of ESCC cells to CP, CI and CM. In addition, knockdown of PCAT6 inhibited mRNA and protein expression levels of CSC-related markers (SOX2, SOX9, OCT4, NANOG). It indicates that PCAT6 exists in ESCC as an oncogene.
We also performed GSEA and found that PCAT6 may affect ESCC proliferation and metastasis by regulating the JAK / STAT pathway. The JAK/STAT axis, a stress inflammatory signalling network, regulates multiple biological processes like CP, CD, CA, and immune response [24]. For example, the lncRNA LINC00669 promotes nasopharyngeal cancer CP and CI by activating the JAK/STAT network [25]. The lncRNA BRCAT54 inhibits tumourigenesis of NSCLC by interacting with RPS9 to regulate the JAK/STAT pathway [26], indicating that lncRNA is associated with the JAK/STAT pathway. The results showed that the expression of JAK/STAT pathway-related proteins was reduced after PCAT6 knockdown, indicating that PCAT6 inhibited the activation of JAK/STAT signaling pathway.
Notably, STAT3 is a CSC-related marker that regulates stemness and metastasis of tumour cells [27]. Wang et al. [28] reported that STAT3 influences the characteristics of stem cells in cervical carcinoma, thereby affecting the occurrence and metastasis of cervical carcinoma. Tian et al. [29] reported that AP markedly abrogated the DOX-triggered stemness and metastasis of osteosarcoma cells by suppressing Sox2 via the STAT3 network. Therefore, based on the mechanism of the JAK/STAT pathway, we speculate that when the kinase function of JAK2 is activated, it first leads to the self-phosphorylation of JAK2; subsequently, phosphorylated JAK2 can phosphorylate STAT3 as the substrate of JAK2 kinase and downstream signalling molecules to form activated STAT3. Then, activated STAT3 enters the nucleus and binds to DNA to regulate the expression of CSC-related markers, affecting tumour stemness. Taken together, the JAK/STAT axis contributes to multiple biological functions like CP, CD, and CA, which may be achieved by affecting tumour stemness.
In conclusion, in most tumors, the expression of PCAT6 is related to the prognosis, clinical characteristics and immune infiltration level of tumor patients. PCAT6 is highly expressed in ESCC and its high expression is associated with poor prognosis. Functionally, PCAT6 promotes CP, CI and CM in ESCC EC109 cells. Mechanistically, PCAT6 promotes ESCC progression by activating the JAK/STAT signaling pathway. It indicates that PCAT6 may be a potential biomarker for ESCC treatment.
Data availability statement
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Sung H, et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209–49.
Niu C, et al. Risk factors for esophageal squamous cell carcinoma and its histological precursor lesions in China: A multicenter cross-sectional study. BMC Cancer. 2021;21(1):1034.
Li W, Liu J, Zhao H. Identification of a nomogram based on long non-coding RNA to improve prognosis prediction of esophageal squamous cell carcinoma. Aging (Albany NY). 2020;12(2):1512–26.
Hu X, et al. circGSK3β promotes metastasis in esophageal squamous cell carcinoma by augmenting β-catenin signaling. Mol Cancer. 2019;18(1):160.
Kong X, et al. LncRNA-CDC6 promotes breast cancer progression and function as ceRNA to target CDC6 by sponging microRNA-215. J Cell Physiol. 2019;234(6):9105–17.
Tan YT, et al. LncRNA-mediated posttranslational modifications and reprogramming of energy metabolism in cancer. Cancer Commun (Lond). 2021;41(2):109–20.
Zhang YX, et al. LncRNA TUC338 promotes invasion of lung cancer by activating MAPK pathway. Eur Rev Med Pharmacol Sci. 2018;22(2):443–9.
Jiang F, et al. PCAT6 May Be a Whistler and Checkpoint Target for Precision Therapy in Human Cancers. Cancers (Basel). 2021;13(23):6101.
Shi SB, et al. PCAT6 may be a new prognostic biomarker in various cancers: a meta-analysis and bioinformatics analysis. Cancer Cell Int. 2021;21(1):370.
Wang S, et al. The Role of lncRNA PCAT6 in Cancers. Front Oncol. 2021;11: 701495.
Lang C, et al. m(6) A modification of lncRNA PCAT6 promotes bone metastasis in prostate cancer through IGF2BP2-mediated IGF1R mRNA stabilization. Clin Transl Med. 2021;11(6): e426.
Xia W, et al. LncRNA PCAT6 aggravates the progression of bladder cancer cells by targeting miR-513a-5p. Eur Rev Med Pharmacol Sci. 2020;24(19):9908–14.
Kong FR, et al. LncRNA PCAT6 promotes occurrence and development of ovarian cancer by inhibiting PTEN. Eur Rev Med Pharmacol Sci. 2019;23(19):8230–8.
Huang W, et al. Long noncoding RNA PCAT6 inhibits colon cancer cell apoptosis by regulating anti-apoptotic protein ARC expression via EZH2. Cell Cycle. 2019;18(1):69–83.
Liang W, et al. Overexpression of LINC00691 promotes the proliferation and invasion of gastric cancer cells via the Janus kinase/signal transducer and activator of transcription signalling pathway. Int J Biochem Cell Biol. 2020;123: 105751.
Xu Y, et al. PCAT6 participates in the development of gastric cancer through endogenously competition with microRNA-30. Eur Rev Med Pharmacol Sci. 2018;22(16):5206–13.
Tan X, et al. The cancer-testis long non-coding RNA PCAT6 facilitates the malignant phenotype of ovarian cancer by sponging miR-143-3p. Front Cell Dev Biol. 2021;9: 593677.
Ma Z, et al. LncRNA PCAT6 accelerates the progression and chemoresistance of cervical cancer through up-regulating ZEB1 by sponging miR-543. Onco Targets Ther. 2020;13:1159–70.
Ma W, et al. LncRNA FER1L4 suppressed cancer cell growth and invasion in esophageal squamous cell carcinoma. Eur Rev Med Pharmacol Sci. 2018;22(9):2638–45.
Li D, et al. LncRNA NLIPMT inhibits tumorigenesis in esophageal squamous-cell carcinomas by regulating miR-320/survivin axis. Cancer Manag Res. 2020;12:12603–12.
Wang B, Liang T, Li J. Long noncoding RNA LINC01296 is associated with poor prognosis in ESCC and promotes ESCC cell proliferation, migration and invasion. Eur Rev Med Pharmacol Sci. 2018;22(14):4524–31.
Liu H, Zhen Q, Fan Y. LncRNA GHET1 promotes esophageal squamous cell carcinoma cells proliferation and invasion via induction of EMT. Int J Biol Markers. 2017;32(4):e403–8.
Gao GD, et al. LncRNA CASC9 promotes tumorigenesis by affecting EMT and predicts poor prognosis in esophageal squamous cell cancer. Eur Rev Med Pharmacol Sci. 2018;22(2):422–9.
Hu X, et al. The JAK/STAT signaling pathway: from bench to clinic. Signal Transduct Target Ther. 2021;6(1):402.
Qing X, et al. LINC00669 insulates the JAK/STAT suppressor SOCS1 to promote nasopharyngeal cancer cell proliferation and invasion. J Exp Clin Cancer Res. 2020;39(1):166.
Yang W, et al. LncRNA BRCAT54 inhibits the tumorigenesis of non-small cell lung cancer by binding to RPS9 to transcriptionally regulate JAK-STAT and calcium pathway genes. Carcinogenesis. 2021;42(1):80–92.
Bradshaw A, et al. Cancer stem cell hierarchy in glioblastoma multiforme. Front Surg. 2016;3:21.
Wang H, et al. STAT3 influences the characteristics of stem cells in cervical carcinoma. Oncol Lett. 2017;14(2):2131–6.
Tian ZC, Wang JQ, Ge H. Apatinib ameliorates doxorubicin-induced migration and cancer stemness of osteosarcoma cells by inhibiting Sox2 via STAT3 signalling. J Orthop Translat. 2020;22:132–41.
Funding
This work was supported by Scientific Research Projects of the Key scientific research projects of universities in Anhui Province [Grant No. 2022AH051235], Health Commission of Anhui Province [Grant No. AHWJ2021b059], postgraduate scientific research project of Universities in Anhui Province [Grant No. YJS20210552] and National Innovation and Entrepreneurship Training for University of PRC [Grant no. 202210368059].
Author information
Authors and Affiliations
Contributions
QL, YSK and BC had full access to all of the data in the manuscript and take responsibility for the integrity of the data and the accuracy of the data analysis. Concept and design: all authors. Acquisition, analysis, and interpretation of data: all authors. Manuscript drafting: QL and YSK. Critical revision of the manuscript for important intellectual content: all authors. Statistical analysis: XXT and XQZ. Supervision: BC. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Ethics statement
Not applicable.
Competing interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Luo, Q., Kuang, Y., Tao, X. et al. Pan-cancer analysis of PCAT6 and its effect on oesophageal squamous cell carcinoma cell proliferation and migration. Med Oncol 40, 125 (2023). https://doi.org/10.1007/s12032-023-01982-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-023-01982-2