Abstract
Immune signal transduction is crucial to the body’s defense against viral infection. Recognition of pathogen-associated molecular patterns by pattern recognition receptors (PRRs) activates the transcription of interferon regulators and nuclear factor-κB (NF-κB); this promotes the release of interferons and inflammatory factors. Efficient regulation of type I interferon and NF-κB signaling by members of the mitogen-activated protein (MAP) kinase kinase kinase (MAP3K) family plays an important role in antiviral immunity. Elucidating the specific roles of MAP3K activation during viral infection is essential to develop effective antiviral therapies. In this review, we outline the specific regulatory mechanisms of MAP3Ks in antiviral immunity and discuss the feasibility of targeting MAP3Ks for the treatment of virus-induced diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mitogen-activated protein kinases (MAPKs) are serine/threonine protein kinases involved in cell physiology, cytopathology, and various illnesses, including cancer [1]. The three-layer MAPK cascade includes MAP kinase kinase kinases (MAPKKKs or MAP3Ks), MAP kinase kinases (MAPKKs or MAP2Ks), and MAP kinases (MAPKs) [2,3,4,5]. MAP3Ks function as links in signal transmission, providing specificity for stimulus-dependent activation of the MAP2K-MAPK pathway, through distinct protein-protein interactions and phosphorylation of signal effectors [6]. Among the 24 identified MAP3Ks, MAP3K1-21, A-Raf, B-Raf, and C-Raf (Raf-1) belong to the most diverse subfamilies in the MAPK signaling cascade [7, 8]. Different MAP3Ks can phosphorylate the same MAP2K and the same MAP3K can phosphorylate different MAP2Ks (Table 1). MAP3Ks serve as tissue nodes or "hubs" in integrating cellular responses to offer specificity in MAPK activation and functional responses [54].
In the innate immune system, pathogen-associated molecular patterns from viruses are mainly recognized by three PRRs, toll-like receptors (TLRs), retinoic acid-inducible gene I (RIG-I)-like receptors (RLRs), and DNA receptors. These will then trigger an immune response pathway that activates type I interferon (IFN-I) and NF-κB signaling for antiviral immunity [55, 56]. IFN-I signaling is strictly regulated. Failure to trigger IFN-I expression can result in severe inflammation, whereas prolonged IFN-I production can contribute to the development of autoimmune diseases [57, 58]. NF-κB can trigger the expression of inflammatory mediators, including cytokines, chemokines, and cell adhesion molecules, thereby exerting antiviral effects [59].
MAP3Ks play several roles during viral infection and have reciprocal effects on virus survival. After viral infection, MAP3Ks regulate the production of interferons, inflammatory mediators, and other antiviral substances through antiviral pathways, which are essential to the host’s defense against viral invasion. Elucidating the functions of MAP3Ks during viral infection may help improve our understanding of viral illnesses and facilitate the development of efficient antiviral medications and techniques. Here, we provide an overview of the processes and functions of MAP3Ks in terms of IFN-I and NF-κB signaling. Additionally, we investigated the potential therapeutic benefits of targeting MAP3Ks in conditions caused by viruses such as human immunodeficiency virus type 1 (HIV-1) and other viruses. This review summarizes recent advances in the field and enables the translation of the current relevant knowledge into therapeutic discoveries.
Mechanisms underlying the regulation of MAP3Ks activity
Upstream MAP4Ks, oxidative stress, inflammatory cytokines, medications, and pressure are the major activators of MAP3Ks. The unhindered signaling, defects in which are strongly linked to many disorders like cancer and inflammatory diseases, is determined by MAP3Ks activity. MAP3K regulates the level of its own activity through a variety of mechanisms, such as regulation of transcriptional abundance, allosteric regulation, and post-translational modification (Table 1).
The abundance of MAP3Ks is regulated at the transcriptional level, and its expression can be induced in a stimulus-dependent manner. MAP3K8 abundance is lower in cells not stimulated by VEGF-A, and it increases after stimulation. Continuous induction has the opposite effect, resulting in a decrease in MAP3K8 expression [24]. Similarly, the transcriptional levels of MAP3Ks are negatively regulated. The mRNA abundance of RAF1, MAP3K2, and MAP3K7 decreases as the temperature increases in heat-stimulated cells [11].
Some MAP3Ks can regulate kinase activity in another important way: allosteric regulation. A key event in RAF activation is RAF dimerization mediated by scaffold and chaperone 14-3-3. In addition, 14-3-3 binds to the catalytic domain of Apoptosis signal-regulating kinase-1 (ASK1) and inhibits its kinase activity [18].
MAP3Ks are phosphokinases; their post-translational modification, particularly phosphorylation, is critical for their activity regulation. The regulation of all MAP3Ks activities is mediated by phosphorylation. Autophosphorylation on Thr-575 activates MAP3K1 [60]. MAP3K2 and MAP3K3 are activated by lipopolysaccharide (LPS)-induced phosphorylation at MAP3K2 Ser-519 and MAP3K3 Ser-526, respectively [61]. Next to phosphorylation, ubiquitination is the most common post-translational modification of MAP3K. In the case of TNFα, MAP3K7 is polyubiquitinated by a Lys48 linkage at K72 and then degraded through the proteasome pathway [23]. An atypical E3 ligase zinc finger protein 91 primarily mediates the synthesis of the Lys63-linked ubiquitinated stabilizing protein, which allows MAP3K14 to activate itself [39]. In addition, MAP3K7 is acetylated to prevent self-phosphorylation and activation [62].
The several types of mechanisms underlying the regulation of MAP3Ks kinase activity make it possible to elicit suitable responses in various circumstances, including the antiviral innate immune response.
IFN-I signaling pathway
After virus infection, the host recognizes viral nucleic acid via different PRRs and activates different signaling pathways according to the type of nucleic acid [63, 64] (Fig. 1).
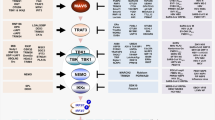
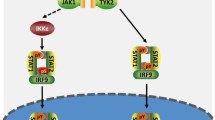
MAP3Ks in the IFN-I pathway. When viral nucleic acids bind to pattern recognition receptors, the corresponding adaptor proteins are recruited to activate three major signaling pathways: RIG-I, cGAS-STING, and TLR. RIG-I binds to MAVS via caspase recruitment domain (CARD)-CARD interaction; cGAS recognizes DNA in the cytosol and is activated to produce cGAMP, which then binds to and activates STING; and TLR3 transmits signals via TRIF. These can activate the downstream protein kinase TBK1 and further phosphorylate IRF3/7. TLR7/8/9 recruit MyD88 and IRAK to form the MyD88 signaling complex, which can activate IRF5/7 and induce downstream IFN-I synthesis and related IFN-stimulating factors. MAP3Ks upregulate or downregulate the signaling are indicated in yellow and red backgrounds, respectively
The RLR family can recognize viral RNA in the cytoplasm. RIG-I recognizes viral double-stranded RNA (dsRNA) and 5′-triphosphate-RNA, while melanoma differentiation-associated gene 5 (MDA5) recognizes long dsRNA. Laboratory of genetics and physiology 2 (LGP2) is mainly involved in the recognition and interaction of RNA with RIG-I and MDA5, but also in signal transduction [65]. Activated RIG-I and MDA5 interact with mitochondrial antiviral-signaling protein (MAVS); when stimulated by MDA5 or RIG-I CARD, MAVS oligomerizes and further recruits multiple tumor necrosis factor (TNF) receptor-associated factor (TRAF) proteins (such as TRAF2, TRAF3, and TRAF6), which are necessary for the activation of TANK-binding kinase 1 (TBK1) and inhibitor of nuclear factor kappa-B kinase ε (IKKε). Ultimately, these kinases phosphorylate interferon regulatory factor 3/7 (IRF3/7), which in turn leads to the production of IFN-I and interferon-stimulated genes (ISGs) [66,67,68,69].
TLRs are the most widely studied PRRs. Humans have ten different TLRs (TLR1–10), most of which can sense RNA viral infections [70]. TLR2 typically functions by forming heterodimers with other TLRs, such as TLR1 and TLR6, to sense virus proteins [71, 72]. In the endosomal compartment, TLR3 recognizes viral dsRNA [73], whereas TLR7/8 recognizes viral single-stranded RNA [74]. All TLRs, except TLR3, activate interleukin-1 (IL-1) receptor-associated kinases (IRAKs) through myeloid differentiation factor 88 (MYD88), which in turn activates TRAF6. TLR3 recruits TIR domain adaptor molecule 1 (TICAM-1, also known as TRIF) [75, 76]. IKK complex, NF-kappa-B essential modulator (NEMO), and TBK1 are activated after a series of signal transductions, and they further activate transcription factors such as IRF3/7 and NF-κB to promote the release of IFN-I and other inflammatory factors.
Viral DNA is primarily recognized by cyclic GMP-AMP (cGAMP) synthase (cGAS) [77]. cGAS then synthesizes cGAMP to deactivate stimulator of interferon genes (STING) on the endoplasmic reticulum, causing it to transfer to the Golgi apparatus, thereby activating the TBK1-IRF3 signaling axis and triggering the production of IFN-I and other inflammatory factors. Absent in melanoma 2 (AIM2) [78], dead-box helicase 41 (DDX41) [79], and TLR9 [64, 80], among others, can also sense viral DNA.
MAP3Ks in IFN-I signaling pathway
Many MAP3Ks affect the function of important IFN-I signaling molecules, thereby regulating IFN-I production (Fig. 1). Here, we focus on how a host can use MAP3Ks to regulate the IFN level for an antiviral immune response. Table 2 summarizes the list of MAP3Ks targeting key molecules in the main innate immune pathways mediated by RNA and DNA viruses.
Raf family
The Raf family is the most studied subfamily of MAP3Ks and includes A-Raf, B-Raf, and C-Raf (Raf-1), which selectively activate MAP2K1 and MAP2K2 [93, 94]. Raf-1 kinase is required for TLR8 and dendritic cell (DC)-specific intercellular adhesion molecule 3-grabbing non-integrin (DC-SIGN)-mediated signaling [95]. In addition, HIV-1 [96] and IFN-β [97] can activate Raf-1. In DCs, Raf-1 limits antiviral IFN-I responses during HIV-1 infection. Raf-1 depletion or pharmacological inhibition of Raf-1 with the small-molecule inhibitor GW5074 could induce the transient expression of IFN-β [81]. Mechanistically, Raf-1 activates mammalian sterile 20-like kinase 1 (MST1), which further phosphorylates polo-like kinase 1 (PLK1) at Thr210 and facilitates its interaction with MAVS, thus blocking MAVS-mediated signaling. In addition, there is an increased viral replication in A-Raf- or Raf-1-deficient cell lines during early vesicular stomatitis virus (VSV) infection [83]. This implies that A-Raf and Raf-1 are also involved in VSV-induced innate immunity.
MAP3K1
MAP3K1 is required for MAPK signal transmission by activating MAP2K1/4/7. MAP3K1 regulates the proliferation of invariant natural killer T cells and promotes the generation of T helper 2 (Th2) cytokines [98]; in addition, it is essential for B cell proliferation and antibody production [99]. Endogenous MAP3K1 is necessary for IFN-I production by cytoplasmic dsRNA [100]. Suppression of MAP3K1 substantially decreases poly(I:C)-induced IFN-β mRNA expression by inhibiting the activation of RIG-I and MAVS. Meanwhile, co-transfection of MAP3K1 and IRF3 synergistically increases IRF3-promoter activity. Real-time polymerase chain reaction analysis demonstrated that MAP3K1 knockdown with various siRNA oligonucleotides considerably lowers IFN-α/β levels. Mechanistically, MAP3K1 interacts with TRAF6, further stimulating the downstream IKK complex and MAP2Ks, thus playing a regulatory role in IFN-I production.
MAP3K2
MAP3K2 primarily transmits signals through MAP2K4/5/7 [101]. MAP3K2-regulated intestinal stromal cells are identified as a new type of intestinal mesenchymal stromal cells that depend on MAP3K2 to regulate intestinal stem cell ecology and protect against intestinal damage [102]. In addition, MAP3K2 is an effector of tumor cell-secreted epidermal growth factor receptor (EGFR) in macrophages, which reduces the host innate antiviral immunity [84]. MAP3K2 phosphorylates IRF3 at Ser173 without involving extracellular-signal-regulated kinase 5 (ERK5) or c-Jun N-terminal kinase (JNK), increases polyubiquitination via the K33-linkage, and inhibits virus-induced IRF3 dimerization and nuclear translation, which further reduces the production of IFN-β and IFN-stimulated genes. In addition, Map3k2-deficient mice are more resistant to viral infection and exhibit reduced viral loads compared to wild-type (WT) mice [84]. Even without the EGFR trigger, MAP3K2 modulates innate antiviral immunity by regulating its activity in different ways. However, further studies are required to better understand the underlying mechanisms.
MAP3K3
In addition to MAP3K2, MAP3K3 can also activate MAP2K5 [103] and regulate vascular malformations [104], kidney diseases [105], and immune activation [106]. Following the cellular activation of the TLR7/9 signaling pathway to induce various types of IFN-Is, MAP3K3 is a potent stimulator of IRF7. Most notably, IRF7, induced by IFN-Is, is a key regulator of virus-driven IFN-I production [107]. MAP3K3 overexpression strongly activates IRF7, resulting in the induction of IFN-Is, whereas TLR7/9 ligand activation results in decreased innate immune responses in cells lacking MAP3K3. Interaction between MAP3K3 and IRF7 induces interferon production, which results in IRF7 phosphorylation at multiple sites [90]. In vivo, MAP3K3 knockdown reduces IFN-I induction and increases the susceptibility to herpes simplex virus type 1 (HSV-1) infection. Endogenous MAP3K3 binds to and phosphorylates IRF7 upon the binding of TLR9 to its specific ligand CpG DNA. These findings implicate the existence of crosstalk between MAP3K activation and IFN-I induction in the endosomal TLR pathway.
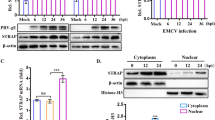
MAP3K5 (ASK1)
MAP3K5 (also known as apoptosis signal-regulating kinase 1; ASK1), MAP3K6 (ASK2), and MAP3K15 (ASK3) are the commonly known members of the ASK family. ASK1, the most studied of the three ASKs, regulates cell survival primarily via the MAP2K3/6-JNK and MAP2K4/7-p38 pathways. ASK2 is extremely similar to ASK1, especially in its kinase domain, and forms a heterologous complex with ASK1, which is necessary for the stability and activity of the ASK2 protein. Therefore, ASK2 functions only as a MAP3K when ASK1 is present [21, 108]. In addition, ASK3 plays a significant role in both osmotic shock and stress [109]. ASK1 not only mediates oxidative stress signaling but also plays a role in innate immunity. ASK1 knockdown in mice enhances the propagation of influenza A virus (IAV), a member of the Orthomyxoviridae family, and reduces IFN-I production in the lungs. Additionally, the increase in IFN-β mRNA abundance in response to encephalomyocarditis virus (EMCV) and newcastle disease virus (NDV) infections was blocked by ASK1 knockdown [87]. This emphasizes the importance of ASK1 as an antiviral protein in vivo. In contrast, ASK2 is required for ASK1-dependent apoptosis but not for induction of IFN-β expression. Furthermore, IAV replication in the lungs is increased in mice with ASK1 or ASK2 gene deletions. Therefore, ASK1/2 is involved in antiviral defense mechanisms; however, ASK2 is a critical regulator of apoptosis rather than IFN-I responses.
Proinflammatory cytokines, including IL-1, IL-6, and TNF-α, are also produced via the RLR pathway. These cytokines regulate the permeability of the endothelium, which lines blood arteries and recruits blood cells and plasma proteins to infection sites. ASK1-/- macrophages infected with the virus display lower levels of IL-6, suggesting the involvement of ASK1 in the induction of these inflammatory cytokines [87].
F-box-only protein 21 (Fbxo21) plays a critical role in regulating the innate antiviral response by facilitating Lys29-linkage and activating ASK1. ASK1 without the Lys29-linkage cannot rescue the innate antiviral response in Map3k5-/- RAW264.7 cells, suggesting that Fbxo21-mediated ubiquitination and ASK1 activation are required for the innate antiviral responses [85].
MAP3K7 (TAK1)
MAP3K7 (also known as transforming growth factor-β-activated kinase 1, TAK1) is a key factor mediating the signal transduction of IL-1 [90], transforming growth factor-beta (TGF-β) [110, 111], and TLRs [112]. IRF3 is the true substrate of JNK, and upstream kinase TAK1 promotes the phosphorylation of IRF3 by JNK [113]. JNK1/2 can directly catalyze the phosphorylation of Ser173 in IRF3. Co-expression of TAK1 significantly increases the activity of this kinase. However, TAK1 cannot directly phosphorylate IRF3.
TAK1 inhibition results in a slight decrease in IFN-β levels after poly(I:C) treatment of WT bone marrow-derived macrophages (BMDMs); however, the difference is not statistically significant [114]. Treatment with TAK1 inhibitors significantly reduces IFN-β induction after poly(I:C) stimulation in IRAK1-/- and IKKε-/- BMDMs compared to that in DMSO-treated IRAK1-/- and IKKε-/- BMDMs. These findings imply that the enzymatic activity of TAK1 is important for regulating IFN-I responses downstream of TLR3 by IRAK1.
MAP3K8 (TPL2)
MAP3K8 (tumor progression locus 2; TPL2) is functionally similar to MAP3K1 in the MAPK cascade signal, but, in most cases, its function does not involve the activation of MAP2K7 [115]. The TPL2-ERK signaling pathway and TLR 2/4/7 directly activate the inflammatory axis and positively regulate ERK phosphorylation and early TNF secretion [116]. In addition, TPL2 positively regulates p38α and p38δ in neutrophils [117]. TPL2 regulates IFN activation through IRF3, which leads to increased replication of foot-and-mouth disease virus (FMDV) [88]. However, IFN-α, myxovirus resistance 2, and CXC motif chemokine ligand 10 expression are significantly reduced in Tpl2-deficient mice, with no detectable change in IRF3 phosphorylation. FMDV capsid protein VP1 inhibits TPL2 phosphorylation at Thr290, a critical functional site of TPL2 that promotes IRF3-mediated activation of the IFN-β signaling pathway. Similar results are obtained when cells are stimulated with poly(I:C) and Sendai virus (SeV) instead of FMDV.
The MAP3K8 (Sluggish) mutation causes splicing errors in the Map3k8-encoded Tpl2 transcript. MAP3K8Sluggish/Sluggish mice are resilient to mouse cytomegalovirus (MCMV) infection but extremely susceptible to group B streptococcus infection [92]. CpG-B-induced IFN-I production is significantly reduced in peritoneal macrophages isolated from homozygous MAP3K8Sluggish/Sluggish mice, whereas LPS- and poly(I:C)-induced IFN-I production remains unaffected. Thus, TPL2 is important for IFN-I production in peritoneal macrophages, especially in response to MyD88-dependent TLR signaling.
MAP3K14 (NIK)
MAP3K14 (also known as NF-κB-inducing kinase, NIK) is a helpful regulator of the DNA virus-activated cGAS-STING pathway [91]. STING and NIK colocalize and interact with each other. NIK relies on its kinase activity and functions in concert with STING to boost IFN-β transcription [91]. NIK dysfunction has serious consequences for pro-immune activation during lymphocytic choriomeningitis virus (LCMV) and vesicular stomatitis virus (VSV) infections. Indeed, viral replication is reduced in the spleen tissue of WT and Map3k14aly/aly mice after infection with either virus strain, and serum IFN levels are considerably reduced [86]. The aly mutation results in an amino acid substitution (G855R) in the Nik code. Aly/aly mice are completely devoid of lymph nodes and Peyer’s patches, thus, they are highly susceptible to viral infections [86, 118]. NIK suppresses hepatitis C virus (HCV) replication in Huh-7 cells [119]. In addition, NIK is a component of the NF-κB-inducing signaling cascade, independent of any specific MAPK cascade [120].
Other MAP3Ks
Following a systematic evaluation of multiple genomic datasets, RNA interference (RNAi) screening tests demonstrated that MAP3K11 deletion drastically lowers RIG-I signaling-mediated IFN production [89]. Increased HIV-1 transcription by MAP3K11 is dependent on its kinase activity, and the corresponding MAP3K11 siRNA reduces HIV-1 infection by 40% [82].
Several MAP3Ks that regulate IFN-I signaling have been identified; however, each host MAP3K has a unique set of functions that vary depending on the virus and cell type. Further research is required to determine the antiviral roles of MAP3Ks in IFN-I signaling.
NF-κB signaling pathway
NF-κB is a major transcriptional regulator in antiviral immunity and is activated after viral infection. In the previous chapters, we mentioned that after viral nucleic acid is recognized by PRR, the NF-κB pathway can be effectively activated, but it is not complete. Here, we introduce in detail two NF-κB pathways that play key roles in cells: the canonical NF-κB pathway and the non-canonical NF-κB pathway. Interestingly, these two pathways are dominated by two respective MAP3Ks (Fig. 2) [121, 122].
MAP3Ks in the NF-κB pathway. Canonical NF-κB pathway: TAK1 phosphorylates and activates the NEMO-IKKα-IKKβ complex, and the activated IKKβ targets IκB for proteasomal degradation, thereby translocating p50-RelA to the nucleus and inducing NF-κB transcription. Non-canonical NF-B pathway: NIK phosphorylates and activates IKKα, and then recruits IKKα to p100, which then phosphorylates and degrades p100 by the proteasome. Finally, p52 bound to RelB is translocated to the nucleus to regulate the transcription of NF-κB. Through phosphorylation of the IKKα subunit, NIK also promotes canonical NF-κB pathway activation
In the canonical NF-κB pathway, MAP3K7, also known as TAK1, occupies a dominant position. After the viral nucleic acid binds to the TLR, TAK1 is activated. TAK1 is an upstream kinase that activates the IKKα, IKKβ, and NEMO complex. The phosphorylation of inhibitor of NF-κB (IκB) by IKK occurs on two serine residues, and proteasomal degradation occurs, resulting in NF-κB dimers (including RelA/p50) being rapidly released. Eventually, the constitutively activated dimer translocates to the nucleus where it induces the transcription of NF-κB target genes [123].
Activation of the non-canonical NF-κB pathway is dependent on another MAP3K: NIK. Activated NIK phosphorylates the IKK complex, which is notable for containing only two IKKα subunits. Upon activation, IKKα directly phosphorylates p100, leading to the processing of p100 into p52 and, finally, nuclear translocation of the RelB-p52 heterodimer [124].
It is noteworthy that both DNA virus and RNA virus can activate NF-κB transcription in two ways. This will be elaborated in the next chapter.
MAP3Ks in NF-κB signaling pathway
TAK1 and NIK are the main kinases mediating the two NF-κB pathways, while other MAP3Ks still play a role in NF-κB signal transduction. Interestingly, many viruses can activate two NF-κB pathways in different ways, which has nothing to do with whether it is a DNA or RNA virus. Here, we not only outline how other MAP3Ks mediate NF-κB signaling (Fig. 2) but also elaborate on how the host regulates NF-κB activity through MAP3Ks after virus invasion, and then exerts antiviral effects (Table 3).
The phosphorylation and subsequent proteolysis of IκB is an important mechanism for NF-κB activation. MAP3K1 transfection causes IκB degradation and activates NF-κB [137]. In addition, MAP3K1 participates in NF-κB signaling by activating IKKα/β [138]. MAP3K3 and MAP3K2 regulate the formation of IκBα:NF-κB and IκBβ:NF-κB complexes [139]; MAP3K3 can also phosphorylate Ser177 and Ser181 sites in IKK2 (IKKβ) [140]. Interestingly, thousand and one-amino acid protein kinase 2 (TAOK2), another MAP3K, inhibits the TAK1-mediated activation of IKKα [141].
TAK1 plays a crucial role in HIV-1 infection; it increases HIV-1 replication by modifying NF-κB signaling. Viral protein R (Vpr), produced by HIV-1, stimulates the association between TAK1 and TAK1-binding protein 3 (TAB3), as well as TAK1 polyubiquitination and phosphorylation. This in turn activates NF-κB and increases long terminal repeat (LTR)-dependent viral gene expression. Activation of HIV-1 LTR promoter by Vpr is reduced when TAK1 expression is knocked down [125]. Transmembrane glycoprotein gp41 (gp41CD) of both HIV and Simian immunodeficiency virus (SIV), which is crucial for viral replication, activates NF-κB via TAK1 [126]. Sterile α motif and histidine-aspartate domain-containing protein 1 (SAMHD1), an HIV-1 restriction factor, prevents TAK1 phosphorylation, thereby inhibiting the transcription of NF-κB and HIV-1 replication [142], in VSV glycoprotein-pseudotyped single-cycle HIV-1. This demonstrates the restrictive role of SAMHD1 in preventing the viral protein activation of NF-κB.
NF-κB is crucial to the immunological response to HSV infection; however, the mechanism underlying the activation of NF-κB by HSV remains unclear. Alternative kinase activities cannot be ruled out; however, inactivation of TAK1, MAP3K1, or NIK mutants decreases NF-κB activation during HSV infection [134]. In addition, phosphorylation of the three MAP3Ks is elevated during HSV infection. Lu et al. [135] screened for the unique short (Us) region of HSV-2, revealing that HSV-2 Us2 activates NF-κB signaling. Specifically, Us2 interacts with TAK1 to activate TAK1 and downstream genes, thereby activating NF-κB. In mice, Us2 also increases the phosphorylation of TAK1, IKKβ, and IκB.
Severe acute respiratory syndrome-associated coronavirus 2 (SARS-CoV-2), the cause of an active pandemic, stimulates NF-κB and causes the release of numerous inflammatory factors, through its nucleocapsid (N) protein. TAK1 and IKK complexes are attracted to and activated following the liquid-liquid phase separation of the N protein, which in turn triggers the transcription of NF-κB [128]. Another study showed that nonstructural protein 6 (NSP6) and open reading frame 7a (ORF7a) of SARS-CoV-2 interacted with TAK1 dependent on their own ubiquitination, and knockout of TAK1 eliminated NSP6 and ORF7a at the activation of NF-κB [129].
Several other viruses can modify the activation state of NF-κB via TAK1. Hepatitis B virus (HBV) mRNA is translated into hepatitis B surface antigen (HBsAg), which specifically binds to TAK1 and TAB2 and prevents TAK1 phosphorylation, thus inhibiting activation of the NF-κB signaling pathway and evading immune system detection [132]. After viral infection, upregulated TAP1 binds to TAK1 and prevents NF-κB signaling. TAK1 boosts NF-κB activation induced by the respiratory syncytial virus (RSV) [130]. Collectively, TAK1 regulates NF-κB signaling following viral infection, suggesting its potential as a viable therapeutic target for associated viral infections.
Viral infection is significantly impacted by NF-κB signaling, which is mediated by NIK. HIV-1 Tat protein promotes NF-κB activation and accelerates IκB degradation through NIK [127]. RSV infection enhances NIK expression and promotes interactions among NIK, IKK, and p52, which strengthens NF-κB activation [131]. The reduction in nuclear accumulation of p52 and blocking of RSV processing of p100 by NIK knockdown indicates that NIK is crucial for RSV-induced NF-κB activation. In addition, nuclear localization of NIK is essential for HBV-induced NF-κB activation [133]. Nuclear localization signal (NLS)-containing NIK is imported into the nucleus after IFN-γ treatment; however, NLS-free NIK remains in the cytoplasm, suggesting that IFN-γ suppresses NF-κB activation by promoting NIK accumulation in the nucleus. Latent infectious membrane protein 1 (LMP1) of the Epstein-Barr virus (EBV) activates NF-κB through its C-terminal activation domains 1 and 2 (CTAR1 and CTAR2). First, CTAR1 specifically recruits TRAF3 and NF-κB via NIK- and IKKα-induced p100 processing. Subsequently, CTAR2 preferentially recruits TRAF6, which activates the IKKα/β/γ complex after activating TAK1, and phosphorylates and degrades IκBα, eventually leading to NF-κB activation [136].
Therefore, NF-κB signaling is critically dependent on the regulatory mechanisms of MAP3Ks during the viral infection cycle. After infecting host cells, viruses manipulate MAP3K activity in various ways, thereby altering NF-κB signaling and affecting antiviral immunity. Exploring new therapeutic strategies targeting MAP3K to inhibit viral replication and disease pathogenesis requires elucidating virus-MAP3K-NF-κB interactions.
MAP3K family: a potential target for antiviral therapies
MAP3Ks provide prospective therapeutic targets for the widespread diagnosis and treatment of viral infections owing to recent developments in our understanding of the function of MAP3Ks in controlling antiviral immune signals. According to this review, it is not difficult to find that MAP3Ks play a key role in HIV-1 virus infection. In addition, MAP3Ks-related inhibitors were shown to have the perfect mechanism for the treatment of HIV-1-related diseases. Therefore, targeting MAP3Ks may lead to the development of effective drugs against HIV-1 infection and related diseases, which we summarized in detail. Moreover, MAP3Ks have also been targeted for the treatment of other viruses with good results.
MAP3Ks as a therapeutic strategy for HIV-1-related diseases
The HIV-1 epidemic is a public health crisis in countries and regions worldwide, even though antiretroviral therapy has greatly reduced the infection rate of HIV-1 and prolonged the lives of HIV-1 patients [143]. Some patients still show HIV-1-associated neurocognitive disorder (HAND), which greatly affects the quality of their lives. Studies have pointed out that HIV-1 proteins, gp120 and Tat, activate Mixed lineage kinase 3 (MLK3) in neurons, leading to the death of neurons and the production of inflammatory factors with neurotoxic effects. Furthermore, either kinase inactivation or the pharmacological inhibition of MLK3 activity protects neuronal survival, suggesting that MLK3 activity may determine the development of HAND [144, 145]. Bodner et al. [146] first discovered that the first-generation MLK inhibitor CEP-1347 dose-dependently protects hippocampal neurons from HIV-1 gp120-induced neurotoxicity. However, Bodner et al. studied only gp120, a neurotoxin, on a single hippocampal neuron; Sui et al. [144] further found that HIV-1 gp120 and Tat can also cause microglial apoptosis by activating MLK3. Interestingly, CEP-1347 treatment also prevented the activation of gp120 and Tat in monocytes. In 2010, Eggert et al. [147] further verified the neuronal protective effect of CEP-1347 in models of NeuroAIDS. These results all suggest that inhibiting MLK3 is a potential strategy for the treatment of HAND. Regrettably, due to the structure of CEP-1347, it is not conducive to the penetration of the blood-brain barrier, which led to its failure to show efficacy during Phase II clinical trials for the treatment of Parkinson’s disease. In addition, the poor pharmacokinetic and metabolic characteristics have become evidence that CEP-1347 has no efficacy [148]. A novel MLK3 inhibitor, URMC-099, has been shown to protect microglia from phagocytosis, lowers inflammation induced by HIV-1 Tat protein, and reduces HAND-associated symptoms [149] (Fig. 3A). In addition, it has also been shown that URMC-099 reverses HIV-1 Nef protein-mediated sequestration of transcription factor EB in the cytoplasm, induces autophagy and lysosomal biogenesis, and encourages the accumulation of co-administered, nano-formulated antiretroviral medication in persistently infected macrophages [145] (Fig. 3B).
Schematic representation of a working model of drugs against HIV-1-related diseases. A Following HIV-1 infection, microglia activate and retract their branches. The action of HIV-1 viral proteins eventually leads to synaptic dendrite damage and cell death. B Following HIV-1 infection, HIV-1 Nef protein sequesters transcription factor EB (TFEB) in the cytoplasm and inhibits lysosomal biogenesis and autophagy, thus preventing nanoART from accumulating in the cells and eventually leading to viral replication. Both these phenomena are suppressed by the administration of URMC-099. C Treatment with an ASK1 inhibitor recovers the fused podocyte foot process and elevated inflammatory cytokines levels, in the HIVAN mice model. MTORC1, the mechanistic target of rapamycin complex 1; LC3, light chain 3
Chronic kidney disease is a common condition among patients with HIV-1, and HIV-1-associated nephropathy (HIVAN) rapidly leads to kidney failure. The ASK1 inhibitor, GS-444217 significantly inhibits ASK1 and downstream MAPK activation. Treatment with GS-444217 reduces HIVAN-associated damage and improves renal function. Therefore, targeting ASK1 is promising for the development of novel treatments against HICAN (Fig. 3C) [150].
MAP3Ks as a therapeutic strategy for other virus-related diseases
The coronavirus disease 2019 epidemic is a global health challenge, and it is a long and complicated process to re-develop drugs to market. Drug repurposing was mentioned as an effective solution against SARS-CoV-2 (Fig. 4) [151].
Schematic representation of two working models of anti-SARS-CoV-2 drugs. A Inhibiting the virus–host combination prevents viral invasion and achieves an antiviral effect. B Inhibiting relevant targets of the transduction pathway of SARS-CoV-2 prevents viral invasion and achieves an antiviral effect. SBD, substrate-binding domain; NBD, nucleotide-binding domain; csBIP, cell surface binding immunoglobin protein
Vemurafenib and PLX-4720 are inhibitors of the B-Raf mutation V600 [152, 153]. Vemurafenib interacts with the nucleotide-binding domain of the cell surface protein BiP, thus limiting viral replication by neutralizing viral binding [154]. In contrast, in another study, vemurafenib caused a time-dependent increase in the expression of angiotensin-converting enzyme 2 (ACE2), which may contribute to SARS-CoV-2 infection [155]. PLX-4720 interacts with receptor-interacting serine/threonine-protein kinase 1 (RIPK1) during SARS-CoV-2 infection [153]. RIPK1 is activated by nsp12 and regulates the expression of ACE2 to promote viral transmission. SARS-CoV-2 enters the human body via the ACE2 receptor, and its high expression increases the body’s susceptibility to SARS-CoV-2. Sorafenib and another multikinase inhibitor, regorafenib, also exert antiviral efficacy [156, 157], and anti-SARS-CoV-2 activity. Sorafenib can decrease the activity of the nidovirus RdRp-associated nucleotidyltransferase domain (NiRAN) of SARS-CoV-2 RdRp by interacting with the aspartate residue of the anticipated active site [158, 159]. Regorafenib represents a potential drug candidate for blocking the interaction between the viral receptor-binding domain S1 and ACE2 [160]. Another Raf inhibitor, Dabrafenib, was found to inhibit SARS-CoV-2 infection in vitro [161, 162]. Collectively, these drugs may be rapidly applied to the clinical treatment of SARS-CoV-2, but their specific mechanisms are still unknown.
EBV expresses latent membrane protein 1 (LMP1), and LMP1-transgenic mice exhibit hyperproliferation of oncogenic cells, inflammation, or malignancies. Effective treatment of EBV-related tumors requires an understanding of LMP1 network signaling [163, 164]. LMP1 activates TPL2 via IKK2 and causes TPL2 phosphorylation, thus activating JNK. When treated with the TPL2 inhibitor, TC-S7006, the capacity of LMP1 to activate JNK is lost, leading to the death of LMP1-dependent cancer cells [164]. Therefore, TPL2 is a potential new target for EBV-induced cancers.
Acute respiratory distress syndrome (ARDS) and pneumonia may be caused by H5N1 viral infection, which is a highly pathogenic avian influenza A virus, that disrupts the alveolar epithelial barrier [165]. Specifically, the H5N1 virus stimulates the expression of the E3 ubiquitin ligase Itch by activating TAK1 and downstream MAPKs. TAK1 knockdown or the use of TAK1 inhibitors restores the expression levels of occludin alveolar junction proteins, which were ubiquitinated and degraded [166]. Thus, TAK1 suppression could provide a target for the development of anti-H5N1 viral infection measures [166].
MAP3Ks offer a therapeutic window for antiviral treatment and MAP3K regulation is a possible antiviral therapeutic approach (Table 4). However, there are some issues to be addressed. RAF inhibitors have the potential to suppress SARS-CoV-2 infection; however, RAF-MAPK is a crucial component of the body’s perception of external pressure stimuli. TPL2 and TAK1 are essential for NF-κB signaling; however, their inhibition could compromise the immune system and facilitate additional infections. Therefore, the negative effects of MAP3K inhibition require careful evaluation.
Discussion and perspectives
MAPKs, not just MAP3Ks, play an important role in regulating IFN-I production. MAP4K1 inhibits IFN-I production by targeting TBK1/IKKε [175], while p38 MAPK reportedly inhibits STING activation by increasing phosphorylation of USP21 at Ser538, thus further controlling the IFN-I pathway [176]. JNK activation is crucial for the proper functioning of the IFN-I pathway [113, 177]. These findings underscore the importance of MAPKs in IFN-I production and could lead to the discovery of new antiviral targets.
MAP3Ks inhibitors are effective against HIV-1 infection and related diseases, and numerous MAP3Ks play vital roles against SARS-CoV-2 and other viruses. An in-depth understanding of the functions of MAP3Ks in the innate immune response is critical to prevent and treat virus-related illnesses. Targeting the MAP3K family of proteins is a promising therapeutic strategy for treating viral infections and associated diseases. Future research should focus on determining the antiviral mechanisms of existing drugs and translate new research findings into antiviral drug development strategies. We believe that elucidating MAP3Ks-virus interactions will resolve the current challenges and improve human health; this is supported by the comprehensive list references reviewed here.
References
Gerits N, Kostenko S, Moens U. In vivo functions of mitogen-activated protein kinases: conclusions from knock-in and knock-out mice. Transgenic Res. 2007;16:281–314. https://doi.org/10.1007/s11248-006-9052-0.
Burotto M, Chiou VL, Lee JM, Kohn EC. The MAPK pathway across different malignancies: a new perspective. Cancer. 2014;120:3446–56. https://doi.org/10.1002/cncr.28864.
Du W, Hu H, Zhang J, Bao G, Chen R, Quan R. The mechanism of MAPK signal transduction pathway involved with electroacupuncture treatment for different diseases. Evid Based Complement Alternat Med. 2019;2019:8138017. https://doi.org/10.1155/2019/8138017.
Liang YJ, Yang WX. Kinesins in MAPK cascade: How kinesin motors are involved in the MAPK pathway? Gene. 2019;684:1–9. https://doi.org/10.1016/j.gene.2018.10.042.
Wang XL, Yuan K, Zhang W, Li SX, Gao GF, Lu L. Regulation of circadian genes by the MAPK pathway: implications for rapid antidepressant action. Neurosci Bull. 2020;36:66–76. https://doi.org/10.1007/s12264-019-00358-9.
Winter-Vann AM, Johnson GL. Integrated activation of MAP3Ks balances cell fate in response to stress. J Cell Biochem. 2007;102:848–58. https://doi.org/10.1002/jcb.21522.
Cuarental L, Sucunza-Saenz D, Valino-Rivas L, Fernandez-Fernandez B, Sanz AB, Ortiz A, et al. MAP3K kinases and kidney injury. Nefrologia. 2019;39:568–80. https://doi.org/10.1016/j.nefro.2019.03.004.
Cargnello M, Roux PP. Activation and function of the MAPKs and their substrates, the MAPK-activated protein kinases. Microbiol Mol Biol Rev. 2011;75:50-83. https://doi.org/10.1128/MMBR.00031-10.
Salem SM, Hamed AR, Mosaad RM. MTDH and MAP3K1 are direct targets of apoptosis-regulating miRNAs in colorectal carcinoma. Biomed Pharmacother. 2017;94:767–73. https://doi.org/10.1016/j.biopha.2017.07.153.
Chadee DN, Yuasa T, Kyriakis JM. Direct activation of mitogen-activated protein kinase kinase kinase MEKK1 by the Ste20p homologue GCK and the adapter protein TRAF2. Mol Cell Biol. 2002;22:737–49. https://doi.org/10.1128/MCB.22.3.737-749.2002.
Enomoto A, Fukasawa T, Terunuma H, Nakagawa K, Yoshizaki A, Sato S, et al. Decrease in MAP3Ks expression enhances the cell death caused by hyperthermia. Int J Hyperthermia. 2022;39:200–8. https://doi.org/10.1080/02656736.2021.2024281.
Li H, Wang N, Jiang Y, Wang H, Xin Z, An H, et al. E3 ubiquitin ligase NEDD4L negatively regulates inflammation by promoting ubiquitination of MEKK2. EMBO Rep. 2022;23:e54603. https://doi.org/10.15252/embr.202254603.
Zheng Y, Liu H, Kong Y. miR-188 promotes senescence of lineage-negative bone marrow cells by targeting MAP3K3 expression. FEBS Lett. 2017;591:2290–8. https://doi.org/10.1002/1873-3468.12720.
Adams DG, Sachs NA, Vaillancourt RR. Phosphorylation of the stress-activated protein kinase, MEKK3, at serine 166. Arch Biochem Biophys. 2002;407:103–16. https://doi.org/10.1016/s0003-9861(02)00464-2.
Yang LX, Gao Q, Shi JY, Wang ZC, Zhang Y, Gao PT, et al. Mitogen-activated protein kinase kinase kinase 4 deficiency in intrahepatic cholangiocarcinoma leads to invasive growth and epithelial-mesenchymal transition. Hepatology. 2015;62:1804–16. https://doi.org/10.1002/hep.28149.
Halfter UM, Derbyshire ZE, Vaillancourt RR. Interferon-gamma-dependent tyrosine phosphorylation of MEKK4 via Pyk2 is regulated by annexin II and SHP2 in keratinocytes. Biochem J. 2005;388:17–28. https://doi.org/10.1042/BJ20041236.
Yi S, Chen K, Zhang L, Shi W, Zhang Y, Niu S, et al. Endoplasmic reticulum stress is involved in stress-induced hypothalamic neuronal injury in rats via the PERK-ATF4-CHOP and IRE1-ASK1-JNK pathways. Front Cell Neurosci. 2019;13:190. https://doi.org/10.3389/fncel.2019.00190.
Obsilova V, Obsil T. The 14-3-3 proteins as important allosteric regulators of protein kinases. Int J Mol Sci. 2020;21 https://doi.org/10.3390/ijms21228824.
Zhang P, Wang PX, Zhao LP, Zhang X, Ji YX, Zhang XJ, et al. The deubiquitinating enzyme TNFAIP3 mediates inactivation of hepatic ASK1 and ameliorates nonalcoholic steatohepatitis. Nat Med. 2018;24:84–94. https://doi.org/10.1038/nm.4453.
Thomson EM, Williams A, Yauk CL, Vincent R. Impact of nose-only exposure system on pulmonary gene expression. Inhal Toxicol. 2009;21(Suppl 1):74–82. https://doi.org/10.1080/08958370902962309.
Takeda K, Shimozono R, Noguchi T, Umeda T, Morimoto Y, Naguro I, et al. Apoptosis signal-regulating kinase (ASK) 2 functions as a mitogen-activated protein kinase kinase kinase in a heteromeric complex with ASK1. J Biol Chem. 2007;282:7522–31. https://doi.org/10.1074/jbc.M607177200.
Ray Chaudhuri N, Ghosh DS. Allosteric boost by TAB1 on the TAK1 kinase favorably sculpts the thermodynamic landscape of activation. J Chem Inf Model. 2023;63:224–39. https://doi.org/10.1021/acs.jcim.2c00778.
Fan Y, Shi Y, Liu S, Mao R, An L, Zhao Y, et al. Lys48-linked TAK1 polyubiquitination at lysine-72 downregulates TNFalpha-induced NF-kappaB activation via mediating TAK1 degradation. Cell Signal. 2012;24:1381–9. https://doi.org/10.1016/j.cellsig.2012.02.017.
Fearnley GW, Abdul-Zani I, Latham AM, Hollstein MC, Ladbury JE, Wheatcroft SB, et al. Tpl2 is required for VEGF-A-stimulated signal transduction and endothelial cell function. Biol Open. 2019;8 https://doi.org/10.1242/bio.034215.
Gutmann S, Hinniger A, Fendrich G, Druckes P, Antz S, Mattes H, et al. The crystal structure of cancer osaka thyroid kinase reveals an unexpected kinase domain fold. J Biol Chem. 2015;290:15210–8. https://doi.org/10.1074/jbc.M115.648097.
Cho J, Tsichlis PN. Phosphorylation at Thr-290 regulates Tpl2 binding to NF-kappaB1/p105 and Tpl2 activation and degradation by lipopolysaccharide. Proc Natl Acad Sci U S A. 2005;102:2350–5. https://doi.org/10.1073/pnas.0409856102.
DeAizpurua HJ, Cram DS, Naselli G, Devereux L, Dorow DS. Expression of mixed lineage kinase-1 in pancreatic beta-cell lines at different stages of maturation and during embryonic pancreas development. J Biol Chem. 1997;272:16364–73. https://doi.org/10.1074/jbc.272.26.16364.
Kokoszka ME, Kall SL, Khosla S, McGinnis JE, Lavie A, Kay BK. Identification of two distinct peptide-binding pockets in the SH3 domain of human mixed-lineage kinase 3. J Biol Chem. 2018;293:13553–65. https://doi.org/10.1074/jbc.RA117.000262.
Durkin JT, Holskin BP, Kopec KK, Reed MS, Spais CM, Steffy BM, et al. Phosphoregulation of mixed-lineage kinase 1 activity by multiple phosphorylation in the activation loop. Biochemistry. 2004;43:16348–55. https://doi.org/10.1021/bi049866y.
Phelan DR, Loveland KL, Devereux L, Dorow DS. Expression of mixed lineage kinase 2 in germ cells of the testis. Mol Reprod Dev. 1999;52:135–40. https://doi.org/10.1002/(SICI)1098-2795(199902)52:2<135::AID-MRD3>3.0.CO;2-N.
Phelan DR, Price G, Liu YF, Dorow DS. Activated JNK phosphorylates the c-terminal domain of MLK2 that is required for MLK2-induced apoptosis. J Biol Chem. 2001;276:10801–10. https://doi.org/10.1074/jbc.M008237200.
Byrnes KA, Phatak P, Mansour D, Xiao L, Zou T, Rao JN, et al. Overexpression of miR-199a-5p decreases esophageal cancer cell proliferation through repression of mitogen-activated protein kinase kinase kinase-11 (MAP3K11). Oncotarget. 2016;7:8756–70. https://doi.org/10.18632/oncotarget.6752.
Yu J, Feng Y, Wang Y, An R. Aryl hydrocarbon receptor enhances the expression of miR-150-5p to suppress in prostate cancer progression by regulating MAP3K12. Arch Biochem Biophys. 2018;654:47–54. https://doi.org/10.1016/j.abb.2018.07.010.
Huntwork-Rodriguez S, Wang B, Watkins T, Ghosh AS, Pozniak CD, Bustos D, et al. JNK-mediated phosphorylation of DLK suppresses its ubiquitination to promote neuronal apoptosis. J Cell Biol. 2013;202:747–63. https://doi.org/10.1083/jcb.201303066.
Li Z, Liu Y, Hou Y, Li Z, Chen C, Hao H, et al. Construction and function analysis of the LncRNA-miRNA-mRNA competing endogenous RNA network in autoimmune hepatitis. BMC Med Genomics. 2022;15:270. https://doi.org/10.1186/s12920-022-01416-4.
Sakuma H, Ikeda A, Oka S, Kozutsumi Y, Zanetta JP, Kawasaki T. Molecular cloning and functional expression of a cDNA encoding a new member of mixed lineage protein kinase from human brain. J Biol Chem. 1997;272:28622–9. https://doi.org/10.1074/jbc.272.45.28622.
Fry CS, Nayeem SZ, Dillon EL, Sarkar PS, Tumurbaatar B, Urban RJ, et al. Glucocorticoids increase skeletal muscle NF-kappaB inducing kinase (NIK): links to muscle atrophy. Physiol Rep. 2016;4 https://doi.org/10.14814/phy2.13014.
Shen C, Liu H, Wang X, Lei T, Wang E, Xu L, et al. Importance of incorporating protein flexibility in molecule modeling: a theoretical study on type I(1/2) NIK inhibitors. Front Pharmacol. 2019;10:345. https://doi.org/10.3389/fphar.2019.00345.
Jin X, Jin HR, Jung HS, Lee SJ, Lee JH, Lee JJ. An atypical E3 ligase zinc finger protein 91 stabilizes and activates NF-kappaB-inducing kinase via Lys63-linked ubiquitination. J Biol Chem. 2010;285:30539–47. https://doi.org/10.1074/jbc.M110.129551.
Chen FC, Brozovich FV. Gene expression profiles of vascular smooth muscle show differential expression of mitogen-activated protein kinase pathways during captopril therapy of heart failure. J Vasc Res. 2008;45:445–54. https://doi.org/10.1159/000126735.
Ultanir SK, Yadav S, Hertz NT, Oses-Prieto JA, Claxton S, Burlingame AL, et al. MST3 kinase phosphorylates TAO1/2 to enable Myosin Va function in promoting spine synapse development. Neuron. 2014;84:968–82. https://doi.org/10.1016/j.neuron.2014.10.025.
Yang Q, Li Y, Wang Y, Qiao X, Liu T, Wang H, et al. The circRNA circSIAE inhibits replication of coxsackie virus B3 by targeting miR-331-3p and thousand and one amino-acid kinase 2. Front Cell Infect Microbiol. 2021;11:779919. https://doi.org/10.3389/fcimb.2021.779919.
Xia Y, Caputo M, Cansby E, Anand SK, Sutt S, Henricsson M, et al. STE20-type kinase TAOK3 regulates hepatic lipid partitioning. Mol Metab. 2021;54:101353. https://doi.org/10.1016/j.molmet.2021.101353.
Zach S, Felk S, Gillardon F. Signal transduction protein array analysis links LRRK2 to Ste20 kinases and PKC zeta that modulate neuronal plasticity. PLoS One. 2010;5:e13191. https://doi.org/10.1371/journal.pone.0013191.
Boehme SA, Franz-Bacon K, Ludka J, DiTirro DN, Ly TW, Bacon KB. MAP3K19 is overexpressed in COPD and is a central mediator of cigarette smoke-induced pulmonary inflammation and lower airway destruction. PLoS One. 2016;11:e0167169. https://doi.org/10.1371/journal.pone.0167169.
Xu WH, Zhang JB, Dang Z, Li X, Zhou T, Liu J, et al. Long non-coding RNA URHC regulates cell proliferation and apoptosis via ZAK through the ERK/MAPK signaling pathway in hepatocellular carcinoma. Int J Biol Sci. 2014;10:664–76. https://doi.org/10.7150/ijbs.8232.
Yang J, Shibu MA, Kong L, Luo J, BadrealamKhan F, Huang Y, et al. Design, synthesis, and structure-activity relationships of 1,2,3-triazole benzenesulfonamides as new selective leucine-zipper and sterile-alpha motif kinase (ZAK) inhibitors. J Med Chem. 2020;63:2114–30. https://doi.org/10.1021/acs.jmedchem.9b00664.
Gotoh I, Adachi M, Nishida E. Identification and characterization of a novel MAP kinase kinase kinase. MLTK. J Biol Chem. 2001;276:4276–86. https://doi.org/10.1074/jbc.M008595200.
Li Y, Zuo H, Wang H, Hu A. Decrease of MLK4 prevents hepatocellular carcinoma (HCC) through reducing metastasis and inducing apoptosis regulated by ROS/MAPKs signaling. Biomed Pharmacother. 2019;116:108749. https://doi.org/10.1016/j.biopha.2019.108749.
Silverman JA, Zurlo J, Watson MA, Yager JD. Expression of c-raf-1 and A-raf-1 during regeneration of rat liver following surgical partial hepatectomy. Mol Carcinog. 1989;2:63–7. https://doi.org/10.1002/mc.2940020203.
Marais R, Light Y, Paterson HF, Mason CS, Marshall CJ. Differential regulation of Raf-1, A-Raf, and B-Raf by oncogenic ras and tyrosine kinases. J Biol Chem. 1997;272:4378–83. https://doi.org/10.1074/jbc.272.7.4378.
Rothman JH, Surriga O, de Stanchina E, Vasudeva SD, Schwartz GK. Obstruction of BRAF(V600E) transcription by complementary PNA oligomers as a means to inhibit BRAF-mutant melanoma growth. Cancer Gene Ther. 2017;24:401–8. https://doi.org/10.1038/cgt.2017.34.
Feng D, Sheng-Dong L, Tong W, Zhen-Xian D. O-GlcNAcylation of RAF1 increases its stabilization and induces the renal fibrosis. Biochim Biophys Acta Mol Basis Dis. 2020;1866:165556. https://doi.org/10.1016/j.bbadis.2019.165556.
Craig EA, Stevens MV, Vaillancourt RR, Camenisch TD. MAP3Ks as central regulators of cell fate during development. Dev Dyn: an official publication of the American Association of Anatomists. 2008;237:3102–14. https://doi.org/10.1002/dvdy.21750.
Wu J, Chen ZJ. Innate immune sensing and signaling of cytosolic nucleic acids. Annu Rev Immunol. 2014;32:461–88. https://doi.org/10.1146/annurev-immunol-032713-120156.
Akira S, Takeda K. Toll-like receptor signalling. Nat Rev Immunol. 2004;4:499–511. https://doi.org/10.1038/nri1391.
Psarras A, Emery P, Vital EM. Type I interferon-mediated autoimmune diseases: pathogenesis, diagnosis and targeted therapy. Rheumatology (Oxford). 2017;56:1662–75. https://doi.org/10.1093/rheumatology/kew431.
Sundaresan B, Shirafkan F, Ripperger K, Rattay K. The role of viral infections in the onset of autoimmune diseases. Viruses. 2023;15 https://doi.org/10.3390/v15030782.
Lawrence T. The nuclear factor NF-kappaB pathway in inflammation. Cold Spring Harb Perspect Biol. 2009;1:a001651. https://doi.org/10.1101/cshperspect.a001651.
Deak JC, Templeton DJ. Regulation of the activity of MEK kinase 1 (MEKK1) by autophosphorylation within the kinase activation domain. Biochem J. 1997;322(Pt 1):185–92. https://doi.org/10.1042/bj3220185.
Zhang D, Facchinetti V, Wang X, Huang Q, Qin J, Su B. Identification of MEKK2/3 serine phosphorylation site targeted by the Toll-like receptor and stress pathways. EMBO J. 2006;25:97–107. https://doi.org/10.1038/sj.emboj.7600913.
Paquette N, Conlon J, Sweet C, Rus F, Wilson L, Pereira A, et al. Serine/threonine acetylation of TGFbeta-activated kinase (TAK1) by Yersinia pestis YopJ inhibits innate immune signaling. Proc Natl Acad Sci U S A. 2012;109:12710–5. https://doi.org/10.1073/pnas.1008203109.
Tong J, Zhang W, Chen Y, Yuan Q, Qin NN, Qu G. The emerging role of RNA modifications in the regulation of antiviral innate immunity. Front Microbiol. 2022;13:845625. https://doi.org/10.3389/fmicb.2022.845625.
Briard B, Place DE, Kanneganti TD. DNA sensing in the innate immune response. Physiology. 2020;35:112–24. https://doi.org/10.1152/physiol.00022.2019.
Jiang QX. Structural variability in the RLR-MAVS pathway and sensitive detection of viral RNAs. Med Chem. 2019;15:443–58. https://doi.org/10.2174/1573406415666181219101613.
Wu B, Hur S. How RIG-I like receptors activate MAVS. Curr Opin Virol. 2015;12:91–8. https://doi.org/10.1016/j.coviro.2015.04.004.
Eisenacher K, Krug A. Regulation of RLR-mediated innate immune signaling--it is all about keeping the balance. Eur J Cell Biol. 2012;91:36–47. https://doi.org/10.1016/j.ejcb.2011.01.011.
Bruns AM, Horvath CM. LGP2 synergy with MDA5 in RLR-mediated RNA recognition and antiviral signaling. Cytokine. 2015;74:198–206. https://doi.org/10.1016/j.cyto.2015.02.010.
Bruns AM, Horvath CM. Antiviral RNA recognition and assembly by RLR family innate immune sensors. Cytokine Growth Factor Rev. 2014;25:507–12. https://doi.org/10.1016/j.cytogfr.2014.07.006.
Wang S, Zhou L, Ling L, Meng X, Chu F, Zhang S, et al. The crosstalk between hippo-YAP pathway and innate immunity. Front Immunol. 2020;11:323. https://doi.org/10.3389/fimmu.2020.00323.
Zong Z, Zhang Z, Wu L, Zhang L, Zhou F. The functional deubiquitinating enzymes in control of innate antiviral immunity. Adv Sci (Weinh). 2021;8:2002484. https://doi.org/10.1002/advs.202002484.
Takeda K, Akira S. TLR signaling pathways. Semin Immunol. 2004;16:3–9. https://doi.org/10.1016/j.smim.2003.10.003.
Le Naour J, Galluzzi L, Zitvogel L, Kroemer G, Vacchelli E. Trial watch: TLR3 agonists in cancer therapy. Oncoimmunology. 2020;9:1771143. https://doi.org/10.1080/2162402X.2020.1771143.
Urcuqui-Inchima S, Cabrera J, Haenni AL. Interplay between dengue virus and Toll-like receptors, RIG-I/MDA5 and microRNAs: Implications for pathogenesis. Antiviral Res. 2017;147:47–57. https://doi.org/10.1016/j.antiviral.2017.09.017.
Luo L, Lucas RM, Liu L, Stow JL. Signalling, sorting and scaffolding adaptors for Toll-like receptors. J Cell Sci. 2019;133 https://doi.org/10.1242/jcs.239194.
Chen L, Zheng L, Chen P, Liang G. Myeloid differentiation primary response protein 88 (MyD88): the central hub of TLR/IL-1R signaling. J Med Chem. 2020;63:13316–29. https://doi.org/10.1021/acs.jmedchem.0c00884.
Hopfner KP, Hornung V. Molecular mechanisms and cellular functions of cGAS-STING signalling. Nat Rev Mol Cell Biol. 2020;21:501–21. https://doi.org/10.1038/s41580-020-0244-x.
Lugrin J, Martinon F. The AIM2 inflammasome: sensor of pathogens and cellular perturbations. Immunol Rev. 2018;281:99–114. https://doi.org/10.1111/imr.12618.
Jiang Y, Zhu Y, Liu ZJ, Ouyang S. The emerging roles of the DDX41 protein in immunity and diseases. Protein Cell. 2017;8:83–9. https://doi.org/10.1007/s13238-016-0303-4.
Medzhitov R. Toll-like receptors and innate immunity. Nat Rev Immunol. 2001;1:135–45. https://doi.org/10.1038/35100529.
Gringhuis SI, Hertoghs N, Kaptein TM, Zijlstra-Willems EM, Sarrami-Forooshani R, Sprokholt JK, et al. HIV-1 blocks the signaling adaptor MAVS to evade antiviral host defense after sensing of abortive HIV-1 RNA by the host helicase DDX3. Nat Immunol. 2017;18:225–35. https://doi.org/10.1038/ni.3647.
Nguyen DG, Yin H, Zhou Y, Wolff KC, Kuhen KL, Caldwell JS. Identification of novel therapeutic targets for HIV infection through functional genomic cDNA screening. Virology. 2007;362:16–25. https://doi.org/10.1016/j.virol.2006.11.036.
Kandasamy RK, Vladimer GI, Snijder B, Muller AC, Rebsamen M, Bigenzahn JW, et al. A time-resolved molecular map of the macrophage response to VSV infection. NPJ Syst Biol Appl. 2016;2:16027. https://doi.org/10.1038/npjsba.2016.27.
Gao L, Wang L, Dai T, Jin K, Zhang Z, Wang S, et al. Tumor-derived exosomes antagonize innate antiviral immunity. Nat Immunol. 2018;19:233–45. https://doi.org/10.1038/s41590-017-0043-5.
Yu Z, Chen T, Li X, Yang M, Tang S, Zhu X, et al. Lys29-linkage of ASK1 by Skp1-Cullin 1-Fbxo21 ubiquitin ligase complex is required for antiviral innate response. Elife. 2016;5 https://doi.org/10.7554/eLife.14087.
Hamdan TA, Bhat H, Cham LB, Adomati T, Lang J, Li F, et al. Map3k14 as a regulator of innate and adaptive immune response during acute viral infection. Pathogens. 2020;9 https://doi.org/10.3390/pathogens9020096.
Okazaki T, Higuchi M, Takeda K, Iwatsuki-Horimoto K, Kiso M, Miyagishi M, et al. The ASK family kinases differentially mediate induction of type I interferon and apoptosis during the antiviral response. Sci Signal. 2015;8:ra78. https://doi.org/10.1126/scisignal.aab1883.
Hao J, Shen C, Wei N, Yan M, Zhang X, Xu G, et al. Foot-and-mouth disease virus capsid protein VP1 antagonizes TPL2-mediated activation of the IRF3/IFN-beta signaling pathway to facilitate the virus replication. Front Immunol. 2020;11:580334. https://doi.org/10.3389/fimmu.2020.580334.
van der Lee R, Feng Q, Langereis MA, Ter Horst R, Szklarczyk R, Netea MG, et al. Integrative genomics-based discovery of novel regulators of the innate antiviral response. PLoS Comput Biol. 2015;11:e1004553. https://doi.org/10.1371/journal.pcbi.1004553.
Cai M, Huang W, Hu X, Chen A, Zhou X. MEKK3 activates IRF7 to trigger a potent type I interferon induction in response to TLR7/9 signaling. Mol Immunol. 2021;134:183–91. https://doi.org/10.1016/j.molimm.2021.03.008.
Parvatiyar K, Pindado J, Dev A, Aliyari SR, Zaver SA, Gerami H, et al. A TRAF3-NIK module differentially regulates DNA vs RNA pathways in innate immune signaling. Nat Commun. 2018;9:2770. https://doi.org/10.1038/s41467-018-05168-7.
Xiao N, Eidenschenk C, Krebs P, Brandl K, Blasius AL, Xia Y, et al. The Tpl2 mutation Sluggish impairs type I IFN production and increases susceptibility to group B streptococcal disease. J Immunol. 2009;183:7975–83. https://doi.org/10.4049/jimmunol.0902718.
Degirmenci U, Wang M, Hu J. Targeting aberrant RAS/RAF/MEK/ERK signaling for cancer therapy. Cells. 2020;9 https://doi.org/10.3390/cells9010198.
Man RJ, Zhang YL, Jiang AQ, Zhu HL. A patent review of RAF kinase inhibitors (2010-2018). Expert Opin Ther Pat. 2019;29:675–88. https://doi.org/10.1080/13543776.2019.1651842.
Gringhuis SI, van der Vlist M, van den Berg LM, den Dunnen J, Litjens M, Geijtenbeek TB. HIV-1 exploits innate signaling by TLR8 and DC-SIGN for productive infection of dendritic cells. Nat Immunol. 2010;11:419–26. https://doi.org/10.1038/ni.1858.
Gringhuis SI, den Dunnen J, Litjens M, van Het Hof B, van Kooyk Y, Geijtenbeek TB. C-type lectin DC-SIGN modulates Toll-like receptor signaling via Raf-1 kinase-dependent acetylation of transcription factor NF-kappaB. Immunity. 2007;26:605–16. https://doi.org/10.1016/j.immuni.2007.03.012.
Simeone E, Scognamiglio G, Capone M, Giannarelli D, Grimaldi AM, Mallardo D, et al. A monocentric phase I study of vemurafenib plus cobimetinib plus PEG-interferon (VEMUPLINT) in advanced melanoma patients harboring the V600BRAF mutation. J Transl Med. 2021;19:17. https://doi.org/10.1186/s12967-020-02680-7.
Suddason T, Anwar S, Charlaftis N, Gallagher E. T-cell-specific deletion of Map3k1 reveals the critical role for Mekk1 and Jnks in Cdkn1b-dependent proliferative expansion. Cell Rep. 2016;14:449–57. https://doi.org/10.1016/j.celrep.2015.12.047.
Gallagher E, Enzler T, Matsuzawa A, Anzelon-Mills A, Otero D, Holzer R, et al. Kinase MEKK1 is required for CD40-dependent activation of the kinases Jnk and p38, germinal center formation, B cell proliferation and antibody production. Nat Immunol. 2007;8:57–63. https://doi.org/10.1038/ni1421.
Yoshida R, Takaesu G, Yoshida H, Okamoto F, Yoshioka T, Choi Y, et al. TRAF6 and MEKK1 play a pivotal role in the RIG-I-like helicase antiviral pathway. J Biol Chem. 2008;283:36211–20. https://doi.org/10.1074/jbc.M806576200.
Chayama K, Papst PJ, Garrington TP, Pratt JC, Ishizuka T, Webb S, et al. Role of MEKK2-MEK5 in the regulation of TNF-alpha gene expression and MEKK2-MKK7 in the activation of c-Jun N-terminal kinase in mast cells. Proc Natl Acad Sci U S A. 2001;98:4599–604. https://doi.org/10.1073/pnas.081021898.
Wu N, Sun H, Zhao X, Zhang Y, Tan J, Qi Y, et al. MAP3K2-regulated intestinal stromal cells define a distinct stem cell niche. Nature. 2021;592:606–10. https://doi.org/10.1038/s41586-021-03283-y.
Chao TH, Hayashi M, Tapping RI, Kato Y, Lee JD. MEKK3 directly regulates MEK5 activity as part of the big mitogen-activated protein kinase 1 (BMK1) signaling pathway. J Biol Chem. 1999;274:36035–8. https://doi.org/10.1074/jbc.274.51.36035.
Castro M, Lavina B, Ando K, Alvarez-Aznar A, Abu Taha A, Brakebusch C, et al. CDC42 deletion elicits cerebral vascular malformations via increased MEKK3-dependent KLF4 expression. Circ Res. 2019;124:1240–52. https://doi.org/10.1161/CIRCRESAHA.118.314300.
Shi L, Zhang Y, Xia Y, Li C, Song Z, Zhu J. MiR-150-5p protects against septic acute kidney injury via repressing the MEKK3/JNK pathway. Cell Signal. 2021;86:110101. https://doi.org/10.1016/j.cellsig.2021.110101.
Zhai C, Cong H, Hou K, Hu Y, Zhang J, Zhang Y, et al. Effects of miR-124-3p regulation of the p38MAPK signaling pathway via MEKK3 on apoptosis and proliferation of macrophages in mice with coronary atherosclerosis. Adv. Clin Exp Med. 2020;29:803–12. https://doi.org/10.17219/acem/121926.
Ning S, Pagano JS, Barber GN. IRF7: activation, regulation, modification and function. Genes Immun. 2011;12:399–414. https://doi.org/10.1038/gene.2011.21.
Nakamura K, Shichita T. Cellular and molecular mechanisms of sterile inflammation in ischaemic stroke. J Biochem. 2019;165:459–64. https://doi.org/10.1093/jb/mvz017.
Suzuki Y, Asami M, Takahashi D, Sakane F. Diacylglycerol kinase eta colocalizes and interacts with apoptosis signal-regulating kinase 3 in response to osmotic shock. Biochem Biophys Rep. 2021;26:101006. https://doi.org/10.1016/j.bbrep.2021.101006.
Xing C, Wang M, Ajibade AA, Tan P, Fu C, Chen L, et al. Microbiota regulate innate immune signaling and protective immunity against cancer. Cell Host Microbe. 2021;29(959-74):e7. https://doi.org/10.1016/j.chom.2021.03.016.
Edlund S, Bu S, Schuster N, Aspenstrom P, Heuchel R, Heldin NE, et al. Transforming growth factor-beta1 (TGF-beta)-induced apoptosis of prostate cancer cells involves Smad7-dependent activation of p38 by TGF-beta-activated kinase 1 and mitogen-activated protein kinase kinase 3. Mol Biol Cell. 2003;14:529–44. https://doi.org/10.1091/mbc.02-03-0037.
Sanchez-Garrido J, Shenoy AR. Regulation and repurposing of nutrient sensing and autophagy in innate immunity. Autophagy. 2021;17:1571–91. https://doi.org/10.1080/15548627.2020.1783119.
Zhang B, Li M, Chen L, Yang K, Shan Y, Zhu L, et al. The TAK1-JNK cascade is required for IRF3 function in the innate immune response. Cell Res. 2009;19:412–28. https://doi.org/10.1038/cr.2009.8.
Bruni D, Sebastia J, Dunne S, Schroder M, Butler MP. A novel IRAK1-IKKepsilon signaling axis limits the activation of TAK1-IKKbeta downstream of TLR3. J Immunol. 2013;190:2844–56. https://doi.org/10.4049/jimmunol.1202042.
Lee HW, Choi HY, Joo KM, Nam DH. Tumor progression locus 2 (Tpl2) kinase as a novel therapeutic target for cancer: double-sided effects of Tpl2 on cancer. Int J Mol Sci. 2015;16:4471–91. https://doi.org/10.3390/ijms16034471.
Liu HY. Down-regulation of miR-144 after Mycobacterium tuberculosis infection promotes inflammatory factor secretion from macrophages through the Tpl2/ERK pathway. Cell Mol Biol. 2016;62:87–93.
Xu D, Matsumoto ML, McKenzie BS, Zarrin AA. TPL2 kinase action and control of inflammation. Pharmacol Res. 2018;129:188–93. https://doi.org/10.1016/j.phrs.2017.11.031.
Shinkura R, Kitada K, Matsuda F, Tashiro K, Ikuta K, Suzuki M, et al. Alymphoplasia is caused by a point mutation in the mouse gene encoding Nf-kappa b-inducing kinase. Nat Genet. 1999;22:74–7. https://doi.org/10.1038/8780.
Schoggins JW, Wilson SJ, Panis M, Murphy MY, Jones CT, Bieniasz P, et al. A diverse range of gene products are effectors of the type I interferon antiviral response. Nature. 2011;472:481–5. https://doi.org/10.1038/nature09907.
Malinin NL, Boldin MP, Kovalenko AV, Wallach D. MAP3K-related kinase involved in NF-kappaB induction by TNF, CD95 and IL-1. Nature. 1997;385:540–4. https://doi.org/10.1038/385540a0.
Pflug KM, Sitcheran R. Targeting NF-kappaB-inducing kinase (NIK) in immunity, inflammation, and cancer. Int J Mol Sci. 2020;21 https://doi.org/10.3390/ijms21228470.
Ghosh S, Dass JFP. Study of pathway cross-talk interactions with NF-kappaB leading to its activation via ubiquitination or phosphorylation: a brief review. Gene. 2016;584:97–109. https://doi.org/10.1016/j.gene.2016.03.008.
Yu H, Lin L, Zhang Z, Zhang H, Hu H. Targeting NF-kappaB pathway for the therapy of diseases: mechanism and clinical study. Signal Transduct Target Ther. 2020;5:209. https://doi.org/10.1038/s41392-020-00312-6.
Sun SC. The non-canonical NF-kappaB pathway in immunity and inflammation. Nat Rev Immunol. 2017;17:545–58. https://doi.org/10.1038/nri.2017.52.
Liu R, Lin Y, Jia R, Geng Y, Liang C, Tan J, et al. HIV-1 Vpr stimulates NF-kappaB and AP-1 signaling by activating TAK1. Retrovirology. 2014;11:45. https://doi.org/10.1186/1742-4690-11-45.
Postler TS, Desrosiers RC. The cytoplasmic domain of the HIV-1 glycoprotein gp41 induces NF-kappaB activation through TGF-beta-activated kinase 1. Cell Host Microbe. 2012;11:181–93. https://doi.org/10.1016/j.chom.2011.12.005.
Li X, Josef J, Marasco WA. Hiv-1 Tat can substantially enhance the capacity of NIK to induce IkappaB degradation. Biochem Biophys Res Commun. 2001;286:587–94. https://doi.org/10.1006/bbrc.2001.5442.
Wu Y, Ma L, Cai S, Zhuang Z, Zhao Z, Jin S, et al. RNA-induced liquid phase separation of SARS-CoV-2 nucleocapsid protein facilitates NF-kappaB hyper-activation and inflammation. Signal Transduct Target Ther. 2021;6:167. https://doi.org/10.1038/s41392-021-00575-7.
Nishitsuji H, Iwahori S, Ohmori M, Shimotohno K, Murata T. Ubiquitination of SARS-CoV-2 NSP6 and ORF7a facilitates NF-kappaB activation. mBio. 2022;13:e0097122. https://doi.org/10.1128/mbio.00971-22.
Dey N, Liu T, Garofalo RP, Casola A. TAK1 regulates NF-KappaB and AP-1 activation in airway epithelial cells following RSV infection. Virology. 2011;418:93–101. https://doi.org/10.1016/j.virol.2011.07.007.
Choudhary S, Boldogh S, Garofalo R, Jamaluddin M, Brasier AR. Respiratory syncytial virus influences NF-kappaB-dependent gene expression through a novel pathway involving MAP3K14/NIK expression and nuclear complex formation with NF-kappaB2. J Virol. 2005;79:8948–59. https://doi.org/10.1128/JVI.79.14.8948-8959.2005.
Deng F, Xu G, Cheng Z, Huang Y, Ma C, Luo C, et al. Hepatitis B surface antigen suppresses the activation of nuclear factor kappa B pathway via interaction with the TAK1-TAB2 complex. Front Immunol. 2021;12:618196. https://doi.org/10.3389/fimmu.2021.618196.
Park SG, Ryu HM, Lim SO, Kim YI, Hwang SB, Jung G. Interferon-gamma inhibits hepatitis B virus-induced NF-kappaB activation through nuclear localization of NF-kappaB-inducing kinase. Gastroenterology. 2005;128:2042–53. https://doi.org/10.1053/j.gastro.2005.03.002.
Mogensen TH, Melchjorsen J, Hollsberg P, Paludan SR. Activation of NF-kappa B in virus-infected macrophages is dependent on mitochondrial oxidative stress and intracellular calcium: downstream involvement of the kinases TGF-beta-activated kinase 1, mitogen-activated kinase/extracellular signal-regulated kinase kinase 1, and I kappa B kinase. J Immunol. 2003;170:6224–33. https://doi.org/10.4049/jimmunol.170.12.6224.
Lu X, Huang C, Zhang Y, Lin Y, Wang X, Li Q, et al. The Us2 gene product of herpes simplex virus 2 modulates NF-kappaB activation by targeting TAK1. Sci Rep. 2017;7:8396. https://doi.org/10.1038/s41598-017-08856-4.
Wu L, Nakano H, Wu Z. The C-terminal activating region 2 of the Epstein-Barr virus-encoded latent membrane protein 1 activates NF-kappaB through TRAF6 and TAK1. J Biol Chem. 2006;281:2162–9. https://doi.org/10.1074/jbc.M505903200.
Lee FS, Hagler J, Chen ZJ, Maniatis T. Activation of the IkappaB alpha kinase complex by MEKK1, a kinase of the JNK pathway. Cell. 1997;88:213–22. https://doi.org/10.1016/s0092-8674(00)81842-5.
Lee FS, Peters RT, Dang LC, Maniatis T. MEKK1 activates both IkappaB kinase alpha and IkappaB kinase beta. Proc Natl Acad Sci U S A. 1998;95:9319–24. https://doi.org/10.1073/pnas.95.16.9319.
Schmidt C, Peng B, Li Z, Sclabas GM, Fujioka S, Niu J, et al. Mechanisms of proinflammatory cytokine-induced biphasic NF-kappaB activation. Mol Cell. 2003;12:1287–300. https://doi.org/10.1016/s1097-2765(03)00390-3.
Yang J, Lin Y, Guo Z, Cheng J, Huang J, Deng L, et al. The essential role of MEKK3 in TNF-induced NF-kappaB activation. Nat Immunol. 2001;2:620–4. https://doi.org/10.1038/89769.
Huangfu WC, Omori E, Akira S, Matsumoto K, Ninomiya-Tsuji J. Osmotic stress activates the TAK1-JNK pathway while blocking TAK1-mediated NF-kappaB activation: TAO2 regulates TAK1 pathways. J Biol Chem. 2006;281:28802–10. https://doi.org/10.1074/jbc.M603627200.
Espada CE, St Gelais C, Bonifati S, Maksimova VV, Cahill MP, Kim SH, et al. TRAF6 and TAK1 contribute to SAMHD1-mediated negative regulation of NF-kappaB signaling. J Virol. 2021;95 https://doi.org/10.1128/JVI.01970-20.
Ghosn J, Taiwo B, Seedat S, Autran B, Katlama C. Hiv. Lancet. 2018;392:685–97. https://doi.org/10.1016/S0140-6736(18)31311-4.
Sui Z, Fan S, Sniderhan L, Reisinger E, Litzburg A, Schifitto G, et al. Inhibition of mixed lineage kinase 3 prevents HIV-1 Tat-mediated neurotoxicity and monocyte activation. J Immunol. 2006;177:702–11. https://doi.org/10.4049/jimmunol.177.1.702.
Saminathan P, Kevadiya BD, Marker DF, Gendelman HE, Gorantla S, Gelbard HA. Broad spectrum mixed lineage kinase type 3 inhibition and HIV-1 persistence in macrophages. J Neuroimmune Pharmacol. 2019;14:44–51. https://doi.org/10.1007/s11481-018-09829-8.
Bodner A, Maroney AC, Finn JP, Ghadge G, Roos R, Miller RJ. Mixed lineage kinase 3 mediates gp120IIIB-induced neurotoxicity. J Neurochem. 2002;82:1424–34. https://doi.org/10.1046/j.1471-4159.2002.01088.x.
Eggert D, Dash PK, Gorantla S, Dou H, Schifitto G, Maggirwar SB, et al. Neuroprotective activities of CEP-1347 in models of neuroAIDS. J Immunol. 2010;184:746–56. https://doi.org/10.4049/jimmunol.0902962.
Ma Q, Gelbard HA, Maggirwar SB, Dewhurst S, Gendelman HE, Peterson DR, et al. Pharmacokinetic interactions of CEP-1347 and atazanavir in HIV-infected patients. J Neurovirol. 2013;19:254–60. https://doi.org/10.1007/s13365-013-0172-z.
Marker DF, Tremblay ME, Puccini JM, Barbieri J, Gantz Marker MA, Loweth CJ, et al. The new small-molecule mixed-lineage kinase 3 inhibitor URMC-099 is neuroprotective and anti-inflammatory in models of human immunodeficiency virus-associated neurocognitive disorders. J Neurosci. 2013;33:9998–10010. https://doi.org/10.1523/JNEUROSCI.0598-13.2013.
Chen A, Xu J, Lai H, D'Agati VD, Guan TJ, Badal S, et al. Inhibition of apoptosis signal-regulating kinase 1 mitigates the pathogenesis of human immunodeficiency virus-associated nephropathy. Nephrol Dial Transplant. 2021;36:430–41. https://doi.org/10.1093/ndt/gfaa198.
Perez-Moraga R, Fores-Martos J, Suay-Garcia B, Duval JL, Falco A, Climent J. A COVID-19 drug repurposing strategy through quantitative homological similarities using a topological data analysis-based framework. Pharmaceutics. 2021;13 https://doi.org/10.3390/pharmaceutics13040488.
Del Bufalo F, Ceglie G, Cacchione A, Alessi I, Colafati GS, Carai A, et al. BRAF V600E inhibitor (vemurafenib) for BRAF V600E mutated low grade gliomas. Front Oncol. 2018;8:526. https://doi.org/10.3389/fonc.2018.00526.
Thirumal Kumar D, Shree Devi MS, Udhaya Kumar S, Sherlin A, Mathew A, Lakshmipriya M, et al. Understanding the activating mechanism of the immune system against COVID-19 by Traditional Indian Medicine: Network pharmacology approach. Adv Protein Chem Struct Biol. 2022;129:275–379. https://doi.org/10.1016/bs.apcsb.2021.11.007.
Zhang Y, Greer RA, Song Y, Praveen H, Song Y. In silico identification of available drugs targeting cell surface BiP to disrupt SARS-CoV-2 binding and replication: drug repurposing approach. Eur J Pharm Sci. 2021;160:105771. https://doi.org/10.1016/j.ejps.2021.105771.
Sinha S, Cheng K, Schaffer AA, Aldape K, Schiff E, Ruppin E. In vitro and in vivo identification of clinically approved drugs that modify ACE2 expression. Mol Syst Biol. 2020;16:e9628. https://doi.org/10.15252/msb.20209628.
Sosman JA, Kim KB, Schuchter L, Gonzalez R, Pavlick AC, Weber JS, et al. Survival in BRAF V600-mutant advanced melanoma treated with vemurafenib. N Engl J Med. 2012;366:707–14. https://doi.org/10.1056/NEJMoa1112302.
Linden T, Hanses F, Domingo-Fernandez D, DeLong LN, Kodamullil AT, Schneider J, et al. Machine learning based prediction of COVID-19 mortality suggests repositioning of anticancer drug for treating severe cases. Artif Intell Life Sci. 2021;1:100020. https://doi.org/10.1016/j.ailsci.2021.100020.
Dwivedy A, Mariadasse R, Ahmad M, Chakraborty S, Kar D, Tiwari S, et al. Characterization of the NiRAN domain from RNA-dependent RNA polymerase provides insights into a potential therapeutic target against SARS-CoV-2. PLoS Comput Biol. 2021;17:e1009384. https://doi.org/10.1371/journal.pcbi.1009384.
Taguchi YH, Turki T. A new advanced in silico drug discovery method for novel coronavirus (SARS-CoV-2) with tensor decomposition-based unsupervised feature extraction. PLoS One. 2020;15:e0238907. https://doi.org/10.1371/journal.pone.0238907.
Awad IE, Abu-Saleh AAA, Sharma S, Yadav A, Poirier RA. High-throughput virtual screening of drug databanks for potential inhibitors of SARS-CoV-2 spike glycoprotein. J Biomol Struct Dyn. 2022;40:2099–112. https://doi.org/10.1080/07391102.2020.1835721.
Wan W, Zhu S, Li S, Shang W, Zhang R, Li H, et al. High-throughput screening of an FDA-approved drug library identifies inhibitors against arenaviruses and SARS-CoV-2. ACS Infect Dis. 2021;7:1409–22. https://doi.org/10.1021/acsinfecdis.0c00486.
Islam T, Rahman MR, Aydin B, Beklen H, Arga KY, Shahjaman M. Integrative transcriptomics analysis of lung epithelial cells and identification of repurposable drug candidates for COVID-19. Eur J Pharmacol. 2020;887:173594. https://doi.org/10.1016/j.ejphar.2020.173594.
Wang LW, Jiang S, Gewurz BE. Epstein-Barr virus LMP1-mediated oncogenicity. J Virol. 2017;91 https://doi.org/10.1128/JVI.01718-16.
Voigt S, Sterz KR, Giehler F, Mohr AW, Wilson JB, Moosmann A, et al. A central role of IKK2 and TPL2 in JNK activation and viral B-cell transformation. Nat Commun. 2020;11:685. https://doi.org/10.1038/s41467-020-14502-x.
Peiris JS, Cheung CY, Leung CY, Nicholls JM. Innate immune responses to influenza A H5N1: friend or foe? Trends Immunol. 2009;30:574–84. https://doi.org/10.1016/j.it.2009.09.004.
Ruan T, Sun Y, Zhang J, Sun J, Liu W, Prinz RA, et al. H5N1 infection impairs the alveolar epithelial barrier through intercellular junction proteins via Itch-mediated proteasomal degradation. Commun Biol. 2022;5:186. https://doi.org/10.1038/s42003-022-03131-3.
Wang J, Kan X, Li X, Sun J, Xu X. Porcine epidemic diarrhoea virus (PEDV) infection activates AMPK and JNK through TAK1 to induce autophagy and enhance virus replication. Virulence. 2022;13:1697–712. https://doi.org/10.1080/21505594.2022.2127192.
Ianevski A, Yao R, Biza S, Zusinaite E, Mannik A, Kivi G, et al. Identification and tracking of antiviral drug combinations. Viruses. 2020;12 https://doi.org/10.3390/v12101178.
Holzberg M, Boergeling Y, Schrader T, Ludwig S, Ehrhardt C. Vemurafenib limits influenza A virus propagation by targeting multiple signaling pathways. Front Microbiol. 2017;8:2426. https://doi.org/10.3389/fmicb.2017.02426.
Benedict A, Bansal N, Senina S, Hooper I, Lundberg L, de la Fuente C, et al. Repurposing FDA-approved drugs as therapeutics to treat Rift Valley fever virus infection. Front Microbiol. 2015;6:676. https://doi.org/10.3389/fmicb.2015.00676.
Michaelis M, Paulus C, Loschmann N, Dauth S, Stange E, Doerr HW, et al. The multi-targeted kinase inhibitor sorafenib inhibits human cytomegalovirus replication. Cell Mol Life Sci. 2011;68:1079–90. https://doi.org/10.1007/s00018-010-0510-8.
Lundberg L, Brahms A, Hooper I, Carey B, Lin SC, Dahal B, et al. Repurposed FDA-Approved drug sorafenib reduces replication of Venezuelan equine encephalitis virus and other alphaviruses. Antiviral Res. 2018;157:57–67. https://doi.org/10.1016/j.antiviral.2018.07.005.
Descamps V, Helle F, Louandre C, Martin E, Brochot E, Izquierdo L, et al. The kinase-inhibitor sorafenib inhibits multiple steps of the Hepatitis C virus infectious cycle in vitro. Antiviral Res. 2015;118:93–102. https://doi.org/10.1016/j.antiviral.2015.03.012.
Uday RVS, Misra R, Harika A, Dolui S, Saha A, Pal U, et al. Dabrafenib, idelalisib and nintedanib act as significant allosteric modulator for dengue NS3 protease. PloS one. 2021;16:e0257206. https://doi.org/10.1371/journal.pone.0257206.
He TS, Huang J, Chen T, Zhang Z, Cai K, Yu J, et al. The kinase MAP4K1 inhibits cytosolic RNA-induced antiviral signaling by promoting proteasomal degradation of TBK1/IKKepsilon. Microbiol Spectr. 2021;9:e0145821. https://doi.org/10.1128/Spectrum.01458-21.
Chen Y, Wang L, Jin J, Luan Y, Chen C, Li Y, et al. p38 inhibition provides anti-DNA virus immunity by regulation of USP21 phosphorylation and STING activation. J Exp Med. 2017;214:991–1010. https://doi.org/10.1084/jem.20161387.
Liang D, Xiao-Feng H, Guan-Jun D, Er-Ling H, Sheng C, Ting-Ting W, et al. Activated STING enhances Tregs infiltration in the HPV-related carcinogenesis of tongue squamous cells via the c-jun/CCL22 signal. Biochim Biophys Acta. 2015;1852:2494–503. https://doi.org/10.1016/j.bbadis.2015.08.011.
Funding
The current work was supported by a special program from the Ministry of Science and Technology of China (2021YFA1101000), the Chinese National Natural Science Funds (U20A20393, 32125016, 31871405, and 31925013), Jiangsu National Science Foundation (19KJA550003), and a project Funded by the Priority Academic Program Development of Jiangsu Higher Education. The funding bodies had no role in study design; in the collection, analysis and interpretation of data; in the writing of the report; and in the decision to submit the article for publication.
Author information
Authors and Affiliations
Contributions
Jizhong Guan and Yao Fan performed the literature search and data analysis; Shuai Wang and Fangfang Zhou drafted and critically revised the work.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guan, J., Fan, Y., Wang, S. et al. Functions of MAP3Ks in antiviral immunity. Immunol Res 71, 814–832 (2023). https://doi.org/10.1007/s12026-023-09401-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12026-023-09401-4